Abstract
Vaccine development against hepatitis C virus (HCV) is hindered by poor understanding of factors defining cross-immunoreactivity among heterogeneous epitopes. Using synthetic peptides and mouse immunization as a model, we conducted a quantitative analysis of cross-immunoreactivity among variants of the HCV hypervariable region 1 (HVR1). Analysis of 26,883 immunological reactions among pairs of peptides showed that the distribution of cross-immunoreactivity among HVR1 variants was skewed, with antibodies against a few variants reacting with all tested peptides. The HVR1 cross-immunoreactivity was accurately modeled based on amino acid sequence alone. The tested peptides were mapped in the HVR1 sequence space, which was visualized as a network of 11,319 sequences. The HVR1 variants with a greater network centrality showed a broader cross-immunoreactivity. The entire sequence space is explored by each HCV genotype and subtype. These findings indicate that HVR1 antigenic diversity is extensively convergent and effectively limited, suggesting significant implications for vaccine development.
Similar content being viewed by others
Introduction
Hepatitis C virus (HCV) is a single-stranded RNA virus belonging to the Flaviviridae family1. HCV infects 3.0% of the world's population and is a major cause of liver disease worldwide2. HCV infection progresses to chronicity in 70%–85% of infected adults3. An estimated 476,000 deaths per year are attributed to hepatitis C2. However, there is no vaccine against HCV and current anti-viral therapy is effective in 50%–60% of patients4. HCV is genetically very heterogeneous and classified into 6 genotypes and numerous subgenotypes5.
Vaccines are among the most efficacious means to control infectious diseases. However, the development of vaccines against highly heterogeneous viruses such as HCV and human immunodeficiency virus (HIV) is considerably hampered by variant-specific neutralizing immune responses. These viruses have seemingly unlimited capacity to rapidly mutate and escape from immune neutralization, thus presenting a major obstacle for formulating broadly protective vaccines6,7. Considering that an estimated 130 million and 33 million individuals are infected worldwide with HCV and HIV, respectively2,7 and that each infected host harbours a large variety of viral variants, the number of viral variants circulating in the world is immense. Developing vaccines against such broad range of viral variants seems a daunting task.
Classical approaches to vaccine development are yet to produce broadly protective vaccines against HCV and HIV6,8. Novel vaccine strategies recently developed to cope with viral antigenic diversity focus either on using epitopes with limited heterogeneity9, generating a concoction of heterogeneous epitopes10,11 or mimotopes12,13, or predicting consensus sequences, center of tree variants or phylogenetic ancestors14,15. These strategies are based on specific assumptions regarding properties of highly heterogeneous epitopes, viz., that immunological specificity is strongly linked to the epitope primary structure, with cross-immunoreactivity (CR) declining with increasing genetic difference between epitopes and that the viral sequence space is shaped by diversifying evolution resulting from an “arms race”16. However, the conditional relevance of these assumptions has not been systematically corroborated.
The most important HCV neutralizing epitope has been mapped in the hypervariable region 1 (HVR1), located at amino acid (aa) positions 384–410 in the structural protein E2. HVR1 sequence variation correlates with neutralization escape and is associated with viral persistence during chronic infection17,18,19,20,21,22. Although some neutralizing epitopes have been discovered in conserved regions of HCV structural proteins23, the variant-specificity of humoral protective responses24,25 points to the essential role played by the variable epitopes in controlling HCV infections.
In the present work, a quantitative analysis of the HVR1 CR, modeled using synthetic peptides and mouse immunization, in conjunction with a network analysis of the HVR1 sequence space showed significant immunological and structural HVR1 convergence. The findings suggest tractability of the HVR1immunological specificity and offer a novel framework for HCV vaccine development, which is applicable to other heterogeneous viruses.
Results
Convergence of HVR1 CR
CR analysis among HVR1 variants using human serum specimens is complicated by the multi-specificity of the humoral response against numerous HCV variants in a given infected host. To overcome this problem, many groups have used mice to study the immune response to the HVR1 epitope26,27,28. Several studies have shown the specific reactivity of HVR1 peptides with sera of infected individuals26,27,29,30,31,32,33. We modeled HVR1 CR using serum specimens from mice immunized with 103 synthetic peptides representing different HVR1 variants (referred to as immunogens). A set of 261 HVR1 peptides was used as antigens in an enzyme immunoassay, of which 102 sequences were also used as immunogens. All HVR1 variants were randomly selected from major branches of a phylogenetic tree constructed using sequences available in GenBank. A total of 165,120 immunoassay reactions were evaluated. Among 26,883 unique immunogen-antigen pairs, 5039 (18.7%) were cross-immunoreactive (Fig. S1 and Table S1).
Many HVR1 variants derived from different genotypes were found to be cross-immunoreactive (Fig. S2). The CR frequency was 16.2% and 20.6% between inter- and intra-genotypic pairs, respectively. This finding points to the independent occurrence of the identical immunological specificity among phylogenetically unrelated variants, indicating the existence of immunological homoplasy. There was an inverse relationship between the Hamming distance and CR frequency among pairs of HVR1 variants (Fig. 1A). Thus, the increase in genetic distance between viral variants was associated with a decline in the capacity of antibody raised against one variant to recognize another variant. However, this distance plot (Fig. 1A) shows two intriguing findings. First, 12.4% of variants differing from each other at 21–25 positions were cross-immunoreactive. Such an extreme difference in aa composition is another strong indication of the independent origin of identical immunological specificities, which suggests convergence of the HVR1 immunological properties. Second, 18.4% of all tested variants did not show self-immunoreactivity. Although most of these peptides showed CR with others (ranging from 0% to 34.9%), the group of immunogens that did not self-react showed a lower average CR level than the group of immunogens that self-reacted (MRPP test34; p = 0.0001).
A) Relationship between CR and genetic distance. The Hamming distance is the number of different aa positions. B) CR distribution. Percentage of peptides found in each CR bin. The numbers below bins show the upper limit of the CR values. C) CR network. Each node is an HVR1 sequence and there is a link between two nodes if the reaction between the two peptides was positive.
The last observation suggests separation between immunogenic and antigenic HVR1 specificities under the experimental conditions. This observation was further extended by analysis of the range of antibody and peptide CR (Fig. 1B). The frequency distributions for immunogens (median 8.8%) and antigens (median 18.4%) were distinctly different (Kolmogorov-Smirnov Test, p < 0.0001); 53.4% of the immunogens reacted with <10% of antigens and only 2.9% of immunogens showed broad immunoreactivity with >90% of peptides, the maximum being 97.7% for HVR1 variant P240 (genotype 1a). These observations show a highly non-uniform CR distribution among HVR1 immunogens. In contrast, the frequency distribution of antigen CR was centered at the average value. None of the antigens reacted with >52.5% of the immunogens (Fig. 1B). This difference between the antigen and immunogen distributions was also reflected in the identification of 2 minimal sets of peptides. One set (n = 3, peptides P032, P240 and P247) collectively elicited antibodies immunoreacting with all 261 antigens. The other set (n = 14) immunoreacted with antibodies against 101 out of 103 immunogens. The observed separation between antigenic and immunogenic properties may be explained by differences in conformational states of the conjugated peptides used for immunization and free peptides used for antibody detection, indicating conformation dependence of the HVR1 antigenic epitopes.
To further evaluate HVR1 CR, we constructed a network where each node is an HVR1 sequence and there is a link between two nodes if the reaction between the two corresponding peptides was positive (Fig. 1C). All HVR1 sequences were connected in a single giant component. Topological analysis showed that this network is a “small world”35 and has a small average shortest path between every pair of reachable vertices (2.04 steps) and a small diameter (5 steps), the maximum shortest path between two vertices. The network also lacks identifiable modules or communities, suggesting an overall homogeneity of the HVR1 immune recognition. These findings emphasize the pervasive incidence of broad CR among HVR1 variants. To reduce bias that may be introduced owing to the asymmetry of this network composed of a different set of immunogens and antigens, we extracted the symmetric subnetwork consisting only of the 102 peptides which were both immunogens and antigens. For this subnetwork, the average CR was 20% and average shortest path distance and diameter were 2.01 and 4, respectively, thus being similar to the total network.
It is important to note that there is no correlation between the CR of a peptide as immunogen and as antigen (Fig. S3). Moreover, only 11% of all links in the subnetwork were bidirectional, indicating that the majority of peptides used in this study had different immunogenic and antigenic specificity. Nonetheless, it was still possible to follow a path from a given variant to almost any other in a few steps, with 89.3% of all possible pairs being reachable. We also calculated the clustering coefficient36, defined as the ratio between the number of links of the neighbors of a node and the number of links they could have if they were a fully connected subgraph or clique, was calculated for each node. The subnetwork had a high local structure with an average clustering coefficient of 0.6658, such a high value being related to the large number of cliques (n = 2,979) in the network. Each clique contained 3–17 nodes (average of 12.72). A clique in the network can be viewed as a set of variants sharing a single immunological specificity.
Convergence of HVR1 sequence space
To study the structure of HVR1 sequence space, we compiled 11,319 HVR1 sequences from 3,172 patients. The full dataset contained sequences of two classes: (i) multiple clonal variants obtained from a single time-point from 176 patients and from 2–14 time-points from 29 patients; (ii) a single consensus sequence from 2,967 independent patients. Of all sequences, 65.4% were obtained from the Los Alamos HCV Sequence Database37 and 34.6% were clonal variants generated in our laboratory using end-point limiting-dilution PCR38. The final dataset included 4757 unique aa sequences: 3432 sequences of genotype 1, 727 of genotype 2, 235 of genotype 3, 103 of genotype 4, 75 of genotype 5, 43 of genotype 6 and 142 of unknown genotypes. When considering only epidemiologically unrelated sequences, the percentage of sequence variation due to differences among genotypes is 4.7 times higher at the nucleotide level (average 20.76%) than at the aa level (4.39%). Even though these average differences among genotypes are statistically significant, the maximum likelihood trees of nucleotide or aa sequences do not show clear genotype-specific clusters (Fig. S4 and S5). The lack of genotype-specific clustering is characteristic only for HVR1, since inclusion of the flanking regions from E1 and/or E2 regions completely restores phylogenetic relationships among HCV sequences.
Taking into consideration a potential convergence among HVR1 aa sequences and the dimensionality problem (see Methods for details), we have studied different ways of portraying this sequence space, giving preference to approaches that could accurately model distances among closely related sequences. The sequence space was visualized using a Pathfinder network (PFNET) based on the entropy-weighted Hamming distances between 4,757 unique HVR1 aa sequences (Fig. 2A). Each unique aa sequence is represented with a node in the PFNET and each link connects nodes of the highest similarity.
A) Location of HVR1 variants from 6 HCV genotypes in the sequence space modeled with PFNET. Each genotype is shown in a different color. B) Exploration of the PFNET by different HVR1 samples. The acute group consists of 34 single time-point samples, the chronic group has 90 single time-point samples, the follow-up group has 29 clusters and the genotype group consisted of 8 clusters. C) Exploration of sequence space by HVR1 variants from 5 follow-up patients. Patient 1001 to 1004 were described in50. Patient 7001 was described in33.
The low level of differentiation among genotypes at the aa level is also evident in the lack of genotype-specific clusters in the PFNET model of sequence space (Fig. 2A), which shows that HVR1 sequences from different HCV genotypes explore the same space at the aa level. Only 3.15% of the variability in path distances among epidemiologically unrelated sequences can be attributed to genotype differences. A similar observation has been made for the two most prevalent subtypes in the PFNET, 1a and 1b. These findings suggest a significant HVR1 convergence among HCV genotypes, with each representing basically the entire HVR1 sequence space. Additionally, we measured the shortest path between any two sequences over the entire PFNET. The diameter and the average shortest path were calculated for populations of intra-host HVR1 variants from (i) single time-points at the acute stage, (ii) single time-points during chronic infection, (iii) multiple time-points follow-up and (iv) variants of different genotypes. Both measures reflect the breadth of distribution of variants in the PFNET. Although there was, in general, a significant difference in both the diameter and average distance among all four sets (MRPP, p = 0.0001), an overlap in these values was observed at different evolutionary levels (Fig. 2B). All pair-wise comparisons of the diameter and average distance between sets were significantly different (Table S2), except between the chronic and follow-up sets (MRPP, p = 0.7305).
Although most of the single time-point samples had a small diameter, some samples contained variants spreading in PFNET far from each other and had a diameter greater than follow-up patients. Another observation was that variants from some chronic cases were found so scattered in the PFNET that they had diameter and average distance overlapping with the measures for some HCV genotypes, thus indicating that HVR1 variants generated during intra-host evolution in some patients may explore the HVR1 space at a scale similar to subgenotypes and genotypes. Fig. 2C shows the range of exploration of the PFNET sequence space by variants from five patients who were followed-up for 8.8 to 26 years. HVR1 variants from patient 1001 (genotype 1a) are represented with 540 sequences collected over a period of 16.0 years at 13 time-points; from patient 1002 (1a) with 497 sequences collected over a period of 18.2 years at 14 time-points; from patient 1003 (1b) with 180 sequences collected over a period of 8.8 years at 5 time-points; from patient 1004 (1a) with 574 sequences collected over a period of 16.0 years at 10 time-points; and from patient 7001 (1a) with 52 sequences collected over a period of 26.0 years at 8 time-points. The variants from each of two patients, 1001 and 7001, were found in different regions of the PFNET. Collectively, all these observations suggest that the HVR1 space is small; it is entirely scanned by each genotype and can be widely traversed by intra-host HVR1 variants. However, although the space is small for the independent accommodation of genotypes and subgenotypes, it is sufficiently large to provide a significant flexibility for the intra-host HVR1 evolution that would facilitate HCV escape from the neutralizing immune responses.
CR Distribution in the HVR1 sequence space
All peptides used in this study were mapped in the PFNET to show the uniform distribution of the tested variants in the modeled sequence space (Fig. 3A). The cross-immunoreactive pairs of sequences were linked to visualize CR distribution in this network. Although this visualization showed that CR occurred among widely scattered peptides rather than being confined to some particular regions of the network, the high density of links precluded the detection of any meaningful structure in this distribution. Further analysis of topological parameters of the PFNET showed that the closeness centrality, measured as the average shortest path from a given node to all other nodes in the network, is significantly associated with the CR range of the peptides (Fig. 3B). The average CR of immunogens with the highest centrality is 3.37 times greater than for immunogens with the lowest centrality (MRPP, p = 0.0099). A similar, albeit less distinct, trend was observed for the antigens, with an average CR that was 1.64 times greater for the most central than for the least central (MRPP, p = 0.0099). This observation suggests that the HVR1 sequence space as modeled with PFNET is associated with CR and, therefore, links HCV evolution with specific immunological recognition of HVR1 epitopes.
A) PFNET map of the experimentally tested HVR1 variants and their CR. The sequences used as both immunogen and antigen are shown in red (n = 103), the sequences used only as antigen are shows in blue (n = 261) and the sequence used only as immunogen is shown in purple (n = 1). The light green lines link cross-immunoreactive variants. B) HVR1 CR peptides according to their PFNET centrality. All nodes were divided into 5 bins starting from most peripheral (left) to most central (right). The numbers below bins show the upper limit of the closeness centrality values. The standard error of the mean is shown as black bars.
Predictive model of HVR1 CR
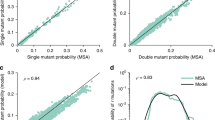
The significant functional and structural convergence observed in this study indicates that the same HVR1 properties evolved frequently and independently in the occupied sequence space. Such regular occurrence of the limited number of immunological specificities renders them tractable and highly amenable to predictive modeling. Therefore, we generated a classification model in the form of a decision tree associating HVR1 sequence and CR. This simple model predicted CR based on sequence of the immunogen and antigen pairs (Fig. S6). A 10-fold cross-validation showed that the average testing accuracy was 92.5% (S.D. 0.4%), with the average sensitivity and specificity being 99.7% (S.D. 0.2%) and 85.3% (S.D. 0.7%), respectively. Using this model, we predicted global cross-immunorectivity among all possible pairs of sequences in the PFNET (n = 2.26E+07). A strong inverse relationship between the distance of two peptides and the predicted CR frequency was found (Fig. 4A). Likewise, the distributions of the predicted and observed cross-immunorectivity are similar both for antigens and immunogens (Fig. 4B). Although most of the immunogens had a low predicted breadth of immunoreactivity, we found 73 sequences (1.53% of all analyzed) with predicted CR of > 99% in different parts of the PFNET (Fig. S7 and S8). These variants may be viewed as highly cross-immunoreactive HVR1 mimotopes, thereby supporting vaccine development strategies that rely on discovery of broadly immunoreactive epitopes12,13,14.
A) Relationship between predicted CR and hamming distance. B) Predicted global CR distribution among 4,757 HVR1 variants. The average global CR of the immunogenes is 18.1% (S.E. = 0.3) and of antigens is 18.1% (S.E. = 0.1). C) Predicted local CR distribution among 4,757 HVR1 variants. The average local CR of the immunogenes is 27.4% (S.E. = 0.5) and of antigens is 28.6% (S.E. = 0.5).
Since the experimentally tested peptides represent a small fraction of all HVR1 sequences, they are substantially different from each other and cannot be used to evaluate CR among very closely related variants that may be expected at a single time-point in individual patients. However, the predicted data can be used to assess such local CR by calculating for each HVR1 variant the CR percentage to its 5 closest genetic sequences in the full HVR1 dataset (Fig. 4C). Local CR is generally higher than global CR because the closer two peptides are, the more likely they will cross-immunoreact. Although both local and global CR distributions were similar for immunogens, the antigen distributions were strikingly different, with the local distribution being considerably skewed (Fig. 4C). However, most HVR1 variants were predicted to be largely non-immunoreactive with each other (52.8% of the immunogens and 39.3% of the antigens do not cross-immunoreact locally). This finding is consistent with the suggested role of HVR1 in HCV escape from immune responses during human infection17,18,19,20,21.
The predicted CR was compared between aforementioned acutely infected (34 patients, 130 unique aa sequences) and chronically infected patients (90 patients, 799 unique aa sequences). The average global CR was significantly lower (MRPP, p = 0.0155) in the acutely infected patients (average = 10.60; S.E. = 1.84) than in the chronically infected patients (average = 17.10; S.E. = 1.46). This finding is consistent with previous observations made for HCV-infected patients27,39. The data shown here suggest, however, that the lower immunogenic CR in the acute-phase specimens is related to HVR1 properties and not only to the lower heterogeneity of intra-host populations usually found in acutely infected patients. Additionally, all these observations indicate that the predictive model captures HVR1 CR not only under the specific experimental conditions but also reflects some HVR1 immunological properties existing during human HCV infections. Nevertheless, considering the conformation dependence of the HVR1 CR (Fig. 1) discussed above, it should be noted that the observed specificity of immunological reactions will vary depending on the experimental conditions. A high accuracy of the decision tree model (Fig.S5) indicates that the HVR1 structural properties are strongly linked to specificity of immunoreactivity, suggesting the use of such models to cope with HCV heterogeneity for vaccine design. However, these models should be generated with consideration of the experimental conditions and HVR1 presentation in different protein contexts.
Discussion
Development of vaccines against HCV and other genetically heterogeneous viruses is hindered by lack of understanding of the factors that define the relationship between genetic variability and immunological specificity of antigenic epitopes. Elucidation of CR among variable antigenic epitopes is crucial for vaccine development. In the present work, we have modeled the HVR1 CR and its distribution in sequence space and formulated a novel concept of antigenic convergence for highly heterogeneous antigenic epitopes.
CR among HVR1 variants has been readily observed in early HCV experiments22,26,29,30. Using well characterized antibodies, several studies confirmed these early observations and showed that the CR is most probably associated with the HVR1 structural constraints26,27,28. These findings prompted a search for natural variants31 and mimotopes13 with a broad HVR1 CR. However, these studies did not succeed in establishing a quantitative association of CR to HVR1 heterogeneity. For example, Jackson et al31 conducted an extensive examination of CR (maximum = 31.1%; mean = 9.53%) among a number of HVR1 peptides but found no firm connection of immunological reactivity to sequence. Detection of CR (maximum = 83.3%; mean = 36.7%) among HVR1 variants derived from different HCV genotypes40,41 and the highly variable degree of immunoreactivity of sera with peptides containing HVR1 variants from the same patients27,41 clearly illustrates the lack of a simple association between sequence similarity and CR.
It must be noted that CR is essential but not sufficient for cross-neutralization. Nevertheless, given that immune recognition is a necessary step in viral neutralization, the modeling of CR among highly variable antigenic epitopes constitutes a critical starting point toward overcoming genetic diversity for development of an efficacious HCV vaccine. Here, we made several important observations useful for harnessing CR. First, the HCV HVR1 sequence space was found to be explored in its entirety by different HCV genotypes and subtypes, thus indicating a smaller size of the space than can be expected from the number of existing HCV strains. Also, it supports the use of HVR1 sequence variants from a single subtype or genotype for vaccine development. Second, in agreement with previous studies, we found a broad CR among genetically distant HVR1 variants, including variants from different HCV genotypes40,41. The skewed CR distribution implies that many HVR1 variants act as natural mimotopes. We showed that antibodies against a few HVR1 variants were immunoreactive with all tested peptides. Thus, in addition to the sequence space limitations, the number of HVR1 immunological specificities is also lower than can be expected from the number of HCV strains. The findings support vaccine development strategies that rely on discovery of broadly cross-immunoreactive epitopes12,13,14. Third, predictive models generated in this study showed for the first time that the HVR1 CR can be predicted based on sequence alone and there is a strong association between CR and HVR1 structure. Such in silico models should significantly facilitate a rational exploitation of HVR1 CR for vaccine development. Fourth, the general separation between antigenic and immunogenic properties suggests that selection of antigens for vaccine development should not be solely based on identifying HVR1 variants with broad antigenic reactivity; rather, immunogenic reactivity should be assessed. Fifth, the observed immunological convergence is defined by the strong conformational dependence of HVR1 epitopes, implying that the specificity of HVR1 CR may vary depending on the model system used for epitope presentation. Therefore, translation of the specific CR results from a model, for example based on synthetic peptides and mice immunization, to human vaccine development must be carefully approached. The HVR1 CR should be modeled at conditions approximating the actual vaccine application.
Although all observations of CR have been made in this study using a model system of synthetic peptides and mouse immunization, antigenic convergence is most probably a common property of highly heterogeneous antigenic epitopes. This conclusion is strongly supported by the recently reported identification of the specific antigenic reactivity among 10,000 random-sequence peptides to all tested antibodies42,43. The observations of the statistically significant association between CR frequency and sequence space centrality, or the stage of HCV infection as described in this paper strongly suggest that the model presented here captured a general distribution of immunological specificities among HVR1 variants.
It is commonly expected that CR rapidly declines with increasing genetic difference between epitopes. This assumption taken in the context of the expanding viral sequence space shaped by diversifying evolution is generally considered as a basis for immune escape and provides the major framework for vaccine development against highly heterogeneous viruses. The data obtained in this study indicate that this assumption is strongly consistent with the CR observed among closely related HVR1 variants usually found in each infected patient. However, frequent CR among genetically distant HVR1 variants significantly reduces the inverse relationship between the genetic distance and CR (Fig. 1A), indicating that the number of immunological specificities distributed across the entire HVR1 sequence space is limited. This restriction has different bearing on the intra- and inter-host evolution. The sequence space available to each HCV variant provides ample opportunities for the successful intra-host HVR1 evolution, facilitating immune escape during chronic infection. However, the HVR1 convergence significantly limits inter-host HCV evolution. This reduction in the genetic and immunological HCV diversity presents a different prospect on vaccine development than the concept of continues diversifying viral evolution.
The HCV HVR1 convergence observed in this study should be viewed as the natural extension of diversifying evolution that occurs in a limited sequence space. Owing to strong structural and functional constraints16,44,45,46, the HVR1 sequence space is severely restricted in size. The expansion under recurring selection pressures can result in “exhaustion” of the limited sequence space so generating conditions for frequent occurrences of homoplasy47. The data obtained here suggest that extensive convergence is a principal factor affecting the distribution of immunological properties among HVR1 variants. As a result, identical immunological specificities are repeatedly being observed among genetically distant variants, contributing to widespread epitope mimicry in the HVR1 sequence space. In contrast to diversification, convergence makes HVR1 variability tractable. Thus, the homoplastic nature of the HVR1 diversity turns the extensive genetic variability from drawback to opportunity and offers a novel and more general framework for development of a viable hepatitis C vaccine.
The interaction between virus and host is frequently compared to an “arms race”16, an idea rooted in diversifying viral evolution driven by antagonistic relationships with the host. Although such a process might explain viral escape from host immune responses, it is not instructive for vaccine development. A game of chess is a more useful metaphor describing HCV evolution. The chessboard symbolizes a limited intra-host phenotypic space available to HCV, which is composed of character states (immunological specificities) represented as squares. Each square may be occupied by different figures (genetic variants), resulting in homoplasy. HCV frequently cheats, making several mutation-moves at once and rapidly traversing across the entire space-board, a strategy that results in persistent infection. With a very similar “game” played during each infection, convergence is ingrained in HCV evolution.
Focus on sequence differences rather than on shared characteristics among viral variants hinders the discovery of phenotypic convergence. Convergent patterns of HCV immune responses pave the way for new prevention strategies that exploit the homoplastic nature of HCV sequence space. The concepts of convergence and tractability of diverse immunological specificities presented here should be applicable to other genetically heterogeneous viruses such as HIV and influenza virus, so contributing to the rational design of vaccines against these infections.
Methods
Synthetic peptides
A set of 262 HVR1 sequences was selected from Genbank sequences as described in Results, with 192 sequences belonging to genotype 1, 26 to genotype 2, 35 to genotype 3, 2 to genotype 4, 1 to genotype 5 and 6 to genotype 6. A control set of 29 synthetic peptides derived from proteins encoded by unrelated hepatitis delta virus and hepatitis G virus. These irrelevant peptides of different sizes were used as negative controls accounting for non-specific immunoreactivity of mouse serum specimens in all enzyme immunoassay experiments. All peptides were synthesized using standard f-moc chemistry. Quality control of the peptides was performed by Matrix-assisted laser desorption/ionization-time-of-flight (MALDI-TOF) mass spectrometry and high-performance liquid chromatography (HPLC). All peptides used in these experiments were similar in their quality.
Immunization of mice
For immunization of mice with synthetic peptides, 103 HVR1 peptides were first conjugated with BSA as a carrier protein. Each conjugated peptide was diluted with 1× PBS to a final concentration of 1 mg/ml and mixed with an equal volume of the adjuvant TiterMax (CytRx Corp, Norcross, GA). Each group of three 4–6 week-old female Balb-C mice was injected intraperitoneally with 100 µl of the resulting emulsion (50 µg of peptide) containing a single BSA-conjugated HVR1 peptide. Two weeks after first injection, a booster was administered with 50 µg BSA-conjugated peptide (without adjuvant). Two weeks after booster mice were bled and serum specimens were obtained.
Enzyme immunoassay (EIA)
Synthetic peptides representing 261 HVR1 variant sequences of 31 aa in length obtained from GenBank and 29 unrelated peptides were used as antigens in the assay. Wells of 384-well high binding plates were coated with 0.25 µg HVR1 peptides in a volume of 25 µl/well in PBS overnight at RT. Each well was coated with a different peptide and 16 wells were coated with 1.25 µg BSA as a control on immunoreactivity. After incubation, wells were washed five times with 100 µl PBS/0.05% Tween 20 (PBST) and 25µl of serum diluted 1∶400 in PBST/10% normal goat serum (NGS) was added. After incubating the plates at 37°C for 90 min, wells were washed again five times with PBST and 25 µl horseradish peroxidase-labeled anti-mouse IgG antibody diluted 1∶50,000 in PBST/10% NGS was added to each well. After washing five times with PBST, 25 µl of 1-Step Ultra TMB-ELISA was added for color development for 30 min at RT, followed by 25 µl 2M H2SO4 to stop the reaction and plates were read at OD450.
Negative controls
The reproducibility of experimental conditions between tests conducted at the same day or at different days is very important for accurate mathematical modeling of results. To control for inter-plate variation in each experiment, each 384-well plate contained 16 BSA-coated wells, 8 of which were tested against a 1∶30,000 dilution of pool of mice sera immunized with BSA and 8 were tested against a 1∶15,000 dilution. The inter-plate variation was low (S.D. 5.47%). The reactivity between serum and peptide was considered positive if it showed a signal/cut-off value >1 calculated for two types of negative controls.
-
i
Negative controls A: A cut-off was calculated for each peptide, based on its reactivity with 35 sera obtained from mice each immunized with different irrelevant peptides conjugated to BSA using the protocol described above. Cut-off value was calculated as the average of negative controls plus 3.5 times of the standard deviation. The average cut-off for all peptides was 0.07547.
-
ii
Negative control B: A cut-off was calculated for each serum, based on its reactivity with 29 different unrelated peptides. Cut-off value was calculated as the average of negative controls plus 3.5 times of the standard deviation. The average cut-off for all sera was 0.08567.
The reactivity of each immunogen and antigen pair was scored as 1 if one or more of the 3 immunized mice showed a positive reaction. Supplementary figure S1A shows the number of mice with a positive reaction between each tested immunogen and antigen. The percentage of cross-immunoreactive reactions with one or more mice is 18.74%, with two or more mice is 7.09% and with three mice is 2.43%. The average agreement between mice immunized with the same BSA-conjugated peptide was 88.78 (S.D. 14.98).
Network construction and analysis
A CR network is a directed graph where each vertex is a HVR1 sequence and there is a link between two nodes if the reaction between the two peptides was positive. Several topological measures of this network were calculated using the PAJEK software48. The CR network is difficult to visualize due to its high density of links, therefore, we used the k-shell decomposition method to disentangle the hierarchical structure of the network49.
Minimal cover set
The smallest set of immunogens that can have positive reactions with the whole set of antigens was considered as the minimal cover set. This set is a binary integer optimization problem, which was solved using a linear programming (LP)-based branch-and-bound algorithm implemented in MATLAB (MathWorks, Natick, MA).
Data from acute and chronic samples
For the determination of diameter and CR differences between acute and chronic stages of HCV, two groups of samples were used: (A) the acute group included 130 different aa sequences from 34 single time-point samples33,50,51,52; (B) the chronic group included 799 different aa sequences from 90 single time-point samples tested in our laboratory53.
Sequence space model
The whole dataset contained sequences of two classes: (A) multiple clone variants obtained in our laboratory from single time-point samples from 176 patients using the end-point limited-dilution (EPLD) real-time PCR38 and obtained from published data on 23 samples51; and multiple clones obtained from the follow-up specimens (2–14 time-points) from 29 patients50,52,54,33; and (B) sequences obtained by consensus sequencing from 2967 unrelated patients, which were recovered from the Los Alamos HCV Sequence Database37 during early 2008. Recombinant and chimeric sequences as well as patented sequences and sequences obtained from non-human hosts were excluded from analysis.
All sequences can be viewed as points in an informational space known as Sequence space. We have studied different ways of portraying this sequence space, taking into consideration two important problems: aa convergence and the dimensionality curse. (i) aa convergence: different aa could have very similar physicochemical properties or fulfill the same function in this particular segment and, therefore, it is desirable to build a sequence space that takes into account these differences. For HVR1, we assume that some positions are less variable because changes affect more its structure and function than changes in highly variable positions. The entropy of each aa position was calculated for the whole dataset and these entropy values were used to create weights in the calculation of the Hamming distances between every pair of sequences. (ii) Dimensionality curse: One of the most important goals in visualizing data is to get a sense of how near or far points are from each other. However, as dimensionality increases, the distance from a given point to the nearest point approaches the distance to the farthest point55. Distances in sequence space lose discriminatory power very rapidly. Accordingly, analysis was focused on the accurate portrayal of local relationships. We have created a model of sequence space using PFNET56, which in essence prunes a dense network. PFNETs have the ability to derive more accurate local structures than other algorithms where the resulting relationships between neighboring points are often significantly different from the original data56. The network generation procedure incorporates two parameters: (1) the r parameter defines the metric used for computing the distance of paths. (2) the q parameter constrains the number of indirect proximities examined in generating the network. The network with the minimum number of links is obtained when q = n−1 and r = ∞, i.e., PFNET(n−1,∞).
Analysis of molecular variance (AMOVA)
Each genotype was considered a subpopulation of the HVR1 sequence space and distances among these subpopulations were further explored by AMOVA as implemented in ARLEQUIN57. The genetic structure was analyzed with consideration of the molecular differences between sequences in addition to differences in their frequencies. Different types of distances were used: The nucleotide Hamming, aa Hamming and path distance (the number of steps in the shortest path connecting the pair of peptides in the PFNET). Significance levels of the variance components were estimated after 10,000 permutations.
Multi-response Permutation Procedure (MRPP)
MRPP is a non-parametric permutation test for testing the hypothesis of no difference between two or more groups of entities34, as implemented in the program BLOSSOM58. The distribution of values under the null hypothesis of no difference between groups was obtained by permutating the grouping labels (n = 10,000).
Ethics Statement
(i) Mice experiments: This study was carried out in strict accordance with the recommendations of the Guide for the Care and Use of Laboratory Animals of the National Institutes of Health. Protocols for mouse immunization experiments have been approved by the Centers for Disease Control and Prevention Animal Care and Use Committee (1403KHUMOUC). All efforts were made to minimize suffering. (ii) Human subjects: Blood specimens of 4 follow-up patients were acquired with written informed consent and all research involving human participants were approved by the institutional review board (Centers for Disease Control and Prevention, IRB#1428). Description of the NHANES III study can be found in53. Clinical investigations have been conducted according to the principles expressed in the Declaration of Helsinki.
References
Choo, Q. et al. Isolation of a cDNA Clone Derived From a Bloodborne Non-A, Non-B Viral Hepatitis Genome. Science 244, 359–362 (1989).
Torresi, J., Johnson, D. & Wedemeyer, H. Progress in the development of preventive and therapeutic vaccines for hepatitis C virus. J Hepatol 54, 1273–1285 (2011).
Alberti, A., Chemello, L. & Benvegnu, L. Natural History of Hepatitis C. .J Hepatol 31, 17–24 (1999).
Bowen, D. & Walker, C. Adaptive immune responses in acute and chronic hepatitis C virus infection. Nature 436, 946–952 (2005).
Simmonds, P. Genetic diversity and evolution of hepatitis C virus – 15 years on. J Gen Virol 85, 3173–3188 (2004).
Houghton, M. Prospects for prophylactic and therapeutic vaccines against the hepatitis C viruses. Immunol Rev. 239, 99–108 (2011).
McBurney, S. & Ross, T. Viral sequence diversity: challenges for AIDS vaccine designs. Expert Rev Vaccines 7, 1405–1417 (2008).
Rerks-Ngarm, S. et al. Vaccination with ALVAC and AIDSVAX to prevent HIV-1 infection in Thailand. N Engl J Med 361, 2209–2220 (2009).
Yang, O. Candidate vaccine sequences to represent intra- and inter-clade HIV-1 variation. PLoS One 4, e7388. (2009).
Kang, K. et al. Synthetic antigens representing the antigenic variation of human hepatitis C virus. Viral Immunol 23, 497–508 (2010).
Yusim, K. et al. Genotype 1 and global hepatitis C T-cell vaccines designed to optimize coverage of genetic diversity. J Gen Virol 91, 1194–1206 (2010).
El-Attar, L., Partidos, C. & Howard, C. A peptide mimotope of hepatitis C virus E2 protein is immunogenic in mice and block human anti-HCV sera. J Med Virol 82, 1655–1665 (2010).
Roccasecca, R. et al. Mimotopes of the hyper variable region 1 of the hepatitis C virus induce cross-reactive antibodies directed against discontinuous epitopes. Mol Immunol 38, 485–492, S0161589001000840 [pii] (2001).
Hamano, T. et al. Determination of HIV type 1 CRF01_AE gag p17 and env-V3 consensus sequences for HIV/AIDS vaccine design. AIDS Res Hum Retroviruses 20, 337–340 (2004).
Arenas, J. et al. Hepatitis C virus quasi-species dynamics predict progression of fibrosis after liver transplantation. J Infect Dis 189, 2037–2046 (2004).
Sheridan, I., Pybus, O., Holmes, E. & Klenerman, P. High-resolution phylogenetic analysis of hepatitis C virus adaptation and its relationship to disease progression. J Virol 78, 3447–3454 (2004).
Van Doorn, L. et al. Sequence evolution of the hypervariable region in the putative envelope region E2/NS1 of hepatitis C virus is correlated with specific humoral immune responses. J Virol 69, 773–778 (1995).
Eckels, D., Zhou, H., Bian, T. & Wang, H. Identification of antigenic escape variants in an immunodominant epitope of hepatitis C virus. Int Immunol 11, 577–583 (1999).
Pavio, N. & Lai, M. The Hepatitis C Virus Persistence: How To Evade The Immune System? J Biosci 3, 287–304 (2003).
Isaguliants, M. Hepatitis C virus clearance: the enigma of failure despite an impeccable survival strategy. Curr Pharm Biotechnol 4, 169–183 (2003).
Lopez-Labrador, F. et al. Genetic variability of hepatitis C virus NS3 protein in human leukocyte antigen-A2 liver transplant recipients with recurrent hepatitis C. .Liver Transpl 10, 217–227 (2004).
Puntoriero, G. et al. Towards a solution for hepatitis C virus hypervariability: mimotopes of the hypervariable region 1 can induce antibodies cross-reacting with a large number of viral variants. EMBO J 17, 3521–3533 (1998).
Law, M. et al. Broadly neutralizing antibodies protect against hepatitis C virus quasispecies challenge. Nat Med 14, 25–27 (2008).
Farci, P. et al. Prevention Of Hepatitis C Virus Infection In Chimpanzees By Hyperimmune Serum Against The Hypervariable Region 1 Of The Envelope 2 Protein. Proc Natl Acad Sci USA 93, 15394–15399 (1996).
Choo, Q. et al. Vaccination of chimpanzees against infection by the hepatitis C virus. Proc Natl Acad Sci U S A. 91, 1294–1298. (1994).
Scarselli, E. et al. Occurrence of antibodies reactive with more than one variant of the putative envelope glycoprotein (gp70) hypervariable region 1 in viremic hepatitis C virus-infected patients. J Virol 69, 4407–4412 (1995).
Mondelli, M. U. et al. Antibody responses to hepatitis C virus hypervariable region 1: evidence for cross-reactivity and immune-mediated sequence variation. Hepatology 30, 537–545 (1999).
Cerino, A. et al. Monoclonal antibodies with broad specificity for hepatitis C virus hypervariable region 1 variants can recognize viral particles. J Immunol 167, 3878–3886 (2001).
Lesniewski, R. R. et al. Hypervariable 5′-terminus of hepatitis C virus E2/NS1 encodes antigenically distinct variants. J Med Virol 40, 150–156 (1993).
da Silva, L. C. et al. Long term follow-up and patterns of response of ALT in patients with chronic hepatitis NANB/C treated with recombinant interferon-alpha. Rev Inst Med Trop Sao Paulo 37, 239–243 (1995).
Jackson, P., Petrik, J., Alexander, G. J., Pearson, G. & Allain, J. P. Reactivity of synthetic peptides representing selected sections of hepatitis C virus core and envelope proteins with a panel of hepatitis C virus-seropositive human plasma. J Med Virol 51, 67–79, 10.1002/(SICI)1096-9071(199701)51:167::AID-JMV11>3.0.CO;2-1 [pii] (1997).
Hjalmarsson, S., Blomberg, J., Grillner, L., Pipkorn, R. & Allander, T. Sequence evolution and cross-reactive antibody responses to hypervariable region 1 in acute hepatitis C virus infection. J Med Virol 64, 117–124 (2001).
von Hahn, T. et al. Hepatitis C virus continuously escapes from neutralizing antibody and T-cell responses during chronic infection in vivo. Gastroenterology 132, 667–678 (2007).
McCune, B. & Grace, J. Analysis of ecological communities. (MjM Software Design, 2002).
Albert, R. & Barabasi, A. Statistical mechanics of complex networks. Reviews of Modern Physics 74, arXivcond-mat/0106096v1 (2002).
Watts, D. & Strogatz, S. Collective dynamics of small-world networks. Nature 393, 440–442 (1998).
Kuiken, C., Yusim, K., Boykin, L. & Richardson, R. The Los Alamos hepatitis C sequence database. Bioinformatics 21, 379–384 (2005).
Ramachandran, S., Xia, G. L., Ganova-Raeva, L. M., Nainan, O. V. & Khudyakov, Y. End-point limiting-dilution real-time PCR assay for evaluation of hepatitis C virus quasispecies in serum: performance under optimal and suboptimal conditions. J Virol Methods 151, 217–224 (2008).
Scotta, C. et al. Influence of specific CD4+ T cells and antibodies on evolution of hypervariable region 1 during acute HCV infection. Journal of Hepatology 48, 216–228 (2008).
Hattori, M. et al. Broadly reactive antibodies to hypervariable region 1 in hepatitis C virus-infected patient sera: relation to viral loads and response to interferon. Hepatology 27, 1703–1710 (1998).
Yoshioka, K. et al. Humoral immune response to the hypervariable region of hepatitis C virus differs between genotypes 1b and 2a. J Infect Dis 175, 505–510 (1997).
Halperin, R., Stafford, P. & Johnston, S. Exploring Antibody Recognition of Sequence Space through Random-Sequence Peptide Microarrays. Mol Cell Proteomics 10 (2011).
Legutki, J., Magee, D., Stafford, P. & Johnston, S. A general method for characterization of humoral immunity induced by a vaccine or infection. 28, 4529–4537 (2010).
Penin, F. et al. Conservation of the conformation and positive charges of hepatitis C virus E2 envelope glycoprotein hypervariable region 1 points to a role in cell attachment. J Virol 75, 5703–5710 (2001).
McAllister, J. et al. Long-term evolution of the hypervariable region of hepatitis C virus in a common-source-infected cohort. J Virol 72, 4893–4905 (1998).
Smith, D. Evolution of the hypervariable region of hepatitis C virus. J Viral Hepat 6, 41–46 (1999).
Wagner, P. Exhaustion of morphologic character states among fossil taxa. Evolution 54, 365–386 (2000).
Batagelj, V. & Mrvar, A. in Graph Drawing Software Mathematics and Visualization (eds M. Juenger & P. Mutzel) 77–103 (Springer, 2003).
Alvarez-hamelin, I., Dall'Asta, L. & Vespignani, A. k-core decomposition: a tool for the visualization of large scale networks. Advances in Neural Information Processing Systems 18 (2006).
Ramachandran, S. et al. Temporal Variations in the Hepatitis C Virus Intra-Host Population During Chronic Infection. J virol, 10.1128/JVI.02204-10 (2011).
Herring, B., Tsui, R. & Peddada, L. Wide Range of Quasispecies Diversity During Primary Hepatitis C Virus Infection. J virol 79 4340–4346 (2005).
Farci, P. et al. The outcome of acute hepatitis C predicted by the evolution of the viral quasispecies. Science 288, 339–334 (2000).
Nainan, O. et al. Hepatitis C virus genotypes and viral concentrations in participants of a general population survey in the United States. Gastroenterology 131, 478–484. (2006).
Farci, P. et al. Evolution of hepatitis C viral quasispecies and hepatic injury in perinatally infected children followed prospectively. Proc Natl Acad Sci U S A 103, 8475–8480 (2006).
Beyer, K., Goldstein, J., Ramakrishnan, R. & Shaft, U. in Proceedings of the 7th International Conference on Database Theory (eds C. Beeri & P. Buneman) 217–235 (Springer-Verlag, Jerusalem, Israel., 1999).
Schvaneveldt, R., Durso, F. & Dearholt, D. Network structures in proximity data. The Psychology of learning and motivation 24, 249–283 (1989).
Schneider, S., Roessli, D. & Excoffier, L. ARLEQUIN version 2000: Software for population genetic data analysis. (2000).
Cade, B. & Richards, J. User Manual For BLOSSOM Statistical Software. Midcontinent Ecological Science Center US Geological Survey Fort Collins, Colorado (2001).
Acknowledgements
Chrys Lynberg for computational support; Yulin Lin for providing HVR1 sequences; Guo-Liang Xia for database management; and Livia Rossi for help in genotyping. NOP was supported by CONACyT Mexico project 2004-C01-421.
Author information
Authors and Affiliations
Contributions
Y.K. and D.S.C. designed the study; D.H., J.Y., N.O.P. and S.R. performed laboratory experiments; D.S.C. and Z.D. developed the sequence space model; D.S.C., Z.D. and Y.K. analyzed data; D.S.C. and Y.K. wrote the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supporting information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareALike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Campo, D., Dimitrova, Z., Yokosawa, J. et al. Hepatitis C Virus Antigenic Convergence. Sci Rep 2, 267 (2012). https://doi.org/10.1038/srep00267
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep00267
This article is cited by
-
Evaluation of a mimotope of the Rickettsia outer membrane protein A (OmpA) as an antigen in enzyme-linked immunosorbent assay to detect rickettsiosis in capybaras (Hydrochoerus hydrochaeris), horses (Equus caballus), and opossums (Didelphis sp.)
Experimental and Applied Acarology (2023)
-
Role of HVR1 sequence similarity in the cross-genotypic neutralization of HCV
Virology Journal (2020)
-
Transmissibility of intra-host hepatitis C virus variants
BMC Genomics (2017)
-
GHOST: global hepatitis outbreak and surveillance technology
BMC Genomics (2017)
-
Design, Synthesis and Structure-Activity Relationship Optimization of Lycorine Derivatives for HCV Inhibition
Scientific Reports (2015)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.