Abstract
Non-aureus staphylococci (NAS) are implicated in many healthcare-acquired infections and an understanding of the genetics of antimicrobial resistance is important in relation to both clinical intervention and the role of NAS as a reservoir of resistance genes. Gap statement: The burden of antimicrobial resistance in NAS, particularly to clinically relevant antimicrobials, is under-recognised. We sourced 394 NAS isolates from clinical samples, healthy human volunteers, animals and type cultures and subjected them to minimum inhibitory concentration (MIC) testing by agar dilution using eight antimicrobials. Cefoxitin was used to screen for methicillin resistance, as it stimulates the expression of mecA in S. aureus. We performed whole genome sequencing on 366 isolates and analysed these genotypically for the presence of genetic mechanisms responsible for the phenotypic levels of reduced antimicrobial susceptibility. We observed 175 sequenced isolates with a MIC ≥ 4 µg/ml to cefoxitin, of which 50% did not harbour a known mec homologue. Eight clinical NAS isolates displayed high daptomycin MICs (>4 µg/ml), with no known mechanism identified. Differences in MICs against erythromycin were attributable to the presence of different resistance genes (msrA and ermC). In total, 49% of isolates displayed reduced susceptibility to three or more of the antimicrobials tested. The widespread presence of reduced antimicrobial susceptibility in NAS is concerning. An increased likelihood of harder-to-treat infections caused directly by NAS with acquired resistance genes has clinical implications for AMR detection, the horizontal resistance gene pool and the management of patients.
Similar content being viewed by others
Introduction
The non-aureus staphylococci (NAS) represent an important source of nosocomial disease, including prosthetic joint infection (PJI), infective endocarditis and infection in pre-term babies1. In the UK, over 215,000 joint replacements (hip, knee and shoulder) took place in 2016, with a year-on-year increase of 4%2. Of these replacements, 1.5% require surgical revision due to infection2. These infections are most commonly caused by Staphylococcus spp., and attributed to NAS in ~31% of cases across Europe3. In our local hospital, the Norfolk and Norwich University Hospital (NNUH), 50% of isolates identified in suspected PJI are NAS.
In clinical microbiology, staphylococci are classified using the coagulase test, with coagulase-positive samples overwhelmingly identified as S. aureus and coagulase-negative samples grouped together under the term coagulase-negative staphylococci (CoNS). CoNS is therefore the term found in antimicrobial surveillance data. However, since coagulase-negative S. aureus strains exist (as do coagulase-positive strains of other staphylococcal species), we use the term “non-aureus staphylococci” (NAS) to encompass all staphylococci which are not S. aureus, including the commonly isolated S. epidermidis, S. haemolyticus and S. capitis species, regardless of coagulase activity.
There is currently an intense focus upon the presence and spread of bacterial antimicrobial resistance, typified in S. aureus by methicillin resistance (MRSA). While the body of literature in antimicrobial resistance research is growing for staphylococci, NAS data remains eclipsed by the focus on S. aureus. Studies investigating antimicrobial resistance (AMR) in NAS4 suggest that 45% of NAS harbour methicillin resistance5, and that NAS may be resistant to a larger number of antimicrobial classes than S. aureus5,6 but comprehensive analyses are missing. We aimed to address this point here by curating a diverse collection of NAS and correlating mechanisms of antibiotic resistance with MICs.
Results and discussion
Our NAS collection comprised over 30 species of Staphyloccocus, including at least 10 isolates of S. epidermidis, S. capitis, S. haemolyticus, S. hominis, S. saprophyticus, S. simulans and S. warneri (Table 1). Isolates were collected over a 4-year period from 2013 to 2016.
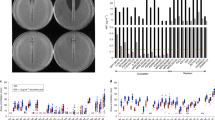
The range of antimicrobials tested was selected based on clinical relevance and availability (Supplementary Table 1). Observed MIC distributions per antimicrobial are shown in Fig. 1 which demonstrates how susceptibility varies within this NAS collection. Erythromycin, tetracycline and gentamicin all displayed bimodal distributions, with erythromycin indicating an additional population of very high MICs. The other five antibiotics displayed Gaussian distributions.
Cefoxitin screening does not correlate with mecA presence in clinically relevant NAS
Cefoxitin is used to screen for methicillin resistance in S. aureus as it induces mecA and mecC expression7. However, while methicillin-resistant S. aureus (MRSA) has a high public profile, much less is known about methicillin-resistant NAS (MRNAS). EUCAST guidelines state that for MRSA “cefoxitin is a very sensitive and specific marker of mecA/mecC-mediated methicillin resistance including in heterogeneous expressing strains and is the agent of choice”8. In this collection, we found 194/394 (49%) displayed reduced susceptibility to cefoxitin with MICs ≥ 4 µg/ml (Supplementary Table 1). The vast majority of these isolates were from clinical samples (FOX Fig. 1, Supplementary Table 1) but analysis at the nucleotide level (Supplementary Tables 2 and 3) indicated that only 88 out of 175 (50 %) sequenced isolates with an MIC ≥ 4 µg/ml harboured a known mecA. MecA is extremely well-conserved and analysis at the amino acid level yielded the same results (Supplementary Table 4). Other mec elements were also identified (e.g. mecC, mecI and mecR1) but only ever in addition to mecA. Breaking this down by species, 20/21 S. saprophyticus isolates with cefoxitin MIC ≥ 4 µg/ml harboured no mecA (Fig. 2). No mecA was detected in eleven species with high cefoxitin MICs and for S. hominis, S. warneri and S. haemolyticus, the percentage of the population that exhibited MIC ≥ 4 µg/ml with no mecA was between 28% and 67% (Fig. 2). Our results support cefoxitin as a good indicator of mecA presence in S. epidermidis9, but suggest that it performs poorly in the less common, but still clinically relevant NAS. In addition, we observed 14 cases where the presence of mecA did not result in a MIC ≥ 4 µg/ml. These were re-tested (alongside 6 others) under conditions designed to encourage mecA expression (see the “Methods” section) and resulted in increased MICs in all isolates, however, 7/20 remained <4 µg/ml (Supplementary Table 1). An alternative mechanism for high FOX MICs could be beta-lactamase production, however, blaZ was the only beta-lactamase detected in the collection, and there were >30 isolates where high FOX MICs were observed in the absence of mecA and blaZ (Supplementary Table 2).
Reduced susceptibility only partly explained by known mechanisms
When exposed to gentamicin and tetracycline, isolates could broadly be divided into two populations, displaying susceptible or reduced susceptibility phenotypes (Fig. 1 GEN and TET). In isolates displaying an MIC ≥ 1 µg/ml for gentamicin, 49/130 isolates harboured aac(6’)-le-aph(2”)-la (Table 2) which is associated with gentamicin resistance in Enterococcus10,11 but has also been observed in Staphylococcus12,13. A total of 12 isolates had a match for aph(3’)IIIa, but only five of them were associated with reduced susceptibility.
Six isolates that contained aac(6’)-le-aph(2”)-la displayed susceptible MICs, making them the equivalent of major errors (MEs) in public health terms, as the isolates were genotypically resistant but phenotypically susceptible14. Accordingly, the 81/130 isolates with reduced susceptibility (≥1 µg/ml) that harboured no aac(6’)-le-aph(2”)-la represented the equivalent of very major errors (VMEs) as they were genotypically susceptible but phenotypically resistant14. This is highly suggestive of novel mechanisms of resistance and was a feature of other antimicrobials tested (Fig. 3). It is possible that assessing antibiotic susceptibility on agar may have produced different MICs. This is mitigated by two aspects: firstly, our study design incorporated 13.5% replication in the MIC assay giving additional confidence to the results as all repeats were within 2-fold MIC, and secondly, even if the MIC value per isolate was different on agar, the spread of MICs would be highly likely to remain and require mechanisms to explain reduced susceptibility.
Percentage of known genetic mechanisms identified in sequenced NAS isolates with reduced susceptibility. Number of isolates with reduced susceptibility per antimicrobial is given in parentheses. Known mechanisms found per antimicrobial are shown in italics. FOX cefoxitin, GEN gentamicin, TET tetracycline, ERY erythromycin, DAP daptomycin, VAN vancomycin, RIF rifampicin and TEC teicoplanin.
To identify whether efflux pumps might play a role in these phenotypes, we assessed the ARIBA output for the staphylococcal-specific norABC, mgrA, mepR and qac genes15. In the sequenced NAS collection, 168/378 (44.4%) contained norA, however of these less than two-thirds (75/130) had reduced susceptibility to gentamicin.
According to the Comprehensive Antimicrobial Resistance Database [CARD]16, tetK is by far the most common tetracycline resistance mechanism in S. aureus and S. epidermidis (10–20%), followed by tetL (<1%) and tetM (<1%). This was borne out in our NAS collection, where 48/148 (32.4%) isolates with MICs ≥ 2 µg/ml of tetracycline contained tetK, as compared to 5/223 (2.2%) with MICs below 2 µg/ml. One animal isolate with an MIC of 16 µg/ml carried tetL and one clinical isolate with an MIC of 64 µg/ml carried tetM; neither had any other tetracycline resistance genes. Again, this demonstrated that 98/148 isolates displayed a reduced susceptibility phenotype that did not associate with a known resistance determinant, indicative of uncharacterised resistance mechanisms.
The distribution of erythromycin phenotypes was more complex. With this antimicrobial, we observed both susceptible isolates and those with reduced susceptibility, but the latter appeared to consist of two populations, one with MICs between ≥2–256 µg/ml and one with MICs ≥ 512 µg/ml (Fig. 1 ERY). We had sequence data available from 135 of the ≥2–256 µg/ml population and 66 of the ≥512 µg/ml population, and identified the presence of a resistance gene (ermA, ermC, msrA) in 65.9% (89/135) of the ≥2–256 µg/ml population and 55/66 (83.3 %) of the ≥512 µg/ml population (Table 2). Our results indicated that the presence of ermC rather than msrA was the major cause of MICs exceeding 256 µg/ml. Although rare (n = 4), harbouring both genes resulted in a MIC ≥ 512 µg/ml in three cases and 128 µg/ml in the other. In isolates with a MIC of ≥ 512 µg/ml, qac was identified 36 times. In 23 of these cases, ermC was also present; qac was only found twice with no other known erythromycin resistance mechanisms present. A total of 101 isolates with a MIC ≥ 2 µg/ml did not contain qac.
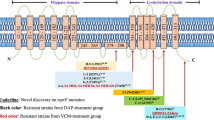
For daptomycin, approximately half the collection displayed reduced susceptibility (MIC ≥ 1 µg/ml, Fig. 1 DAP and Supplementary Table 1). A small subset, comprising eight isolates from clinical samples only, displayed MICs ≥ 4 µg/ml; such high MICs to daptomycin have not been previously reported, according to the European Committee on Antimicrobial Susceptibility Testing (EUCAST) and The British Society for Antimicrobial Chemotherapy (BSAC) surveillance data. These MICs were repeated a second time and confirmed. This is concerning given that daptomycin is a current therapeutic choice for treating soft tissue infections caused by NAS17. Seven of these isolates were sequenced and our ARIBA analysis (Supplementary Table 3) indicated only a single S. epidermidis isolate contained genes implicated in daptomycin resistance: gshF and liaFRS with an MIC of 1 µg/ml, the remaining 169 isolates with an ≥1 µg/ml MIC did not harbour any of these genes. SNP mutations in mprF and rpoC are associated with daptomycin resistance in S. aureus but none of these were identified in the NAS collection18,19. More recently, SNPs in walK have also been associated with daptomycin resistance in S. aureus and S. epidermidis20. The three S. aureus SNPs are present in CARD and were not identified in our collection. The V500F mutation from S. epidermidis20 was also not identified in our S. epidermidis with DAP MICs ≥ 4 µg/ml. Whilst walK was identified by protein BLAST as present across the NAS collection (as expected for an essential gene21), sequence variation was observed at the protein level which prevents SNPs observed in S. aureus or S. epidermidis being extrapolated to all NAS. We, therefore, conclude that there are potentially novel daptomycin resistance mechanisms present in these strains.
Higher MICs found in clinical samples
Vancomycin is a treatment option for prosthetic joint infection, and 94% of isolates had a MIC below 4 µg/ml (Fig. 1 VAN). However, of the 24 isolates with reduced susceptibility, 22 (92%) came from clinical samples and only 2/24 were found in healthy volunteers. This is indicative of a wider trend, where isolates associated with clinical samples had significantly higher MICs (p < 0.005) than non-clinical isolates for cefoxitin, erythromycin, gentamicin, tetracycline, daptomycin and vancomycin (Supplementary Fig. s). Given the importance of NAS in nosocomial infections, this is a worrying prospect both in terms of what is present in the clinic and also the possibility of AMR gene transfer into organisms more capable of causing infection, including S. aureus. In addition, no known mechanisms of resistance were identified for vancomycin, rifampicin or teicoplanin (Fig. 3 and Supplementary Table 3).
Over half of the NAS collection displayed susceptibility to multiple antimicrobials
Out of all the isolates tested, 48% (192/394) had reduced susceptibility to three or more antimicrobials. Twenty-five isolates had reduced susceptibility to six antimicrobials, and three isolates had reduced susceptibility to seven antimicrobials; of these 24/25 and 3/3 were isolated from clinical samples (Supplementary Table 1). The implications of these are difficult-to-treat infections and potentially a large reservoir of staphylococcal resistance genes within the patient under antimicrobial treatment.
Animal isolates have similar MIC distributions to human isolates
It is generally acknowledged that the presence of reduced susceptibility in microorganisms isolated from domesticated animals can impact public health if those organisms also cause infection in humans22,23. In our collection, there were 40 NAS isolated from domesticated animals (7 NCTC strains), of which we obtained genome sequences from 23. Animals included were cats, dogs, cattle and sheep and although in much fewer numbers than the human isolates in the collection, the animal isolates displayed very similar MIC distributions and harboured corresponding genetic mechanisms. This does not rule out the possibility that animals could be a reservoir of AMR for staphylococci.
Conclusion
Genome analysis of isolates displaying MICs to cefoxitin of ≥4 µg/ml indicated that approximately half harboured the mecA element. The absence of mecA from the other half suggests that other mechanisms are likely present. This was apparent across many of the antimicrobials tested as between 0% and 65% of phenotypic resistance in clinical isolates could be attributed to known resistance mechanisms. The remaining 35–100% suggests that there are potentially numerous unknown mechanisms underpinning NAS resistance, which warrant further investigation.
Methods
NAS collection
Under NHS Research Ethics Committee approval, the Norwich Biorepository banks blood, solid tissue and bacterial isolates from the NNUH and research institutes on the Norwich Research Park, including the University of East Anglia (UEA), and makes these available to the research community. This enabled us to assemble a collection of 380 NAS from (a) clinical specimens which were isolated from suspected NAS PJI infections (229, NNUH), (b) healthy human volunteers (114 skin swabs from adults at UEA), and (c) animal samples (33, UEA) with five having no source recorded. Animal isolates were taken from healthy domestic dogs, cats, sheep and cows. An additional 14 strains of NAS from the National Collection of Type Cultures (NCTC) were supplied by Public Health England.
Isolates were identified at NNUH using MALDI-TOF (Bruker) to the species level (Table 1). All strains were cultured overnight on TSA plates (Oxoid), checked for contamination and purified. Once purified, the NAS collection was stored as glycerol stocks to be screened for their antimicrobial susceptibility. Staphylococcus aureus NCTC 12973 was used as a control.
Susceptibility testing
To assay the entire NAS collection, five deep well 96-well microplates (VWR) were prepared with 1 ml TSB (Oxoid) per well. Glycerol stocks were used to inoculate the corresponding well. Per plate, one well was designated as a sterility control (TSB only) and one well was inoculated with the S. aureus control. After inoculation, plates were sealed and incubated at 37 oC at 180 rpm for a minimum of 10 h. The experimental design enabled 13.5% of the collection to be tested in duplicate; MIC data were compared and then tabulated (Supplementary Table 1).
Following standard BSAC guidelines (version 14) at the time, iso-sensitest agar (Oxoid) was prepared and sterilised. Antimicrobial stocks were added to obtain the desired final concentrations once the media had cooled to <50 oC. For daptomycin, Ca2+ was also added at 50 µg/ml. The agar antimicrobial mixture was then poured into sterile rectangular plates (Fisher Scientific) and dried.
Per strain, a 1:10 dilution of overnight culture was transferred to a 96-well plate and the OD600 was measured. An average OD600 was calculated for each column, which was then diluted to approximately OD600 0.6 to generate an inoculum plate for susceptibility testing.
Using a 96-pin multi-point inoculator (Denley), ~1 µl of inoculum per isolate was stamped onto the agar containing antimicrobials, from the lowest concentration to the highest. Between inoculum plates, the pins were washed in 70% ethanol for 30 s and allowed to dry before stamping on an antimicrobial-free plate to confirm sterility. Washes were also carried out between antimicrobials using sterile water. All stamped plates were incubated at 37 oC.
Isolates found to have reduced susceptibility to daptomycin had their MICs determined for a second time by spotting 10 μl of culture onto TSA plates containing various daptomycin concentrations (supplemented with Ca2+ at 50 µg/ml). To increase mecA expression, 14 isolates which contained mecA but on initial testing showed susceptibility to cefoxitin (MIC < 4 µg/ml) were re-tested on Mueller Hinton Agar with 3% NaCl added alongside a further 6 isolates. Overnight cultures were diluted in PBS and 5 µl spots containing 104 cells were spotted onto plates which were incubated at 35 oC.
Test MIC ranges (in µg/ml) were as follows: daptomycin 0.25–2, erythromycin 0.125–256, gentamicin 0.016–64, rifampicin 0.004–0.064, teicoplanin 0.25–16, tetracycline 0.25–256 and vancomycin 1–4, cefoxitin 0.25–4 µg/ml (based upon published work9). Isolates were considered to have reduced susceptibility to the specified antibiotic if they displayed the following MICs: ≥4 µg/ml (cefoxitin, teicoplanin, vancomycin); ≥2 µg/ml (tetracycline, erythromycin); ≥1 µg/ml (gentamicin, daptomycin); ≥0.06 µg/ml (rifampicin).
Statistical comparison of clinical and non-clinical isolates
Using Prism (GraphPad, San Diego, USA, v 5.04), a Mann–Whitney test was performed (non-parametric test, two-tailed with Gaussian approximation) to compare the MIC of clinical and non-clinical isolates. Statistical significance was given to a p-value < 0.05.
DNA extraction and sequencing
Overnight cultures derived from single colonies were pelleted and resuspended in lysis buffer (Qiagen), transferred to 2 ml lysis matrix B tubes (MPBio) and subjected to bead beating for 15 min at 30 Hz (Tissuelyser II, Qiagen) with RNAse A added. DNA was extracted according to the QiaCube HT protocol with an additional 30 min incubation at 65 oC after proteinase K addition and eluted into Tris–10 mM HCl.
Libraries for sequencing were prepared using the Nextera XT DNA Library Prep Protocol and sequenced on the Illumina MiSeq or NextSeq with a loading concentration of 1.8 picomolar.
Genome analysis
The raw reads were subject to FastQC quality control24, adapters were trimmed using Trimmomatic [v 0.39]25 using the supplied NexteraXT adapter sequences. In some cases, read normalisation was performed using BBNorm [v35.85]26 to remove low coverage contamination. The lowest coverage cutoff level parameter used was dependent on the total coverage of the sequence since some sequencing runs had a high difference in coverage level across the run. Finally, reads were concatenated if they originated from Illumina NextSeq since this platform produces eight reads per sample, four forward and four reverse. A total of 364 samples passed QC and were suitable for downstream analysis. These reads were used as described below. Sequences are available from the European Nucleotide Archive, under project PRJEB31403.
To determine which antimicrobial resistance genes and associated individual mutations were present in each of our 364 NAS genomes, reference gene sequences were downloaded from CARD v2.0.016 and used as input to ARIBA v2.13.227 which generates local assemblies from sequence reads and reports back which reference genes (and individual mutations) are identified, with a minimum percent identity cut off at 90% (Supplementary Table 5). For genes where ‘partial’ or ‘interrupted’ was reported, this was not considered sufficient evidence for intact gene presence. The tabulated results were evaluated for gene and mutation presence/absence relative to MIC per antimicrobial. Twelve NCTC sequences were downloaded as genome assemblies from the European Nucleotide Archive (accessions: SAMEA4364213; SAMEA4364214; SAMEA4384234; SAMEA4384058; SAMEA4384237; SAMEA4384064; SAMEA4412661; SAMEA4384059; SAMEA4384235; SAMEA4384060; SAMEA4384339; SAMEA4384403) and analysed by ABRIcate v0.9.728 using CARD v2.0.016 as the reference database with a minimum DNA coverage of 90%. NCTC 13831 and 13837 sequence data was not available at the time of sequencing and therefore these isolates were sequenced as described above for the main NAS collection. Protein level conservation was assessed using BLAST v2.10.1 against the NCBI AMR database. Hits were recorded for greater than 40% identity at the protein level over 80% of the query and subject sequence.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
Sequences are available from the European Nucleotide Archive, under project PRJEB31403.
References
Becker, K., Heilmann, C. & Peters, G. Coagulase-negative Staphylococci. Clin. Microbiol. Rev. 27, 870–926 (2014).
National Joint Registry 14th Annual Report 2017, NJR (Wales National Joint Registry for England, Northern Ireland and the Isle of Man, 2017).
Lamagni, T. Epidemiology and burden of prosthetic joint infections. J. Antimicrob. Chemother. 69, i5–i10 (2014).
Osman, K. et al. Prevalence of the antibiotic resistance genes in coagulase-positive-and negative-Staphylococcus in chicken meat retailed to consumers. Front. Microbiol. 7, (2016) https://doi.org/10.3389/fmicb.2016.01846.
Aggarwal, V. et al. Organism profile in periprosthetic joint infection: pathogens differ at two arthroplasty infection referral centers in Europe and in the United States. J. Knee Surg. 27, (2014) https://doi.org/10.1055/s-0033-1364102.
Morgenstern, M. et al. Antibiotic resistance of commensal Staphylococcus aureus and coagulase-negative Staphylococci in an international cohort of surgeons: a prospective point-prevalence study. PLoS ONE 11, e0148437 (2016).
Skov, R. et al. Phenotypic detection of mecC-MRSA: cefoxitin is more reliable than oxacillin. J. Antimicrob. Chemother. 69, 133–135 (2014).
EUCAST. EUCAST Guidelines for Detection of Resistance Mechanisms and Specific Resistances of Clinical and/or Epidemiological Importance, Version 2.0, http://www.eucast.org (2017).
Swenson, J. et al. Correlation of cefoxitin MICs with the presence of mecA in Staphylococcus spp. J. Clin. Microbiol. 47, 1902–1905 (2009).
Ferretti, J., Gilmore, K. & Courvalin, P. Nucleotide sequence analysis of the gene specifying the bifunctional 6'-aminoglycoside acetyltransferase 2"-aminoglycoside phosphotransferase enzyme in Streptococcus faecalis and identification and cloning of gene regions specifying the two activities. J. Bacteriol. 167, 631–638 (1986).
Chow, J. Aminoglycoside resistance in enterococci. Clin. Infect. Dis. 31, 586–589 (2000).
Perreten, V. et al. Clonal spread of methicillin-resistant Staphylococcus pseudintermedius in Europe and North America: an international multicentre study. J. Antimicrob. Chemother. 65, 1145–1154 (2010).
Schmitz, F.-J. et al. The prevalence of aminoglycoside resistance and corresponding resistance genes in clinical isolates of staphylococci from 19 European hospitals. J. Antimicrob. Chemother. 43, 253–259 (1999).
Neuert, S. et al. Prediction of phenotypic antimicrobial resistance profiles from whole genome sequences of non-typhoidal Salmonella enterica. Front. Microbiol. 9, 592 (2018).
Sapula, S. & Brown, M. Antimicrobial drug efflux pumps in Staphylococcus aureus. In Efflux-Mediated Antimicroboal Resistance in Bacteria: Mechanisms, Regulation and Clinical Implications (eds Li, X. Z., Elkins, C. & Zgurskaya, H.) Ch. 7 (2016).
Jia, B. et al. CARD 2017: expansion and model-centric curation of the comprehensive antibiotic resistance database. Nucleic Acids Res. 4, D566–D573 (2017).
White, B. & Seaton, R. A. Complicated skin and soft tissue infections: literature review of evidence for and experience with daptomycin. Infect. Drug Res. 4, 115 (2011).
Sabat, A. et al. Daptomycin resistant Staphylococcus aureus clinical strain with novel non-synonymous mutations in the mprF and vraS genes: a new insight into daptomycin resistance. Front. Microbiol. 9, 2705 (2018).
Friedman, L., Alder, J. D. & Silverman, J. A. Genetic changes that correlate with reduced susceptibility to daptomycin in Staphylococcus aureus. Antimicrob. Agents Chemother. 50, 2137–2145 (2006).
Yang, S., Mishra, N. N., Rubio, A. & Bayer, A. S. Additional routes to Staphylococcus aureus daptomycin resistance as revealed by comparative genome sequencing, transcriptional profiling, and phenotypic studies. Antimicrob. Agents Chemother. 57, 5658–5664 (2013).
Dubrac, S., Boneca, I. G., Poupel, O. & Msadek, T. New insights into the WalK/WalR (YycG/YycF) essential signal transduction pathway reveal a major role in controlling cell wall metabolism and biofilm formation in Staphylococcus aureus. J. Bacteriol. 189, 8257–8269 (2007).
Landers, T., Cohen, B., Wittum, T. E. & Larson, E. L. A review of antibiotic use in food animals: perspective, policy, and potential. Public Health Rep. 127, 4–22 (2012).
Woolhouse, M., Ward, M. & van Bunnik, B. Antimicrobial resistance in humans, livestock and the wider environment. Philos. Trans. R. Soc. B 370, 20140083 (2015).
Babraham Bioinformatics. FastQC: a Quality Control Tool for High Throughput Sequence Data. http://www.bioinformatics.babraham.ac.uk/projects/fastqc/.
Bolger, A., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Brian- Joint Genome Institute, BBMap short read aligner, and other bioinformatic tools. https://sourceforge.net/projects/bbmap/.
Hunt, M. et al. ARIBA: rapid antimicrobial resistance genotyping directly from sequencing reads. Microb. Genom. 3, e000131 (2017).
Seeman, T. ABRicate: Mass Screening of Contigs for Antiobiotic Resistance Genes (GitHub, 2016).
Acknowledgements
We thank David Livermore and Iain McNamara for productive discussions on antimicrobial usage. This work was funded by the Orthopaedics Trust (registered charity 1110248).
Author information
Authors and Affiliations
Contributions
Initial sequencing method development was carried out by R. Clifford and A. Correia. R. Clifford and R. Dean were involved in the isolate collection. H. Felgate and E. Gray performed MIC testing. Further laboratory testing was carried out by H. Felgate. Analysis of data was performed by H. Felgate, L.C. Crossman and G.C. Langridge. G.C. Langridge and J. Wain were involved in method development and scientific discussions. The initial manuscript was written by H. Felgate and G.C. Langridge; all authors were involved in editing and have approved the final version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Felgate, H., Crossman, L.C., Gray, E. et al. Known mechanisms cannot account for a third of reduced susceptibility in non-aureus staphylococci. npj Antimicrob Resist 1, 15 (2023). https://doi.org/10.1038/s44259-023-00008-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s44259-023-00008-1