Abstract
Although ion channels are attractive targets for drug discovery, the systematic screening of ion channel-targeted drugs remains challenging. To facilitate automated single ion-channel recordings for the analysis of drug interactions with the intra- and extracellular domain, we have developed a parallel recording methodology using artificial cell membranes. The use of stable lipid bilayer formation in droplet chamber arrays facilitated automated, parallel, single-channel recording from reconstituted native and mutated ion channels. Using this system, several types of ion channels, including mutated forms, were characterised by determining the protein orientation. In addition, we provide evidence that both intra- and extracellular amyloid-beta fragments directly inhibit the channel open probability of the hBK channel. This automated methodology provides a high-throughput drug screening system for the targeting of ion channels and a data-intensive analysis technique for studying ion channel gating mechanisms.
Similar content being viewed by others
Introduction
Membrane proteins, such as transmembrane receptors and ion channels, are major molecular targets for commercial drug therapies1,2. Compared to the successes achieved in the efficacies of drugs targeting transmembrane receptors, such as G protein-coupled receptors, only 30% of the drug-targeted ion channels predicted by in silico gene expression analyses are being utilised in current therapies3. When searching for drugs that target ion channels, it is important to identify the site of drug action and to determine whether the site is intra- or extracellular. This type of topological information facilitates the determination of the duration of drug administration, thereby reducing side effects4,5. Moreover, single-channel recording of point-mutated ion channels is essential to determine how each drug affects channel conformation. However, topological (intra- and extracellular) single ion-channel recording is labour intensive and requires highly skilled researchers; even then, it is difficult to acquire an extensive data set that can be used to reveal biologically relevant molecular mechanisms. Therefore, an automated parallel measurement technique for single-channel recording is required for efficient drug discovery or ion-channel analysis.
In general, two electrophysiological methodologies can be used to examine the effect of a ligand on the gating mechanism of an ion channel: first, the conventional inside-out and outside-out patch-clamp method using living cells6,7 and second, the artificial bilayer lipid membrane (BLM) method8,9,10,11,12,13. Single-channel recordings have not been achieved for either method using an automated system. For the patch-clamp method, an auto-patch system has been commercialised but does not provide inside-out/outside-out recordings to identify the intra- and extracellular effects of drugs at the single-molecule level14. In contrast, the BLM method of ion channel analysis provides a platform for achieving the equilibrium recognition of the intra- or extracellular topology using purified materials at the single-channel level15. Although several attempts to create the parallel formation of BLMs have been reported16,17,18,19,20,21, single-channel recording for identifying membrane topology has not been achieved in the parallel system due to the low stability and reproducibility of BLMs. Therefore, these conventional methods present challenges in the preparation of a reproducible bilayer and its stability for use in automated, parallel recording22.
Here, we report an automated, parallel methodology for single ion-channel recording based on Droplet Contact Method (DCM)23. In this method, a BLM is formed at the interface between contacting droplets submerged in an oil/lipid mixture (Fig. 1a, b), which is recently referred to as Droplet Interface Bilayer (DIB)24. To enhance the stability of the BLM, we reduced the area of the BLM by placing a polymer sheet with microsized pores in between the droplets (Fig. 1c). This improvement allows us to form highly stable and reproducible BLMs. Our system has advantages over the previous system in simultaneously satisfying the abilities as following: i) intensive data acquisition, ii) single-channel analysis and iii) analysis of topologically identified targets. Using this method, we performed single-channel recordings on several types of ion channels with automation for intensive data acquisition and we identified whether these channels had intra- or extracellular sites of drug action. We also demonstrated that amyloid-beta fragments (Aβ) directly inhibits the opening of a Ca2+-dependent K+ (hBK) channel at both the intra- and extracellular domains.
Automated, parallel ion channel-recording system using the droplet contact method.
(a) Formation of a bilayer lipid membrane by the droplet-contact method. Two lipid monolayers are brought into contact with one another and spontaneously form a bilayer. (b) The PDW chip for the simple preparation of lipid bilayers. (c) Schematic illustration of the PDW chip. (d) Schematic diagram of the PDW chip connected to a patch-clamp amplifier. (e) An arrayed 16-ch PDW chip connected to a multi-patch-clamp amplifier. (f) Automated, parallel bilayer single-channel recording system. The injection robot automatically prepares the bilayer lipid membranes, reconstitutes the ion channels and controls the drug application.
Results
Automated lipid bilayer preparation using an injection robot
A double-well chip (Fig. 1b–d) was used for the ion channel recordings in conjunction with the droplet contact method as previously reported23. In this approach, lipid/oil and buffer solutions are sequentially deposited onto a double-well chip to prepare the BLM; this method has been broadly used as a simple means of BLM formation24,25. Here, the stability of the lipid membrane on the chip was improved for use in the automated system (as discussed below). To decrease the membrane capacitance and to increase its mechanical stability, the contact area of the lipid monolayer was reduced using a poly(chloro-p-xylylene) (parylene) separator that was patterned with micropores26,27. The parylene film was integrated into the double-well chip to produce a parylene double-well (PDW) chip (Fig. 1c). Therefore, the BLMs that formed in the PDW chip were mechanically stable as described in the following section. Due to the high stability of the BLMs, we were able to construct parallel BLMs using an injection robot; the resulting arrayed BLM device, which consists of a 16-channel (16-ch) PDW chip, is shown in Figure 1e and f and Supplementary Movie 1.
Stability and noise examination of BLMs prepared by the droplet contact method
We first estimated the mechanical stability of the BLMs. The mechanical stability of a BLM prepared by the traditional method cannot be tested because it is too fragile28,29. However, due to the robustness of the BLMs formed with the DCM, mechanical stability was evaluated using a mechanical oscillator. The survival rate of the BLMs in the double-well (DW) chip lacking the parylene separator and the PDW chip as a function of the vibration frequency is depicted in Figure 2a. The survival rate of the DW chip was less than 40% at a vibration of 30 Hz. In contrast, the BLMs in the PDW chip maintained their form within this frequency range. The resonant frequency of the BLMs in the PDW chip was calculated to be 1.3 kHz (Supplementary Text 6). In the range of ambient vibration frequencies encountered in the daily environment (<100 Hz)30, the BLMs in the PDW chip were mechanically stable.
(a) Mechanical stability in the DW and PDW chips as determined with a mechanical vibrator. The survival rate was defined as the number of stable BLMs/16-ch for 30 s (n = 4, mean ± S.D.). (b) Comparison of the power density spectra of αHL open-channel currents between the DW and PDW chips. The PDW chip showed significantly reduced noise in the lipid bilayer. (c) The long-term stability of BLMs in the PDW chip. Channel opening events are presented as a function of days. The channel signals of alamethicin were observed for 14 days.
Electrical current noise is an important aspect of artificial cell membrane experiments31. Figure 2b shows the power spectra of the mean current noise of the α-haemolysin (αHL) open channel signal in the DW chip and the PDW chip. Due to the small membrane capacitance of the PDW chip, the current noise at lower frequencies (<100 Hz) was two orders of magnitude lower for the PDW chip than for the DW chip. The overall membrane capacitance was reduced with the reduction of the bilayer area, which resulted in the reduction in the current noise.
The long-term stability of the BLMs on the PDW chip was also measured by obtaining alamethicin channel signals (Fig. 2c). Since this stability is required for long-term experiments such as cell-free expression experiment, the evaluation of the long-term stability is important. The signal event frequency was approximately 10 Hz in this experiment (Supplementary Text 6). Although the event frequency was slightly reduced compared with the initial state, the channel signals were observed for 14 days; compared to previous reports, this period is the longest reported for BLM stability32,33.
Introducing ion channels into the planar lipid bilayer using a vesicle fusion technique
Vesicle fusion is an efficient methodology for channel reconstitution in planar lipid bilayers34. The single-channel current of the purified and reconstituted ion channel on the surface of a proteoliposome can be observed using this method. The process of vesicle fusion is described in Supplementary Text 5. The fusion rate of the vesicle containing the hBK channel was defined as So (So = signal-obtaining chamber/16-ch) and it was used to optimise vesicle fusion in the PDW array (n = 16, N = 3). We found that So increased with increasing hydrostatic pressure (Supplementary Text 7) at the interface between the droplets (Fig. 3a). This result can be explained because the area of the bilayer formed in the droplet interface is increased by the high pressure, thereby increasing the probability of vesicle fusion. Meanwhile, in the higher pressure region, the BLMs were ruptured within 120 min. The optimal pressure was determined to be 14 N m−1. So was stable at 0.6 under these conditions after 120 min.
The optimisation of vesicle fusion.
(a) Comparison of the hBK channel signal rate (signals/16-ch) at the first signal observation and after 120 min with various hydrostatic pressures at the droplet interface (N = 3, n = 16, mean ± S.D.). (b) The signal rate was increased with increasing concentrations of proteoliposomes. The single-channel signals overlapped and could not be discriminated at levels higher than the optimal concentration (N = 3, n = 16, mean ± S.D.).
The other important factor for vesicle fusion is the concentration of the vesicle. Figure 3b shows the concentration dependence of So. The curve relative to the concentration was convex upward. At levels under 10−5 mg/mL lipid, the probability of fusion between the vesicle and the planar bilayer should be enhanced by increasing the vesicle concentration. In contrast, So was decreased at levels higher than the apical concentration. The current signals overlapped due to an excess of vesicles that fused after the initial vesicle fusion. Therefore, So was decreased in the high-concentration range. The highest So value (more than 0.9) was obtained at a lipid concentration of 10−5 mg/mL. In addition, the addition of cholesterol to the BLM decreased So (Supplementary Text 7).
Previously, few studies of vesicle fusion probability have been made on the basis of quantitative data due to the requirement of labour-intensive experiments. Here, the vesicle fusion conditions were optimised based on extensive data that were obtained using the automated BLM method. The optimised condition of the channel reconstitution was utilised in the following experiments.
Verification of the automated system for measuring several ion channels and a mutant channel
To test the device, toxin channels (αHL and alamethicin), K+ channels (hBK and Wild-Type (WT)KcsA) and a mutant channel (KcsA mutant (E71A)) were reconstituted in the 16-ch PDW chip using the automated system (Supplementary Text 5). E71 is an amino acid residue of the pore helix region of KcsA and its corresponding mutant (denoted E71A here) has been used in studies on channel-gating mechanisms35. The currents that passed through each ion channel were observed in parallel at the single-channel level under various recording conditions. Figures 4a–e show the current signals and histograms of five different proteins at the single-channel level. The I–V curves from the hBK, WT-KcsA and E71A channels were measured from a single run of the automated system (Figs. 4f and g). Compared to the WT-KcsA channel, E71A displayed an increased channel open probability (Po) (Figs. 4d and e) and a decreased outward rectification due to a reduced energy barrier for ion entry into the selective filter at negative holding potentials15. These differences between the WT and the mutant are consistent with previous reports36. The application of this methodology to gene-targeted ion channels will provide essential information regarding drug-targeted amino acid residues that contribute to the molecular basis of drug action at specific sites on ion channels.
Examples of single-channel current recordings using the automated system.
(a–e) Current traces (left) and corresponding point histograms (right) from (a) α-hemolysin (holding potential (Vhold) = +50 mV), (b) alamethicin (Vhold = +150 mV), (c) hBK (Vhold = +60 mV), (d) WT KcsA and (e) E71A (Vhold = +100 mV). The arrows indicate the channel closed state. (f, g) I–V curves of the hBK channel (f) and KcsA channels (WT and E71A, g). The data are shown as the mean ± S.D. (n > 3 each).
Massive data acquisition for the hBK channel using automated and parallel single-channel recordings
Parallel recordings of the hBK channel were performed using the 16-ch PDW chip. The hBK channel controls firing patterns in neurons and modulates the tone of blood vessels37. The current recordings from the PDW chip are shown in Figure 5a. Rapid open-and-close transitions associated with channel gating were observed at the single-channel level from all of the channels in the PDW chip. Because this parallel measurement easily facilitated a high degree of reproducibility, we obtained numerous single-channel recordings (n = 102) and used the large amount of data to analyse the channel conductance and dwell time (Figs. 5b and c). The large amount of data obtained ensured reliability in the measurement of these biophysical phenomena. Furthermore, a particular channel orientation could be observed by controlling the solution contents. For the hBK channels, the current signals were observed from oriented channels, in which the Ca2+-binding site was directed toward a Ca2+ solution that was added to one of the compartments of the PDW chip. The characteristics of the oriented hBK channels were investigated and the results are shown in Supplementary Figure S4. The fundamental properties of the hBK channels, which were measured with the automated parallel system, were similar to previously reported results obtained with live cells and BLMs38,39.
The performance of the 16-ch recording system.
(a) Parallel recordings of the hBK channel obtained using the automated system. Single-channel currents were observed from all 16 channels of the arrayed PDW chip. The arrows indicate the channel closed state. (b, c) Histograms of the conductance and dwell time of hBK channels obtained using numerous single-channel recordings (n = 102, each recording period > 20 s). The holding potential was +60 mV.
Topologically identified hBK channel inhibition using the automated system
The greatest advantage of the automated system is that it can be used to examine the direct inhibition of purified channels from both membrane topological directions (intracellular and extracellular) while excluding interactions with cytosolic components. Here, we demonstrate the ability to analyse direct interactions between the ion channel and ligands that are applied from either the intra- or extracellular side. Iberiotoxin (IbTX) is a toxin for K+ channels and inhibits the hBK channel from only the extracellular side by decreasing Po40. This extracellular inhibition was observed at the single-channel level in our system (Supplementary Fig. S6c, d, Supplementary Text 9). The dose-dependent inhibition of IbTX on Po was evaluated to determine the IC50 and the Hill coefficient, which were 5.6 nM and 2.1, respectively (Fig. 6a). In addition, the inhibition of hBK by the L-type calcium channel blocker verapamil was also examined. The dissociation constant (kd) for verapamil was determined to be 6.3 μM (Fig. 6b, Supplementary Text 9). These measurements were consistent with previous reports on the inhibitory effects of IbTX and verapamil on reconstituted hBK channels40,41. This type of pharmacological analysis using our automated system has great potential for aiding in the determination of drug activity.
Amyloid-beta fragments directly inhibited a single hBK channel
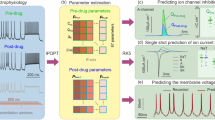
We investigated the topological effects of an amyloid-beta fragment (Aβ1-42) on hBK at the single-channel level. Figures 7a and c show the current recordings obtained from the hBK channel before and after the injection of Aβ1-42 on the intracellular side. The Po value decreased immediately after the injection. A similar reduction was also observed following an extracellular application (Fig. 7d). The effect of Aβ1-42 on the lipid bilayer itself was also examined because amyloid proteins can form a channel structure in negatively charged membranes (DOPE)42. However, Aβ1-42 did not affect the neutral membrane used in this study (EggPC) as shown in Figure 7b. This experiment demonstrates another advantage of this artificial system: the lipid content can be controlled to directly observe channel activities without disturbing the membrane. Figures 7e and f show the current histograms that were obtained before and after the application of Aβ1-42 to a single hBK channel from the intracellular (Fig. 7e) and extracellular (Fig. 7f) sides. In both cases, the open channel states were inhibited by Aβ1-42. The effects of Aβ1-42 on channel conductance are shown in Figure 7g; the same buffer solution without Aβ1-42 was used as a control (Aβ (-)). The conductance of the hBK channel did not change upon Aβ1-42 application. In contrast, Po decreased upon Aβ1-42 application from both directions (Fig. 7h).
Automated single hBK channel inhibition by Aβ1-42 from both the intra- and extracellular sides.
(a) Effect of Aβ1-42 on single hBK channels before and after the addition of Aβ1-42 to the extracellular side. (b) Effect of Aβ1-42 application on the lipid bilayer (EggPC) itself. (c, d) The intra- (c) and extracellular side (d) effects of the application of Aβ1-42 to the hBK channel. (e, f) Point current histograms for the hBK channel obtained before (top) and after (bottom) the application of Aβ1-42 from the intracellular (e) or extracellular side (f) direction. (g, h) Effects of Aβ1-42 on the channel conductance (g) and open probability (h) (n = 12, mean ± S.D.). The significance was assessed by t-tests: *P < 0.05, **P < 0.01.
Discussion
Our automated methodology represents a powerful means to acquire a massive amount of data regarding topologically identified single ion channels. Using this method, we investigated hBK channel inhibition with respect to membrane topology (intracellular or extracellular) by using the amyloid-beta fragment (Aβ1-42). Abnormal BK channel activity has been recently reported in the pyramidal cells of Alzheimer's disease model mice43. However, the effect of Aβ on the hBK channel remains unclear for two reasons. First, it is unknown whether Aβ affects the BK channel directly or via another in vivo inhibitory cascade. Second, it is unknown whether Aβ affects the channel from the intra- or extracellular direction. Although extracellular deposits of Aβ in the cortical neuropil and blood vessel walls in the brain are classical histopathological features of Alzheimer's disease, emerging evidence from transgenic mice and human patients has revealed that Aβ also accumulates intracellularly in neurons44. Therefore, we used our system to measure the inhibitory response of Aβ1-42 applied to topologically identified hBK channels. It was revealed that Aβ1-42 directly inhibits the hBK channel from both the intra- and extracellular sides. These results suggest that in neurons, Aβ1-42 directly inhibits the hBK channel at both the intra- and extracellular domains. A similar inactivation by Aβ1-42 may also occur with other ion channels, such as extracellular Aβ-sensitive ion channels45,46; this process may represent one cause of neuronal dysfunction in the brain of Alzheimer's disease patients.
Taken together, our automation system readily revealed the topological and direct effects of various compounds, including pharmacological agents and Aβ1-42, on ion channels at the single-molecule level. In the DCM system, the inner solution of the droplet can also be exchanged to perform drug application assays47. Moreover, automated ion-channel screening using the droplet contact method may also be used to analyse mutated ion channels, in which the determination of the site of drug action is necessary for personal gene therapy. Further comprehensive studies with this automated ion channel system will contribute to the systematic discovery of drugs that target ion channels.
Methods
Automated lipid bilayer preparation by the droplet contact method and ion channel introduction into BLM
The following method was used to form the bilayer lipid membrane (BLM) by the droplet contact method: 1) An EggPC/n-decane (20 mg/mL) solution (6 to 9 uL) was initially deposited into each well. 2) The aqueous buffer and the same buffer containing the water-soluble membrane proteins or proteoliposomes (15 to 20 uL) were then deposited into each well (Supplementary Fig. S2a). The lipid monolayers contact each other and form a BLM within 10 minutes of the addition of the buffer solution. The BLMs can be formed in the array device using an automated injection robot (epMotion 5070, Eppendorf). A movie of the automated preparation of BLMs is provided (Movie S1).
We defined the two droplets that sat on the recording and the ground electrodes as droplets A and B, respectively. A symmetrical buffer solution (in mM: 1000 KCl, 2 KH2PO4, 8 K2HPO4, 1 EDTA, pH 7.4) was used in both droplets A and B for the water-soluble toxin channels (αHL and alamethicin). αHL and alamethicin were dissolved in droplet A at 600 and 10 nM, respectively. In contrast, the reconstitution of the hBK channels and KcsAs was more difficult because the channels are water-insoluble and retain their conformation only within a lipid bilayer membrane. Hence, the reconstitution of these channels was conducted using vesicle fusion. The process of vesicle fusion involves the following steps: i) A vesicle (the ion channel-expressing proteoliposome) approaches a BLM (Supplementary Fig. S4a). ii) The vesicle and the lipid bilayer fuse with each other when the vesicle attaches to the bilayer surface. To enhance the fusion event, the osmotic pressure between the inside and outside of the vesicle was controlled in this experiment (Supplementary Fig. S4c). iii) The channel signals are measured through Ag/AgCl electrodes after the reconstitution. The experimental conditions used for the simultaneous and parallel recordings are described in Supplementary Text 5. In the binding experiments, a small volume of solution containing the inhibitors can be injected into the droplet after determining the single-channel currents. This type of kinetic experiment was impossible using previous microdrop methods.
Ion channel current recordings and data analysis
The channel current signals were detected using a multi-patch-clamp amplifier (JET-Bilayer, Tecella) with a 1 kHz low-pass filter at a sampling frequency of 5 kHz (unless otherwise noted) at 23 ± 1°C. Most of the experiments on the ion channels, including the pharmacological tests, were performed using a K-recording buffer (in mM: 140 KCl, 0.1 CaCl2, 10 HEPES, pH 7.0) or an Na-recording buffer (in mM: 120 NaCl, 0.1 CaCl2, 10 HEPES, pH 7.0). The current analysis was performed using the pCLAMP ver. 10.6 (Molecular Devices, Sunnyvale) and Igor Pro 6.2 (Wavemetrics, Oregon) software programs. The single-channel analysis of the ion channels is described below. The channel recordings were conducted for 2 h after injecting the proteoliposome. If the BLM was ruptured, it could be reformed by repairing the BLM using a hydrophobic needle. The channel signals typically began approximately 1 to 10 min after injecting the droplets. At least 20 s of continuous recording was used to estimate the channel open probability (P0). The durations of open and closed times were measured with a half-amplitude threshold analysis using the pCLAMP software program. Occasional long closed periods (shut intervals > 5 s) were excluded because of extremely low activity. To promote a decreased signal/noise ratio, an off-line digitised low-pass filter (<500 Hz) and the baseline offset were used. The results are given as the mean ± S.D.
In case of the topology identification of the channels, the channel which is oriented undesired direction can be discriminated with changing the voltage polarization because of the rectifying. For instance, using voltage-dependent KcsA channel (as shown in Fig. 4), the oppositely oriented channel shows the inappropriate conductance under applying the opposite voltage. As the result, we can identify the channel orientation by the channel conductance. On the other hand, if the channel does not have the rectifying, the topology can be recognized by observing the direction of the blocker or the opener application. In addition, the specific orientation can be maintained owing to the conical shape of the transmembrane domain, which matches the high curvature of small vesicle (<500 nm dia.)48. The orientation is discriminated from the direction of the vesicle fusion.
References
Drews, J. Drug discovery: A historical perspective. Science 287, 1960–1964 (2000).
Overington, J. P., Al-Lazikani, B. & Hopkins, A. L. Opinion - How many drug targets are there? Nature Reviews Drug Discovery 5, 993–996 (2006).
Terstappen, G. C. & Reggiani, A. In silico research in drug discovery. Trends in Pharmacological Sciences 22, 23–26 (2001).
Macpherson, L. J. et al. Noxious compounds activate TRPA1 ion channels through covalent modification of cysteines. Nature 445, 541–545 (2007).
Rajendran, L., Knoelker, H.-J. & Simons, K. Subcellular targeting strategies for drug design and delivery. Nature Reviews Drug Discovery 9, 29–42 (2010).
Hamill, O. P., Marty, A., Neher, E., Sakmann, B. & Sigworth, F. J. Improved patch-clamp techniques for high-resolution current recording from cells and cell-free membrane patches. Pflugers Archiv-European Journal of Physiology 391, 85–100 (1981).
Neher, E. Correction for liquid junction potentials in patch clamp experiments. Methods Enzymol. 207, 123–131 (1992).
Latorre, R. & Alvarez, O. Voltage-dependent channels in planar lipid bilayer membranes. Physiological Reviews 61, 77–150 (1981).
Montal, M. & Mueller, P. Formation of bimolecular membranes from lipid monolayers and a study of their electrical properties. Proc. Natl. Acad. Sci. U.S.A. 69, 3561–3566 (1972).
Ide, T. & Yanagida, T. An artificial lipid bilayer formed on an agarose-coated glass for simultaneous electrical and optical measurement of single ion channels. Biochem. Biophys. Res. Commun. 265, 595–599 (1999).
Ervin, E., Kawano, R., White, R. & White, H. Simultaneous alternating and direct current readout of protein ion channel blocking events using glass nanopore membranes. Anal. Chem. 80, 2069–2076 (2008).
Bayley, H. & Martin, C. R. Resistive-pulse sensing - From microbes to molecules. Chem. Rev. 100, 2575–2594 (2000).
Heron, A. J., Thompson, J. R., Mason, A. E. & Wallace, M. I. Direct detection of membrane channels from gels using water-in-oil droplet bilayers. J. Am. Chem. Soc. 129, 16042–16047 (2007).
Dunlop, J., Bowlby, M., Peri, R., Vasilyev, D. & Arias, R. High-throughput electrophysiology: an emerging paradigm for ion-channel screening and physiology. Nature Reviews Drug Discovery 7, 358–368 (2008).
Rotem, D., Mason, A. & Bayley, H. Inactivation of the KcsA potassium channel explored with heterotetramers. J. Gen. Physiol. 135, 29–42 (2010).
Syeda, R., Holden, M. A., Hwang, W. L. & Bayley, H. Screening Blockers Against a Potassium Channel with a Droplet Interface Bilayer Array. J. Am. Chem. Soc. 130, 15543–15548 (2008).
El-Arabi, A. M., Salazar, C. S. & Schmidt, J. J. Ion channel drug potency assay with an artificial bilayer chip. Lab Chip 12, 2409–2413 (2012).
Baaken, G., Ankri, N., Schuler, A. K., Ruhe, J. & Behrends, J. C. Nanopore-Based Single-Molecule Mass Spectrometry on a Lipid Membrane Microarray. Acs Nano 5, 8080–8088 (2011).
Zagnoni, M., Sandison, M. E. & Morgan, H. Microfluidic array platform for simultaneous lipid bilayer membrane formation. Biosensors & Bioelectronics 24, 1235–1240 (2009).
Andersson, M. et al. Voltage-induced gating of the mechanosensitive MscL ion channel reconstituted in a tethered lipid bilayer membrane. Biosensors & Bioelectronics 23, 919–923 (2008).
Suzuki, H. & Takeuchi, S. Microtechnologies for membrane protein studies. Anal. Bioanal. Chem. 391, 2695–2702 (2008).
Farre, C. & Fertig, N. HTS techniques for patch clamp-based ion channel screening - advances and economy. Expert Opinion on Drug Discovery 7, 515–524 (2012).
Funakoshi, K., Suzuki, H. & Takeuchi, S. Lipid bilayer formation by contacting monolayers in a microfluidic device for membrane protein analysis. Anal. Chem. 78, 8169–8174 (2006).
Bayley, H. et al. Droplet interface bilayers. Mol. Biosyst. 4, 1191–1208 (2008).
Thapliyal, T., Poulos, J. L. & Schmidt, J. J. Automated lipid bilayer and ion channel measurement platform. Biosensors & Bioelectronics 26, 2651–2654 (2011).
Kawano, R., Osaki, T., Sasaki, H. & Takeuchi, S. A Polymer-Based Nanopore-Integrated Microfluidic Device for Generating Stable Bilayer Lipid Membranes. Small 6, 2100–2104 (2010).
Kawano, R. et al. Rapid Detection of a Cocaine-Binding Aptamer Using Biological Nanopores on a Chip. J. Am. Chem. Soc. 133, 8474–8477 (2011).
Needham, D., McIntosh, T. J. & Lasic, D. D. Repulsive interactions and mechanical stability of polymer-grafted lipid-membranes. Biochim. Biophys. Acta 1108, 40–48 (1992).
Mayer, M., Kriebel, J. K., Tosteson, M. T. & Whitesides, G. M. Microfabricated teflon membranes for low-noise recordings of ion channels in planar lipid bilayers. Biophys. J. 85, 2684–2695 (2003).
Steinberg, D. S. Vibration analysis for electronic equipment, Edn. 3rd. (John Wiley & Sons, New York; 2000).
Wonderlin, W. F., Finkel, A. & French, R. J. Optimizing planar lipid bilayer single-channel recordings for high-resolution with rapid voltage steps. Biophys. J. 58, 289–297 (1990).
Le Pioufle, B., Suzuki, H., Tabata, K. V., Noji, H. & Takeuchi, S. Lipid bilayer microarray for parallel recording of transmembrane ion currents. Anal. Chem. 80, 328–332 (2008).
Hirano-Iwata, A. et al. Free-Standing Lipid Bilayers in Silicon Chips-Membrane Stabilization Based on Microfabricated Apertures with a Nanometer-Scale Smoothness. Langmuir 26, 1949–1952 (2010).
Labarca, P. & Latorre, R. Insertion of ion channels into planar lipid bilayers by vesicle fusion. Methods Enzymol. 207, 447–463 (1992).
Cordero-Morales, J. F. et al. Molecular determinants of gating at the potassium-channel selectivity filter. Nat. Struct. Mol. Biol. 13, 311–318 (2006).
Hirano, M., Onishi, Y., Yanagida, T. & Ide, T. Role of the KcsA Channel Cytoplasmic Domain in pH-Dependent Gating. Biophys. J. 101, 2157–2162 (2011).
Butler, A., Tsunoda, S., McCobb, D. P., Wei, A. & Salkoff, L. Mslo, a complex mouse gene encoding maxi calcium-activated potassium channels. Science 261, 221–224 (1993).
Sun, X. P., Schlichter, L. C. & Stanley, E. F. Single-channel properties of BK-type calcium-activated potassium channels at a cholinergic presynaptic nerve terminal. Journal of Physiology-London 518, 639–651 (1999).
Yuan, C., O'Connell, R. J., Jacob, R. F., Mason, R. P. & Treistman, S. N. Requiation of the gating of BKCa channel by lipid bilayer thickness. J. Biol. Chem. 282, 7276–7286 (2007).
Giangiacomo, K. M., Garcia, M. L. & McManus, O. B. Mechanism of iberiotoxin block of the large-conductance calcium-activated potassium channel from bovine aortic smooth-muscle. Biochemistry 31, 6719–6727 (1992).
Harper, A. A., Catacuzzeno, L., Trequattrini, C., Petris, A. & Franciolini, F. Verapamil block of large-conductance Ca-activated K channels in rat aortic myocytes. J. Membr. Biol. 179, 103–111 (2001).
Arispe, N., Rojas, E. & Pollard, H. B. Alzheimer disease amyloid beta protein forms calcium channels in bilayer membranes: blockade by tromethamine and aluminum. Proc. Natl. Acad. Sci. U.S.A. 90, 567–571 (1993).
Yamamoto, K. et al. Suppression of a Neocortical Potassium Channel Activity by Intracellular Amyloid-beta and Its Rescue with Homer1a. J. Neurosci. 31, 11100–11109 (2011).
LaFerla, F. M., Green, K. N. & Oddo, S. Intracellular amyloid-beta in Alzheimer's disease. Nat. Rev. Neurosci. 8, 499–509 (2007).
Good, T. A., Smith, D. O. & Murphy, R. M. beta-amyloid peptide blocks the fast-inactivating K+ current in rat hippocampal neurons. Biophys. J. 70, 296–304 (1996).
Qi, J. S., Ye, L. & Qiao, J. T. Amyloid beta-protein fragment 31-35 suppresses delayed rectifying potassium channels in membrane patches excised from hippocampal neurons in rats. Synapse 51, 165–172 (2004).
Tsuji, Y. et al. Droplet-based lipid bilayer system integrated with microfluidic channels for solution exchange. Lab Chip 13, 1476–1481 (2013).
Yanagisawa, M., Iwamoto, M., Kato, A., Yoshikawa, K. & Oiki, S. Oriented Reconstitution of a Membrane Protein in a Giant Unilamellar Vesicle: Experimental Verification with the Potassium Channel KcsA. J. Am. Chem. Soc. 133, 11774–11779 (2011).
Acknowledgements
We thank Yoshimi Komaki and Yoshimi Nozaki for technical assistance with the experiments and Amy Hsiao for useful comments on the manuscript. This work was partially supported by a Grant-in-Aid for Scientific Research on Innovative Areas (No. 23107534), for Challenging Exploratory Research (No. 24655072), for Young Scientists (A) (No. 25708024) and for Scientific Research (A) (No. 25246071) from MEXT in Japan.
Author information
Authors and Affiliations
Contributions
R. K. and S. T. conceived the study design. R. K., Y. T. and M. H. performed the experiments and analysed the data. T. O. and K. K. contributed to the device concept and fabrication. T. I. and M. N. provided considerable advice on the initial direction of the research. R. K., K. S. and S. T. wrote the paper. All of the authors discussed the results and commented on the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Materials
Supplementary Information
Supplementary Video
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Kawano, R., Tsuji, Y., Sato, K. et al. Automated Parallel Recordings of Topologically Identified Single Ion Channels. Sci Rep 3, 1995 (2013). https://doi.org/10.1038/srep01995
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep01995
This article is cited by
-
De novo design of a nanopore for single-molecule detection that incorporates a β-hairpin peptide
Nature Nanotechnology (2022)
-
3D Artificial Cell Membranes as Versatile Platforms for Biological Applications
BioChip Journal (2022)
-
Development of an automated system to measure ion channel currents using a surface-modified gold probe
Scientific Reports (2021)
-
Photolithographic Fabrication of Micro Apertures in Dry Film Polymer Sheets for Channel Recordings in Planar Lipid Bilayers
The Journal of Membrane Biology (2019)
-
Electrophysiological measurement of ion channels on plasma/organelle membranes using an on-chip lipid bilayer system
Scientific Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.