Abstract
Although two classes of antivirals, NA inhibitors and M2 ion channel blockers, are licensed for influenza treatment, dual resistant mutants, including highly pathogenic H5N1 viruses, have appeared. Alternative treatment options are, therefore, needed. Influenza A viral RNA (vRNA) transcription/replication is a promising target for antiviral development, since it is essential for virus replication. Accordingly, an efficient and reliable method to identify vRNA transcription/replication inhibitors is desirable. Here, we developed a cell-based screening system by establishing a cell line that stably expresses influenza viral ribonucleoprotein complex (vRNP). Compound library screening using this cell line allowed us to identify a compound that inhibits vRNA transcription/replication by using reporter protein expression from virus-like RNA as a readout and virus replication in vitro. vRNP-expressing cells have potential as a simple and convenient high-throughput screening (HTS) system, and, thus, are promising to identify vRNA transcription/replication inhibitors for various RNA viruses, especially for primary screens.
Similar content being viewed by others
Introduction
To control influenza virus outbreaks, we have two options: vaccines and antivirals. The effect of vaccines is, however, limited to virus strains that are antigenically closely related to the vaccine reference virus. To combat new influenza viruses with vaccination, such as the recently emerged swine-origin pandemic (H1N1) 2009 virus, vaccine production took 3–6 months. Therefore, we are highly dependent on antiviral compounds for the treatment and prevention of influenza, particularly in pandemic situations. Although two classes of anti-influenza virus agent, M2 ion channel blockers1 and neuraminidase inhibitors2, are currently licensed, resistance to both types of antivirals has already been found among seasonal H1N13, highly-pathogenic avian H5N14,5 and pandemic (H1N1) 20096,7,8 viruses. There is, therefore, an urgent need to develop novel antivirals that target steps in the virus life cycle other than the M2 ion channel or neuraminidase activity.
Influenza viruses are members of the family Orthomyxoviridae and are classified into three types: influenza A, B and C viruses. All three pandemic viruses in the last century, as well as the currently circulating H5N1 viruses, are type A viruses, which possess eight-segmented negative-sense RNAs as their genome9. Each viral RNA (vRNA) is transcribed and replicated by forming viral ribonucleoprotein complexes (vRNPs) together with heterotrimeric viral polymerase subunit proteins (PA, PB1 and PB2) and nucleoprotein (NP): these four viral proteins are necessary and sufficient for vRNA transcription and replication in cultured cells10. In the virus-infected cell, vRNPs from incoming viruses are transported into the nucleus, where vRNA transcription and replication take place [for a review, see reference11]. Negative-sense vRNA, whose noncoding regions at the 3′ and 5′ ends serve as a promoter for viral polymerase-mediated RNA synthesis, are transcribed into complementary RNA (cRNA) and mRNA. The synthesized cRNA, which is positive-sense but not capped at the 5′ end and polyadenylated at the 3′ end, subsequently acts as the template for vRNA replication. vRNA transcription/replication is essential for virus replication and is thus an attractive influenza antiviral target.
Recently, high-throughput screening (HTS) of compound libraries has enhanced target-based drug discovery12. Although the original HTS system was developed for cell-free assays a decade ago, rapid technical innovations have enabled its application to cell-based assays13. Cell-based compound library screening is ideal for uncovering agents that target influenza vRNA transcription/replication, because various known and unknown cellular components are involved in these processes, together with multiple viral proteins and vRNA. In addition, drug efficacy must be evaluated in cells. Furthermore, cell-based assays have the advantage over cell-free assays of eliminating cytotoxic, membrane-impermeable, or intracellularly inactivate agents from a large library of drug candidates.
Influenza vRNPs can be transiently reconstituted in cells by using plasmid transfection; however, the efficiency and, therefore, gene expression levels greatly fluctuate among cells. Such conditions are not suitable for compound screening assays, since they do not permit consistent and reproducible results. Therefore, we sought to establish a cell line that stably expresses the five components of influenza vRNP (i.e., a virus-like RNA, PB1, PB1, PA and NP). To this end, we used retroviral vectors that facilitate the efficient integration and stable expression of multiple genes of interest and established a vRNP-expressing cell line. Our vRNP-expressing cell line represents a simple, convenient and reliable HTS system for the identification of influenza vRNA transcription/replication inhibitors.
Results
Retroviral vector-mediated influenza vRNP formation
To evaluate whether a retroviral vector could be used to produce functional vRNPs, we transduced human embryonic kidney-derived 293 cells with a retroviral vector expressing a GFP-encoding influenza virus-like RNA under the control of the human RNA polymerase I (PolI) promoter and the mouse PolI terminator (Figure 1A). Simultaneous transfection with four plasmids for the expression of the influenza viral polymerase subunits and NP resulted in GFP expression 48 h post-transduction, whereas no GFP expression was detected in cells transduced with the vector alone (Figure 1B). To confirm that the virus-like RNA was indeed expressed from the integrated retroviral vector, an envelope protein-uncoated retroviral vector with the virus-like RNA transcription cassette was prepared and transduced into 293 cells. Simultaneous transfection with the four expression plasmids for the viral polymerase subunits and NP resulted in limited GFP expression 48 h post-transduction (Supplementary Figure S1). These results indicate that retroviral vector transduction leads to the formation of functional influenza vRNP in cells.
Establishment of 293 cells stably expressing influenza vRNP by retroviral vectors.
(A) A schematic diagram of a retroviral vector for virus-like RNA. The virus-like RNA encoding GFP comprises the 3′ noncoding region of NP vRNA (3′ NCR), the GFP open reading frame in the negative sense and the 5′ noncoding region of NP vRNA (5′ NCR), expressed under the control of the human PolI promoter (PPolI) and the mouse PolI terminator (TPolI). 5′ LTR, murine leukemia virus (MLV) 5′ long terminal repeat; Ψ, MLV packaging signal; Δ3′ LTR, MLV U3 region-deleted 3′ long terminal repeat. (B) Virus-like vRNA expression by retroviral vector transduction. 293 cells were transduced with the retroviral vector for the expression of the GFP-encoding virus-like RNA. Simultaneously, the cells were transfected with plasmids for the expression of the polymerase subunits (PB2, PB1 and PA) and NP (right panels) or with nothing further (left panels). Forty-eight hours later, GFP expression (lower panels) was examined by fluorescence microscopy. The upper panels show bright-field images. Scale: 50 μm.
Establishment and characterization of influenza vRNP-expressing cells
Next, to establish a cell clone constitutively expressing vRNP, we generated a retroviral vector for the expression of a vRNA encoding a puromycin resistance gene. We then co-transduced 293 cells with this vector and the four additional retroviral vectors expressing PB2, PB1, PA and NP and cultured in the presence of puromycin. Although a total of 300 cell clones exhibited puromycin-resistance, most grew slowly. One clone, however, proliferated reasonably well and we designated it 293vRNP-Puro.
To validate the expression of the four viral proteins (i.e., PB2, PB1, PA and NP) and the virus-like RNA encoding puromycin resistance gene, total RNAs were extracted from 293vRNP-Puro and the parental 293 cells and subjected to RT-PCR with specific primers. Whereas no RT-PCR product for the vRNP components was detected in 293 cells, mRNAs for each viral protein (Supplementary Figure S2A) and negative sense (i.e. vRNA sense) RNA corresponding to the puromycin resistance vRNA (Supplementary Figure S2B) were detected in 293vRNP-Puro cells. Further, to demonstrate the functionality of the four viral proteins in cells, 293vRNP-Puro cells were transduced with adenovirus vector AdV/PolI-GFP14, which expresses a virus-like RNA encoding GFP gene, at a multiplicity of infection of 10. GFP expression was observed in most of the transduced cells, whereas no GFP expression was detected in mock-transduced cells (Supplementary Figure S2C). These results indicate that the vRNA encoding puromycin-resistance gene and viral proteins required for vRNA transcription are stably expressed in 293vRNP-Puro cells and suggest that the puromycin resistance of 293vRNP-Puro cells was conferred by the puromycin resistant gene product from the vRNP.
Influenza vRNP-expressing cell-based screening assay for influenza vRNA transcription/replication inhibitors
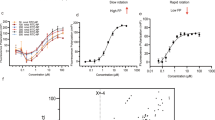
To test 293vRNP-Puro cells for their applicable to a cell-based screening assay for influenza vRNA transcription/replication inhibitors, we used favipiravir (also called T-705; 6-fluoro-3-hydroxy-2-pyrazinecarboxamide), which inhibits influenza virus RNA polymerase activity15,16,17. The inhibitory effect of favipiravir on influenza vRNA transcription/replication was confirmed by a plasmid-based minigenome assay18; favipiravir inhibited vRNA transcription/replication in dose-dependent manner (Supplementary Figure S3A) and the 50% inhibitory concentration (IC50) of favipiravir under these conditions was 110.1 μM. We then assessed the effect of favipiravir on the viability of 293vRNP-Puro cells in the presence or absence of puromycin. Given the time required for cytotoxicity induction with puromycin, the cells were plated at relatively low concentration (2,000 cells per well on 384-well plates) and cultured for a relatively long time (120 h). We added various concentrations of favipiravir to the cells 24 h after seeding and incubated them for a further 96 h. Cell viability was measured by use of the CellTiter-Glo® Luminescent Cell Viability Assay (Promega, Madison, WI), which can be readily applied to an HTS format. Under these HTS-optimized conditions, favipiravir reduced the viability of 293vRNP-Puro cells grown in puromycin-containing medium in a dose-dependent manner, whereas the cell viability in the absence of puromycin was unaffected by favipiravir (Supplementary Figure S3B). These results suggest that the puromycin-resistance of 293vRNP-Puro cells is dependent on vRNA transcription/replication. The IC50 calculated from the 293vRNP-Puro cell-based assay was 1285 μM and thus the detection sensitivity of the cell-based assay was about 12-fold lower than that of the plasmid-based assay using luciferase activity originating from virus-like RNA as a readout.
Despite this lower detection sensitivity, 293vRNP-Puro cells clearly have potential to provide a simple and convenient HTS system to identify factors that affect influenza vRNA replication/transcription. To test the system's ability to select compounds that inhibit influenza vRNA transcription/replication, 4,160 compounds from three commercially available compound libraries, LOPAC1280 Navigator (Sigma-Aldrich, St. Louis, MO), the Spectrum Collection (Microsource Discovery Systems, Inc., Gaylordsville, CT) and the Prestwick Chemical Library (Prestwick Chemical, Inc., Illkirch, France), were screened on the basis of 293vRNP-Puro cell viability in the presence and absence of puromycin (Supplementary Figure S4A). After two rounds of screening, 11 compounds that exhibited more than 90% and less than 5% cell viability in the normal and puromycin-containing media, respectively, were identified as candidate influenza vRNA transcription/replication inhibitors (Supplementary Figure S4B). The Z′ values, a statistical parameter used to evaluate and validate the performance and robustness of HTS assays19, for the first and second screenings were 0.51–0.82 and 0.71–0.88, respectively, indicating the robustness of the 293vRNP-Puro cell-based screening assay [Z′ values of > 0.5 are considered acceptable for HTS systems19].
Identification of an influenza vRNA transcription/replication inhibitor
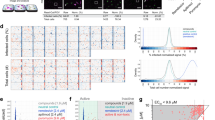
To assess the inhibitory effect of the selected compounds on influenza vRNA transcription/replication, the plasmid-based minigenome assay was performed in the presence of the compounds; since two of the 11 compounds (compound IDs, B21 and D10) were then not available, we tested the remaining nine. While none of the compounds tested exhibited cytotoxicity in 293 cells (Figure 2A), five exhibited a significant inhibitory effect on minigenome expression (Figure 2B), suggesting anti-influenza virus activity. We, therefore, assessed the inhibitory effect of the five compounds on in vitro replication of two influenza virus strains, a laboratory strain A/WSN/33 (H1N1) and a pandemic (H1N1) 2009 strain A/California/04/09 (H1N1) (Figure 2C). One of the compounds tested, D12, 3beta-acetoxydeoxodihydrogedunin (Figure 3A), significantly impaired virus replication. This compound did not exhibit any inhibitory effect on the replication of either influenza B or Sendai virus (Figure 3B). These results indicate that this compound inhibits influenza virus replication by affecting vRNA transcription/replication.
In vitro characterization of influenza vRNA transcription/replication inhibitor candidates.
(A) Cytotoxicity of the identified compounds. 293 cells were cultured with the indicated compound (10 μM each: FP, favipiravir) or DMSO (Ctrl) for 48 h. Cell viability was measured by using the CellTiter-Glo® Luminescent Cell Viability Assay (Promega). Error bars indicate standard deviations of triplicate experiments. (B) Effects of identified compounds on influenza virus transcription/replication in a minigenome assay. 293 cells were transfected with plasmids for the expression of a virus-like RNA encoding the firefly luciferase gene together with expression plasmids for PB2, PB1, PA, NP and Renilla luciferase, which served as an internal control. Simultaneously, the indicated compound (10 μM each: FP, favipiravir) or DMSO (Ctrl) was added to the cells. Twenty-four hours later, luciferase activity was measured by use of the Dual-Luciferase® Reporter Assay System (Promega). Error bars indicate standard deviations of triplicate experiments. Statistical significance was assessed by use of the Student's t-test: *, P < 0.05. (C) Effects of identified compounds on influenza virus replication. 293 cells were infected with A/WSN/33 (H1N1) or A/California/04/09 (H1N1) at a multiplicity of infection of 0.001. One hour later, the indicated compound (10 μM each: FP, favipiravir) or DMSO (Ctrl) was added to the cells. Virus titers in the culture supernatants at 48 h post-infection were determined by plaque assays in MDCK cells. Error bars indicate standard deviations of triplicate experiments. Statistical significance was assessed by use of the Student's t-test: *, P < 0.05.
Characterization of 3beta-acetoxydeoxodihydrogedunin.
(A) Structure of 3beta-acetoxydeoxodihydrogedunin. (B) Effects of identified compounds on Sendai and influenza B viruses replication. 293 cells were infected with influenza B (left) or Sendai (right) virus at a multiplicity of infection of 0.001. One hour later, 10 μM 3beta-acetoxydeoxodihydrogedunin (3beta) or DMSO was added to the cells. Virus titers in the culture supernatants at 48 h post-infection were determined by plaque assays in MDCK cells. (C–F) Effects of 3beta-acetoxydeoxodihydrogedunin on the in vitro growth kinetics of influenza virus. Human embryonic kidney-derived 293 (C), human lung adenocarcinoma A549 (D), canine kidney-derived MDCK (E), or chicken embryonic fibroblast DF1 cells (F) were infected with A/WSN/33 (H1N1) at a multiplicity of infection of 0.001. One hour later, 10 μM 3beta-acetoxydeoxodihydrogedunin (3beta) or DMSO was added to the cells. Virus titers in the culture supernatants were determined at the indicated time points by means of plaque assays in MDCK cells. Error bars indicate standard deviations of entirely separate triplicate experiments. Statistical significance was assessed by use of the Student's t-test: *, P < 0.05.
To further assess the in vitro efficacy of 3beta-acetoxydeoxodihydrogedunin against influenza virus, the growth kinetics of A/WSN/33 (H1N1) in the presence or absence of the compound were determined in the following cell lines: human embryonic kidney-derived 293, human lung adenocarcinoma A549, canine kidney-derived MDCK and chicken embryonic fibroblast DF1 cells (Figure 3C–F). Virus growth in all of the cells tested was significantly impaired and/or delayed, especially at early time points, indicating the broad inhibitory effect of 3beta-acetoxydeoxodihydrogedunin on influenza virus replication. In 293 cells, the IC50 of this compound against A/WSN/33 (H1N1) replication was 44.6 nM and the 50% cytotoxicity concentration was over 50 μM. These results suggest that 3beta-acetoxydeoxodihydrogedunin is a promising inhibitor of influenza A vRNA transcription/replication.
Mechanism of action of 3beta-acetoxydeoxodihydrogedunin
3beta-acetoxydeoxodihydrogedunin is a derivative of gedunin, which is a limonoid from the Bangladeshi mangrove tree X. granatum. Some gedunin-derivatives inhibit the cellular chaperone heat shock protein 90 (Hsp90)20, although this inhibitory activity has not been reported for 3beta-acetoxydeoxodihydrogedunin. Therefore, to test whether 3beta-acetoxydeoxodihydrogedunin inhibits Hsp90 activity, we assessed its effect on the expression of two Hsp90 client proteins, the serine/threonine kinase AKT and the receptor tyrosine kinase HER2. In the presence of 3beta-acetoxydeoxodihydrogedunin, AKT and HER2 were degraded in a compound dose-dependent manner, whereas the expression levels of Hsp90 and beta-actin (as internal controls) were not affected (Figure 4). Geldanamycin, a well-known Hsp90 inhibitor that inhibits influenza virus replication21, also prevented the maturation of the Hsp90 client proteins (Figure 4). These results suggest that 3beta-acetoxydeoxodihydrogedunin inhibits Hsp90 activity.
Effect of 3beta-acetoxydeoxodihydrogedunin on Hsp90 client proteins.
293 cells were treated with 3beta-acetoxydeoxodihydrogedunin (3beta; 2, 10, or 50 μM), geldanamycin (GA; 1 μM), or DMSO for 48 h. The expression levels of the serine/threonine kinase AKT and the receptor tyrosine kinase HER2, both of which are Hsp90 client proteins and of Hsp90 and beta-actin were analyzed by means of western blotting.
Discussion
Here, we developed a cell-based compound screening system for influenza A vRNA transcription/replication inhibitors by establishing a human cell line stably expressing influenza vRNP components: four viral proteins (PB2, PB1, PA and NP) together with a virus-like RNA (Figure 1 and Supplementary Figures S1–4). The reconstituted vRNP may not perfectly reflect the nature of the infection process. Further, the detection sensitivity of our 293vRNP-puro cell-based system was about 12-fold lower than that of the plasmid-based assay (Supplementary Figure S3). However, since the screening read-out is cell viability and further modifications (e.g., plasmid transfection and virus infection) are not necessary, our vRNP-expressing cell line represents a simple, convenient and reliable HTS system. Given its low detection sensitivity and the possibility that the hits based on this screening system may interfere with the PolI transcription machinery, our cell line-based HTS system may be more suitable for primary screens of antiviral agents.
By using a cell-based compound screening system, we identified 3beta-acetoxydeoxodihydrogedunin as a candidate inhibitor of influenza A vRNA transcription/replication. This compound inhibited minigenome expression (Figure 2B) and virus replication (Figure 2C). Gedunin and Gedunin-like limonoids have inhibitory activity against HIV (http://home.ncifcrf.gov/mtdp/Catalog/compounds/309912.html), parasites22,23,24 and insects25,26,27. Our results show that 3beta-acetoxydeoxodihydrogedunin, as well as some other gedunin-derivatives, inhibits the activity of the Hsp90 chaperone. Hsp90 plays a role in nuclear import and assembly of the influenza viral polymerase complex by interacting with PB1 and PB228 and stimulates in vitro influenza vRNA synthesis29. Other Hsp90 inhibitors, such as geldanamycin and its derivative 17-AAG, also impair influenza virus replication21, attesting to the potential of 3beta-acetoxydeoxodihydrogedunin as an anti-influenza agent.
Our 293vRNP-puro cells represent the first cell line to simultaneously express all five components of the influenza vRNP (i.e., a virus-like RNA, PB1, PB1, PA and NP), although cell lines expressing only the reporter gene-encoding virus-like RNA30,31,32 or the four viral proteins33 have been reported previously. One of the key steps in this process was the generation of a retroviral vector that possesses the PolI transcription cassette for influenza virus-like RNA expression (Figure 1A). PolI-driven vRNA expression has been applied to several negative-strand RNA viruses including biosafety level 4 agents such as Ebolavirus34 and Crimean-Congo hemorrhagic fever virus35. vRNP (or nucleocapsid)-expressing cell lines does not require highly-contained facilities to handle infectious viruses. Our retroviral vector-based strategy thus has the potential to accelerate development of antivirals against various RNA viruses by providing promising HTS systems for vRNA transcription/replication inhibitors.
The drug resistance gene on the virus-like RNA allowed us to easily establish a cell line expressing multiple foreign genes and to screen many compounds based on cell viability. Replacing the drug resistance gene with a reporter gene (e.g., GFP and luciferase genes) would increase the detection sensitivity, representing an alternative strategy to obtain the desired cell clones and maintain the expression of all of the foreign genes.
In conclusion, we developed a simple, convenient and reliable compound screening system for vRNA transcription/replication inhibitors that is based on a vRNP-expressing cell line. By using this system, we identified a promising candidate inhibitor of influenza A vRNA transcription/replication.
Methods
Cells and viruses
Plat-GP cells, which highly and stably express retroviral structure genes derived from murine leukemia virus (MLV) except for the env gene36 were kindly provided by T. Kitamura (University of Tokyo). These cells, as well as human embryonic kidney-derived 293 cells, 293T and 293vRNP-Puro cells (established in this study), were maintained in Dulbecco's modified Eagle's medium supplemented with 10% fetal calf serum. Madin-Darby canine kidney (MDCK) cells were maintained in Eagle's minimal essential medium (MEM) supplemented with 5% newborn calf serum. All cells were cultured at 37°C in 5% CO2. For maintenance of Plat-GP cells and 293vRNP-Puro cells, 10 μg/ml blasticidin and 2 μg/ml puromycin were added to the culture media, respectively. A laboratory strain A/WSN/33 (H1N1) and a pandemic (H1N1) 2009 strain A/California/04/09 (H1N1) were generated by plasmid-based reverse getetics10 in 293T cells and propagated in MDCK cells. B/Lee/40 and Sendai (Enders strain; kindly provided by Allan Portner, St. Jude Children's Research Hospital, Memphis, TN) viruses were propagated in chicken embryonated eggs.
Plasmid construction and transfection
To produce a retroviral vector for the expression of a virus-like RNA encoding GFP gene, we cloned the cDNA corresponding to the transcriptional region in pPolI-GFP37 [i.e., the cDNA corresponding to the 3′ noncoding ends of NP vRNA, the GFP open reading frame and the 5′ noncoding region of NP vRNA flanked by the human RNA polymerase I (PolI) promoter and the mouse PolI terminator] into a retroviral packaging plasmid pMXΔU3, which was based on the retroviral packaging plasmid pMX38 and whose U3 region of the 3′ long terminal repeat (LTR) was deleted to create so-called self-inactivation retroviral vectors39, resulting in pMXΔU3/PolI-GFP. Similarly, we cloned the transcriptional region in pPolIHA(0)Puro(0), which expresses HA vRNA encoding the puromycin-resistance gene, into pMXΔU3, resulting in pMXΔU3/PolI-HA(0)Puro(0). To produce retroviral vectors for the expression of the viral polymerase subunits (PB2, PB1 and PA) and nucleoprotein (NP), the cDNAs corresponding to the open reading frames of each protein of the influenza virus strain A/WSN/33 (H1N1) were cloned into pMX. All plasmid constructs were sequenced to ensure that no undesirable mutations were introduced. Plasmid transfections into 293 or Plat-GP cells were performed with TransIT-293 reagent (Mirus, Madison, WI) according to the manufacturer's instructions.
Retroviral vector production and transduction
MLV-based retroviral vectors were produced by co-transfection of pMX (or pMXΔU3) encoding the gene of interest and the expression plasmid for the G glycoprotein of vesicular stomatitis virus (VSVG)40 into Plat-GP cells. At 48 h post-transfection, culture supernatants were collected, clarified through a 0.45 μm filter and stored at −80°C. For retroviral vector transduction, 5-fold-diluted retroviral vectors with fresh medium were replaced with the culture media of 293 cells.
RNA extraction and reverse transcription-PCR (RT-PCR)
Total RNAs were extracted from 293 and 293vRNP-Puro cells by using the Illustra RNAspin mini RNA Isolation Kit (GE Healthcare, Buckinghamshire, UK) according to the manufacturer's instructions. The extracted RNA was reverse transcribed using SuperScriptTM III reverse transcriptase (Invitrogen, Carlsbad, CA) and either oligo(dT)16 primer (Supplementary Figure S2A) or random hexamer primer (Supplementary Figure S2B). The former and latter cDNAs were amplified by PCR with primers specific for the 3′ and 5′ ends of the open reading frames of each protein (Supplementary Figure S2A) and with primers specific for the 3′ and 5′ noncoding ends of HA vRNA (Supplementary Figure S2B), respectively. In both cases, a partial fragment of the human glyceraldehyde-3-phosphate dehydrogenase (hGAPDH) gene, corresponding to 598 bps, was amplified with the specific primers as an internal control.
Plasmid-mediated reporter assay for vRNA transcription/replication activity
A plasmid-based minigenome assay was performed as described previously18. Briefly, 293 cells were co-transfected with plasmids for the expression of viral polymerase complex proteins (i.e., PB2, PB1, PA and NP) and a firefly luciferase gene-encoding influenza viral minigenome along with pGL4.74[hRluc/TK] (Promega), which expresses Renilla luciferase and served as an internal control. At 24 h post-transfection, luciferase activity was measured by use of the Dual-Luciferase® Reporter Assay System (Promega).
Cell viability
Cell viability was assessed by using the CellTiter-Glo® Luminescent Cell Viability Assay (Promega) according to the manufacturer's instructions. Briefly, a volume of CellTiter-Glo® reagent equal to that of the culture media was added to 293vRNP-Puro cells cultured on color-walled 384-well plates. Plates were shaken on a plate shaker for two min to induce cell lysis, incubated at room temperature for ten min and subjected to luminescence measurement.
Statistical analysis
Statistical significance in cell viability, relative luciferase activity and virus titer was assessed by use of the Student's t-test. The Z′ value, a statistical parameter used to evaluate and validate the performance and robustness of HTS assays19, was calculated for each assay plate as follows: Z′ = 1 – (3 × SD of positive control + 3 × SD of negative control)/|mean of positive control – mean of negative control|, where SD represents the standard deviation. DMSO and staurosporine (Sigma-Aldrich), which is an apoptosis inducer, served as the positive and negative control, respectively. A Z′ value between 0.5 and 1.0 is considered robust enough for an HTS assay.
Western blotting
293 cells treated with 3beta-acetoxydeoxodihydrogedunin, geldanamycin, or DMSO for 48 h were lysed with Passive Lysis Buffer (Promega). Proteins in the lysates were separated by sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) and transferred to nitrocellulose membranes by using a iBlot dry blotting system (Invitrogen). The serine/threonine kinase AKT, receptor tyrosine kinase HER2, Hsp90 and beta-actin were detected by monoclonal antibodies against AKT (clone 9Q7, Pierce), HER2 (clone e2-4001 + 3B5, Thermo Scientific), Hsp90 (clone AC88, abcam) and beta-actin (clone AC-15, Sigma-Aldrich), respectively.
References
Davies, W. L. et al. Antiviral Activity of 1-Adamantanamine (Amantadine). Science 144, 862–863 (1964).
Hayden, F. G. Perspectives on antiviral use during pandemic influenza. Philos Trans R Soc Lond B Biol Sci 356, 1877–1884 (2001).
Tamura, D. et al. Oseltamivir-resistant influenza a viruses circulating in Japan. J Clin Microbiol 47, 1424–1427 (2009).
de Jong, M. D. et al. Oseltamivir resistance during treatment of influenza A (H5N1) infection. N Engl J Med 353, 2667–2672 (2005).
Le, Q. M. et al. Avian flu: isolation of drug-resistant H5N1 virus. Nature 437, 1108 (2005).
Centers for Disease Control and Prevention. Oseltamivir-resistant novel influenza A (H1N1) virus infection in two immunosuppressed patients - Seattle, Washington, 2009. MMWR. Morbidity and mortality weekly report 58, 893–896 (2009).
Leung, T. W., Tai, A. L., Cheng, P. K., Kong, M. S. & Lim, W. Detection of an oseltamivir-resistant pandemic influenza A/H1N1 virus in Hong Kong. J Clin Virol 46, 298–299 (2009).
Le, Q. M. et al. A community cluster of oseltamivir-resistant cases of 2009 H1N1 influenza. N Engl J Med 362, 86–87 (2010).
Palese, P. a. S. M. L. in Fields virology (ed Knipe D. M. and Howley P. M.) 1647–1689 (Lippincott-Raven Publishers, 2007).
Neumann, G. et al. Generation of influenza A viruses entirely from cloned cDNAs. Proc Natl Acad Sci U S A 96, 9345–9350 (1999).
Neumann, G., Brownlee, G. G., Fodor, E. & Kawaoka, Y. Orthomyxovirus replication, transcription and polyadenylation. Curr Top Microbiol Immunol 283, 121–143 (2004).
Hertzberg, R. P. & Pope, A. J. High-throughput screening: new technology for the 21st century. Current opinion in chemical biology 4, 445–451 (2000).
Johnston, P. A. & Johnston, P. A. Cellular platforms for HTS: three case studies. Drug discovery today 7, 353–363 (2002).
Ozawa, M., Goto, H., Horimoto, T. & Kawaoka, Y. An adenovirus vector-mediated reverse genetics system for influenza A virus generation. J Virol 81, 9556–9559 (2007).
Furuta, Y. et al. In vitro and in vivo activities of anti-influenza virus compound T-705. Antimicrob Agents Chemother 46, 977–981 (2002).
Furuta, Y. et al. Mechanism of action of T-705 against influenza virus. Antimicrob Agents Chemother 49, 981–986 (2005).
Sidwell, R. W. et al. Efficacy of orally administered T-705 on lethal avian influenza A (H5N1) virus infections in mice. Antimicrob Agents Chemother 51, 845–851 (2007).
Ozawa, M. et al. Contributions of two nuclear localization signals of influenza A virus nucleoprotein to viral replication. J Virol 81, 30–41 (2007).
Zhang, J. H., Chung, T. D. & Oldenburg, K. R. A Simple Statistical Parameter for Use in Evaluation and Validation of High Throughput Screening Assays. J Biomol Screen 4, 67–73 (1999).
Hieronymus, H. et al. Gene expression signature-based chemical genomic prediction identifies a novel class of HSP90 pathway modulators. Cancer cell 10, 321–330 (2006).
Chase, G. et al. Hsp90 inhibitors reduce influenza virus replication in cell culture. Virology 377, 431–439 (2008).
MacKinnon, S. et al. Antimalarial activity of tropical Meliaceae extracts and gedunin derivatives. J Nat Prod 60, 336–341 (1997).
Bickii, J., Njifutie, N., Foyere, J. A., Basco, L. K. & Ringwald, P. In vitro antimalarial activity of limonoids from Khaya grandifoliola C.D.C. (Meliaceae). J Ethnopharmacol 69, 27–33 (2000).
Hay, A. E. et al. Limonoid orthoacetates and antiprotozoal compounds from the roots of Pseudocedrela kotschyi. J Nat Prod 70, 9–13 (2007).
Nathan, S. S., Kalaivani, K. & Murugan, K. Effects of neem limonoids on the malaria vector Anopheles stephensi Liston (Diptera: Culicidae). Acta tropica 96, 47–55 (2005).
Koul, O., Multani, J. S., Singh, G., Daniewski, W. M. & Berlozecki, S. 6beta-hydroxygedunin from Azadirachta indica. Its potentiation effects with some non-azadirachtin limonoids in neem against lepidopteran larvae. J Agric Food Chem 51, 2937–2942 (2003).
Cespedes, C. L., Calderon, J. S., Lina, L. & Aranda, E. Growth inhibitory effects on fall armyworm Spodoptera frugiperda of some limonoids isolated from Cedrela spp. (Meliaceae). J Agric Food Chem 48, 1903–1908 (2000).
Naito, T., Momose, F., Kawaguchi, A. & Nagata, K. Involvement of Hsp90 in assembly and nuclear import of influenza virus RNA polymerase subunits. J Virol 81, 1339–1349 (2007).
Momose, F. et al. Identification of Hsp90 as a stimulatory host factor involved in influenza virus RNA synthesis. J Biol Chem 277, 45306–45314 (2002).
Groseth, A., Feldmann, H., Theriault, S., Mehmetoglu, G. & Flick, R. RNA polymerase I-driven minigenome system for Ebola viruses. J Virol 79, 4425–4433 (2005).
Flick, R., Flick, K., Feldmann, H. & Elgh, F. Reverse genetics for crimean-congo hemorrhagic fever virus. J Virol 77, 5997–6006 (2003).
Morita, S., Kojima, T. & Kitamura, T. Plat-E: an efficient and stable system for transient packaging of retroviruses. Gene Ther 7, 1063–1066 (2000).
Neumann, G., Watanabe, T. & Kawaoka, Y. Plasmid-driven formation of influenza virus-like particles. J Virol 74, 547–551 (2000).
Onishi, M. et al. Applications of retrovirus-mediated expression cloning. Exp Hematol 24, 324–329 (1996).
Yu, S. F. et al. Self-inactivating retroviral vectors designed for transfer of whole genes into mammalian cells. Proc Natl Acad Sci U S A 83, 3194–3198 (1986).
Shimojima, M. et al. Tyro3 family-mediated cell entry of Ebola and Marburg viruses. J Virol 80, 10109–10116 (2006).
Lutz, A. et al. Virus-inducible reporter genes as a tool for detecting and quantifying influenza A virus replication. J Virol Methods 126, 13–20 (2005).
Hossain, M. J. et al. Establishment and characterization of a Madin-Darby canine kidney reporter cell line for influenza A virus assays. J Clin Microbiol 48, 2515–2523 (2010).
Zhang, J. et al. Identification of novel virus inhibitors by influenza A virus specific reporter cell based screening. Antiviral Res 93, 48–54 (2012).
Nakamura, Y., Oda, K. & Nakada, S. Growth complementation of influenza virus temperature-sensitive mutants in mouse cells which express the RNA polymerase and nucleoprotein genes. J Biochem 110, 395–401 (1991).
Acknowledgements
We thank Qiao Xue and Lisa Burley for technical assistance and Susan Watson for editing the manuscript. This work was supported, in part, by Grants-in-Aid for Specially Promoted Research and for Scientific Research, by a Contract Research Fund for the Program of Founding Research Centers for Emerging and Reemerging Infectious Diseases, by ERATO (Japan Science and Technology Agency), by the Special Coordination Funds for Promoting Science and Technology from the Ministry of Education, Culture, Sports, Science and Technology of Japan and by National Institute of Allergy and Infectious Diseases Public Health Service research grants, USA. MO was partly supported by a Japan Society for the Promotion of Science Postdoctoral Fellowship for Research Abroad. None of the funding sources had any role in the design or conduct of the study, in the collection, management, analysis, or interpretation of the data, or in the preparation, review, or approval of the manuscript.
Author information
Authors and Affiliations
Contributions
MO and YK designed the study; MO, MH, MS, HG, SW, YH and NP performed the experiments; YF, NP and MH provided reagents/materials; MO and YK analyzed the data and wrote the manuscript.
Ethics declarations
Competing interests
YK has received speaker's honoraria from Chugai Pharmaceuticals, Novartis, Daiichi-Sankyo, Toyama Chemical, Wyeth and GlaxoSmithKline; grant support from Chugai Pharmaceuticals, Daiichi Sankyo Pharmaceutical and Toyama Chemical; is a consultant for Theraclone and Crucell and is a founder of FluGen.
Electronic supplementary material
Supplementary Information
Supplementary Information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Ozawa, M., Shimojima, M., Goto, H. et al. A cell-based screening system for influenza A viral RNA transcription/replication inhibitors. Sci Rep 3, 1106 (2013). https://doi.org/10.1038/srep01106
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep01106
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.