Abstract
Cork oak (Quercus suber) is native to southwest Europe and northwest Africa where it plays a crucial environmental and economical role. To tackle the cork oak production and industrial challenges, advanced research is imperative but dependent on the availability of a sequenced genome. To address this, we produced the first draft version of the cork oak genome. We followed a de novo assembly strategy based on high-throughput sequence data, which generated a draft genome comprising 23,347 scaffolds and 953.3 Mb in size. A total of 79,752 genes and 83,814 transcripts were predicted, including 33,658 high-confidence genes. An InterPro signature assignment was detected for 69,218 transcripts, which represented 82.6% of the total. Validation studies demonstrated the genome assembly and annotation completeness and highlighted the usefulness of the draft genome for read mapping of high-throughput sequence data generated using different protocols. All data generated is available through the public databases where it was deposited, being therefore ready to use by the academic and industry communities working on cork oak and/or related species.
Design Type(s) | genome assembly |
Measurement Type(s) | whole genome sequencing assay • sequence annotation |
Technology Type(s) | DNA sequencing • RNA sequencing |
Factor Type(s) | |
Sample Characteristic(s) | Quercus suber • pollen • leaf • xylem • bark • phellem |
Machine-accessible metadata file describing the reported data (ISA-Tab format)
Similar content being viewed by others
Background & Summary
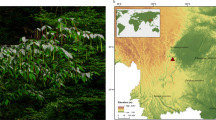
Quercus suber is an evergreen tree, commonly known as cork oak, which is native to the western Mediterranean Basin, especially southwest Europe, where it occurs in the coastal regions. Cork oak has the rare characteristic of producing a continuous and renewable cork layer, which has fine physical and chemical properties that make it highly profitable for industrial uses.
The cork oak is the basis of an ecological system, known as “montado” that is unique in the world, contributing to the survival of many native species of fauna and flora and preventing desertification in vulnerable areas. Over the last few years, a decline in cork oak populations has been observed, eventually as a result of agriculture intensification, biotic stresses, fires and climate changes. If this trend is not stopped, serious negative impacts may arise at the ecosystem, social and economic levels.
The availability of a fully sequenced and annotated genome is essential to support many of the studies needed to answer fundamental questions about cork oak biology and interactions with the environment, and about cork formation/production, as well as to develop the tools required to address the problems presently affecting this species. In fact, despite the economic relevance of cork production, no genetic breeding/selection schemes have so far been established in this species, thus compromising the development of genetically superior trees, able to produce high quality cork, displaying enhanced resistance to the (a)biotic stress factors they are exposed to and with potential to adapt to new forestry management practices that may be implemented in the future.
The development of genetic markers in cork oak is an extremely important field, which will greatly benefit from the availability of a reference genome. For example, cork quality, perhaps the most important trait in cork oak production, cannot be properly evaluated before at least 40 years of growth, due to the specificities of the production cycle. This extremely long cycle represents a textbook scenario in which genetic markers associated with better cork quality can be highly beneficial. The same reasoning can be applied to other relevant traits, such as resistance to drought, diseases, pests, or developmental modifications important to face climate changes.
A comprehensive characterization of the cork oak transcriptome was previously produced1, which increased the amount of genomic resources available for cork oak and allowed transcriptomic analyses of several biological systems, such as the response to ectomycorrhizal symbiosis establishment2, response to drought3 and flower and fruit development4,5.
To assist and enhance the development of the solutions needed for the future of cork oak production we have generated a draft version of the cork oak genome (Data Citation 1 and Data Citation 2).
The genome size of cork oak, a diploid (2n=24) species, was estimated, using flow cytometry, to be 934 Mb6. In the present study we used a combination of Paired-End (PE) and Mate-Pair (MP) libraries sequenced using the Illumina platform to generate a draft genome assembly with an estimated genome size of 953.3 Mb, which is a very close match to the previous estimate. The bioinformatics pipeline involved a de novo genome assembly step, followed by scaffolding, gap filling and removal of heterozygous regions. The cork oak draft genome is distributed over 23,344 scaffolds, even though the vast majority of the assembly is represented in a considerable smaller number of larger scaffolds (approximately 94.6% of the assembled genome present in the 4,730 scaffolds longer than 10,000 bp).
The structural annotation of the genome yielded 79,752 genes, with complete open reading frames, and 83,814 transcripts. The number of transcripts with a valid functional annotation varied with the database used, and the maximum number was 69,218, when searching against InterPro signatures, which represented 82.6% of the total. Finally, using a validation approach based on the RNA-Seq data available for five cork oak tissues, a total of 33,658 predicted genes could be confirmed and classified as high confidence genes, since they presented assembled transcripts within the genome annotation coordinates.
Methods
Selection of the target individual for genome sequencing
Interspecific hybridization is a widespread phenomenon in plants, which also occurs among species of the Quercus genus. In fact, hybridization between Quercus suber and Quercus Ilex, mainly Quercus ilex subsp. rotundifolia, another species widely distributed in the Mediterranean region, is fairly common, since both are found in mixed stands where they cohabit. Even though some of these hybrids maintain the ability to produce cork, only a pure cork oak tree can produce high quality cork. Thus, in order to select the tree to sequence, a total of 28 trees, derived from four different locations, were selected, based on historical records that demonstrated their ability to produce very high quality cork. These individuals fulfilled an estimated median age of more than 50 years, were at least 50 meters apart (to reduce chances of selecting possibly related individuals), and were not planted in rows, to ensure that they were naturally grown trees. Leaves were collected from these trees for DNA extraction. Then, all individuals were genotyped with 16 microsatellite markers, to estimate the degree of homozygosity, since a higher level is preferable for a genome sequencing project. The microsatellite markers used were selected from available literature, and included markers developed in Quercus mongolica7, Quercus petraea8, Quercus robur9, Quercus macrocarpa10, Quercus myrsinifolia11, Castanea sativa12 and two additional markers developed for cork oak at CEBAL and INIAV (published here). This information is summarized in Table 1.
Multiplex PCR amplification was conducted in a GeneAmp PCR system 2700 (Applied Biosystems) and PCR products were analyzed on an ABI Prism 310 capillary electrophoresis system (Applied Biosystems). Size alignment and quality control were analyzed on Genescan 3.7 (Applied Biosystems). The number of homozygous loci per tree ranged from four to nine, the latter being observed in three cork oaks located at Herdade dos Leitões (Montargil, Ponte de Sor, Portugal). From these three individuals the HL8 cork oak was considered the most robust tree and selected for genome sequencing.
Extraction of nucleic acids and high-throughput sequencing
Leaves collected from the HL8 cork oak tree were used for DNA extraction. To reduce contamination with chloroplast and mitochondrial DNA nuclei were first isolated using the CelLytic™ PN Isolation/Extraction Kit (Sigma) and nuclear DNA was then extracted using the innuPREP Plant DNA Kit (AnalytiK Jena).
All Illumina DNA sequencing was performed at the Beijing Genomics Institute (BGI), and included a combination of PE and MP libraries (Data Citation 2), sequenced using different Illumina sequencers (Table 2).
A total of 10,560,988,448 reads were generated. The raw reads were filtered by removing adapter sequences and reads containing undetermined nucleotides (N’s). The reads that remained in the dataset were further processed with Sickle13, which trimmed/removed the low quality reads, using as parameters minimum quality (equal or larger than 20) and minimum length (80% of the read length). To remove reads derived from the chloroplast and mitochondrial genomes the high quality reads were mapped, with BWA using the BWA-mem algorithm14, against eight mitochondrial and eight chloroplast genomes. These genomes were selected from the species more closely related to cork oak for which these organelle genomes were available at the time this step was executed. The NCBI Reference Sequence numbers for each of these genomes are indicated in Table 3.
The reads that did not map against any of the 16 genomes were kept and formed the set of Illumina DNA data that was used for all downstream analyses, which comprised 8,544,112,336 reads (80.9% of the initial number of reads). Fig. 1 provides a detailed description of the pre-processing workflow.
A total of five tissues were sampled from the HL8 cork oak tree, in order to generate the RNA-Seq data (Data Citation 2) needed for the genome annotation stage (Table 4). These tissues included pollen, leaf, xylem, inner bark and phellem. Total RNA was extracted from all tissues using the Plant/fungi Total RNA purification kit (Norgen). All Illumina RNA-Sequencing was performed using the HiSeq 4000 system, with a read length of 100 nucleotides and a paired-end sequencing protocol.
A total of 1,530,447,601 RNA-Seq reads were produced. These reads were further processed using a similar approach to the one described for the DNA sequence data, which included the application of the same procedures for the removal of reads with adapters or undetermined nucleotides and trimming of the reads. The number of reads kept amounted to 1,438,136,157, which represented 94% of the initial number of reads.
Cork oak genome properties
A k-mer analysis of the PE libraries with read length of 100 bp was performed to characterize the genome sequence using Jellyfish15 with a k length of 23. The k-mer spectrum followed a binomial distribution expected for a heterozygous diploid genome (Fig. 2). The homozygous peak (right peak) was observed at 109x, while the heterozygous peak (left peak) was observed at 55x. There were noticeably more heterozygous k-mers than homozygous k-mers, which indicated a high level of heterozygosity. The diploid genome size was estimated following the approach described by Sork and colleagues16, which included (1) dividing all k-mers under the haploid peak, except the error k-mers (all k-mers represented with a single read), by the haploid coverage depth, (2) dividing all k-mers counted between the haploid coverage peak +1 and the diploid coverage peak by the diploid coverage depth, and (3) summing those values. The estimated cork oak genome size based on sequence data was 968.7 Mb, a value that slightly exceeds the genome size estimated by flow cytometry (934 Mb)6.
Additionally, GenomeScope17 was also used to estimate genome size and heterozygosity, using the k-mer distribution determined with a k-mer size of 23, which provided estimates of 1.06 Gb and 1.62% for genome size and heterozygosity, respectively.
Genome assembly
The Illumina reads were used to generate the genome assembly, a process that was performed in two stages, due to the different read length of the PE libraries. Hence, two independent genome assemblies were produced, one for each type of read length, using Ray18, the assembler that was selected because it provided the best results (less contigs, larger genome assembly, larger N50) after testing several assembly tools. The k-mer sizes used in the assembly were 81 and 121, for the PE libraries with a read length of 100 bp and 150 bp, respectively. The best k-mer size was estimated with SGA preqc19.
The contigs that resulted from each assembly were integrated into a single assembly using GARM20, in order to obtain a set of contigs common to both individual assemblies and identify the unique contigs from each assembly.
The latter set of contigs was used as the reference on which all PE reads were mapped using BWA with the BWA-mem algorithm14, in order to differentiate the contigs representing valid genome regions from the erroneous contigs possibly derived from the assembly step. The mapping results were analyzed for coverage using BEDtools21. We considered as valid all the contigs which displayed a minimum length of 1,000 bp, at least 95% of the length covered with mapped reads, and that 95% of the regions with mapped reads had a minimum coverage of 50x. The contigs that passed these filters were added to the set of contigs common to both assemblies, which formed the assembly used to proceed with scaffolding of the genome. The total number of contigs that were used as the reference for the scaffolding step was 168,041, and comprised 939,042,321 bp of sequence (Table 5).
The mate-pair read dataset was mapped to the assembly using BWA with the BWA-mem algorithm14. The mapped reads were filtered for quality (minimum quality score of 10) and subsequently used for scaffolding the genome using BESST22, using the mate-pair libraries in ascending order relative to their respective insert size. In order to reduce the number of gaps two rounds of gap closing were performed using SOAP de novo Gap Closer23,using in each round a different set of two PE libraries from all available insert sizes (170 bp, 300 bp, 500 bp, 800 bp). Given the high heterozygosity of the cork oak genome, a final step targeting the removal of possible alternative heterozygous scaffolds was performed, using Redundans24, on an assembly that contained 44,287 scaffolds and comprised 1,000,388,124 bp. A total of 47.1 Mb of assembled sequence was removed, which represented a 4.7% decrease, while 20,943 scaffolds were discarded, a decrease of 47.3%. These results represent an approximate haplotype duplication level for the cork oak draft genome sequence.
The final assembly of the cork oak draft genome contained 23,344 scaffolds greater than 1,000 bp, which represented an assembly length of 953.3 Mb (Table 6). The percentage of undetermined nucleotides was 2%. The vast majority of the assembled genome was contained in the larger scaffolds. For instance, approximately 94.6% of the assembled genome was present in the 4,730 scaffolds longer than 10,000 bp. Similarly, the 2,022 scaffolds with a minimum length of 100 Kb contained 823.7 Mb of the genome, which represented 86.4%. These were clear indications that the majority of the fragmentation observed in the draft genome was related to shorter scaffolds, which may be due to the high heterozygosity and unsolved repeats of the cork oak genome. The N50 observed was 465.2 Kb, while the longest scaffold was 2,284,287 bp in length.
Genome annotation
Augustus25 was employed to predict gene models in cork oak. To achieve best performance, species-specific parameters were generated by training Augustus with a subset of cork oak genes. To generate this subset, Maker26 was run with transcripts obtained from the five HL8 RNA-Seq tissues assembled with a Star27 and StringTie28 pipeline and protein sequences from Arabidopsis thaliana. The run with Maker was stopped when a reasonable number of genes had been predicted, which resulted in a total of 1,268 genes selected to train Augustus. In order to maximize annotation accuracy several sources of external hints were produced to enhance the Augustus gene predictions. Initially, a custom repeat library was generated with RepeatModeler29, which was then used to perform the repeat masking of the genome with RepeatMasker30. Augustus repeat hints were obtained based on the RepeatMasker output. In addition, protein hints were generated based on the genome coordinates where CEGMA31 conserved proteins mapped (please check the section “Technical Validation”). Exonic and intronic hints were produced based on the STAR mappings of the HL8 RNA-Seq libraries from four tissues (leaf, xylem, inner bark, phellem) using Augustus utilities (bam2hints, wig2hints). Finally, Augustus was run by adding the option to also report alternative transcripts when they were suggested by hints.
The number of genes annotated for the cork oak draft genome was 79,752. The final list included only the genes for which a start and stop codon was detected. Additionally, a total of 83,814 transcripts were identified, an indication of the presence of alternative splicing events in the cork oak transcriptome.
Functional annotation was performed for the set of 83,814 transcripts using different sources of information. Detection of homologies in the predicted gene models was performed using BLASTP against the databases NCBI-nr and Swiss-Prot32. Additionally, eggNOG-mapper33 was used to assign orthologies based on precomputed phylogenies of Viridiplantae organisms from the eggNOG database34. Last, InterProscan35 was used to detect protein domains, Gene Ontology terms and KEGG mappings from the InterPro database36.
The percentage of functionally annotated transcripts varied from 55.6% to 82.5%, for the SwissProt and InterPro databases, respectively (Table 7). The percentage of transcripts with a valid functional annotation is highly dependent on the information deposited in the publicly available databases used in this type of analyses. Moreover, considering that cork oak is a species with unique biological features, it is also possible that some genes unique to cork oak were identified. Consequently, these genes did not have any matches in the databases used, and the characterization of their function will require additional studies specifically targeting this goal. Overall, a total of 40,599 transcripts (37,724 genes) were annotated in all databases, which is a promising number, especially when considering that the SwissProt database contains manually curated protein sequences.
The percentage of the genome covered by repeat elements was 11.96%, for a total of 113,971,428 bp of the 953.3 Mb contained in the assembled cork oak draft genome (934.2 Mb after excluding the regions with Ns). These results were obtained running RepeatMasker using the eudicotyledons subset of RepBase, and are summarized in Table 8. Matches for retroelements accounted for 7.59%, while the percentage of DNA transposons and simple repeats was 0.74 and 2.72, respectively. These results are quite similar to the ones obtained for the valley oak genome16, and lower than the pedunculated oak genome, which appears to contain approximately 33% of repetitive elements37.
Code Availability
The execution of this work involved using many software tools, whose versions, settings and parameters are described below.
1) Sickle: minimum quality (equal or larger than 20); minimum length (80% of the read length); 2) BWA: version 0.7.15, default parameters; 3) Jellyfish: version 2.2.6, k-mer size of 23; 4) GenomeScope: parameters used were k-mer length 23; read length 80; maximum k-mer coverage 1000; 5) Ray: version 2.3.1, default parameters, k-mer size of 81 for the assembly with the PE100 reads, k-mer size of 121 for the assembly with the PE150 reads; 6) SGA: version 0.10.14, default parameters; 7) GARM: version 0.7.5, default parameters; 8) BEDtools: version 2.25.0; 9) BESST: version 2.2.5, default parameters; 10) Gap Closer: version 1.12, parameters used were -l 150, in configFile: asm_flags=4; PE100 lib: rd_len_cutoff=90, pair_num_cutoff=7, map_len=45; PE150 libs: rd_len_cutoff=140, pair_num_cutoff=10, map_len=50; 11) Redundans: version 0.13a, with default parameters for homology, --nogapclosing, --noscaffolding, --norearrangements; 12) Augustus: version 3.2.2, parameters: (--protein=on --codingseq=on --cds=on --introns=on -genemodel=partial --softmasking=1 --UTR=off --alternatives-from-evidence=true --uniqueGeneId=true; 13) Maker: version 3.0.0 beta, parameters: est=merged transcripts from stringtie assemblies for 5 tissues, protein=arabidopsis_proteins, rmlib=repeatModelerOutput, est2genome=1, protein2genome=1; run was stopped when 2,574 scaffolds had been scanned, which predicted 2,394 genes genes, subsequently used to train Augustus; 14) Star: version 2.5.2b, parameters: --outSAMattributes All --outFilterIntronMotifs RemoveNoncanonical --outWigType wiggle --outSAMtype BAM SortedByCoordinate --chimSegmentMin 20 --outReadsUnmapped Fastx --outFilterScoreMinOverLread 0.4 --outFilterMatchNminOverLread 0.4 --outFilterMatchNmin 60; 15) StringTie: version 1.3.1b; parameters for first transcriptome assemblies for maker: default parameters; merge (samples per tissue and reference per tissue: -F 0.5 -T 0.5 -i); transcriptome assemblies for annotation validation: -c 5 -j 1.5 -a 20 -m 400 -f 0.2; 16) RepeatModeler: RepeatModeler-open-1.0.8; 17) RepeatMasker: RepeatMasker-open-4.0.6 (with specific specific cork oak library generated with repeatModeler using the scaffolds>5000 bp); 18) CEGMA: version2.5; 19) Blastp: version 2.6.0; 20) NCBI-nr: we used the subset of green plant proteins extracted using taxonomic ID 33090, downloaded on 11th January 2017, for a total of 7,903,686 entries; 21) SwissProt: we used the database version downloaded on 15th March 2017, for a total 593,119 entries, including isoforms; 22) eggNOG mapper: version 0.12.7; 23) eggnog: version 4.5; 24) InterProscan: version 5.22-61.0, with parameters -goterms, --pathways and using an –iprlookup; 25) BUSCO: version 1.22; 26) GMAP: version 2017-06-20, parameters: -D, -S, -f; 27) LAST: version 914.
Data Records
This Whole Genome Shotgun project has been deposited at DDBJ/ENA/GenBank (Data Citation 1). Raw read files are available at NCBI Sequence Read Archive (Data Citation 2).
Technical Validation
Assessing the completeness of the genome assembly and annotation
We used CEGMA31 and BUSCO38, tools that investigate the presence of highly conserved orthologous genes in a genome assembly, to perform a first evaluation of the quality of the cork oak draft genome. CEGMA contains a total of 248 core eukaryotic genes. BUSCO was run over the plant set, which contains a total of 956 orthologue groups, defining Arabidopsis as the model species for the gene prediction performed by Augustus within the BUSCO pipeline. The results showed that the vast majority of the core genes/orthologues was present in the genome assembly (98.8% CEGMA, 95.6% BUSCO), and with complete matches (98.0% CEGMA, 94.9% BUSCO), which evidenced a good quality of the draft genome, in particular of the gene content and transcriptome. The number of complete orthologs found to be duplicated using BUSCO was 130, which represented 13.6% of the total (14.3% of the complete orthologs), percentages that are lower than the ones determined for the valley16 and pedunculated37genomes (52 and 49%, respectively).
Structural annotation validation with RNA-Seq data
Predicted gene models were evaluated by running Stringtie over the RNA-Seq libraries, using the Augustus annotation as the reference and more stringent settings for the identification of transcripts. High confidence tags were assigned to the reference genes for which there were assembled transcripts within the genome coordinates determined during the annotation step. A total of 33,658 predicted genes were confirmed as high confidence genes with the analysis of the RNA-Seq data, which represented 42.2% of the 79,752 predicted genes. It should be emphasized that this validation procedure was limited to the genes expressed in the five tissues for which RNA-Seq data was available. Hence, assuming that transcriptomic data derived from different cork oak tissues and organs will likely contain a distinct set of expressed genes, it is reasonable to assume that more genes would be validated, emphasizing the quality of the genome annotation produced.
Read mapping against the cork oak draft genome
The availability of several sets of cork oak high-throughput sequence data, produced with different types of sequencing protocols and in diverse individuals, enabled investigating the performance of the draft genome sequence in read mapping experiments, since these represent the analysis step expected to be most commonly used by other researchers. The available reads were retrieved from NCBI’s Sequence Read Archive (SRA) and were derived from RNA-Seq (generated in the 454 and Illumina platforms) and small RNA-Seq experiments, including the response of cork oak roots to drought3, the transcriptomic analysis of male and female flowers4, the comparison of good and bad quality cork samples39, the response of oak roots to the establishment of ectomycorrhizal symbiosis2, the dynamics of cork oak somatic embryogenesis (unpublished results) and miRNA profiling in leaf and cork tissues40. Additionally, the RNA-Seq reads derived from the five cork oak tissues were also used, even though this read dataset was derived from samples collected in the tree (HL8) used for genome sequencing. The Illumina RNA-Seq reads were mapped to the draft genome using Star, while GMAP41 and BWA-mem were used to map the RNA-Seq 454 and small RNA-Seq reads, respectively. Since the Illumina reads were produced using a paired-end sequencing protocol, the number of mapped reads was defined as the reads that were mapped as proper pairs, following the information specific to each PE library, while for the 454 reads only the reads that mapped to a unique genome location were considered. The percentage of mapped reads ranged from 65.6% to 89.6%, for the SRR1012034 and SRX2239662datasets, respectively (Table 9).
Comparison with other Quercus genomes
Recently, the draft genome sequences for two other species of the Quercus genus were released, which included pedunculated oak (Quercus robur)37 and valley oak (Quercus lobata)16. A comparison of the assembly metrics of the cork oak draft genome with these two genomes indicated a similar number of scaffolds longer than 2 kb (15,058 cork oak; 17,910 pedunculate oak; 18,512 valley oak), even though the total genome assembly included in these scaffolds differed between species (941.0 Mb cork oak; 1.34 Gb pedunculated oak; 1.15 Gb valley oak). The N50 scaffold size of the cork oak assembly was 465.2 kb, while for pedunculated oak and valley oak the N50 values were 260 kb and 278.1 kb, respectively. Moreover, the heterozygosity estimates for cork oak (1.62%) and valley oak (1.25%) indicate that Quercus genomes indeed display high levels of heterozygosity. Additionally, the similarity level between the cork oak and the other two Quercus genomes, defined as the percentage of the cork oak genome that aligned with the other Quercus genomes, was determined aligning the valley and pedunculated oak genomes to the cork oak genome, using LAST42. The results showed that at the nucleotide level the similarity between the cork oak and valley and pedunculated oak was estimated to be 36.7 and 58.7%, respectively.
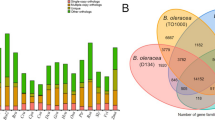
As this analysis did not discriminate between gene and non-gene space, a similar analysis was performed using the set of BUSCO orthologs found in each of the Quercus species. BUSCO was run with the same parameters described before over the valley and pedunculated oak genomes, in order to obtain the set of orthologs identified within their genomes. For this analysis two sets of orthologs were used, which included the orthologs found in common between cork oak and valley oak and the ones found in common between cork oak and pedunculated oak. In both cases, only the orthologs defined as “Complete” and “Duplicated” were used. As the number of copies of the orthologs between the three species was different only the largest set for each ortholog identifier was considered for the analysis. The final set for the Quercus suber vs. Quercus lobata comparison contained a total of 880 orthologs, while the set of Quercus suber vs. Quercus robur contained a total of 890 orthologs. After aligning the valley and pedunculated oak orthologs against the ones from cork oak, the similarity at the nucleotide level was estimated to be 78.8% and 81.8%, respectively. Additionally, the orthologs in common between valley and pedunculated oaks (895) were also aligned using the valley oak as a reference, which resulted in an estimated similarity of 85.3%. Previously, the identity between the valley and pedunculated oak genomes was determined to be 93.1% (ref. 16). When focusing on the set of conserved orthologs identified by BUSCO, the three Quercus species display a high percentage of similarity between them. While the valley and pedunculated oaks present more similarity between them, cork oak is genetically more distant from these two species.
Even though the unique genome characteristics of each species must be considered, the differences observed on the genome assembly metrics were not substantial and may be related to the sequence datasets and assembly strategies followed in each study. A more comprehensive comparison will be possible once higher quality genome assemblies for these, and possibly other, Quercus species become available.
Final considerations
These results obtained in this study fully validate the usefulness of the assembled cork oak draft genome for several different types of genomic studies in this species. These resources will help cork oak becoming a useful model for studying Quercus species, for instance for genome evolution studies, and it will also greatly enhance the development and execution of current and novel research lines tackling the main concerns regarding cork oak and the cork industry. Additional work, carried by our research team, is currently underway, in order to develop an improved version of the cork oak genome and generate a detailed molecular characterization of the mechanisms underlying cork formation.
Additional information
How to cite this article: Ramos, A. M. et al. The draft genome sequence of cork oak. Sci. Data 5:180069 doi: 10.1038/sdata.2018.69 (2017).
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
References
Pereira-Leal, J. B. et al. A comprehensive assessment of the transcriptome of cork oak (Quercus suber) through EST sequencing. BMC Genomics 15, 371 (2014).
Sebastiana, M. et al. Oak root response to ectomycorrhizal symbiosis establishment: RNA-seq derived transcript identification and expression profiling. PLoS ONE 9, e98376 (2014).
Magalhães, A. P. et al. RNA-seq and gene network analysis uncover activation of an ABA-dependent signalosome during the cork oak root response to drought. Front. Plant Sci. 6, 1195 (2016).
Rocheta, M. et al. Comparative transcriptomic analysis of male and female flowers of monoecious Quercus suber. Front. Plant Sci. 5, 599 (2014).
Miguel, A. et al. Characterization of the cork oak transcriptome dynamics during acorn development. BMC Plant Biol. 15, 158 (2015).
Zoldos, V., Papes, D., Brown, S. C., Panaud, O. & Siljak-Yakovlev, S. Genome size and base composition of seven Quercus species: inter- and intra-population variation. Genome 41, 162–168 (1998).
Ueno, S. & Tsumura, Y. Development of ten microsatellite markers for Quercus mongolica var. crispula by database mining. Conserv. Genet. 9, 1083–1085 (2008).
Steinkellner, H. et al. Identification and characterization of (GA/CT)n-microsatellite loci from Quercus petraea. Plant Mol. Biol. 33, 1093–1096 (1997).
Kampfer, S., Lexer, C., Glössl, J. & Steinkellner, H. Characterization of (GA)n Microsatellite Loci from Quercus Robur. Hereditas 129, 183–186 (2004).
Dow, B. D., Ashley, M. V. & Howe, H. F. Characterization of highly variable (GA/CT) n microsatellites in the bur oak, Quercus macrocarpa. Theor. Appl. Genet. 91 (1995).
Isagi, Y. & Suhandono, S. PCR primers amplifying microsatellite loci of Quercus myrsinifolia Blume and their conservation between oak species. Mol. Ecol. 6, 897–899 (1997).
Sebastiani, F., Carnevale, S. & Vendramin, G. G. A new set of mono- and dinucleotide chloroplast microsatellites in Fagaceae. Mol. Ecol. Notes 4, 259–261 (2004).
Joshi, N. A. & Fass, J. N. Sickle: A sliding-window, adaptive, quality-based trimming tool for FastQ files (Version 1.33). Github https://github.com/najoshi/sickle (2011).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM (2013).
Marçais, G. & Kingsford, C. A fast, lock-free approach for efficient parallel counting of occurrences of k-mers. Bioinformatics 27, 764–770 (2011).
Sork, V. L. et al. First draft assembly and annotation of the genome of a California endemic oak Quercus lobata Née (Fagaceae). G3 (Bethesda). doi:10.1534/g3.116.030411 (2016).
Vurture, G. W. et al. GenomeScope: Fast reference-free genome profiling from short reads. in Bioinformatics 33, 2202–2204 (2017).
Boisvert, S., Laviolette, F. & Corbeil, J. Ray: Simultaneous assembly of reads from a mix of high-throughput sequencing technologies J. Comput. Biol. 17, 1519–1533 (2010).
Simpson, J. T. & Durbin, R. Efficient de novo assembly of large genomes using compressed data structures. Genome Res. 22, 549–556 (2012).
Soto-Jimenez, L. M., Estrada, K. & Sanchez-Flores, A. GARM: genome assembly, reconciliation and merging pipeline. Curr. Top. Med. Chem. 14, 418–424 (2014).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Sahlin, K., Vezzi, F., Nystedt, B., Lundeberg, J. & Arvestad, L. BESST--efficient scaffolding of large fragmented assemblies. BMC Bioinformatics 15, 281 (2014).
Luo, R. et al. SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler. Gigascience 1, 18 (2012).
Pryszcz, L. P. & Gabaldón, T. Redundans: an assembly pipeline for highly heterozygous genomes. Nucleic Acids Res. 44, e113–e113 (2016).
Stanke, M., Steinkamp, R., Waack, S. & Morgenstern, B. AUGUSTUS: a web server for gene finding in eukaryotes. Nucleic Acids Res. 32, W309–W312 (2004).
Campbell, M. S., Holt, C., Moore, B. & Yandell, M. in Current Protocols in Bioinformatics 48, 4.11.1-4.11.39 (John Wiley & Sons, Inc., 2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 33, 290–295 (2015).
Smit, A.F.A. & Hubley, R. RepeatModeler Open-1.0. 2008-2015. Available at http://www.repeatmasker.org.
Smit, A.F.A., Hubley, R. & Green, P. RepeatMasker Open-4.0. 2013-2015. Available at http://www.repeatmasker.org.
Parra, G., Bradnam, K. & Korf, I. CEGMA: a pipeline to accurately annotate core genes in eukaryotic genomes. Bioinformatics 23, 1061–1067 (2007).
The UniProt Consortium. UniProt: the universal protein knowledgebase. Nucleic Acids Res. 45, D158–D169 (2017).
Huerta-Cepas, J. et al. Fast genome-wide functional annotation through orthology assignment by eggNOG-mapper. Mol. Biol. Evol. 34, 2115–2122 (2017).
Huerta-Cepas, J. et al. eggNOG 4.5: a hierarchical orthology framework with improved functional annotations for eukaryotic, prokaryotic and viral sequences. Nucleic Acids Res. 44, D286–D293 (2016).
Jones, P. et al. InterProScan 5: genome-scale protein function classification. Bioinformatics 30, 1236–1240 (2014).
Finn, R. D. et al. InterPro in 2017—beyond protein family and domain annotations. Nucleic Acids Res 45, D190–D199 (2017).
Plomion, C. et al. Decoding the oak genome: public release of sequence data, assembly, annotation and publication strategies. Mol. Ecol. Resour. 16, 254–265 (2016).
Simão, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212 (2015).
Teixeira, R. T., Fortes, A. M., Pinheiro, C. & Pereira, H. Comparison of good- and bad-quality cork: application of high-throughput sequencing of phellogenic tissue. J. Exp. Bot. 65, 4887–4905 (2014).
Chaves, I., Lin, Y.-C., Pinto-Ricardo, C., Van de Peer, Y. & Miguel, C. miRNA profiling in leaf and cork tissues of Quercus suber reveals novel miRNAs and tissue-specific expression patterns. Tree Genet. Genomes 10, 721–737 (2014).
Wu, T. D. & Watanabe, C. K. GMAP: a genomic mapping and alignment program for mRNA and EST sequences. Bioinformatics 21, 1859–1875 (2005).
Kiełbasa, S. M., Wan, R., Sato, K., Horton, P. & Frith, M. C. Adaptive seeds tame genomic sequence comparison. Genome Res. 21, 487–493 (2011).
Data Citations
GenBank PKMF00000000 (2018)
NCBI Sequence Read Archive SRP111728 (2017)
Acknowledgements
This study was funded by project InAlentejo ALENT-07-0224-FEDER-001754 (Genosuber - Sequenciação do genoma do sobreiro). Financial support for AMR was provided by Investigador FCT (Fundação para a Ciência e a Tecnologia) project IF/01015/2013/CP1209/CT0001. Financial support for PB and AU was provided by projects InAlentejo ALENT-07-0224-FEDER-001754 and IF/01015/2013/CP1209/CT0001. Funding was also provided by FCT project UID/AGR/00115/2013, and the FCT research unit UID/Multi/04551/2013 (GREEN-IT Bioresources for Sustainability). FCT funding of PMB (SFRH/BPD/86742/2012), and the IF grants of NS (IF/01126/2012), IAA (IF/00960/2013) and CM (IF/01168/2013) are also acknowledged. Financial support for ICh was provided by a postdoc scholarship from the EU project ELIXIR-EXCELERATE (H2020, ID 676559).
The authors would like to thank Fundação João Lopes Fernandes for providing all plant material.
Author information
Authors and Affiliations
Contributions
M.M.O., C.P.R., C.F., J.M., F.S., N.S., C.M.M. and S.G. conceived the study. S.G., M.M.O., C.P.R., J.M., F.S. and C.F. contributed to secure funding. F.S., D.M., J.M., M.C.V., F.N., I.C., P.M.B., M.M.O., I.Ch., C.M.M., S.G., L.R. and T.C. identified field material and collected biological samples. F.S., P.M.B., D.M. and J.B.G. performed laboratorial experiments. A.U. and P.B. performed bioinformatics analyses. A.M.R. supervised the bioinformatics analyses. A.M.R., A.U., P.B., P.M.B., M.M.O., C.P.R., C.M.M., S.G., N.S., I.A.A., I.Ch., C.E., J.M. and F.S. discussed and interpreted the results. A.M.R., A.U. and P.B. wrote the manuscript. P.M.B., M.M.O., N.S., C.E., C.F., I.A.A., C.P.R., C.M.M., S.G. and I.Ch. revised the manuscript. A.M.R. and S.G. coordinated the project. These authors are co-second authors: P.M.B., T.C., I.C. and F.S.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
ISA-Tab metadata
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/ The Creative Commons Public Domain Dedication waiver http://creativecommons.org/publicdomain/zero/1.0/ applies to the metadata files made available in this article.
About this article
Cite this article
Ramos, A., Usié, A., Barbosa, P. et al. The draft genome sequence of cork oak. Sci Data 5, 180069 (2018). https://doi.org/10.1038/sdata.2018.69
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/sdata.2018.69
This article is cited by
-
ContScout: sensitive detection and removal of contamination from annotated genomes
Nature Communications (2024)
-
ddRAD-seq generated genomic SNP dataset of Central and Southeast European Turkey oak (Quercus cerris L.) populations
Genetic Resources and Crop Evolution (2024)
-
Genome-wide analysis of the NAC gene family and functional verification of the DcNAC043s in Dendrobium catenatum
Plant Growth Regulation (2024)
-
Key triggers of adaptive genetic variability of sessile oak [Q. petraea (Matt.) Liebl.] from the Balkan refugia: outlier detection and association of SNP loci from ddRAD-seq data
Heredity (2023)
-
Transcriptome Analysis of Persian Oak (Quercus brantii L.) Decline Using RNA-seq Technology
Biochemical Genetics (2023)