Abstract
In γ-proteobacterial species, such as Escherichia coli, the Arc (anoxic redox control) two-component system plays a major role in mediating the metabolic transition from aerobiosis to anaerobiosis, and thus is crucial for anaerobic growth but dispensable for aerobic growth. In Shewanella oneidensis, a bacterium renowned for respiratory versatility, Arc (SoArc) primarily affects aerobic growth. To date, how this occurs has remained largely unknown although the growth defect resulting from the loss of DNA-binding response regulator SoArcA is tryptone-dependent. In this study, we demonstrated that the growth defect is in part linked to utilization of oligopeptides and di-tripeptides, and peptide uptake but not peptide degradation is significantly affected by the SoArcA loss. A systematic characterization of major small peptide uptake systems manifests that ABC peptide transporter Sap and four proton-dependent oligopeptide transporters (POTs) are responsible for transport of oligopeptides and di-tripeptides respectively. Among them, Sap and DtpA (one of POTs) are responsive to the SoarcA mutation but only dtpA is under the direct control of SoArcA. We further showed that both Sap and DtpA, when overproduced, improve growth of the SoarcA mutant. While the data firmly establish a link between transport of oligopeptides and di-tripeptides and the SoarcA mutation, other yet-unidentified factors are implicated in the growth defect resulting from the SoArcA loss.
Similar content being viewed by others
Introduction
Two-component systems (TCSs) are employed by prokaryotes to respond to constantly changing environmental conditions. In Escherichia coli, the Arc (anoxic redox control) system (EcArc) regulates gene expression in response to redox conditions1,2. EcArc consists of a transmembrane sensor kinase EcArcB and a DNA binding response regulator EcArcA1. Under anaerobic or microaerobic respiratory conditions, EcArcB undergoes autophosphorylation by sensing the redox state of quinone pool, eventually resulting in phosphorylated EcArcA (EcArcA-P) at the 54th Asp residue through a phospho-relay mechanism3,4. EcArcA-P functions primarily as a global repressor of aerobic metabolic pathways, the tricarboxylic acid (TCA) cycle in particular. As a result, the EcArc system is crucial for anaerobic growth but dispensable for aerobic growth in E. coli5,6. Besides, as a global regulatory system, EcArc is implicated in diverse biological processes with a regulon consisting of hundreds of target genes7,8.
Shewanella, a group of facultative Gram-negative anaerobes renowned for their remarkable respiratory abilities, have become a research model for bacterial physiology9. As respiration is the predominant means for energy production in these bacteria, global regulators mediating metabolic transition in response to the availability of different electron acceptors (EAs) have been investigated for decades and many surprising observations have been made, especially in S. oneidensis, which is the most intensively studied representative of the genus.
The Arc system in S. oneidensis (SoArc) is atypical as the role of the sensor kinase is fulfilled by two proteins, ArcS (or ArcB1) and HptA10,11,12. Unlike EcArcB, which senses changes in the quinone composition, ArcS unlikely detects quinol species because it does not have the redox-sensitive cysteine residues of EcArcB1,2. Instead, ArcS has a periplasmic CaChe domain, which may function to sense yet-unknown extracellular signals11. This difference probably underpins the distinct influences of the two Arc systems on growth: EcArc is of the highest importance for micro-aerobic and anaerobic growth whereas SoArc plays a critical role in aerobic growth13. Despite this, activation of the SoArc system is similar to its E. coli counterpart through phospho-relay and 15-bp DNA-binding motifs for Arc proteins are highly conserved6,11,13,14,15. Interestingly, E. coli and S. oneidensis ArcA regulons differ from each other substantially; only 6 out of at least 50 members are shared13. Given that these two bacteria are rather close phylogenetically and there are more than 2,000 genes in common in E. coli and S. oneidensis genomes1,13, the difference is surprising. As a result, it is conceivable that the observed growth defect is not due to interference with expression of genes encoding established members of the EcArcA regulon16,17.
Further analyses of the SoarcA null mutant reveal that the growth defect is substantial in rich media such as Lysogeny broth (LB) but diminished in minimal media13,16,18. The growth defect re-emerges with addition of tryptone, suggesting that the SoarcA mutation may compromise the efficiency of oligopeptide metabolism18. Moreover, the SoArcA loss also results in a severely impaired cell envelope through yet-unknown mechanisms19. Given that SoArc is atypical and implicated in many biological processes clearly different from those revealed from bacteria hosting a canonical one such as E. coli, this regulatory system can serve as an ideal model to study respiration control for environmental microorganisms. The goal of this study was to unravel the mechanisms for the growth defect of the SoarcA mutant. We established that the major systems for oligopeptide (ATP peptide transporter Sap) and di-tripeptide (proton-dependent oligopeptide transporters, POTs) transport are subject to SoArcA regulation. Although these transporting systems are clearly implicated in promotion of aerobic growth on short peptides, only one of di-tripeptide transporter is under the direct control of SoArcA. As the growth defect of the SoarcA mutant could not be fully corrected by manipulated production of either system, we propose that the SoarcA mutation impairs growth through multi-fold complex impacts in physiology.
Results
The arcA mutation compromises peptide utilization in S. oneidensis
Our preliminary analysis had demonstrated that the growth defect of the S. oneidensis arcA mutant in complex medium LB is dependent on tryptone18. As the SoarcA mutant cultivated in mineral medium MS with lactate as the carbon source (dubbed as MS-L throughout this study) is not defective with respect to aerobic growth (Figs 1A, S1A), this study was initiated by systematically assessing effects of tryptone on growth of the wild-type and ∆SoarcA strains in MS-L with tryptone addition. Clearly, growth of the wild-type increased with tryptone when its concentrations were no more than 0.2%, and became less sensitive to further augment to 0.5% in tryptone amounts (Fig. 1B). By generation times for 2 h during the exponential phase (Table S1), there was a 1.5-fold difference in growth rates between cells grown with 0.25% tryptone and without. In contrast, the ∆SoarcA strain was not responsive to the treatment with tryptone at 0.2% or lower, and grew slightly faster with 0.5% tryptone (Fig. 1C). These data verify that the SoarcA mutation compromises the growth- promoting effect of tryptone. Therefore, we reasoned that utilization of oligopeptides and/or amino acids is impaired by the mutation because tryptone is abundant in such nutrients, a result of incomplete enzymatic hydrolysis of casein.
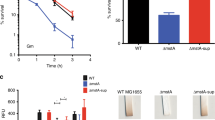
Comparative analysis of growth of the wild-type (WT) and the arcA mutant of S. oneidensis. (A) Growth on LB plates. Cultures of indicated strains prepared to contain approximately 109 cfu/ml were regarded as the undiluted (dilution factor, 0), which were subjected to 10-fold series dilution. Five microliters of each dilution was dropped on indicated agar plates. Results were recorded after incubation of 18 h and 30 h for LB and MS-L respectively. Successful complementation by expressing a copy of the arcA gene in trans (ArcA) has been performed and reported repeatedly before. Data for cultures grown in liquid LB were shown in Fig. S1A. (B,C) Growth of WT and ∆arcA in MS-L without and with tryptone (T). Concentrations of tryptone were given in subscript. (D) Growth of WT in MS-L containing 0.5% oligopeptide (O-P) mixture, di-tripeptide (DT-P) mixture, or both (O/DT-P, refer to Table 1 for O-P and DT-P information). (E) Growth of ∆arcA in MS-L without or with 0.5% O/DT-P. Experiments were performed independently at least 5 times, and data were presented as the average and error bars representing standard errors.
To test this, impacts of tryptone, casamino acids, and an amino acid mixture on growth of the wild-type and ∆SoarcA strains were compared. In contrast to tryptone, neither casamino acids nor the amino acid mixture could elicit a significant difference in growth rates of both strains (Fig. S1B,C). This observation rules out the possibility that amino acids are an important factor responsible for the growth defect of the ∆SoarcA strain. Subsequently, we tested effects of oligopeptides (O-P, peptides of 4 and more residues, Table 1) of different lengths on growth of the wild-type and ∆SoarcA strains. Additions of a variety of oligopeptides only modestly improved growth of the wild-type (compared to that in MS-L) but showed no detectable effects on growth of the ∆SoarcA strain (Fig. 1D,E). Similar results were obtained with a mixture of di-tripeptides (DT-P) (Table 1). When both O-P and DT-P (O/DT-P) were present, growth of the wild-type was further accelerated whereas the ∆SoarcA strain remained unaffected. These data therefore suggest that the SoarcA mutant is defective in utilization of short peptides.
The ∆SoarcA strain is heavily defective in growing on O/DT-P
Growth monitored above was in fact mainly supported by lactate, the carbon source in MS-L, which interferes with the contribution of the peptide mixtures under test. As an attempt to show the growth defect of the SoarcA mutant more clearly, growth on O/DT-P, casamino acids, and an amino acid mixture as carbon sources was assessed (Fig. 2A, 2B, 2C). Both the wild-type and ∆SoarcA strains were able to grow in MS with each of these carbon sources at concentrations up to 0.5% (Fig. S2). The amino acid mixture at all testing concentrations supported these two strains to grow comparably (Fig. 2C), manifesting that the SoarcA mutation does not have a significant impact on utilization of amino acids. In contrast, O/DT-P at all testing concentrations were not only more effective than the amino acid mixture in supporting growth, but more importantly, distinguished from the wild-type from the SoarcA mutant with respect to growth rate (Fig. 2A). Clearly, the difference in growth rates between them was substantial (Table S1). In the case of casamino acids, the growth difference between the two strains was negligible, noticeable, and significant with 0.1%, 0.2%, and 0.5% respectively (Fig. 2B). This further supports the impact of the SoarcA mutation in utilization of O/DT-P because small peptides make up ∼6% of casamino acids (250–500 (molecular weight), ∼6%, BD BactoTM).
Defects of the S. oneidensis arcA mutant in aerobic growth are associated with peptide metabolism. (A) Growth of the wild-type (WT) and ∆arcA strains in MS with the peptide mixture (O/DT-P, 0.2%). (B) Growth the wild-type and ∆arcA strains in MS with casamino acids (CA, 0.2% and 0.5%). (C) Growth of the the wild-type and ∆arcA strains MS with the amino acid mixture (AA, 0.5%). More data for growth of WT and ∆arcA on these carbon sources were given in Fig. S2. Experiments were performed independently at least 5 times, and data were presented as the average and error bars representing standard errors.
Peptide digestion is not significantly affected by the SoarcA mutation
Utilization of peptides by the cell depends on two distinct steps, import from the outside and digestion into amino acids. To evaluate the contribution of peptide digestion to the growth difference between the wild-type and ∆SoarcA strains, we measured activity of metalloaminopeptidases, which are predominant in short peptide hydrolysis20. Both the wild-type and ∆SoarcA strains were grown in MS-L with tryptone to the mid-log phase and cells were collected by centrifugation. The pelleted cells were resuspended and disrupted by sonication with external cooling on ice, and after removal of cellular debris by centrifugation, cell extracts were assayed for aminopeptidase activity using fluorescent peptide-AMC (7-amido-4-methylcoumarin) substrates21. As shown in Fig. 3A, the amount of hydrolyzed peptide of the cell extracts from both the wild-type and ∆SoarcA strains linearly increased with time. The difference in peptidase activity between two strains, by normalized total protein amounts of each sample, was statistically insignificant. To verify that the hydrolysis is mainly attributed to metalloaminopeptidases, activities in the wild-type and ∆SoarcA cell extracts were assayed in the presence of 1,10-phenanthroline (OPT), a metal chelator functioning as unspecific and potent metalloprotease inhibitor21. Clearly, OPT inhibited the hydrolysis of peptide-AMC strongly; at 1 mM the maximum inhibition was achieved. In addition, we examined the effect of serineproteases on peptide hydrolysis. In the presence of 1 mM AEBSF (4-(2-aminoethlyl)benzenesulfony fluride), a broad spectrum and irreversible inhibitor for serineproteases22, the amount of fluorescence in the wild-type cell extracts was also affected (Fig. S3), suggesting a role of serineproteases in peptide digestion. However, given that the impact of AEBSF was substantially weaker than that of OPT, it is apparent that metalloaminopeptidases are a major contributing factor for peptide digestion. Importantly, the data obtained from the wild-type and ∆SoarcA strains were similar. Additionally, we performed the droplet assay to assess susceptibilities of the wild-type and ∆SoarcA strains to OPT and found that they were similar on MS-L plates containing tryptone (Fig. 3B). These data thus rule out the possibility that peptide hydrolysis is critically responsible for the growth defect of the ∆SoarcA strain.
Contribution of peptide digestion to the growth defect of the S. oneidensis arcA mutant is negligible. (A) Metallaminopeptidase activity in cell extracts. An aliquot of cell extracts (of indicated strains grown to the mid-log phase in MS-L containing tryptone (MS-L-T)) was mixed with fluorescent peptide-AMC substrate. In parallel, a second aliquot was mixed with the substrate and OPT, a metalloprotease inhibitor. Fluorescence emission due to proteolytic cleavage of the substrate was measured in a microplate reader. (B) OPT susceptibility assay as performed the same as described in Fig. 1A on MS-L-T plates. In both panels, experiments were performed at least three times, with representative results or the average with error bars representing standard errors.
Selectivity of Sap and POTs in peptide transport
In bacteria, oligopeptide transport is predominantly mediated by transporters of three families: the proton-dependent oligopeptide transporter (POT), the ATP-binding cassette (ABC) peptide transporter, and the electrochemical potential-driven oligopeptide transporter (OPT)23,24. In the S. oneidensis genome, genes encoding POT and ABC peptide transporter system but not OPT are present25. For confirmation, the S. oneidensis proteome was screened for homologues of transporters in these three families whose oligopeptide transport activity has been established, including ABC peptide transporter systems of E. coli (OppABCDF)26 and Pseudomonas aeruginosa (DppABCDF)27, as well as a variety of POTs and OPTs typified by E. coli DtpA (formerly YdgR or TppB) and DtpB (formerly YhiP)28,29, and Shewanella pealeana Spea_3867 (Spe1)23. BLASTp screening revealed an ABC peptide transporter system, SapABCDF, and four POTs, SO_0002, SO_1277, SO_1505, and SO_3195 (Table 2). Among them, SO_0002 and SO_1277 have been proven to be di-tripeptide transporters biochemically30,31, and therefore are named as DtpA and DtpB respectively.
To determine selectivity of Sap and POT peptide transport, we constructed deletion strains for each of them. While all four POT genes are transcribed from single-gene operons, a single operon, sapABCDF, encodes the ABC peptide transporter system. In MS-L containing tryptone, ∆sap (a strain lacking the entire sap operon) mutants showed a significant defect in growth, implicating that Sap plays a major role in transporting short peptides contained in tryptone (Fig. 4A). For complementation, we manipulated expression of the sap operon by using IPTG-inducible promoter Ptac32. Defects of ∆sap in growth in tryptone-containing MS-L were corrected by expressing the missing sap genes in trans with IPTG ranging from 0.05 to 0.5 mM (Fig. 4A). According to previous calibration of the Ptac promoter within pHGE, 0.5 mM IPTG confers the activity of the Ptac promoter approximately 2000 Miller units33,34,35, more than 6 times of that of the sap promoter in the wild-type (refer to Fig. 6B). Therefore, these data, in addition to validate the correlation between the mutation and phenotypes, also indicate that the Sap peptide transporter produced in the wild-type is sufficient for importing short peptides contained in tryptone to support full growth on lactate as the carbon source. The selectivity of Sap was then investigated with O-P and DT-P as the carbon source. While the ∆sap strain grew only slightly slower than the wild-type on DT-P, it could barely do so on O-P (Fig. 4B,C). Expectedly, growth of the ∆sap strain on O-P was restored by expressing the sap operon in the presence of 0.05 mM IPTG. Moreover, we found that growth on O-P, unlike that in the tryptone-containing MS-L, further increased with IPTG concentrations (up to 0.5 mM), manifesting enhanced O-P uptake by overproduced Sap (Fig. 4B). On the contrary, Sap in overabundance had little effect on growth of the ∆sap strain on DT-P (Fig. 4C). These data indicate that the Sap system is primarily for uptake of peptides with 4 amino acid residues or larger.
Characterization of Sap systems in S. oneidensis. (A) Growth of ∆sap and its complemented strains in MS-L or MS-L-T. Expression of sap on a plasmid (psap) was driven by IPTG-inducible promoter Ptac. With IPTG ranging from 0.05 to 0.5 mM, results were similar, with data from 0.5 mM being shown. Growth of the ∆sap strain in MS-L, similar to that of WT, was not shown for clarity. (B) Growth of ∆sap and its complemented strains in MS-O-P. (C) Growth of ∆sap and its complemented strains in MS-DT-P. In both B and C, numbers represent IPTG concentrations used for induction. Experiments were performed independently at least 5 times, and data were presented as the average and error bars representing standard errors.
Characterization of POT systems in S. oneidensis. (A) Growth of the wild-type and ∆pot in MS-O-P. (B) Growth of the wild-type and ∆pot strains in MS-DT-P. Expression of dtpA on a plasmid (pdtpA) was driven by IPTG-inducible promoter Ptac. Numbers represent IPTG concentrations used for induction. (C) Alafosfolin susceptibility determined by disk diffusion assay. Ten µl of 2 mg/ml alafosfolin was added to filter paper discs (8 mm) on cultures of indicated strains. Plates were incubated at 30 °C for 24 h. Asterisks not associated with a bracket represent statistically significant difference compared to the wild-type value (*P < 0.05; **P < 0.01; ***P < 0.001). Asterisks associated with the bracket represent the difference between the two linked strains. Experiments were performed independently at least 5 times, and data were presented as the average and error bars representing standard errors.
In the case of POTs, depletion of each POT alone did not significantly affect growth of the wild-type in tryptone-containing MS-L (Fig. S4). Although this observation implies that POTs are not critical in growth improvement by tryptone when lactate is used as the carbon source, it may be a result of functional overlapping among the four putative POTs. To test this, a strain (∆pot) devoid of all four POT genes was constructed and characterized with peptides as the carbon source. Compared to the wild-type, this strain displayed normal and substantially impaired growth on O-P and DT-P as the carbon source, respectively (Fig. 5A,B). The dependence of DT-P import on POTs was further verified by assessing the susceptibility of the ∆pot strain to alafosfolin, a di-peptide analog antibiotic that is a good substrate for POTs28: the ∆pot strain exhibited substantially elevated resistance whereas the ∆sap strain was almost unaffected (Fig. 5C). For complementation, we manipulated expression of the dtpA gene by using IPTG-inducible promoter Ptac. Effects of dtpA expression on growth of the POT-deficient mutant on di-tripeptides were evident: significantly improved with 0.01 mM IPTG and fully restored to the wild-type level with 0.05 mM (Fig. 5B). DtpA in overproduction (0.5 mM IPTG) was able to further enhance growth on di-tripeptides and increased susceptibility of the ∆pot strain to alafosfolin (Fig. 5B,C). These data clearly manifest that POTs dictate transport of di-tripeptides, and that these POTs, at least some of them, can functionally complement one another.
Peptide transport has a role in explaining the ∆SoarcA growth defect
To determine whether these peptide transporters are implicated in the growth defect of the ∆SoarcA strain, we compared expression of their coding genes in the wild-type and the mutant with qRT-PCR. Cells of both strains grown with the short-peptide mixture as the carbon source to the mid-log phase were used. Analysis of transcripts of these genes revealed that the sap and dtpA genes displayed significantly reduced transcription in the ∆SoarcA strain, approximately by 30% and 3-fold respectively, while the remaining three genes were transcribed comparably in these two strains (Fig. 6A). We then utilized an integrative lacZ reporter system for confirmation36. DNA fragments of ∼400 bp upstream of the coding sequences for these operons, which are sufficiently long to cover entire promoters for most of operons, were amplified and cloned into the reporter vector pHGEI01. The resulting vectors, verified by sequencing, were introduced into relevant S. oneidensis strains for integration and subsequent removal of the antibiotic marker37. From similarly prepared cells, we surprisingly found that the differences in expression of the sap and dtpA genes were enhanced, and dtpB and SO_1505 were also modestly down-regulated in the mutant (Fig. 6B). These data indicate that at least sap and dtpA are subject to positive regulation of SoArcA, although it is clear that more investigations are needed to address the difference in data revealed by these two methods.
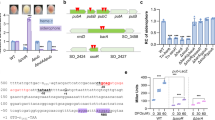
Peptide transport is partially accountable for the S. oneidensis ∆arcA growth defect. (A) Expression levels revealed by qRT-PCR assay. The wild-type and ∆arcA strains grown to the mid-log phase in MS containing 0.5% tryptone were used for the assay. The averaged values for each gene under test were normalized to that of the recA gene, giving to relative abundance (RA) of transcripts. Asterisks indicate statistically significant difference (*P < 0.05; **P < 0.01; ***P < 0.001). (B) Promoter activity assay. Indicated promoters were used to drive expression of the full-length E. coli lacZ gene within an integrative system. β-galactosidase activities in the mid-log phase cells were determined and presented as Miller Units. (C,D) Growth of the wild-type and ∆arcA strains in MS-O-P or MS-DT-P. Overexpression of sap and dtpA was achieved as described above. Numbers represent IPTG concentrations used for induction. Experiments were performed independently at least 5 times, and data were presented as the average and error bars representing standard errors.
To assess whether the down-regulated expression of sap or dtpA is responsible for the growth difference between the wild-type and ∆SoarcA strains on O-P and DT-P, we monitored growth of the ∆SoarcA strain with sap and dtpA expressed at different levels. Consistent with compromised production of Sap and POTs, the SoarcA mutant exhibited significantly impaired growth on O-P and DT-P (Fig. 6C,D). Importantly, the growth difference between the wild-type and ∆SoarcA strains on O-P and DT-P decreased with forced expression of sap and dtpA respectively. In the presence of 0.05 mM IPTG, these transporters significantly improved growth of the SoarcA mutant on the respective carbon source (Fig. 6C,D), contrasting undetectable effects on the growth of the wild-type (Fig. S5). These data support that the SoarcA mutant suffers from impairment in peptide import. In addition, we found that the SoarcA mutant with overproduced Sap and DtpA by 0.5 mM IPTG grew at further accelerated rates, but it still was inferior to the wild-type (Fig. 6C,D). For comparison, the same treatments enable ∆sap and ∆pot strains to grow faster than the wild-type (Figs 4B, 5B). Thus, while these data indicate that peptide transport has a role in determining growth of the SoarcA mutant, other factors are involved.
dtpA is under the direct control of SoArcA in S. oneidensis
The promoter region of dtpA (encoding one of POTs) but not the sap operon has been predicted to have an SoArcA-binding site (GGTAAATAGATGTAA; E. coli and S. oneidensis consensus, GTTAATTAAATGTTA)13,14. However, the confidence for this site is rather low, based on its weight value (6.5) obtained from genome screening. To test whether these genes are under the direct control of SoArcA, electrophoretic motility shift assay (EMSA) was used with the recombinant S. oneidensis His6-tagged SoArcA protein produced in E. coli and purified as described before13,14. As phosphorylation of SoArcA is required for its specific binding, only SoArcA (SoArcA-P) phosphorylated by carbamoyl phosphate was used. The DNA fragments, approximately 300 bp in length centered by the predicted SoArcA-binding motif (if available) of the genes to be tested, were amplified with 33P end-labeled primers. Among the tested PCR sequences in EMSA with phosphorylated SoArcA, dtpA showed SoArcA(-P) binding activity (Fig. 7A). In contrast, retardation of the sequences of sap and genes for other two POTs was not observed.
Only dtpA is under the direct control of ArcA in S. oneidensis. (A) EMSA assay. Experiments were performed in the presence of 0, 1, or 2 μM ArcA-P and 2–5 nM radiolabeled promoter DNA. 0.2 μg/μl poly(dI·dC) was used in all these binding reactions to block non-specific interactions. The phosphorylation of the ArcA protein was done with carbamoyl phosphate. The images shown were cropped from different gels. (B) B1H assay. Experiments were carried out as described in Materials and Methods. Colonies for the positive interactions were from M9 salt agar plates containing 25 mg/ml chloramphenicol and 12.5 mg/ml tetracycline with or without 3-amino-1,2,4-triazole (3-AT) after 24 h incubation and confirmed by streaking colonies on plates containing both 3-AT and streptomycin (12.5 mg/ml). PSO1661 and P16S were used as positive and negative controls. Experiments were performed independently at least 5 times, with representative results or the average with error bars representing standard errors.
Interactions in vivo between SoArcA and dtpA and sap promoter region sequences were then tested with bacterial one-hybrid (B1H) analysis, which has proven to be a robust technique applicable to a wide variety of different transcriptional factor families and has been frequently used in S. oneidensis33,38,39. This is enabled by the fact that E. coli arcB expression restores the wild-type phenotype of an arcS/hptA deletion in S. oneidensis and that, vice versa, upon ectopic production ArcS/HptA can functionally complement the loss of EcarcB in E. coli10,11,12. In the B1H system, vectors containing ‘bait’ (DNA) and ‘target’ (DNA-binding regulator) are co-transformed into reporter cells and positive interaction between them allows growth in the presence of 3-amino-1,2,4-triazole (3-AT). When cultured under aerobic conditions, growth was detected only from the positive control pair SoArcA/PSO_1661 (Promoter region of SO_1661)13,14 with 20 colonies on average (Fig. 6B). Given that similar studies with other regulators/cognate DNAs usually generate ∼300 colonies33,39, we reasoned that this low efficacy is likely due to that i) EcArcA proteins may compete for the target DNA and ii) most of SoArcA proteins are not phosphorylated under test conditions. To test the first possibility, the EcarcA gene was knocked out from the reporter strain and the resulting mutant was verified by genetic complementation with respect to increased sensitivity to many dyes such as toluidine blue and crystal violet40,41 (Fig. S6A). However, with this strain the B1H assay gave out similar results (Fig. S6B), indicating that the interference from EcArcA is negligible. To improve phosphorylation, we cultivated cells under fumarate-respiring conditions. In the absence of oxygen, more than 200 colonies were obtained from the reporter strains with PSO_1661/SoArcA, either with or without the EcarcA gene (Figs 6B, S6B). Importantly, the reporter strains containing PdtpA/SoArcA was able to form 50 or so colonies on 3-AT plates whereas Psap/SoArcA failed to support growth on 3-AT. These data, collectively, indicate that SoArcA directly regulates transcription of dtpA but not genes for other peptide transporters in S. oneidensis.
Discussion
The Arc system is among the most intensely investigated regulatory systems in γ-proteobacteria and its physiological function(s) known to date is largely derived from studies on the E. coli paradigm. However, as a TCS, the Arc system is rather unusual. First, the Arc system is only known to exist in four γ-proteobacteria families: Alteromonaceae, Enterobacteraceae, Vibrionaceae, and Pasterurellaceae, although its components, OmpR-family regulators and hybrid sensor histidine kinases, are common in bacteria1. Second, unlike most of TCSs, the components of all Arc systems, including both the typical (ArcB-ArcA) and the atypical (without a full length ArcB), are never encoded by genes in a single operon, not even in proximity in most cases42. Third, the atypical Arc system, as in S. oneidensis, is in fact quite common among the Alteromonads1.
Although regarded to be a global regulator with pleiotropic impacts in physiology, the canonical Arc system functions primarily as a global repressor of aerobic metabolic pathways2,43. However, this may be one side of the regulatory effects about Arc systems. There are only several overlaps in the ArcA-controlled regulons of E. coli and S. oneidensis, indicating that the physiological function of S. oneidensis ArcA is substantially different from that of the E. coli counterpart13. The best example is growth under aerobic conditions. In contrast to the E. coli paradigm, which appears to have a rather limited role during aerobic growth, the SoArc system is critical6,13,44. However, the growth phenotype resulting from the SoArcA loss is conditional, unlike the defect in cell envelope, another major defect which is independent of growth parameters tested.
The dependence of the growth phenotype on tryptone indicates that it is associated with nutrients, oligopeptides in particular18. Following this lead, in this study we examined roles of peptide transporters and peptidases in S. oneidensis. Peptidases, including both metalloaminopeptidases and serineproteases, are dispensable for the growth defect resulting from the SoArcA loss, ruling out the possibility that peptide hydrolysis is critically involved. On the contrary, oligopeptide transport is implicated. ABC peptide transporter consisting of SapABCDF and four POTs are primarily responsible for uptake of oligopeptides and di-tripeptides respectively. By using both qRT-PCR and lacZ-reporters, we concluded that expression of dtpA (encoding one of POTs) and sap is significantly down-regulated in the absence of SoArcA. Importantly, both DtpA and Sap, when manipulatedly produced, are able to improve growth of the SoarcA mutant, supporting a link between the growth defect resulting from the SoArcA loss and peptide transport. Despite this, it is clear that there are other factors implicated in the growth defect of the SoarcA mutant.
We also presented evidence from both in vitro and in vivo experiments to show that transcription of dtpA but not the sap operon is directly mediated by SoArcA. The interaction between SoArcA and the dtpA promoter region appears weak, likely due to low conservation of its SoArcA-binding site as suggested before13,14. In S. oneidensis and E. coli, it has been demonstrated that phosphorylation is essential for biological activity of ArcA and ArcA can be phosphorylated under anaerobic as well as aerobic conditions8,10,11,12,15,19,45. However, studies from us and others have argued that SoArcA proteins exist mostly in the phosphorylated form when cells grow aerobically11,13,15, contrasting the scenario observed in E. coli because the histidine kinase domain of its EcArcB is activated under anaerobic conditions45,46. Results of the B1H analysis performed in E. coli certainly support this proposal. In E. coli cells grown aerobically, positive interactions were detected only between SoArcA and DNA fragments imbedded with a highly conserved binding motif (SO_1661). Under oxygen-free conditions, more EcArcB became activated, which in turn phosphorylated more SoArcA. As a result, SoArcA exhibited stronger binding to the SO_1661 promoter, and more importantly, it was able to interact with the dtpA promoter region, validating that SoArcA directly regulates transcription of the dtpA gene in vivo.
During the investigation, we noted a significant difference in expression levels of genes for peptide transporters obtained from qRT-PCR and lacZ-reporters. With the lacZ-reporters, all five operons were found to be down-regulated in the mutant whereas qRT-PCR only differentiated dtpA and sap. We speculate that this may be due to compromised translation efficacy in the absence of the SoArc system. Translation is carried out by ribosome and there is a linear relation between the ribosome mass fraction and the growth rate47. This understanding resonates with our previous findings that the SoarcA mutation concertedly down-regulates the majority of components in protein synthetic machinery13,18. In addition, a remarkable number of translation-associated proteins, such as translation initiation and elongation factors and tRNA synthases, are present in less amounts in the ∆SoarcA strain than the wild-type with respect to transcripts and proteins. On the contrary, the SoArcA loss has no effect on expression of most of the transcription machinery members. We are working to test this notion.
The Arc system of E. coli was initially recognized to confer resistance to dyes such as toluidine blue O and methylene blue, photosensitizers that facilitate generation of reactive oxygen species in the presence of light40,48. However, the underlying mechanism for this phenotype remains largely elusive, implying that it may be unexpectedly complex41,49. In parallel, we previously showed that the cell envelope of S. oneidensis is impaired in the absence of the Arc system. A whole-genome screening with dual-effect transposon constructs, which are able to both inactivate genes by interruption and overexpress genes after the insertion site, fails to identify suppressors for the phenotype19,50. The failure suggests that the observed envelope defect, the same as the growth defect, unlikely due to altered expression of single gene/operon, is a result of multi-fold complex impacts in physiology. We continue our effort to identify the important proteins for both the growth and envelope defects as they are the key to better understand the physiological significance of this atypical Arc system.
Materials and Methods
Bacterial strains, plasmids, and culture conditions
A list of all bacterial strains and plasmids used in this study was given in Table 3. Information for primers used in this study was given in Table S2. Chemicals are from Sigma-Aldrich Co. unless otherwise noted. E. coli and S. oneidensis strains under aerobic conditions were grown in Lysogeny Broth (LB, Difco, Detroit, MI) medium, which was modified to contain tryptone (10 g/L), yeast extract (5 g/L), and NaCl (5 g/L), at 37 °C and 30 °C for genetic manipulation. When needed, the growth medium was supplemented with chemicals at the following concentrations: 2,6-diaminopimelic acid (DAP), 0.3 mM; ampicillin sodium, 50 µg/ml; kanamycin sulfate, 50 µg/ml; and gentamycin sulfate; 15 µg/ml.
For all other purposes, cells of S. oneidensis strains grew in MS mineral medium [KCl, 1.34 mM; NaH2PO4, 5 mM; Na2SO4, 0.7 mM; NaCl, 52 mM; piperazine-N,N = -bis(2-ethanesulfonic acid) (PIPES), 3 mM; NH4Cl, 28 mM; MgSO4, 1 mM; CaCl2, 0.27 mM; FeCl2, 3.6 µM, pH 7.0]51. When supplemented with sodium lactate (30 mM by default), or each of the following (0.5% by default): tryptone, casamino acids, an oligopeptide mixture, a di-tripeptide mixture, an oligo- and di-tri-peptide mixture, and an amino acid mixture (Table 1), media were named as MS-L, MS-T, MS-CA, MS-O-P, MS-DT-P, MS-O/DT-P, and MS-AA, respectively. The amino acid mixture was composed of 500 µg/ml of each alanine, cysteine, glycine, histidine, aspartic acid, glutamic acid, phenylalanine, asparagine, glutamine, methionine, leucine, isoleucine, proline, serine, threonine, lysine, and valine, 50 µg/ml of tryptophan and tyrosine. The MS basal medium was sterilized by in an autoclave, and required supplements sterilized by filtration were added before usage. Fresh medium was inoculated with overnight cultures grown from a single colony by 1:100 dilution, and growth was determined by recording the optical density of cultures at 600 nm (OD600). For anaerobic growth, mid-log phase aerobic cultures were pelletted by centrifugation, purged with nitrogen, suspended in fresh medium containing 20 mM fumarate as the electron acceptor prepared anaerobically to an OD600 of ∼0.02.
Mutant construction and complementation
In-frame deletion strains derived from E. coli MG1655 and S. oneidensis MR-1 were constructed by the Red recombination deletion method and the att-based Fusion PCR method, respectively52,53. In the case of S. oneidensis, two fragments flanking the target gene were amplified by PCR with outside primer containing attB and the gene specific sequence and inside primer containing the linker and the gene specific sequence, which were joined by the second round of PCR with two outside primers. The fusion fragments were introduced into plasmid pHGM01 by using Gateway BP clonase II enzyme mix (Invitrogen) according to the manufacturer’s instruction, resulting in mutagenesis vectors, which were maintained in E. coli DAP auxotroph WM3064. The vectors were subsequently transferred into relevant S. oneidensis strains via conjugation. Integration of the mutagenesis constructs into the chromosome was selected by resistance to gentamycin and confirmed by PCR. Verified transconjugants were grown in LB broth in the absence of NaCl and plated on LB containing 10% sucrose. Gentamycin-sensitive and sucrose-resistant colonies were screened by PCR for deletion of the target gene. Mutants were verified by sequencing the mutated regions.
Growth of S. oneidensis mutation strains generated in this study was measured by recording the optical density at 600 nm (OD600) values in triplicate with the wild-type as the control in MS- based media as mentioned above51. For genetic complementation of the mutants and inducible gene expression, genes of interest generated by PCR were placed under the control of Isopropyl β-D-1-thiogalactoside (IPTG)-inducible promoter Ptac within pHGE-Ptac32. After verification by sequencing, the resultant vectors in E. coli DAP auxotroph WM3064 were transferred into the relevant strains via conjugation.
Peptidase activity assays
Fluorescence peptidase assays were performed using L-Leu-AMC substrate21. Reaction mixtures contained 20 mM Tris-Cl (pH 7.5), 10 μM L-Leu-AMC, and 150 μl of extracts of cells grown in MS-L-T to the mid-log phase. The peptidase activity was monitored at 460 nm with a Synergy 2 Pro200 Multi-Detection (Microplate Reader (Tecan) by detection of 7-amino-4-methylcoumarin (MCA) released from the carboxyl terminus of N-terminally blocked peptides. Negative controls (no enzyme) were run in triplicate to account for thermal degradation of substrates as the background. One unit of protease activity was defined as the amount of enzyme required to release 1 μmol of MCA per min per mg protein. Protein concentration was determined with a bicinchoninic acid assay kit with bovine serum albumin as a standard according to the manufacturer’s instructions (Pierce Chemical).
Droplet assays
Droplet assays were employed to evaluate growth inhibition on plates54. Cells of the mid-log phase (were collected by centrifugation and adjusted to 109 cfu/ml (colony forming unit), which was set as the undiluted (dilution factor 0). Ten-fold series dilutions were prepared with fresh medium indicated. Five microliters of each dilution was dropped onto LB or MS-L-T plates containing required agents as indicated in the figure. The plates were incubated for 24 h or longer before being read. All experiments were conducted at least three times.
Disc diffusion assays
Disc diffusion assays were done similarly to those done previously39. Briefly, overnight cultures were diluted into LB and grown to the mid-log phase. One hundred microliters of cultures was spread onto an agar plate containing required chemicals. Paper discs (diameter, 8 mm) containing 10 μl of 2 mg/ml alafosfolin were placed on top of the agar. The plates were incubated at 30 °C for 16 h prior to analysis. The diameters of the zones of clearing (zones of inhibition, in millimeters) were measured. All assays were done in triplicate using independent cultures, and the resulting zones of inhibition were averaged.
Analysis of gene expression
For qRT-PCR, S. oneidensis cells were grown in MS containing 0.5% tryptone with the required additives to the mid-log phase and collected by centrifugation, and RNA extraction was performed using the RNeasy minikit (Qiagen) as described before55. RNA was quantified by using a NanoVue spectrophotometer (GE Healthcare). The analysis was carried out with an ABI7300 96-well qRT-PCR system (Applied Biosystems). The expression of each gene was determined from three replicates in a single real-time qRT-PCR experiment. The Cycle threshold (CT) values for each gene of interest were averaged and normalized against the CT value of the recA gene, whose abundance is relatively constant during the log phase. Relative abundance (RA) of each gene was presented.
Activity of target promoters was assessed using a single-copy integrative lacZ reporter system as described previously36. Briefly, fragments containing the sequence upstream of the target operon were amplified, cloned into the reporter vector pHGEI01, and verified by sequencing. The resultant vector in E. coli WM3064 was then transferred by conjugation into relevant S. oneidensis strains, in which it integrated into the chromosome and the antibiotic marker was removed subsequently37. Cells grown to the mid-log phase under conditions specified in the text and/or figure legends were collected and β-galactosidase activity was determined by monitoring color development at 420 nm using a Synergy 2 Pro200 Multi-Detection Microplate Reader (Tecan) presented as Miller units.
Electrophoretic motility shift assay (EMSA)
Expression and purification of His-tagged S. oneidensis ArcA has been described before13,19. Phosphorylation of purified SoArcA was performed in buffer containing 100 mM Tris/HCl (pH 7.0), 10 mM MgCl2, 125 mM KCl, 50 mM dilithium carbamoyl phosphate for 60 minutes at room temperature. The probes used for EMSA were prepared by PCR with 33P end-labeled primers13. The binding reaction was performed with ∼25-50 fmol (∼2-5 nM) labeled probes and various amounts of protein in 12 µl binding buffer containing 100 mM Tris/HCl (pH 7.4), 20 mM KCl, 10 mM MgCl2, 2 mM DTT, 0.2 μg/μl poly(dI·dC), and 10% glycerol at 15 °C for 60 minutes and resolved on pre-run 4.8% polyacrylamide native gels. Band shifts were visualized by autoradiography.
Bacterial one-hybrid (B1H) assay
B1H system was used to investigate DNA-protein interaction in vivo in E. coli cells38. Briefly, plasmid constructs were created by cloning the bait ‘DNA’ and target ‘the SoarcA gene’ in the pBXcmT and pTRG vectors and verified by sequencing. The resultant plasmids were used to co-transform BacterioMatch II Validation Reporter Competent Cells on M9 salt agar plates containing 25 mg/ml chloramphenicol and 12.5 mg/ml tetracycline with or without 3-amino-1,2,4-triazole (3-AT). DNA fragments of ∼300 bp for SO_1661 and the 16s rRNA gene promoters (PSO1661 and P16s respectively) were used as positive and negative controls. The plates were incubated for 24 h and then moved to room temperature for an additional 16 h (the colonies indicating positive interaction usually appeared between 18 and 24 h). For the assay under anaerobic conditions, plates were incubated in an anaerobic glovebox (Coy Manufacturing Co.) until the colonies were fully developed. The positive interactions were confirmed by streaking colonies on plates containing both 3-AT and streptomycin (12.5 mg/ml).
Other analyses
Homologues of proteins of interest were identified via a BLASTp search of the NCBI’s nonredundant protein database, using the amino acid sequence as the query. Experimental values were subjected to statistical analyses and presented as means ± standard error of the mean (SEM). Student’s t-test was performed for pairwise comparisons of groups.
References
Green, J. & Paget, M. S. Bacterial redox sensors. Nat Rev Micro. 2, 954–966 (2004).
Dong, Y., Wan, F., Yin, J. & Gao, H. Ecological roles of Arc signal transduction system revealed by evolutionary genetics analysis. J Bacteriol Mycol. 1, 8 (2014).
Lynch, A. S. & Lin, E. C. Transcriptional control mediated by the ArcA two-component response regulator protein of Escherichia coli: characterization of DNA binding at target promoters. J Bacteriol. 178, 6238–6249 (1996).
Georgellis, D., Kwon, O. & Lin, E. C. C. Quinones as the redox signal for the Arc two-component system of bacteria. Science. 292, 2314–2316 (2001).
Iuchi, S. & Lin, E. C. Mutational analysis of signal transduction by ArcB, a membrane sensor protein responsible for anaerobic repression of operons involved in the central aerobic pathways in Escherichia coli. J Bacteriol. 174, 3972–3980 (1992).
Perrenoud, A. & Sauer, U. Impact of global transcriptional regulation by ArcA, ArcB, Cra, Crp, Cya, Fnr, and Mlc on glucose catabolism in Escherichia coli. J Bacteriol. 187, 3171–3179 (2005).
Liu, X. & De, W. P. Probing the ArcA-P modulon of Escherichia coli by whole genome transcriptional analysis and sequence recognition profiling. J Biol Chem. 279, 12588–12597 (2004).
Salmon, K. A. et al. Global gene expression profiling in Escherichia coli k12: effects of oxygen availability and arca. J Biol Chem. 280, 15084–15096 (2005).
Fredrickson, J. K. et al. Towards environmental systems biology of Shewanella. Nat Rev Micro. 6, 592–603 (2008).
Gralnick, J. A., Brown, C. T. & Newman, D. K. Anaerobic regulation by an atypical Arc system in Shewanella oneidensis. Mol Microbiol. 56, 1347–1357 (2005).
Lassak, J., Henche, A.-L., Binnenkade, L. & Thormann, K. M. ArcS, the cognate sensor kinase in an atypical Arc system of Shewanella oneidensis MR-1. Appl Environ Microbiol. 76, 3263–3274 (2010).
Shroff, N. P., Charania, M. A. & Saffarini, D. A. ArcB1, a homolog of Escherichia coli ArcB, regulates dimethyl sulfoxide reduction in Shewanella oneidensis MR-1. J Bacteriol. 192, 3227–3230 (2010).
Gao, H., Wang, X., Yang, Z., Palzkill, T. & Zhou, J. Probing regulon of ArcA in Shewanella oneidensis MR-1 by integrated genomic analyses. BMC Genomics. 9, 42 (2008).
Wang, X. H. et al. A high-throughput percentage-of-binding strategy to measure binding energies in DNA-protein interactions: application to genome-scale site discovery. Nucleic Acids Res. 36, 4863–4871 (2008).
Lassak, J., Bubendorfer, S. & Thormann, K. M. Domain analysis of ArcS, the hybrid sensor kinase of the Shewanella oneidensis MR-1 Arc two-component system, reveals functional differentiation of its two receiver domains. J Bacteriol. 195, 482–492 (2013).
Gao, H. et al. Physiological roles of ArcA, Crp, and EtrA and their interactive control on aerobic and anaerobic respiration in Shewanella oneidensis. PLoS One. 5, e15295 (2010).
Charania, M. A. et al. Involvement of a membrane-bound class III adenylate cyclase in regulation of anaerobic respiration in Shewanella oneidensis MR-1. J Bacteriol. 191, 4298–4306 (2009).
Yuan, J., Wei, B., Lipton, M. & Gao, H. Impact of ArcA loss in Shewanella oneidensis revealed by comparative proteomics under aerobic and anaerobic conditions. Proteomics. 12, 1957–1969 (2012).
Wan, F. et al. Impaired cell envelope resulting from arcA mutation largely accounts for enhanced sensitivity to hydrogen peroxide in Shewanella oneidensis. Sci Rep. 5, 10228 (2015).
Lowther, W. T. & Matthews, B. W. Metalloaminopeptidases: common functional themes in disparate structural surroundings. Chem Rev. 102, 4581–4608 (2002).
Wildeboer, D., Jeganathan, F., Price, R. G. & Abuknesha, R. A. Characterization of bacterial proteases with a panel of fluorescent peptide substrates. Anal Biochem. 384, 321–328 (2009).
Madala, P. K., Tyndall, J. D. A., Nall, T. & Fairlie, D. P. Update 1 of: proteases universally recognize beta strands in their active sites. Chem Rev. 110, PR1-PR31 (2010).
Gomolplitinant, K. M. & Saier, M. H. Evolution of the oligopeptide transporter family. J Membrane Biol. 240, 89–110 (2011).
Saier, M. H. et al. The transporter classification database (TCDB): recent advances. Nucleic Acids Res. 44, D372–D379 (2016).
Heidelberg, J. F. et al. Genome sequence of the dissimilatory metal ion-reducing bacterium Shewanella oneidensis. Nat Biotech. 20, 1118–1123 (2002).
Abouhamad, W. N., Manson, M., Gibson, M. M. & Higgins, C. F. Peptide transport and chemotaxis in Escherichia coli and Salmonella typhimurium: characterization of the dipeptide permease (Dpp) and the dipeptide-binding protein. Mol Microbiol. 5, 1035–1047 (1991).
Pletzer, D. et al. High-throughput screening of dipeptide utilization mediated by the ABC transporter DppBCDF and its substrate-binding proteins DppA1-A5 in Pseudomonas aeruginosa. PLoS One. 9, e111311 (2014).
Weitz, D. et al. Functional and structural characterization of a prokaryotic peptide transporter with features similar to mammalian PEPT1. J Biol Chem. 282, 2832–2839 (2007).
Bippes, C. A. et al. Peptide transporter DtpA has two alternate conformations, one of which is promoted by inhibitor binding. Proc Natl Acad Sci USA 110, E3978–E3986 (2013).
Guettou, F. et al. Selectivity mechanism of a bacterial homolog of the human drug-peptide transporters PepT1 and PepT2. Nat Struct Mol Biol. 21, 728–731 (2014).
Newstead, S. Recent advances in understanding proton coupled peptide transport via the POT family. Curr Opin Struct Biol. 45, 17–24 (2017).
Luo, Q., Dong, Y., Chen, H. & Gao, H. Mislocalization of Rieske protein PetA predominantly accounts for the aerobic growth defect of tat mutants in Shewanella oneidensis. PLoS One. 8, e62064 (2013).
Shi, M., Gao, T., Ju, L., Yao, Y. & Gao, H. Effects of FlrBC on flagellar biosynthesis of Shewanella oneidensis. Mol Microbiol. 93, 1269–1283 (2014).
Jin, M., Zhang, Q., Sun, Y. & Gao, H. NapB in excess inhibits growth of Shewanella oneidensis by dissipating electrons of the quinol pool. Sci Rep. 6, 37456 (2016).
Meng, Q., Liang, H. & Gao, H. Roles of multiple KASIII homologues of Shewanella oneidensis in initiation of fatty acid synthesis and in cerulenin resistance. Biochem Biophys Acta. 1863, 1153–1163 (2018).
Fu, H., Jin, M., Ju, L., Mao, Y. & Gao, H. Evidence for function overlapping of CymA and the cytochrome bc1 complex in the Shewanella oneidensis nitrate and nitrite respiration. Environ Microbiol. 16, 3181–3195 (2014).
Fu, H. et al. Crp-dependent cytochrome bd oxidase confers nitrite resistance to Shewanella oneidensis. Environ Microbiol. 15, 2198–2212 (2013).
Guo, M. et al. Dissecting transcription regulatory pathways through a new bacterial one-hybrid reporter system. Genome Res. 19, 1301–1308 (2009).
Jiang, Y. et al. Protection from oxidative stress relies mainly on derepression of OxyR-dependent KatB and Dps in Shewanella oneidensis. J Bacteriol. 196, 445–458 (2014).
Roeder, W. & Somerville, R. L. Cloning the trpR gene. Mol Gen Genet. 176, 361–368 (1979).
Alvarez, A. F., Malpica, R., Contreras, M., Escamilla, E. & Georgellis, D. Cytochrome d but not cytochrome o rescues the toluidine blue growth sensitivity of arc mutants of Escherichia coli. J Bacteriol. 192, 391–399 (2010).
Capra, E. J. & Laub, M. T. Evolution of two-component signal transduction systems. Annu Rev Microbiol. 66, 325–347 (2012).
Park, D. M., Akhtar, M. S., Ansari, A. Z., Landick, R. & Kiley, P. J. The bacterial response regulator ArcA uses a diverse binding site architecture to regulate carbon oxidation globally. PLoS Genet. 9, e1003839 (2013).
Alexeeva, S., de Kort, B., Sawers, G., Hellingwerf, K. J. & de Mattos, M. J. T. Effects of limited aeration and of the ArcAB system on intermediary pyruvate catabolism in Escherichia coli. J Bacteriol. 182, 4934–4940 (2000).
Rolfe, M. D. et al. Transcript profiling and inference of Escherichia coli K-12 ArcA activity across the range of physiologically relevant oxygen concentrations. J Biol Chem. 286, 10147–10154 (2011).
Malpica, R., Franco, B., Rodriguez, C., Kwon, O. & Georgellis, D. Identification of a quinone-sensitive redox switch in the ArcB sensor kinase. Proc Natl Acad Sci USA 101, 13318–13323 (2004).
Scott, M., Gunderson, C. W., Mateescu, E. M., Zhang, Z. & Hwa, T. Interdependence of cell growth and gene expression: origins and consequences. Science. 330, 1099–1102 (2010).
Iuchi, S. & Lin, E. C. arcA (dye), a global regulatory gene in Escherichia coli mediating repression of enzymes in aerobic pathways. Proc Natl Acad Sci USA 85, 1888–1892 (1988).
Ruiz, J. A., Fernández, R. O., Nikel, P. I., Méndez, B. S. & Pettinari, M. J. dye (arc) mutants: insights into an unexplained phenotype and its suppression by the synthesis of poly (3-hydroxybutyrate) in Escherichia coli recombinants. FEMS Microbiol Lett. 258, 55–60 (2006).
Yin, J. et al. Regulation of nitrite resistance of the cytochrome cbb3 oxidase by cytochrome c ScyA in Shewanella oneidensis. MicrobiologyOpen. 4, 84–99 (2014).
Shi, M., Wan, F., Mao, Y. & Gao, H. Unraveling the mechanism for the viability deficiency of Shewanella oneidensis oxyR null mutant. J Bacteriol. 197, 2179–2189 (2015).
Jin, M. et al. Unique organizational and functional features of the cytochrome c maturation system in Shewanella oneidensis. PLoS One. 8, e75610 (2013).
Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc Natl Acad Sci USA 97, 6640–6645 (2000).
Yin, J., Sun, Y., Mao, Y., Jin, M. & Gao, H. PBP1a/LpoA but Not PBP1b/LpoB Are Involved in Regulation of the Major β-Lactamase Gene blaA in Shewanella oneidensis. Antimicrob Agents Chemother. 59, 3357–3364 (2015).
Yuan, J., Wei, B., Shi, M. & Gao, H. Functional assessment of EnvZ/OmpR two-component system in Shewanella oneidensis. PLoS One. 6, e23701 (2011).
Acknowledgements
This research was supported by Ten Thousand Talent Program of China, by Natural Science Foundation of Zhejiang Province (LZ17C010001), and by the Fundamental Research Funds for the Central Universities.
Author information
Authors and Affiliations
Contributions
H.G. conceived the idea and designed the project. H.L., Y.M. and Y.S. carried out the experiments. H.L., Y.M. and H.G. analyzed data. H.L., Y.M. and H.G. wrote the paper.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liang, H., Mao, Y., Sun, Y. et al. Transcriptional regulator ArcA mediates expression of oligopeptide transport systems both directly and indirectly in Shewanella oneidensis. Sci Rep 9, 13839 (2019). https://doi.org/10.1038/s41598-019-50201-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-50201-4
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.