Abstract
Formation of higher-order structure of nucleic acids (hairpins or loops, for example) may impact not only gene regulation, but also molecular biology techniques and approaches critical for design and production of vectors needed for genetic engineering approaches. In the course of designing vectors aimed to modify the murine Foxd3 locus through homologous recombination in embryonic stem cells, we discovered a 370 nucleotide segment of DNA resistant to polymerase read-through. In addition to sequencing and PCR disruptions, we were unable to use BAC recombineering strategies to exchange sequences within the Foxd3 locus. This segment corresponds to a putative DNA hairpin region just upstream of the 5’ untranslated region of Foxd3. This region is also highly conserved across vertebrate species, suggesting possible functional significance. Our findings provide a cautionary note for researchers experiencing technical challenges with BAC recombineering or other molecular biology methods requiring recombination or polymerase activity.
Similar content being viewed by others
Introduction
Recombination-mediated genetic engineering using bacterial artificial chromosomes (BAC recombineering) is a powerful approach for efficiently generating complex DNA constructs, transgenes and gene targeting vectors 1,2,3,4. Using this method, much larger fragments of DNA can be assembled, allowing for inclusion of distal upstream regulatory elements for analysis and the precise replacement of endogenous regions of the genome. The technique utilizes homologous recombination between short homology fragments in modified strains of E. coli expressing phage recombinase proteins. BAC recombineering also enables relatively rapid generation of complex constructs. We initially aimed to use BAC recombineering to generate DNA constructs with a goal of modifying the Foxd3 locus. In our attempts to produce these vectors, we encountered a conserved stretch of 370 nucleotides (nt) in the 5’ untranslated region (UTR) of the Foxd3 locus that was resistant to polymerase read-through during both PCR and sequencing reactions. In addition, we were also unable to achieve successful recombination over this region.
High guanine-cytosine content (GC content) within DNA can lead to formation of secondary structure through structures such as DNA hairpins or loops. This secondary structure can be prohibitive to polymerase read-through under normal conditions, resulting in abrupt sequencing stops5,6,7,8,9. In addition to empirical evidence 10, there is an abundance of anecdotal evidence (including online troubleshooting guides from multiple sequencing facilities) suggesting that abrupt stops in sequencing reads may also be the result of DNA hairpin structures. The 370 nt sequence we describe is predicted to assemble into secondary structure in the form of a cluster of DNA hairpins. We present a case here demonstrating that such a hairpin, while troublesome for sequencing, may also prohibit BAC recombineering.
Results
Discovery of a polymerase-resistant region of DNA upstream of the Foxd3 coding sequence
Sequencing of a plasmid containing the genomic Foxd3 coding region with flanking DNA (under both normal conditions and conditions for GC-rich templates), revealed that 3’ to 5’ sequencing reads came to an abrupt stop 442 nt upstream (position −442) of the Foxd3 ATG and 5’ to 3’ sequencing reads resulted in a sequencing stop precisely 811 nt upstream (position −811) of the Foxd3 ATG (Figure 1). These two positions define a segment of 370 nt resistant to polymerase read-through (blue rectangle in Figure 1). Multiple primers flanking this segment were used (Primers 1–3 and 11–13 in Figure 1), but sequencing reads extending across this entire region were never obtained, with sequencing stops consistently occurring at the border of this 370 nt sequence. However, when primers that anneal inside the 370 nt sequence were used to sequence out of the region, sequencing reads could easily extend through either border (Primers 4–10 in Figure 1). Using primers within this segment, we were able to sequence across the region and verify that no abnormal intervening sequence was the cause of the sequencing stops. Similar results were obtained when sequencing from a BAC DNA template containing the Foxd3 locus.
A 370 nt region upstream of the Foxd3 coding sequence is resistant to polymerase read-through during PCR or sequencing reactions.
A schematic representing the murine Foxd3 locus shows the location of primers used for sequencing and PCR (arrows labeled 1–13). When primers outside the 370 nt (numbers 1 – 3 and 11 – 13) were used to sequence this region, reads stopped abruptly at position −811 for 5’ to 3’ sequencing or position −442 for 3’ to 5’ sequencing. Sequencing of DNA using primers located within the 370 nt region (numbers 4 – 10) continued successfully through the sequence. The scale bar shows increments of 500 nt, with position 1 indicating the A in the Foxd3 ATG. The 370 nt region is shown in blue. The table details locations of each primer and the positions where polymerase read-through ends abruptly, where applicable. Brackets indicate the positions of homology fragments A through D used for BAC recombineering. Abbreviations: ATG, start codon; nt, nucleotide; Seq, sequencing; TGA, stop codon; UTR, untranslated region.
In parallel, we attempted PCR across this region with genomic DNA extracted from mouse tail biopsies and murine embryonic stem (ES) cells, in addition to DNA from BAC clones or Foxd3-containing plasmids. Without exception, these reactions were unsuccessful under multiple conditions; the PCR presumably failed due to the presence of the polymerase-resistant region. This stretch of sequence is GC-rich (see analysis in Figure 2); therefore we attempted PCR amplification of this region using multiple polymerases and kits with conditions tailored to GC-rich templates (more details in Methods Section). However, these attempts were universally unsuccessful. In contrast, similar to results with DNA sequencing reactions above, when we performed PCR with one primer that annealed within the 370 nt sequence and one that annealed outside the 370 nt sequence (ex. Primers 9 or 10 with a primer external to the 370 nt region), we obtained amplicons that crossed the boundary of the 370 nt segment. Amplification across a boundary was only successful when one of the primers used for PCR was within the 370 nt sequence, suggesting that polymerases primed from external primers could not extend the amplicon through the putative boundary established by the 370 nt segment, but were able to travel along the DNA far enough to meet the strand amplified from the inside outward. We used this strategy to amplify the entire 370 nt sequence in two pieces.
The GC-rich 370 nt region is conserved among multiple vertebrate species and correlates with enrichment of H3K4 methylation.
Three stretches of 370 nt upstream of the Foxd3 coding region are analyzed: the center 370 nt polymerase-resistant region (segment  ) plus 370 nt flanking either side (segments
) plus 370 nt flanking either side (segments  and
and  ). Analysis of GC-content shows that both the 370 nt polymerase-resistant region
). Analysis of GC-content shows that both the 370 nt polymerase-resistant region  and the 3’ 370 nt region
and the 3’ 370 nt region  are highly GC-rich. A schematic representing the murine Foxd3 locus with the location of the regions below (
are highly GC-rich. A schematic representing the murine Foxd3 locus with the location of the regions below ( ,
, and
and  ) is shown in green. Nucleotide BLAST analysis of these three regions is given in the table. Chlorocebus: Chlorocebus aethiops (monkey), Homo: Homo sapiens (human), Gallus: Gallus gallus (chicken), Danio: Danio rerio (zebrafish), Xenopus: Xenopus laevis (frog). Percentage (%) indicates percent of the query (370 nt) that was aligned, while “score” indicates the alignment score assigned by the blastn scoring matrix. The mouse Foxd3 locus is shown within the NCBI DCODE ECR browser identifying regions of homology across species shown by peaks. The 5’ UTR is not predicted in humans, but there is a conserved region of 5’ sequence. The 370 nt region
) is shown in green. Nucleotide BLAST analysis of these three regions is given in the table. Chlorocebus: Chlorocebus aethiops (monkey), Homo: Homo sapiens (human), Gallus: Gallus gallus (chicken), Danio: Danio rerio (zebrafish), Xenopus: Xenopus laevis (frog). Percentage (%) indicates percent of the query (370 nt) that was aligned, while “score” indicates the alignment score assigned by the blastn scoring matrix. The mouse Foxd3 locus is shown within the NCBI DCODE ECR browser identifying regions of homology across species shown by peaks. The 5’ UTR is not predicted in humans, but there is a conserved region of 5’ sequence. The 370 nt region  (large green arrow) is just upstream of the mouse 5’ UTR and is conserved in human, rat, opossum, dog, rhesus monkey and chimpanzee. Blue represents conserved coding sequence, yellow represents conserved UTRs, green represents conserved simple repeats or transposons and red represents conserved intergenic peaks. Tracks from the DCODE ECR Browser for the human FOXD3 locus were aligned with tracks from ENCODE data in the UCSC Genome Browser, demonstrating that region
(large green arrow) is just upstream of the mouse 5’ UTR and is conserved in human, rat, opossum, dog, rhesus monkey and chimpanzee. Blue represents conserved coding sequence, yellow represents conserved UTRs, green represents conserved simple repeats or transposons and red represents conserved intergenic peaks. Tracks from the DCODE ECR Browser for the human FOXD3 locus were aligned with tracks from ENCODE data in the UCSC Genome Browser, demonstrating that region  corresponds to peaks in H3K4me3 in two cell lines and a peak representing open chromatin, indicated by DNase hypersensitivity.
corresponds to peaks in H3K4me3 in two cell lines and a peak representing open chromatin, indicated by DNase hypersensitivity.
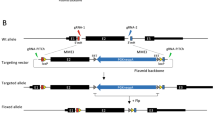
We discovered that this 370 nt sequence also interfered with attempts to recombine sequences overlapping with this region during BAC recombineering. Using traditional BAC recombineering 4 to generate constructs targeting the Foxd3 locus, we generated 4 homology fragments of approximately 500 nt (Fragments A–D shown in Figure 1) to extract 5’ and 3’ homology arms of approximately 8.5 kb (from start of A [−8549 to −8039] to end of B [−547 to 0] and 5 kb (from start of C [3665 to 4216] to end of D [7699 to 8239]), respectively. Fragment B contained the approximately 500 nt immediately upstream of the Foxd3 start codon (ATG), which included the entire 326 nt 5’ UTR (labeled 5’ in Figure 1). The 5’ end of Fragment B partially overlapped with the 370 nt of proposed secondary structure we describe in this report. Although we were able to electroporate a BAC containing genomic Foxd3 sequence into EL350 cells, multiple attempts at BAC recombineering were unsuccessful. While some clones survived antibiotic selection, restriction enzyme digestion analyses of DNA prepared from recombinant clones with appropriate antibiotic resistance resulted in fragment patterns indicating smaller plasmid sizes than expected. Although fragments corresponding to the 3’ arm (between C and D) were found, fragments corresponding to pieces upstream of the 370 nt region were never found in the recombined BAC (between Fragments A and B). When these clones were sequenced, the DNA corresponding to Fragment B matched the expected sequence until position −442, the same position of the precise stop observed in sequencing and PCR, where the subsequent sequence was unreadable.
The 370 nt region is conserved among vertebrate species and highly conserved among mammals
When we examined homologous regions upstream of the Foxd3 coding region in other vertebrates, we observed a significant degree of conservation over this 370 nt segment (Figure 2). Nucleotide BLAST analysis of the 370 nt sequence of the mouse genome revealed conservation with human (Homo), monkey (Chlorocebus), zebrafish (Danio), chicken (Gallus) and frog (Xenopus). In contrast, the same length of sequence immediately upstream (5’) of the 370 nt segment showed no significant conservation and a 370 nt length of sequence immediately downstream (3’) of the 370 nt region of interest showed only conservation with mammals (human and monkey) (Figure 2). Note: for simplicity, we will define these three fragments as  : the 370 upstream of the polymerase resistant region,
: the 370 upstream of the polymerase resistant region,  : the resistant region and
: the resistant region and  : the 370 nt immediately 3’ of the resistant region (diagrammed at the top of Figure 2). All three of these sequences are 5’ of the coding region although
: the 370 nt immediately 3’ of the resistant region (diagrammed at the top of Figure 2). All three of these sequences are 5’ of the coding region although  is within the 5’ UTR. Using the NCBI DCODE software to identify ECRs (Evolutionarily Conserved Regions) upstream of human FOXD3 further demonstrated conservation of this region in mammalian species (Figure 2).
is within the 5’ UTR. Using the NCBI DCODE software to identify ECRs (Evolutionarily Conserved Regions) upstream of human FOXD3 further demonstrated conservation of this region in mammalian species (Figure 2).
The polymerase-resistant 370 nt region is GC-rich and predicted to form secondary structure
The presence of a segment of DNA resistant to polymerase read-through suggested secondary structures such as DNA hairpins 10. Secondary structure that inhibits sequencing or PCR reactions typically occurs in GC-rich regions. Therefore, we analyzed the GC-content of the 370 nt sequence compared to the two 370 nt segments immediately flanking it (regions  and
and  ) (Figure 2). Although the 370 nt sequence is highly GC-rich (61% GC content) in contrast with region
) (Figure 2). Although the 370 nt sequence is highly GC-rich (61% GC content) in contrast with region  (39% GC content), which is relatively GC-poor, its GC character is not unique within the Foxd3 locus, as region
(39% GC content), which is relatively GC-poor, its GC character is not unique within the Foxd3 locus, as region  is also highly GC-rich (71% GC content). To determine if the region of polymerase-resistance was due to the presence of secondary structure in the 370 nt, we analyzed this region with RNAfold secondary structure-predicting software 11 for DNA. We focused on the 1110 nt that includes the three regions discussed here (
is also highly GC-rich (71% GC content). To determine if the region of polymerase-resistance was due to the presence of secondary structure in the 370 nt, we analyzed this region with RNAfold secondary structure-predicting software 11 for DNA. We focused on the 1110 nt that includes the three regions discussed here ( ,
, and
and  ). At 72 degrees Celsius (the extension temperature for polymerase), the 370 nt region of interest (
). At 72 degrees Celsius (the extension temperature for polymerase), the 370 nt region of interest ( ) was predicted by minimum free energy to form a tight cluster of hairpin structures (Figure 3A, boxed region). The nucleotides corresponding to the sequencing stop boundaries of this region (arrowheads in Figure 3B) were each located within a separate long predicted hairpin with the highest base-pairing probability, consistent with the possibility that these long, stable hairpin arms defined the boundary for the 370 nt segment. While the GC-rich 370 nt region
) was predicted by minimum free energy to form a tight cluster of hairpin structures (Figure 3A, boxed region). The nucleotides corresponding to the sequencing stop boundaries of this region (arrowheads in Figure 3B) were each located within a separate long predicted hairpin with the highest base-pairing probability, consistent with the possibility that these long, stable hairpin arms defined the boundary for the 370 nt segment. While the GC-rich 370 nt region  segment was also predicted to form a series of hairpins, it lacked any strong, high base-pairing probability hairpins (Figure 3B). This analysis suggested the involvement of secondary structure as a cause for the precise stops during sequencing and interference during PCR and BAC recombineering of the Foxd3 locus. A second method using UNAfold prediction software, also predicted a hairpin that started at approximately the −811 position (the same 5’ boundary corresponding to a barrier to sequencing and PCR).
segment was also predicted to form a series of hairpins, it lacked any strong, high base-pairing probability hairpins (Figure 3B). This analysis suggested the involvement of secondary structure as a cause for the precise stops during sequencing and interference during PCR and BAC recombineering of the Foxd3 locus. A second method using UNAfold prediction software, also predicted a hairpin that started at approximately the −811 position (the same 5’ boundary corresponding to a barrier to sequencing and PCR).
A predicted, stable DNA hairpin loop exists in the 370 nt polymerase-resistant region.
(A) RNAfold analysis for DNA at 72 degrees Celsius performed on the 1110 bases of nucleotide sequence comprising regions ,
,  and
and  . (B) An enlargement of the region in A outlined with the dashed box. The structure shown is the predicted optimum structure for minimum free energy. Sequence is displayed from 5’ to 3’ counterclockwise. The first 370 nt, region
. (B) An enlargement of the region in A outlined with the dashed box. The structure shown is the predicted optimum structure for minimum free energy. Sequence is displayed from 5’ to 3’ counterclockwise. The first 370 nt, region  (−1181 to −811) has almost no predicted secondary structure. The 370 nt polymerase-resistant region
(−1181 to −811) has almost no predicted secondary structure. The 370 nt polymerase-resistant region  (−811 to −442) corresponds to a tight structure of clustered hairpins. The two long hairpin arms with highest base-pairing probability (orange to red) contain the boundaries of the polymerase-resistant region as marked with black arrowheads at −442 and −811 and bracketed at the bottom of the figure. The rainbow scale indicates a range of base-pairing probability from 0 to 1, violet to red.
(−811 to −442) corresponds to a tight structure of clustered hairpins. The two long hairpin arms with highest base-pairing probability (orange to red) contain the boundaries of the polymerase-resistant region as marked with black arrowheads at −442 and −811 and bracketed at the bottom of the figure. The rainbow scale indicates a range of base-pairing probability from 0 to 1, violet to red.
To determine if the barrier to DNA polymerase through this region was independent of the genomic or plasmid sequence context, we took advantage of resident restriction enzyme sites (Figure 4A). Cutting the plasmid shown in Figure 4A with BspEI removed almost all (348 of 370 nts) of the predicted hairpin sequence (region  shown in orange). When the BspEI fragment was removed from the vector (Figure 4B), we were then able to sequence across the remaining regions. The BspEI fragment was inserted into pBluescript (Figure 4C). The 5’ BspEI site is located 22 nucleotides into the predicted hairpin (region
shown in orange). When the BspEI fragment was removed from the vector (Figure 4B), we were then able to sequence across the remaining regions. The BspEI fragment was inserted into pBluescript (Figure 4C). The 5’ BspEI site is located 22 nucleotides into the predicted hairpin (region  in orange), therefore we would predict that the inserted BspEI fragment would have a disrupted 5’ hairpin boundary, but an intact 3’ hairpin boundary. Consistent with this, we were able to sequence through the hairpin with forward primers upstream of the predicted hairpin, but not with reverse primers downstream of the predicted hairpin. RNAfold analysis of secondary structure of this fragment further suggested that the 5’ hairpin boundary was disrupted (Figure 4D), resulting in a much weaker predicted hairpin than in intact Foxd3 genomic DNA.
in orange), therefore we would predict that the inserted BspEI fragment would have a disrupted 5’ hairpin boundary, but an intact 3’ hairpin boundary. Consistent with this, we were able to sequence through the hairpin with forward primers upstream of the predicted hairpin, but not with reverse primers downstream of the predicted hairpin. RNAfold analysis of secondary structure of this fragment further suggested that the 5’ hairpin boundary was disrupted (Figure 4D), resulting in a much weaker predicted hairpin than in intact Foxd3 genomic DNA.
Disruption or insertion of the polymerase-resistant region alters the barrier to sequencing read-through.
(A–C) Removal of the majority of the sequence corresponding to the predicted hairpin from Foxd3-containing plasmids and insertion of a partially disrupted hairpin sequence into a pBluescript backbone. Successful sequencing read-through is indicated by a single black arrow, whereas unsuccessful read-through is indicated by a black arrow ending in a red X. (A) Schematic of a plasmid containing a 7642 bp EcoRI genomic Foxd3 fragment, showing the location of the EcoRI and BspEI sites. (B) Schematic of a vector generated by removal of the 938 bp and 123 bp BspEI fragments. (C) Schematic of 938 bp BspEI fragment inserted into the EcoRV site of pBluescript. (D) RNAfold secondary structure prediction for the 938 bp BspEI fragment alone indicates disruption of the 5’ hairpin boundary.
An alternative secondary structure particularly common for GC-rich sequences is the G-quadruplex that can form via Hoogstein base-pairing among stretches of guanine nucleotides. These structures occupy the proximal promoter of some genes and correlate with repression of gene expression 12. Although these structures would be an attractive alternative candidate for a conserved polymerase-resistant region, software prediction models of G-quadruplexing did not show any significant potential for G-quadruplexes in the 370 nt region  sequence (data not shown).
sequence (data not shown).
Discussion
Here we describe a DNA sequence of 370 nt resistant to polymerase read-through, resulting in precise DNA sequencing stops and preventing PCR amplification across this region. This region is conserved and predicted to form secondary structure comprised of a cluster of hairpins. Regions of sequence that produce sequencing stops are usually attributed to GC-rich regions or secondary structure, but consideration of such structures in the design of vectors has been generally under-studied and further analyses would be necessary to systematically determine if there is a defined effect of secondary structure on homologous recombination at other loci or with other techniques. The presence of this type of region or secondary structure may be important to consider in other genes, especially if it has the potential to interfere with attempts to amplify these sequences by PCR or execute recombination across these regions, such as during BAC recombineering. We also acknowledge the possibility that the predicted secondary structure could interfere with any of the other components needed for homologous recombination in addition to DNA polymerase13. In previously constructed Foxd3 targeting vectors this area was cloned using restriction enzyme sites located outside the 370 nt sequence, avoiding the need for PCR amplification or homologous recombination in E. coli14,15. Although BAC recombineering and PCR at this specific area of the Foxd3 locus was unsuccessful and the mechanism for hindrance of BAC recombineering is not precisely defined, we show here that another potential strategy to overcome the inability to PCR-amplify specific segments of polymerase-resistant DNA is to start from the middle and generate two amplicons that can later be pieced together. This may be an important alternative for researchers encountering difficulties with these commonly used polymerase-dependent approaches.
Our observation and analysis of conservation of this region is consistent with the possibility that the predicted secondary structure may also have functional significance. This region may be important as part of the Foxd3 promoter or an enhancer element that regulates Foxd3 transcription in specific cell types. Interestingly, aligning the same sequence analyzed within the DCODE ECR Browser with tracks from the ENCODE (Encyclopedia of DNA Elements) Project using the UCSC Genome Browser showed the conserved 370 nt region  corresponded to peaks in trimethylation of Histone 3 Lysine 4 (H3K4me3) and DNase hypersensitivity, both indicators of open chromatin in two mesoderm-derived cell lines (GM12878 and K562) (Figure 2). This correlation could also explain the sequence conservation we observed. It is unknown whether this predicted structure occurs in vivo, where recruitment of specific DNA or histone binding factors could stabilize or eliminate higher-order structures formed in this area. Although further functional analysis of this 370 nt segment is necessary to determine whether it impacts Foxd3 gene expression, we have identified an intriguing conserved element and offer the interesting idea that this potential DNA secondary structure may have functional significance.
corresponded to peaks in trimethylation of Histone 3 Lysine 4 (H3K4me3) and DNase hypersensitivity, both indicators of open chromatin in two mesoderm-derived cell lines (GM12878 and K562) (Figure 2). This correlation could also explain the sequence conservation we observed. It is unknown whether this predicted structure occurs in vivo, where recruitment of specific DNA or histone binding factors could stabilize or eliminate higher-order structures formed in this area. Although further functional analysis of this 370 nt segment is necessary to determine whether it impacts Foxd3 gene expression, we have identified an intriguing conserved element and offer the interesting idea that this potential DNA secondary structure may have functional significance.
Methods
Sequencing
All DNA was sequenced by the Vanderbilt University Medical Center Genome Sciences Sanger DNA Sequencing laboratory using BigDye Terminator chemistry with resolution on an ABI 3730xl DNA Analyzer. In addition to standard sequencing protocols, we used protocols for GC-rich templates, including an increased denaturing temperature and additional enzyme. Primer sequences are listed in Figure 1. Templates used for sequencing included murine genomic Foxd3 fragments subcloned into a pBluescript backbone vector and BAC DNA containing the Foxd3 locus. Two 129S6 BAC clones containing Foxd3 (m355-P15 and m284-J23) were obtained from the RPCI-22 Mouse BAC library 16 (Research Genetics, Inc., now Invitrogen). A 129S7 BAC clone containing Foxd3 (bMQ-388O11) was identified in the Ensembl genome browser and obtained from the Mouse bMQ BAC library (GeneService Ltd.).
PCR
Polymerase chain reaction across the 370 nt region of predicted secondary structure or from within this region was attempted with multiple protocols using different polymerases. This included GoTaq Flexi DNA polymerase (Promega), Pfu Ultra (Stratagene), LA Taq (Takara) and Expand Long Template Kit (Roche). For LA Taq, both GC Buffer I and GC Buffer II were used in an attempt to improve amplification through the polymerase-resistant region. For the Expand Long Template Kit, Systems 1, 2 and 3 were used. DNA templates used for PCR: genomic DNA isolated from tail biopsies from mice of either a predominantly C57B6 or 129S6 genetic background, TL1 ES cells 17 (129S6), two distinct plasmids containing fragments of Foxd3 genomic sequence and BAC DNA as described above.
BAC recombineering
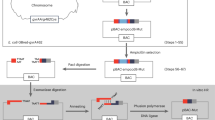
We electroporated a Foxd3-containing BAC into the E. coli strain EL350. We then electroporated an insertion vector containing two 500 bp homology fragments homologous to sequences 5’ of the Foxd3 start codon and 3’ of the Foxd3 stop codon (Figure 1) into the BAC-containing EL350 cells. Recombination events were identified by acquisition of antibiotic growth selection when compared to controls. Recombinant clones were selected and analyzed by PCR and restriction digests to determine whether the insertion sequence had been acquired. A retrieval vector was constructed by PCR-amplifying sequences corresponding to Fragments A and D (Figure 1) from mouse Foxd3 BAC DNA with primers that also added restriction enzyme sites (XbaI and SpeI for Fragment A, SacI and SacII for Fragment B) and cloning them into a modified pBluescript vector containing a diphtheria-toxin cassette (pBS.DT-A, a gift from the laboratory of Dr. Mark Magnuson). We then electroporated some of these clones with this retrieval vector containing homology fragments A and D, approximately 8.5 kb and 4.6 kb from the Foxd3 start and stop codons, respectively. Recombinant clones were selected by acquisition of antibiotic resistance. However, because the first recombineering step did not generate clones with the full desired sequence, even successful recombination at this step could not result in clones with the complete target sequence.
Annotation
The adenine nucleotide of the Foxd3 ATG start codon was designated position 1. Upstream (5’) nucleotides were assigned positions in negative numbers relating their distance from the Foxd3 ATG, whereas downstream (3’) nucleotides were designated by positive numbers indicating distance from the first A of the Foxd3 ATG.
Conservation analysis
NCBI Basic Local Alignment Search Tool (BLAST) analysis (http://blast.ncbi.nlm.nih.gov/Blast.cgi) 17 was performed independently for the 370 nt region of polymerase resistance from the murine genome and the two flanking stretches of 370 nt. The blastn algorithm for somewhat similar sequences was used to query the nr/nt nucleotide collection. Maximum alignment scores were reported for the top vertebrate species. In the case where scores were not given, they are listed as “below threshold”. DCODE ECR browser (http://ecrbrowser.dcode.org) analysis 18 was performed individually on the mouse Foxd3 (July 2007 mouse genome assembly – NCBI build 37/mm9) and human FOXD3 (March 2006 human genome assembly – NCBI build 36.1/hg18) loci. Within the ECR browser, pairwise alignments for conservation analysis were selected for 11 available genomes: chimpanzee (Pan troglodytes), rhesus monkey (Macaca mulatta), mouse (Mus musculus), human (Homo sapiens), rat (Rattus norvegicus), dog (Canis familiaris), cow (Bos taurus), opossum (Monodelphis domestica), chicken (Gallus gallus), frog (Xenopus laevis), zebrafish (Danio rerio) and fugu pufferfish (Takifugu rubripes). For comparison with the mouse genome, the ECR browser window covered an interval of 13,869 nt, from positions 99313783 to 99327651 on mouse chromosome 4 (the Foxd3 transcribed region, including the 5’ and 3’ UTRs, is 99322990 to 99325362). For comparison with the human FOXD3 locus, the ECR browser window covered an interval of 13,786 nt, from positions 63552012 to 63565798 on human Chromosome 1 (FOXD3 transcribed region is 63561318-63563385). Parameters used for analysis and display included an ECR length of 100, ECR similarity of 90, layer height of 55 and a relative coordinate system. Tracks from human ENCODE data aligned to the same interval of FOXD3 (Chromosome 1: 63552012 to 63565798) from the March 2006 assembly of the human genome were visualized within the UCSC genome browser (http://genome.ucsc.edu/cgi-bin/hgGateway). Tracks for H3K4 trimethylation and DNaseI hypersensitivity were selected from the Expression and Regulation track group. H3K4me3 data was obtained from ChIP-Seq experiments in GM12878 and K562 cell lines, while DNaseI hypersensitivity data displayed was from DNase-seq experiments in GM12878 cells; both indicate regions of open chromatin20,21.
Prediction of secondary structure
RNAFOLD analysis was performed with the Vienna RNAfold Webserver (http://rna.tbi.univie.ac.at/cgi-bin/RNAfold.cgi) 11 on the 1110 nt upstream region of the mouse Foxd3 locus comprised of the 370 nt polymerase-resistant region and 370 nt flanking sequence on either side. Input parameters were entered for a linear DNA molecule (DNA parameters from 2004 David Matthews model) at 72 degrees Celsius allowing for dangling energies on both sides of a helix in any case, using minimum free energy (MFE) and partition function fold algorithms 22. The output was visualized as an interactive secondary structure plot. Additional analysis of secondary structure was performed with the UNAFold program for two-state melting/folding located on the mfold/UNAFold webserver (http://mfold.rna.albany.edu/) 23. UNAFold parameters were set for DNA at 72 degrees Celsius, 1M Na+, 0M Mg2+ and for a linear molecule. Due to program limits, only 1000 nucleotides (rather than 1110) were entered; this sequence was the 370 nt region of interest plus 315 nucleotides of flanking sequence on either side.
Construction of plasmids
A 10.6 kb Foxd3 plasmid consisting of a backbone vector and an approximately 7.6 kb EcoRI genomic Foxd3 fragment (containing the entire Foxd3 coding sequence plus surrounding 5’ and 3’ genomic sequence) was digested with the enzyme BspEI, resulting in fragments of approximately 9.6, 0.9 and 0.1 kb that were separated using agarose gel electrophoresis. The 0.9 kb fragment spanned most (348 of 370 nt) of the predicted hairpin sequence and additional downstream sequence. The 9.6 kb band was gel purified and this linear molecule with BspEI ends ligated to form a Foxd3 plasmid missing most of the predicted hairpin. The 0.9 kb fragment containing most of the hairpin was also purified and ligated into the EcoRV site of pBluescript forming a plasmid that contained most of the hairpin.
References
Lee, E. C. et al. A highly efficient Escherichia coli-based chromosome engineering system adapted for recombinogenic targeting and subcloning of BAC DNA. Genomics 73, 56–65 (2001).
Liu, P., Jenkins, N. A. & Copeland, N. G. A highly efficient recombineering-based method for generating conditional knockout mutations. Genome Res 13, 476–484 (2003).
Testa, G. et al. Engineering the mouse genome with bacterial artificial chromosomes to create multipurpose alleles. Nat Biotechnol 21, 443–447 (2003).
Warming, S., Costantino, N., Court, D. L., Jenkins, N. A. & Copeland, N. G. Simple and highly efficient BAC recombineering using galK selection. Nucleic Acids Res 33, e36 (2005).
Gough, J. A. and Murray, N. E. Sequence diversity among related genes for recognition of specific targets in DNA molecules. J. Mol. Biol. 166, 1–19 (1983).
Kieleczawa, J. & Mazaika, E. Optimization of protocol for sequencing of difficult templates. J Biomol Tech 21, 97–102 (2010).
Mizusawa, S., Nishimura, S. & Seela, F. Improvement of the dideoxy chain termination method of DNA sequencing by use of deoxy-7-deazaguanosine triphosphate in place of dGTP. Nucleic Acids Res 14, 1319–1324 (1986).
Ranu, R. S. Relief of DNA polymerase stop(s) due to severity of secondary structure of single-stranded DNA template during DNA sequencing. Anal Biochem 217, 158–161 (1994).
Winship, P. R. An improved method for directly sequencing PCR amplified material using dimethyl sulphoxide. Nucleic Acids Res 17, 1266 (1989).
Weaver, D. T. & DePamphilis, M. L. Specific sequences in native DNA that arrest synthesis by DNA polymerase alpha. J Biol Chem 257, 2075–2086 (1982).
Gruber, A. R., Lorenz, R., Bernhart, S. H., Neubock, R. & Hofacker, I. L. The Vienna RNA websuite. Nucleic Acids Res 36, W70–74 (2008).
Huppert, J. L. Structure, location and interactions of G-quadruplexes. FEBS J 277, 3452–3458 (2010).
Kowalczykowski, S. C., Dixon, D. A., Eggleston, A. K., Lauder, S. D. & Rehrauer, W. M. Biochemistry of homologous recombination in Escherichia coli. Microbiol Rev 58, 401–465 (1994).
Hanna, L. A., Foreman, R. K., Tarasenko, I. A., Kessler, D. S. & Labosky, P. A. Requirement for Foxd3 in maintaining pluripotent cells of the early mouse embryo. Genes Dev 16, 2650–2661 (2002).
Teng, L., Mundell, N. A., Frist, A. Y., Wang, Q. & Labosky, P. A. Requirement for Foxd3 in the maintenance of neural crest progenitors. Development 135, 1615–1624 (2008).
Osoegawa, K. et al. Bacterial artificial chromosome libraries for mouse sequencing and functional analysis. Genome Res 10, 116–128 (2000).
Labosky, P. A., Barlow, D. P. & Hogan, B. L. Mouse embryonic germ (EG) cell lines: transmission through the germline and differences in the methylation imprint of insulin-like growth factor 2 receptor (Igf2r) gene compared with embryonic stem (ES) cell lines. Development 120, 3197–3204 (1994).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25, 3389–3402 (1997).
Ovcharenko, I., Nobrega, M. A., Loots, G. G. & Stubbs, L. ECR Browser: a tool for visualizing and accessing data from comparisons of multiple vertebrate genomes. Nucleic Acids Res 32, W280–286 (2004).
Ku, M. et al. Genomewide analysis of PRC1 and PRC2 occupancy identifies two classes of bivalent domains. PLoS Genet 4, e1000242 (2008).
Boyle, A. P. et al. High-resolution mapping and characterization of open chromatin across the genome. Cell 132, 311–322 (2008).
Zuker, M. & Stiegler, P. Optimal computer folding of large RNA sequences using thermodynamics and auxiliary information. Nucleic Acids Res 9, 133–148 (1981).
Markham, N. R. & Zuker, M. UNAFold: software for nucleic acid folding and hybridization. Methods Mol Biol 453, 3–31 (2008).
Acknowledgements
We are grateful for discussion and support from all of the members of the Labosky lab. We would like to thank Drs. Doug Mortlock and Mark Magnuson for BAC recombineering advice and protocols and the VUMC DNA sequencing core facility for timely and efficient analysis of submitted samples. We would also like to thank Drs. William Tansey and Doug Mortlock for initial comments on the manuscript. This work was funded by grants from the following institutions: NIH - R01 HD036720 to PAL and T32 HD007043 to BLN, American Heart Association - Postdoctoral Fellowship 0725615B to BLN and by Vanderbilt University Medical Center Academic Program Support.
Author information
Authors and Affiliations
Contributions
B.L.N. conceived and performed the experiments and together with P.A.L. carried out data analysis. B.L.N. and P.A.L. co-wrote the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareALike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Nelms, B., Labosky, P. A predicted hairpin cluster correlates with barriers to PCR, sequencing and possibly BAC recombineering. Sci Rep 1, 106 (2011). https://doi.org/10.1038/srep00106
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep00106
This article is cited by
-
Towards practical and robust DNA-based data archiving using the yin–yang codec system
Nature Computational Science (2022)
-
On conflict free DNA codes
Cryptography and Communications (2021)
-
A Characterization of the DNA Data Storage Channel
Scientific Reports (2019)
-
BRAF mutation-specific promoter methylation of FOX genes in colorectal cancer
Clinical Epigenetics (2013)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.