Abstract
The use of small molecules to target miRNAs is a new type of therapy for human diseases, particularly cancers. We proposed a novel high-throughput approach to identify the biological links between small molecules and miRNAs in 23 different cancers and constructed the Small Molecule-MiRNA Network (SMirN) for each cancer to systematically analyze the properties of their associations. In each SMirN, we partitioned small molecules (miRNAs) into modules, in which small molecules (miRNAs) were connected with one miRNA (small molecule). Almost all of the miRNA modules comprised miRNAs that had similar target genes and functions or were members of the same miRNA family. Most of the small molecule modules involved compounds with similar chemical structures, modes of action, or drug interactions. These modules can be used to identify drug candidates and new indications for existing drugs. Therefore, our approach is valuable to drug discovery and cancer therapy.
Similar content being viewed by others
Introduction
miRNAs are single-stranded non-coding RNA molecules that play fundamental roles in multiple biological processes, such as cell differentiation, proliferation, apoptosis and so on1. Many studies have identified alterations of miRNA expression in a variety of human cancers2,3. Furthermore, complex interactions exist among miRNAs, experimental factors and phenotypes, especially drugs and diseases4. And the secondary structures of miRNA, such as stem loops and bulges, are targets to which pharmaceuticals can be applied5. Currently, RNA inhibition agents, which include antisense oligonucleotides, small interfering RNAs and anti-miRNA agents, have been applied to preclinical and clinical studies that have investigated potential therapies for human disorders6. Therefore, targeting over-expressed miRNAs is a potential methodology to develop miRNA-specific drugs for therapeutic purposes. To this end, some recent efforts have focused on miRNA-targeted drug discovery. Santaris has developed SPC3649, which is a potential treatment for hepatitis C infection. As the first miRNA-targeted drug to enter human clinical trials, SPC3649 successfully inhibits miR-122 which is required by the hepatitis C virus for replication7,8. Gumireddy et al. have conducted a primary screen of >1,200 compounds and determined that approximately 100 structurally modified molecules are related to the diazobenzene core structure. They found diazobenzene and its derivatives are effective inhibitors of miR-219.
In addition, miRNAs may affect chemoresistance. Blower et al. have investigated the impact of cellular levels of let-7i, miR-16 and miR-21 on the growth-inhibitory potencies of 14 anticancer compounds and found that miRNAs modulate the sensitivity and resistance to anticancer drugs in substantial ways10. Previous studies have shown that anticancer drugs and miRNAs may have common target genes11. If the target receptors participate in the overall clinical effects of a drug, these receptors may contribute to drug resistance12. Yang et al. have demonstrated that miR-214 induces cisplatin resistance by primarily targeting the PTEN/Akt pathway in human ovarian cancer13.
In summary, investigating the relationships between small molecules (potential drugs) and miRNAs is at the initial stage. Although Calin has suggested that structure-based approaches, such as molecular docking, are useful to identify compounds that target miRNAs5, the three-dimensional structure prediction of miRNA remains a challenge. However, gene expression profiles following drug treatments are available. Approaches that are based on transcriptional responses have been used to discover drug mode of action and identify new uses for existing drugs14. One of the major contributions of this study is the discovery of links between small molecules and miRNAs in 23 different cancers based on the differential expression of miRNA target genes and gene signatures from the gene expression profiles following treatment with small molecules. As a result, we constructed the Small Molecule-MiRNA Networks (SMirNs) for 17 different cancers and identified miRNA and small molecule modules. Using these networks and modules, we predicted drug target miRNAs, drug candidates for cancer therapy and unique clinical applications for known drugs.
Results
Small Molecule-MiRNA Networks (SMirNs) in human cancers
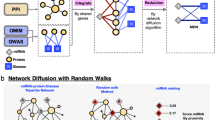
In order to identify the relationships between small molecules and miRNAs in human cancers, the transcriptional responses to 1,309 compounds and the differentially expressed genes in 23 human cancers were obtained from the Connectivity Map (cMap)15 and the unifying caner microarray resource (ONCOMINE)16, respectively. The miRNA target genes were obtained from our integrated database, which included seven widely used miRNA target genes prediction tools. Here, using the hypergeometric test we evaluated the extent to which the miRNA target genes appeared in the differentially expressed genes in cancer. At the significance level of 0.05, we identified 406 cancer-related miRNAs (CRMs) for 23 human cancers. The Gene Ontology (GO) enrichment analysis was used to partition the differentially expressed target genes of one CRM into significant gene ontology modules (GOMs) (see details in Materials and Methods). In each significant GOM, the association between a compound and the miRNA was evaluated by the Kolmogorov-Smirnov (KS) test based on miRNA differentially expressed target genes and transcriptional response of compound, which was represented as a list of genes ranked according to their differential expression. For each CRM in a specific cancer, if a significant (P≤0.05) association between the small molecule and the CRM was detected in at least 80% of significant GOMs, we hypothesized that a biological link existed between the small molecule and the miRNA in this cancer. As a result, 2,199 associations between 226 CRMs and 572 small molecules were discovered in 17 different cancers. The workflow diagram was shown in Figure 1.
We constructed a Small Molecule-MiRNA Network (SMirN) for each cancer (Figure 2). The sizes of the 17 networks exhibited great differences. The minimal one was ovary cancer specific SMirN, which only contained two miRNAs and one small molecule. In contrast, Bone marrow cancer specific SMirN involved 42 miRNAs, 283 small molecules and 981 interactions (Figures 3a and 3b). In all SMirNs, we determined that the majority of miRNAs (75%) were linked to a few small molecules (≤10). The average number of interactions per miRNA was 9.7. However, a few miRNAs, such as miR-125b, miR-139-5p, miR-302b, miR-487a and miR-570, significantly interacted with numerous (>65) small molecules. Similarly, the majority of small molecules (93%) were linked to a few miRNAs (≤10). The average number of interactions per small molecule was 3.8. Trichostatin A (TSA) interacted with the greatest number of miRNAs (216) and would be further analyzed.
The topological properties of SMirNs.
(a) The number of CRMs in each cancer. (b) The number of associated small molecules in each cancer. (c) The frequency of each small molecule in all SMirNs. (d) The frequency of each miRNA in all SMirNs. (e) The frequency of each miRNA-small molecule interaction in all SMirNs.
Next, the frequency of each small molecule in all 17 SMirNs was calculated (Figure 3c). We observed that TSA appeared in all SMirNs, which suggested that TSA may possess broad-spectrum anticancer activities. Previous studies have shown that TSA promotes the expression of some apoptosis-related genes and decreases tumor cells survival. Therefore, TSA may have the ability to inhibit the growth of many cancers17,18. miR-19a, miR-19b and miR-23b were the top 3 miRNAs with the highest frequency in the 17 SMirNs (Figure 3d). miR-19a and miR-19b are members of the miR-17-92 cluster and are potential human onco-miRNAs that play important roles in tumorigenesis19. The aberrant expression of miR-23b has been reported in several cancers20,21. In this study, we found that two target genes (ALDH1A1 and ALDH1A2) of miR-23b were drug targets of tretinoin that was associated with this miRNA in thyroid cancer (Figure 4a). Tretinoin is an anticancer drug that induces cells apoptosis and inhibits cells growth in some cancers22. Thus, our results indicated that miR-23b might be a new potential drug target for cancer therapy because of its ability to regulate the target genes ALDH1A1 and ALDH1A2. In addition, we determined the frequencies of small molecule-miRNA interactions in all SMirNs (Figure 3e). Interestingly, the associations between TSA and miR-19a, miR-19b and miR-23b were the top three interactions ranked based on frequency. The common targets of the three miRNAs were significantly enriched in the MAPK signaling pathway (P = 2.63×10−4, Figure 4b) and the mTOR signaling pathway (P = 1.38×10−3, Figure 4c) by using the hypergeometric test. The MAPK signaling pathway is crucial in the progression of tumor23. The mTOR is involved in human tumorigenesis and activated mTOR provides certain tumor cells with a growth advantage24. In conclusion, we predicted that TSA might affect the genesis and promote the development of multiple tumors by targeting the MAPK and mTOR signaling pathways.
(a) miR-23b and tretinoin, which was associated with this miRNA in thyroid cancer, shared the common target genes, ALDH1A1 and ALDH1A2. (b) The MAPK signaling pathway that was enriched with the common targets of miR-19a, miR-19b and miR-23b. The red nodes represented these common targets. (c) The mTOR signaling pathway that was enriched with the common targets of miR-19a, miR-19b and miR-23b. The red nodes represented these common targets.
In order to globally validate the predicted small molecule-miRNA associations, we manually collected the relevant literatures that investigated the relationships between small molecules and miRNAs from PubMed and Google scholar. Then we tested the extent to which the predicted associations appeared in the known associations using hypergeometric test. As a result, our predictions significantly covered the known associations (P = 1.26×10−12). Especially for the TSA, all but one of the experimentally validated associations were predicted by our approach. TSA is a member of the histone deacetylase inhibitor (HDACi), which is effective to the treatment of human tumors and being tested in clinical trials. Brest et al. investigated the mechanism of action responsible for therapeutic efficiency of HDACi in tumors25. They treated cancer cell lines with TSA and found that TSA treatment induced over-expression of miR-129-5p. The increasing of the miR-129-5p expression was related to antitumor activity of TSA, whereas the inactivation of miR-129-5p significantly blocked the TSA-induced cell death. Chen et al. pointed out that miR-373 was reactivated by the pharmacologic induction of TSA26. Zhang et al. evaluated the effect of TSA to miRNA expression in BxPC-3 human cancer cell27. It was found that miR-200c was over-expressed after TSA treatment by Northern blot analysis.
Biological insights from miRNA modules
In each of the cancer specific SMirNs, we identified miRNA modules, which were defined as groups of miRNAs that were connected with the same compound. According to the previous studies28,29,30,31 and the fraction of the modules with the following properties, the empirical value 5 was set as the threshold of the sizes of the modules. Only modules with at least 5 miRNAs were reserved for further study. Each miRNA module was coded with the name of the central compound. We investigated whether miRNA modules involved biologically similar miRNAs. To this end, we assessed all of the miRNA pairs in each miRNA module using three characteristics: common target genes, functional similarity and same miRNA family. Firstly, the “Meet/Min” score was employed to calculate the rate of co-regulating the same target genes that were differentially expressed in the cancer (see details in Materials and Methods). Secondly, the functional similarity of two miRNAs in the same module was evaluated based on the “Biological Process” category in GO by using Lin D's algorithm32. Finally, miRNA families were extracted from the miRBase database. We found that 90.625% of all miRNA modules involved miRNA pairs with at least one of the above similarities (Supplementary Tables S1 and S2). For example, in the miRNA module TSA of the esophagus cancer (Figure 5), a high rate of co-regulating the same differentially expressed targets and high functional similarity were detected between miR-15b and miR-195 (0.80 and 0.95), miR-15b and miR-424 (0.79 and 0.94). miR-15b and miR-195, miR-520h and miR-520g are the members of the miR-15 and miR-515 miRNA families, respectively.
Biological insights from small molecule modules
Similarly, a small molecule module was defined as a group of (≥5) small molecules that were associated with the same miRNA and was coded with the name of the central miRNA. We further analyzed the compounds in these modules based on similar structures, common drug targets and drug interactions. Firstly, the two-dimensional Tanimoto chemical similarity score was used to calculate the structural similarity between two compounds within a module (see details in Materials and Methods). A Tanimoto score of 0.6 was selected as the threshold33. Secondly, the rate of sharing drug targets was also evaluated using the “Meet/Min” score and the known drug-target relationships were obtained from the DrugBank database34. Thirdly, we considered drug interactions that were extracted from a popular and comprehensive online source of drug information (http://www.drugs.com/).
We determined that most (64.546%) of the small molecule modules involved drug pairs with at least one of the aforementioned similarities (Supplementary Tables S3 and S4). For example, in the small molecule module miR-1207-3p of skin cancer (Figure 5), fluphenazine, thioridazine and prochlorperazine are chemically similar based on the Tanimoto scores. Meanwhile, the three small molecules also shared many common drug targets. These findings were consistent with those of previous studies indicating that chemically similar drugs are likely to have the same targets35. Moreover, a total of 21 known drug interactions, such as sirolimus interacting with rifabutin, were detected in this module. Coadministration of these two small molecules may reduce the plasma concentrations and pharmacologic effects of sirolimus. In the small molecule module miR-768-3p of the bone marrow cancer (Figure 5), camptothecin and irinotecan had similar chemical structures and shared drug targets. Irinotecan is a topoisomerase 1 inhibitor and an antineoplastic agent that has been approved by the U.S. Food and Drug Administration (FDA) for the treatment of cancer34. Camptothecin is a DNA enzyme topoisomerase I inhibitor and an antineoplastic agent in preliminary clinical trials34. The small molecule module miR-139-5p of thyroid cancer (Figure 5) revealed that lomustine and streptozocin have similar chemical structures. Both of them are the FDA-approved drugs against tumors34. It indicated that our method can be used to identify functionally similar small molecules within the same module.
The small molecule modules could be used to predict drug candidates. For example, streptozocin, which can decrease tumor size36, has been used as an FDA-approved antineoplastic agent34. In the small molecule module miR-139-5p of thyroid cancer (Figure 5), 2-deoxy-D-glucose (2DOG) is chemical structure similar with streptozocin. 2DOG is a glycolytic inhibitor that has not be previously included in the DrugBank database. We predicted that 2DOG might exhibit potential anticancer activity. Previous studies have shown that 2DOG can enhance autophagy, which is involved in cancer37. Accelerated glucose uptake for anaerobic glycolysis is an important metabolic change in tumor cells. 2DOG inhibits effectively glucose metabolism and causes tumor cell death38. Some studies have also suggested that the combination of 2DOG and other specific chemotherapeutic agents is more effective against cancer cells than single treatments39. Collectively, 2DOG might be a new drug candidate for cancer therapy.
In addition, these modules were also useful for drug repositioning. It is known that inflammatory cells provide a favorable microenvironment for tumor growth and development. Therefore, anti-inflammatory therapy exhibits beneficial effects in cancer treatments40. Rimexolone is a known anti-inflammatory agent34. Estradiol (E2), which belongs to the small molecule module miR-203 of bone marrow cancer (Figure 5), is an estrogen, anticholesteremic agent and anti-menopausal agent according to the DrugBank database34. E2 is chemically similar to rimexolone. Joshua et al. have reported that E2 attenuates inflammation in the brain of adult male rats with severe burn injury41. These results suggested that E2 might have anti-inflammatory activity.
Discussion
We developed a general procedure to predict the associations between small molecules and miRNAs based on transcriptional responses. In previous studies, gene signatures that are extracted from gene expression profiles have been used to identify associations among drugs and diseases. Here, we predicted the small molecule-miRNA associations based on the transcriptional responses following drug treatment and miRNA regulation.
Using network theory, we analyzed the constructed SMirNs for 17 different cancers. TSA, miR-19a, miR-19b and miR-23b as well as their associations appeared most frequently in the SMirNs. After literature review and functional analysis, we predicted that miR-23b might be a new potential drug or drug target for cancer therapy because of its ability to regulate target genes ALDH1A1 and ALDH1A2. In addition, TSA might affect the development of a broad spectrum of tumors by targeting the MAPK and mTOR signaling pathways.
Furthermore, we grouped compounds and miRNAs into modules separately. In almost all of the miRNA modules, we identified miRNAs with similar biological functions by assessing whether they were members of the same miRNA family, sharing common targets or having GO functional similarities. Most of the compound modules included small molecules with similar structures, common drug targets or drug interactions. As we known, the compounds with similar structures exhibit similar physical and biological properties. Therefore, the compound modules can be used to predict drug candidates and drug repositioning. We identified 2DOG as a potential new drug candidate for cancer therapy. Based on our module, we also proposed that E2, which is a known anticholesteremic and anti-menopausal agent, might have anti-inflammatory activity.
To our knowledge, this is the first study to systematically identify and analyze the relationships between bioactive small molecules and miRNAs in a broad spectrum of different human cancers. Our approach does not require the prior structure information about compound and miRNA, which may be unknown or difficult to obtain, except gene expression profiles of compound treatment and miRNA targets. However, because there are not many researches describing the relationships between small molecules and miRNAs, the validation of predicted associations is still a challenge. After a comprehensive literature-based validation, our predictions significantly appeared in the current experimental knowledge. In addition, lack of transcriptional response data following genome-wide miRNA perturbations limits the reliability of our results. With the development of high-throughput detection techniques and the advance of miRNA research, our approach can directly extend to process the gene expression profiles following drug treatment and miRNA perturbation and to identify precise associations between small molecules and miRNAs. Thus, in our future study, we will further improve our approach and experimentally validate the predicted small molecule-miRNA associations for a specific cancer.
Methods
Datasets
Differentially expressed genes in human cancers
A total of 136 differentially expressed gene sets spanning 23 human cancer tissue types were downloaded from the “cancer versus normal” type of analysis in the ONCOMINE database16. For one cancer, multiple gene sets were integrated as follows: a gene that was over/under-expressed in at least two gene sets was considered as an over/under-expressed gene in this cancer. Thus, we obtained the over- and under-expressed gene sets of 23 cancers. Finally, gene symbols were converted into the probe sets on the Affymetrix Human Genome U133A (HG-U133A) Array for further analysis.
The miRNAs and their targets
Previous studies have demonstrated that a combination of multiple algorithms would increase the predictive accuracy of miRNA targets42,43. Thus, the miRNA targets in this study were gathered from seven algorithms: RNAhybrid44, DIANA-microT45, RNA2246, miRBase Targets47, miRanda48, PicTar49 and TargetScan50. We extracted miRNA-target pairs that were predicted by at least two algorithms. We obtained 289,468 miRNA-target gene pairs for 775 miRNAs. Gene symbols were also converted into probe sets on the HG-U133A Array for further analysis.
Expression profiles in the Connectivity Map database
Small-molecule perturbed genome-wide transcriptional response data were downloaded from the Connectivity Map (cMap, build 02)15. These data comprise more than 7,000 gene expression profiles from human cultured cell lines that are treated with 1,309 bioactive small chemical molecules at varying concentrations, representing 6,100 individual instances. Each instance denotes a treatment and control pair for one small molecule. The list of probe sets is ranked according to the extent of differential expression between the treatment and control cells. An instance has some attributes such as perturbagen name, concentration, cell line and batch etc. Therefore, several instances may be derived from the treatment of cultured cells with a small molecule.
Identifying cancer-related miRNAs
Enrichment analysis is usually used to assess the functional associations of an interesting gene set with GO, KEGG and so on51,52,53. Here, the differentially expressed gene set of one cancer was considered as a functional category, which was similar with the priori defined gene sets of the Molecular Signatures Database (MSigDB), such as CGP (chemical and genetic perturbations) subcollection54. If the target genes of one miRNA were over-represented in the differentially expressed gene set of one cancer, we hypothesized that this miRNA was the cancer-related miRNA (CRM). The significance of the over-representation was evaluated based on the P-value of the hypergeometric test:

where M was the total number of target genes of all miRNAs; N was the number of all differentially expressed miRNA target genes; m was the number of target genes of one miRNA; n was the number of differentially expressed target genes of this miRNA. Using an adjusted P-value (FDR) cutoff of 0.05, we identified CRMs for each of the 23 cancers. The intersections of the target genes of the CRMs and the over/under-expressed genes in the cancer were denoted as the miRNA over/under-expressed target genes in the cancer.
Identifying the small molecules associated with CRMs
The cMap database establishes biological connections among diseases, drugs and genes. In this study, we extended the application of cMap and developed a novel computational framework to identify relationships between small molecules and miRNAs in human cancers.
Determining the enriched gene function module
The results of the global similarity search in cMap were dependent on the probe selection and the number of input probes. To overcome this limitation, Li et al. have proposed a module-based chemical functional similarity search approach55. In this study, we extended the approach to identify the links between small molecules and CRMs. We used GO to group genes into different functional classes. Each GO term was called a GO module (GOM), which represented a set of genes that performed a specific biological function. For differentially expressed targets of a CRM in one cancer, GO enrichment analysis was performed using the hypergeometric test. The P-value denoted whether the differentially expressed targets were enriched in certain GOM and was calculated as follows:

where M was the number of differentially expressed targets of all CRMs for this cancer; N was the number of targets in the GOM of interest; m was the number of differentially expressed targets of the miRNA; n was the number of genes that were annotated to this GOM out of the above m targets. GOMs with P≤0.05 were selected (Biological Process category).
Evaluating the similarity of differentially expressed genes between CRMs regulations and small molecule perturbation
For each significant GOM of a CRM, differentially expressed targets in this GOM were partitioned into two groups based on whether they were over- or under-expressed. The Kolmogorov-Smirnov (KS) scores were calculated for over-expressed (KSup) and under-expressed (KSdown) target genes, respectively:



where t was the number of target genes in the over- or under-expressed target gene set, j represented the jth target gene based on the rank of differential expression; N was the number of all genes in the HG-U133A array. The N genes were ranked according to the extent of differential expression for an instance, the jth element V(j) of a vector V was the position of the jth target gene in the ordered list of whole genes. Then similarity score (S score) was calculated by combining KSup and KSdown, which showed the extent of expression pattern similarity between an expression profile treated with a small molecule at a specific concentration and the miRNA targets in this GOM. S was equal to 0 when KSup and KSdown had the same sign and was equal to KSup-KSdown otherwise.
Identifying the links between the CRMs and small molecules
For each significant GOM, the set of t instances from a perturbagen and the list of all instances were ranked in descending order of the S score. The jth element V(j) of the vector V denoted the position of the jth instance in the set of t instances of this perturbagen in the ordered list of all n instances (j = 1,2,…,t). The enrichment score was defined as follows:



We randomly selected t instances from all n instances and calculated the enrichment score, repeating this procedure for 1,000 times. Using this process, 1,000 fake enrichment scores were obtained. The P-value was the fraction that the absolute value of the fake enrichment score was larger than the absolute value of the real enrichment score. The P-values represented the statistical significance of the small molecule- miRNA associations. For each CRM, if one small molecule with P≤0.05 appeared in at least 80% GOMs, the small molecule was considered to link with the CRM. If no small molecule satisfied the above conditions, the CRM was removed.
Meet/Min score
For one miRNA pair, i and j, their differentially expressed target gene sets were targets(i) and targets(j), respectively. The “Meet/Min” score was calculated as follows:

Namely, the number of the common differentially expressed targets of the two miRNAs was divided by the size of the smaller target set.
Chemical structure similarity calculation
To calculate the structural similarity of small molecules, the MOL file of each small molecule was downloaded from the PubChem compound database56. The small molecules lacking structural information were filtered out. The two-dimensional Tanimoto chemical similarity score was determined using the Small Molecule Subgraph Detector software57.
References
Bartel, D. P. MicroRNAs: genomics, biogenesis, mechanism and function. Cell. 116, 281–297 (2004).
Esquela-Kerscher, A. & Slack, F. J. Oncomirs - microRNAs with a role in cancer. Nat Rev Cancer. 6, 259–269 (2006).
Sevignani, C., Calin, G. A., Siracusa, L. D. & Croce, C. M. Mammalian microRNAs: a small world for fine-tuning gene expression. Mamm Genome. 17, 189–202 (2006).
Yang, Q., Qiu, C., Yang, J., Wu, Q. & Cui, Q. miREnvironment database: providing a bridge for microRNAs, environmental factors and phenotypes. Bioinformatics. 27, 3329–3330 (2011).
Zhang, S., Chen, L., Jung, E. J. & Calin, G. A. Targeting microRNAs with small molecules: from dream to reality. Clin Pharmacol Ther. 87, 754–758 (2010).
Spizzo, R., Rushworth, D., Guerrero, M. & Calin, G. A. RNA inhibition, microRNAs and new therapeutic agents for cancer treatment. Clin Lymphoma Myeloma. 9 Suppl 3, S313–318 (2009).
Pan, Q., Tilanus, H. W., Janssen, H. L. & van der Laan, L. J. Prospects of RNAi and microRNA-based therapies for hepatitis C. .Expert Opin Biol Ther. 9, 713–724 (2009).
Lanford, R. E. et al. Therapeutic silencing of microRNA-122 in primates with chronic hepatitis C virus infection. Science. 327, 198–201 (2010).
Gumireddy, K. et al. Small-molecule inhibitors of microrna miR-21 function. Angew Chem Int Ed Engl. 47, 7482–7484 (2008).
Blower, P. E. et al. MicroRNAs modulate the chemosensitivity of tumor cells. Mol Cancer Ther. 7, 1–9 (2008).
Gusev, Y., Schmittgen, T. D., Lerner, M., Postier, R. & Brackett, D. Computational analysis of biological functions and pathways collectively targeted by co-expressed microRNAs in cancer. BMC Bioinformatics. 8 Suppl 7, S16 (2007).
Bertino, J. R., Banerjee, D. & Mishra, P. J. Pharmacogenomics of microRNA: a miRSNP towards individualized therapy. Pharmacogenomics. 8, 1625–1627 (2007).
Yang, H. et al. MicroRNA expression profiling in human ovarian cancer: miR-214 induces cell survival and cisplatin resistance by targeting PTEN. Cancer Res. 68, 425–433 (2008).
Iorio, F. et al. Discovery of drug mode of action and drug repositioning from transcriptional responses. Proc Natl Acad Sci U S A. 107, 14621–14626 (2010).
Lamb, J. et al. The Connectivity Map: using gene-expression signatures to connect small molecules, genes and disease. Science. 313, 1929–1935 (2006).
Rhodes, D. R. et al. Oncomine 3.0: genes, pathways and networks in a collection of 18,000 cancer gene expression profiles. Neoplasia. 9, 166–180 (2007).
Vanhaecke, T., Papeleu, P., Elaut, G. & Rogiers, V. Trichostatin A-like hydroxamate histone deacetylase inhibitors as therapeutic agents: toxicological point of view. Curr Med Chem. 11, 1629–1643 (2004).
Shankar, S. & Srivastava, R. K. Histone deacetylase inhibitors: mechanisms and clinical significance in cancer: HDAC inhibitor-induced apoptosis. Adv Exp Med Biol. 615, 261–298 (2008).
Zhang, Z. W., An, Y. & Teng, C. B. [The roles of miR-17-92 cluster in mammal development and tumorigenesis]. Yi Chuan. 31, 1094–1100 (2009).
Gottardo, F. et al. Micro-RNA profiling in kidney and bladder cancers. Urol Oncol. 25, 387–392 (2007).
Wang, T. et al. A micro-RNA signature associated with race, tumor size and target gene activity in human uterine leiomyomas. Genes Chromosomes Cancer. 46, 336–347 (2007).
Karabulut, B. et al. Enhancing cytotoxic and apoptotic effect in OVCAR-3 and MDAH-2774 cells with all-trans retinoic acid and zoledronic acid: a paradigm of synergistic molecular targeting treatment for ovarian cancer. J Exp Clin Cancer Res. 29, 102 (2010).
Sos, M. L. et al. Identifying genotype-dependent efficacy of single and combined PI3K- and MAPK-pathway inhibition in cancer. Proc Natl Acad Sci U S A. 106, 18351–18356 (2009).
Yang, Q. & Guan, K. L. Expanding mTOR signaling. Cell Res. 17, 666–681 (2007).
Brest, P. et al. MiR-129-5p is required for histone deacetylase inhibitor-induced cell death in thyroid cancer cells. Endocr Relat Cancer. 18, 711–719 (2011).
Chen, Y. et al. Methyl-CpG binding protein MBD2 is implicated in methylation-mediated suppression of miR-373 in hilar cholangiocarcinoma. Oncol Rep. 25, 443–451 (2011).
Zhang, S., Cai, X., Huang, F., Zhong, W. & Yu, Z. Effect of trichostatin a on viability and microRNA expression in human pancreatic cancer cell line BxPC-3. Exp Oncol. 30, 265–268 (2008).
Li, W. et al. Integrative analysis of many weighted co-expression networks using tensor computation. PLoS Comput Biol. 7, e1001106 (2011).
Jayaswal, V., Lutherborrow, M., Ma, D. D. & Yang, Y. H. Identification of microRNA-mRNA modules using microarray data. BMC Genomics. 12, 138 (2011).
Zheng, S., Tansey, W. P., Hiebert, S. W. & Zhao, Z. Integrative network analysis identifies key genes and pathways in the progression of hepatitis C virus induced hepatocellular carcinoma. BMC Med Genomics. 4, 62 (2011).
Marras, E., Travaglione, A., Chaurasia, G., Futschik, M. & Capobianco, E. Inferring modules from human protein interactome classes. BMC Syst Biol. 4, 102 (2010).
Lin, D. An information-theoretic definition of similarity. Proceeding of the 15th International Conference on Machine Learning, San Francisco. (1998).
Campillos, M., Kuhn, M., Gavin, A. C., Jensen, L. J. & Bork, P. Drug target identification using side-effect similarity. Science. 321, 263–266 (2008).
Wishart, D. S. et al. DrugBank: a knowledgebase for drugs, drug actions and drug targets. Nucleic Acids Res. 36, D901–906 (2008).
Martin, Y. C., Kofron, J. L. & Traphagen, L. M. Do structurally similar molecules have similar biological activity? J Med Chem. 45, 4350–4358 (2002).
Brentjens, R. & Saltz, L. Islet cell tumors of the pancreas: the medical oncologist's perspective. Surg Clin North Am. 81, 527–542 (2001).
Chen, N. & Karantza, V. Autophagy as a therapeutic target in cancer. Cancer Biol Ther. 11, 157–168 (2011).
Aft, R. L., Zhang, F. W. & Gius, D. Evaluation of 2-deoxy-D-glucose as a chemotherapeutic agent: mechanism of cell death. Br J Cancer. 87, 805–812 (2002).
Zhang, F. & Aft, R. L. Chemosensitizing and cytotoxic effects of 2-deoxy-D-glucose on breast cancer cells. J Cancer Res Ther. 5 Suppl 1, S41–43 (2009).
Coussens, L. M. & Werb, Z. Inflammation and cancer. Nature. 420, 860–867 (2002).
Gatson, J. W. et al. Estrogen treatment following severe burn injury reduces brain inflammation and apoptotic signaling. J Neuroinflammation. 6, 30 (2009).
Jayaswal, V., Lutherborrow, M., Ma, D. D. & Yang, Y. H. Identification of microRNA-mRNA Modules using Microarray Data. BMC Genomics. 12, 138 (2011).
Li, X. et al. Dissection of human MiRNA regulatory influence to subpathway. Brief Bioinform. (2011).
Rehmsmeier, M., Steffen, P., Hochsmann, M. & Giegerich, R. Fast and effective prediction of microRNA/target duplexes. RNA. 10, 1507–1517 (2004).
Kiriakidou, M. et al. A combined computational-experimental approach predicts human microRNA targets. Genes Dev. 18, 1165–1178 (2004).
Miranda, K. C. et al. A pattern-based method for the identification of MicroRNA binding sites and their corresponding heteroduplexes. Cell. 126, 1203–1217 (2006).
Griffiths-Jones, S., Saini, H. K., van Dongen, S. & Enright, A. J. miRBase: tools for microRNA genomics. Nucleic Acids Res. 36, D154–158 (2008).
Enright, A. J. et al. MicroRNA targets in Drosophila. Genome Biol. 5, R1 (2003).
Krek, A. et al. Combinatorial microRNA target predictions. Nat Genet. 37, 495–500 (2005).
Lewis, B. P., Shih, I. H., Jones-Rhoades, M. W., Bartel, D. P. & Burge, C. B. Prediction of mammalian microRNA targets. Cell. 115, 787–798 (2003).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc. 4, 44–57 (2009).
Zhou, X. & Su, Z. EasyGO: Gene Ontology-based annotation and functional enrichment analysis tool for agronomical species. BMC Genomics. 8, 246 (2007).
Li, C. et al. SubpathwayMiner: a software package for flexible identification of pathways. Nucleic Acids Res. 37, e131 (2009).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A. 102, 15545–15550 (2005).
Li, Y. et al. Gene expression module-based chemical function similarity search. Nucleic Acids Res. 36, e137 (2008).
Wang, Y. et al. PubChem: a public information system for analyzing bioactivities of small molecules. Nucleic Acids Res. 37, W623–633 (2009).
Rahman, S. A., Bashton, M., Holliday, G. L., Schrader, R. & Thornton, J. M. Small Molecule Subgraph Detector (SMSD) toolkit. J Cheminform. 1, 12 (2009).
Acknowledgements
The authors would like to acknowledge the support of the Funds for Creative Research Groups of The National Natural Science Foundation of China [grant numbers 81121003], the National Natural Science Foundation of China [grant numbers 30800268, 61073136, 30900837, 91029717] and the National High Tech Development Project of China, the 863 Program [grant number 2007AA02Z329].
Author information
Authors and Affiliations
Contributions
X.L and W.J designed the study. M.Z.L, X.W.C and Y.J collected data. W.J, X.W.C and M.Z.L conceived and developed the computational model and analyzed the network. W.L, B.F.L and L.H.W contributed analysis tools. W.J, X.W.C and M.Z.L wrote the article. F.L.M, X.Y.L and X.J.C performed literature-based validation. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareALike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Jiang, W., Chen, X., Liao, M. et al. Identification of links between small molecules and miRNAs in human cancers based on transcriptional responses. Sci Rep 2, 282 (2012). https://doi.org/10.1038/srep00282
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep00282
This article is cited by
-
Noncoding RNA therapeutics — challenges and potential solutions
Nature Reviews Drug Discovery (2021)
-
A review of computational drug repositioning: strategies, approaches, opportunities, challenges, and directions
Journal of Cheminformatics (2020)
-
Systems Pharmacogenomic Landscape of Drug Similarities from LINCS data: Drug Association Networks
Scientific Reports (2019)
-
Identification and profiling of microRNAs expressed in oral buccal mucosa squamous cell carcinoma of Chinese hamster
Scientific Reports (2019)
-
Inferring potential small molecule–miRNA association based on triple layer heterogeneous network
Journal of Cheminformatics (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.