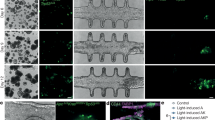
Abstract
Cell type-specific interfaces within living animals will be invaluable for achieving communication with identifiable cells over the long term, enabling applications across many scientific and medical fields. However, biological tissues exhibit complex and dynamic organization properties that pose serious challenges for chronic cell-specific interfacing. A new technology, combining chemistry and molecular biology, has emerged to address this challenge: genetically targeted chemical assembly (GTCA), in which specific cells are genetically programmed (even in wild-type or non-transgenic animals, including mammals) to chemically construct non-biological structures. Here, we discuss recent progress in genetically targeted construction of materials and outline opportunities that may expand the GTCA toolbox, including specific chemical processes involving novel monomers, catalysts and reaction regimes both de cellula (from the cell) and ad cellula (towards the cell); different GTCA-compatible reaction conditions with a focus on light-based patterning; and potential applications of GTCA in research and clinical settings.
Key points
-
Genetically targeted chemical assembly (GTCA) uses cell-specific genetic information to guide the assembly of functional materials in situ.
-
The GTCA toolbox can be expanded through specific chemical processes involving novel monomers, catalysts and reaction conditions or regimes.
-
GTCA allows both building structures from the targeted cell membrane (de cellula) and an alternative approach (ad cellula) for cell-specific attachment of partially synthesized materials.
-
Different GTCA-compatible reaction conditions can be imposed through modulation of light, pH, heat and other signals.
-
The broad GTCA concept can be applied for both fundamental research and the treatment of diseases in the central and peripheral nervous systems.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$99.00 per year
only $8.25 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Acarón Ledesma, H. et al. An atlas of nano-enabled neural interfaces. Nat. Nanotechnol. 14, 645–657 (2019).
Chen, R., Canales, A. & Anikeeva, P. Neural recording and modulation technologies. Nat. Rev. Mater. 2, 16093 (2017).
Rivnay, J., Wang, H., Fenno, L., Deisseroth, K. & Malliaras, G. G. Next-generation probes, particles, and proteins for neural interfacing. Sci. Adv. 3, e1601649 (2017).
Zhang, A., Lee, J.-H. & Lieber, C. M. Nanowire-enabled bioelectronics. Nano Today 38, 101135 (2021).
Luan, L. et al. Recent advances in electrical neural interface engineering: minimal invasiveness, longevity, and scalability. Neuron 108, 302–321 (2020).
Zhang, M., Tang, Z., Liu, X. & Van der Spiegel, J. Electronic neural interfaces. Nat. Electron. 3, 191–200 (2020).
Song, E., Li, J., Won, S. M., Bai, W. & Rogers, J. A. Materials for flexible bioelectronic systems as chronic neural interfaces. Nat. Mater. 19, 590–603 (2020).
Woods, G. A., Rommelfanger, N. J. & Hong, G. Bioinspired materials for in vivo bioelectronic neural interfaces. Matter 3, 1087–1113 (2020).
Berggren, M., Głowacki, E. D., Simon, D. T., Stavrinidou, E. & Tybrandt, K. In vivo organic bioelectronics for neuromodulation. Chem. Rev. 122, 4826–4846 (2022).
Tang, X., Shen, H., Zhao, S., Li, N. & Liu, J. Flexible brain–computer interfaces. Nat. Electron. 6, 109–118 (2023).
Tang, T.-C. et al. Materials design by synthetic biology. Nat. Rev. Mater. 6, 332–350 (2021). This review article on synthetic biology describes biopolymer synthesis from natural building blocks.
Burgos-Morales, O. et al. Synthetic biology as driver for the biologization of materials sciences. Mater. Today Bio 11, 100115 (2021).
Brophy, J. A. N. & Voigt, C. A. Principles of genetic circuit design. Nat. Methods 11, 508–520 (2014).
Ouyang, L., Shaw, C. L., Kuo, C.-C., Griffin, A. L. & Martin, D. C. In vivo polymerization of poly(3,4-ethylenedioxythiophene) in the living rat hippocampus does not cause a significant loss of performance in a delayed alternation task. J. Neural Eng. 11, 026005 (2014). This article reports the first work on in vivo polymerization (not genetically targeted).
Liu, J. et al. Genetically targeted chemical assembly of functional materials in living cells, tissues, and animals. Science 367, 1372–1376 (2020). This article is the first demonstration of GTCA that uses cell-specific genetic information to guide neurons to deposit conductive or insulating polymers in situ.
Zhang, A. et al. Genetically targeted chemical assembly of polymers specifically localized extracellularly to surface membranes of living neurons. Sci. Adv. 9, eadi1870 (2023).This article reports the second-generation GTCA that allows highly localized membrane expression.
Dai, Y. et al. Oxidative polymerization in living cells. J. Am. Chem. Soc. 143, 10709–10717 (2021).
Pieszka, M. et al. Controlled supramolecular assembly inside living cells by sequential multistaged chemical reactions. J. Am. Chem. Soc. 142, 15780–15789 (2020).
Zhang, Y. et al. Controlled intracellular polymerization for cancer treatment. JACS Au 2, 579–589 (2022).
Guimard, N. K., Gomez, N. & Schmidt, C. E. Conducting polymers in biomedical engineering. Prog. Polym. Sci. 32, 876–921 (2007).
Tabba, H. D. & Smith, K. M. Anodic oxidation potentials of substituted pyrroles: derivation and analysis of substituent partial potentials. J. Org. Chem. 49, 1870–1875 (1984).
Berlin, A., Pagani, G. A., Sannicolò, F., Schiavon, G. & Zotti, G. Monomer tailoring to control the redox potentials of conductive polyheterocycles. Polymer 32, 1841–1842 (1991).
Tan, Y. & Ghandi, K. Kinetics and mechanism of pyrrole chemical polymerization. Synth. Met. 175, 183–191 (2013).
Gaupp, C. L. et al. Poly(3,4-ethylenedioxypyrrole): organic electrochemistry of a highly stable electrochromic polymer. Macromolecules 33, 1132–1133 (2000).
Sönmez, G., Schottland, P., Zong, K. & Reynolds, J. R. Highly transmissive and conductive poly[(3,4-alkylenedioxy)pyrrole-2,5-diyl] (PXDOP) films prepared by air or transition metal catalyzed chemical oxidation. J. Mater. Chem. 11, 289–294 (2001).
Kuwabata, S., Ito, S. & Yoneyama, H. Copolymerization of pyrrole and thiophene by electrochemical oxidation and electrochemical behavior of the resulting copolymers. J. Electrochem. Soc. 135, 1691 (1988).
Tewari, A. et al. Soybean peroxidase catalyzed enzymatic synthesis of pyrrole/EDOT copolymers. Macromol. Chem. Phys. 211, 1610–1617 (2010).
John, R. & Wallace, G. Doping–dedoping of polypyrrole: a study using current-measuring and resistance-measuring techniques. J. Electroanal. Chem. 354, 145–160 (1993).
Cruz-Silva, R. et al. Biocatalytic synthesis of polypyrrole powder, colloids, and films using horseradish peroxidase. J. Colloid Interface Sci. 328, 263–269 (2008).
Song, H.-K., Lee, E. J. & Oh, S. M. Electrochromism of 2,2′-azinobis(3-ethylbenzothiazoline-6-sulfonate) incorporated into conducting polymer as a dopant. Chem. Mat. 17, 2232–2233 (2005).
Yoo, J. E. et al. Improving the electrical conductivity of polymer acid-doped polyaniline by controlling the template molecular weight. J. Mater. Chem. 17, 1268–1275 (2007).
Bayer, C. L., Trenchard, I. J. & Peppas, N. A. Analyzing polyaniline-poly(2-acrylamido-2-methylpropane sulfonic acid) biocompatibility with 3T3 fibroblasts. J. Biomater. Sci. Polym. Ed. 21, 623–634 (2010).
Zhang, W., Yang, F. K., Pan, Z., Zhang, J. & Zhao, B. Bio-inspired dopamine functionalization of polypyrrole for improved adhesion and conductivity. Macromol. Rapid Commun. 35, 350–354 (2014).
Xie, C. et al. Electroresponsive and cell-affinitive polydopamine/polypyrrole composite microcapsules with a dual-function of on-demand drug delivery and cell stimulation for electrical therapy. NPG Asia Mater. 9, e358 (2017).
Chalmers, E., Lee, H., Zhu, C. & Liu, X. Increasing the conductivity and adhesion of polypyrrole hydrogels with electropolymerized polydopamine. Chem. Mat. 32, 234–244 (2020).
Bedard, K. & Krause, K.-H. The NOX family of ROS-generating NADPH oxidases: physiology and pathophysiology. Physiol. Rev. 87, 245–313 (2007).
Ma, M. W. et al. NADPH oxidase in brain injury and neurodegenerative disorders. Mol. Neurodegener. 12, 1–28 (2017).
Suh, S. W. et al. Glucose and NADPH oxidase drive neuronal superoxide formation in stroke. Ann. Neurol. 64, 654–663 (2008).
Bankar, S. B., Bule, M. V., Singhal, R. S. & Ananthanarayan, L. Glucose oxidase — an overview. Biotechnol. Adv. 27, 489–501 (2009).
Mayer, A. M. & Staples, R. C. Laccase: new functions for an old enzyme. Phytochemistry 60, 551–565 (2002).
Kausaite, A., Ramanaviciene, A. & Ramanavicius, A. Polyaniline synthesis catalysed by glucose oxidase. Polymer 50, 1846–1851 (2009).
Karamyshev, A. V., Shleev, S. V., Koroleva, O. V., Yaropolov, A. I. & Sakharov, I. Y. Laccase-catalyzed synthesis of conducting polyaniline. Enzyme Microb. Technol. 33, 556–564 (2003).
Ramanavicius, A., Kausaite, A., Ramanaviciene, A., Acaite, J. & Malinauskas, A. Redox enzyme — glucose oxidase — initiated synthesis of polypyrrole. Synth. Met. 156, 409–413 (2006).
Song, H.-K. & Palmore, G. T. R. Conductive polypyrrole via enzyme catalysis. J. Phys. Chem. B 109, 19278–19287 (2005).
Porto de Souza Vandenberghe, L. et al. in Biomass, Biofuels, Biochemicals (eds Singh, S. P. et al.) 11–30 (Elsevier, 2020).
Prescher, J. A. & Bertozzi, C. R. Chemistry in living systems. Nat. Chem. Biol. 1, 13–21 (2005). This review article discusses bioorthogonal chemistry, a tool for genetically targeted conjugation of pre-synthesized materials.
Chyan, W. & Raines, R. T. Enzyme-activated fluorogenic probes for live-cell and in vivo imaging. ACS Chem. Biol. 13, 1810–1823 (2018).
Yang, Y., Lee, P. & Sternson, S. M. Cell type-specific pharmacology of NMDA receptors using masked MK801. eLife 4, e10206 (2015).
Saxon, E. & Bertozzi, C. R. Cell surface engineering by a modified Staudinger reaction. Science 287, 2007–2010 (2000).
Prescher, J. A., Dube, D. H. & Bertozzi, C. R. Chemical remodelling of cell surfaces in living animals. Nature 430, 873–877 (2004).
Cioce, A. et al. Cell-specific bioorthogonal tagging of glycoproteins. Nat. Commun. 13, 6237 (2022).
Schumann, B. et al. Bump-and-hole engineering identifies specific substrates of glycosyltransferases in living cells. Mol. Cell 78, 824–834.e15 (2020).
Fan, X. et al. Cell-type-specific labeling and profiling of glycans in living mice. Nat. Chem. Biol. 18, 625–633 (2022).
Chin, J. W. Expanding and reprogramming the genetic code. Nature 550, 53–60 (2017).
Tian, L. et al. Selective esterase–ester pair for targeting small molecules with cellular specificity. Proc. Natl Acad. Sci. USA 109, 4756–4761 (2012).
Zakeri, B. et al. Peptide tag forming a rapid covalent bond to a protein, through engineering a bacterial adhesin. Proc. Natl Acad. Sci. USA 109, E690–E697 (2012). This article introduces the SpyTag–SpyCatcher system, which is the basis of genetically targeted conjugation using modular protein–peptide interaction systems.
Grenier, V., Daws, B. R., Liu, P. & Miller, E. W. Spying on neuronal membrane potential with genetically targetable voltage indicators. J. Am. Chem. Soc. 141, 1349–1358 (2019).
Bedbrook, C. N. et al. Genetically encoded Spy peptide fusion system to detect plasma membrane-localized proteins in vivo. Chem. Biol. 22, 1108–1121 (2015).
Veggiani, G. et al. Programmable polyproteams built using twin peptide superglues. Proc. Natl Acad. Sci. USA 113, 1202–1207 (2016).
Prakash, R. et al. Two-photon optogenetic toolbox for fast inhibition, excitation and bistable modulation. Nat. Methods 9, 1171–1179 (2012).
Deisseroth, K. Optogenetics: 10 years of microbial opsins in neuroscience. Nat. Neurosci. 18, 1213–1225 (2015).
Marshel, J. H. et al. Cortical layer-specific critical dynamics triggering perception. Science 365, eaaw5202 (2019).
Wojtovich, A. P. & Foster, T. H. Optogenetic control of ROS production. Redox Biol. 2, 368–376 (2014).
Balena, A., Bianco, M., Pisanello, F. & De Vittorio, M. Recent advances on high-speed and holographic two-photon direct laser writing. Adv. Funct. Mater. https://doi.org/10.1002/adfm.202211773 (2023).
Bulina, M. E. et al. A genetically encoded photosensitizer. Nat. Biotechnol. 24, 95–99 (2006).
Zhou, Z., Song, J., Nie, L. & Chen, X. Reactive oxygen species generating systems meeting challenges of photodynamic cancer therapy. Chem. Soc. Rev. 45, 6597–6626 (2016).
Rodríguez-Pulido, A. et al. Assessing the potential of photosensitizing flavoproteins as tags for correlative microscopy. Chem. Commun. 52, 8405–8408 (2016).
Shu, X. et al. A genetically encoded tag for correlated light and electron microscopy of intact cells, tissues, and organisms. PLoS Biol. 9, e1001041 (2011). This article describes miniSOG, the genetically encoded photosensitizer that could enable 3D in vivo photolithography.
Trewin, A. J. et al. Light-induced oxidant production by fluorescent proteins. Free Radic. Biol. Med. 128, 157–164 (2018).
Westberg, M., Holmegaard, L., Pimenta, F. M., Etzerodt, M. & Ogilby, P. R. Rational design of an efficient, genetically encodable, protein-encased singlet oxygen photosensitizer. J. Am. Chem. Soc. 137, 1632–1642 (2015).
Westberg, M., Bregnhøj, M., Etzerodt, M. & Ogilby, P. R. No photon wasted: an efficient and selective singlet oxygen photosensitizing protein. J. Phys. Chem. B 121, 9366–9371 (2017).
Makhijani, K. et al. Precision optogenetic tool for selective single-and multiple-cell ablation in a live animal model system. Cell Chem. Biol. 24, 110–119 (2017).
He, J. et al. A genetically targetable near-infrared photosensitizer. Nat. Methods 13, 263–268 (2016).
Zhang, A., Kadur, C. S., Ramakrishnan, C., Bao, Z. & Deisseroth, K. Genetically-encoded photosensitizers enable light-controlled polymerization on living neuronal membranes. Preprint at bioRxiv https://doi.org/10.1101/2022.12.27.521977 (2022).
Sessler, C. D. et al. Optogenetic polymerization and assembly of electrically functional polymers for modulation of single-neuron excitability. Sci. Adv. 8, eade1136 (2022).
del Campo, A. & Greiner, C. SU-8: a photoresist for high-aspect-ratio and 3D submicron lithography. J. Micromech. Microeng. 17, R81 (2007).
Deisseroth, K. From microbial membrane proteins to the mysteries of emotion. Cell 184, 5279–5285 (2021).
Ferenczi, E. A. et al. Optogenetic approaches addressing extracellular modulation of neural excitability. Sci. Rep. 6, 23947 (2016). This article describes use of proton-pumping rhodopsins to control juxtamembranous pH-dependent channel function; this approach could enable light-controlled pH-regulated GTCA.
Mattis, J. et al. Principles for applying optogenetic tools derived from direct comparative analysis of microbial opsins. Nat. Methods 9, 159–172 (2012).
Pruckmayr, G. & Wu, T. K. Polymerization of tetrahydrofuran by proton acids. Macromolecules 11, 662–668 (1978).
Deisseroth, K. & Hegemann, P. The form and function of channelrhodopsin. Science 357, eaan5544 (2017).
Hatchett, D. W., Josowicz, M. & Janata, J. Acid doping of polyaniline: spectroscopic and electrochemical studies. J. Phys. Chem. B 103, 10992–10998 (1999).
Li, X. et al. Nanotransducers for wireless neuromodulation. Matter 4, 1484–1510 (2021). This review summarizes nanoparticle transducers that convert external fields into different forms of energy, which could enable temperature-regulated GTCA.
Liu, Y., Yi, Z., Yao, Y., Guo, B. & Liu, X. Noninvasive manipulation of ion channels for neuromodulation and theranostics. Acc. Mater. Res. 3, 247–258 (2022).
Chen, S. et al. Near-infrared deep brain stimulation via upconversion nanoparticle-mediated optogenetics. Science 359, 679–684 (2018).
Chen, R. et al. Deep brain optogenetics without intracranial surgery. Nat. Biotechnol. 39, 161–164 (2021).
Carvalho-de-Souza, J. L. et al. Photosensitivity of neurons enabled by cell-targeted gold nanoparticles. Neuron 86, 207–217 (2015).
Rastogi, S. K. et al. Remote nongenetic optical modulation of neuronal activity using fuzzy graphene. Proc. Natl Acad. Sci. USA 117, 13339–13349 (2020).
Jiang, Y. et al. Heterogeneous silicon mesostructures for lipid-supported bioelectric interfaces. Nat. Mater. 15, 1023–1030 (2016).
Jiang, Y. et al. Rational design of silicon structures for optically controlled multiscale biointerfaces. Nat. Biomed. Eng. 2, 508–521 (2018).
Huang, H., Delikanli, S., Zeng, H., Ferkey, D. M. & Pralle, A. Remote control of ion channels and neurons through magnetic-field heating of nanoparticles. Nat. Nanotechnol. 5, 602–606 (2010).
Chen, R., Romero, G., Christiansen, M. G., Mohr, A. & Anikeeva, P. Wireless magnetothermal deep brain stimulation. Science 347, 1477–1480 (2015).
Sebesta, C. et al. Subsecond multichannel magnetic control of select neural circuits in freely moving flies. Nat. Mater. 21, 951–958 (2022).
Marino, A. et al. Piezoelectric nanoparticle-assisted wireless neuronal stimulation. ACS Nano 9, 7678–7689 (2015).
Kim, T. et al. Deep brain stimulation by blood–brain-barrier-crossing piezoelectric nanoparticles generating current and nitric oxide under focused ultrasound. Nat. Biomed. Eng. 7, 149–163 (2023).
Jin, W., Wang, R. & Huang, X. Horseradish peroxidase-catalyzed oxidative polymerization of aniline in bicontinuous microemulsion stabilized by AOT/SDS. J. Mol. Liq. 302, 112529 (2020).
Yamada, M., Nagasaki, S. C., Ozawa, T. & Imayoshi, I. Light-mediated control of gene expression in mammalian cells. Neurosci. Res. 152, 66–77 (2020). This review introduces optically targeted gene expression approaches that could control downstream GTCA-relevant genes.
Yamada, M., Suzuki, Y., Nagasaki, S. C., Okuno, H. & Imayoshi, I. Light control of the Tet gene expression system in mammalian cells. Cell Rep. 25, 487–500.e6 (2018).
Gossen, M. et al. Transcriptional activation by tetracyclines in mammalian cells. Science 268, 1766–1769 (1995).
Yamada, M., Nagasaki, S. C., Suzuki, Y., Hirano, Y. & Imayoshi, I. Optimization of light-inducible Gal4/UAS gene expression system in mammalian cells. iScience 23, 101506 (2020).
Uda, Y. et al. Efficient synthesis of phycocyanobilin in mammalian cells for optogenetic control of cell signaling. Proc. Natl Acad. Sci. USA 114, 11962–11967 (2017).
Jin, X. & Riedel-Kruse, I. H. Biofilm lithography enables high-resolution cell patterning via optogenetic adhesin expression. Proc. Natl Acad. Sci. USA 115, 3698–3703 (2018).
Zhao, F. et al. Light-induced patterning of electroactive bacterial biofilms. ACS Synth. Biol. 11, 2327–2338 (2022).
Lovett-Barron, M. et al. Ancestral circuits for the coordinated modulation of brain state. Cell 171, 1411–1423.e17 (2017).
Lovett-Barron, M. et al. Multiple convergent hypothalamus–brainstem circuits drive defensive behavior. Nat. Neurosci. 23, 959–967 (2020).
Yokogawa, T., Hannan, M. C. & Burgess, H. A. The dorsal raphe modulates sensory responsiveness during arousal in zebrafish. J. Neurosci. 32, 15205–15215 (2012).
Rajan, K., Harvey, C. D. & Tank, D. W. Recurrent network models of sequence generation and memory. Neuron 90, 128–142 (2016).
Perich, M. G. et al. Inferring brain-wide interactions using data-constrained recurrent neural network models. Preprint at bioRxiv, https://doi.org/10.1101/2020.12.18.423348 (2020).
Andalman, A. S. et al. Neuronal dynamics regulating brain and behavioral state transitions. Cell 177, 970–985.e20 (2019).
Deisseroth, K. Circuit dynamics of adaptive and maladaptive behaviour. Nature 505, 309–317 (2014).
Chettih, S. N. & Harvey, C. D. Single-neuron perturbations reveal feature-specific competition in V1. Nature 567, 334–340 (2019).
Kishi, K. E. et al. Structural basis for channel conduction in the pump-like channelrhodopsin ChRmine. Cell 185, 672–689.e23 (2022).
Liu, Y. et al. Morphing electronics enable neuromodulation in growing tissue. Nat. Biotechnol. 38, 1031–1036 (2020).
Li, T. L. et al. Stretchable mesh microelectronics for the biointegration and stimulation of human neural organoids. Biomaterials. 290, 121825 (2022).
Ward, P. J. & English, A. W. Optical stimulation and electrophysiological analysis of regenerating peripheral axons. Bio Protoc. 9, e3281 (2019).
Yalcin, I. et al. The sciatic nerve cuffing model of neuropathic pain in mice. J. Vis. Exp. https://doi.org/10.3791/51608 (2014).
Yizhar, O. et al. Neocortical excitation/inhibition balance in information processing and social dysfunction. Nature 477, 171–178 (2011).
Tatti, R., Haley, M. S., Swanson, O. K., Tselha, T. & Maffei, A. Neurophysiology and regulation of the balance between excitation and inhibition in neocortical circuits. Biol. Psychiatry 81, 821–831 (2017).
Selimbeyoglu, A. et al. Modulation of prefrontal cortex excitation/inhibition balance rescues social behavior in CNTNAP2-deficient mice. Sci. Transl Med. 9, eaah6733 (2017).
Peñagarikano, O. et al. Absence of CNTNAP2 leads to epilepsy, neuronal migration abnormalities, and core autism-related deficits. Cell 147, 235–246 (2011).
Berndt, A. et al. Structural foundations of optogenetics: determinants of channelrhodopsin ion selectivity. Proc. Natl Acad. Sci. USA 113, 822–829 (2016).
Pernelle, G., Nicola, W. & Clopath, C. Gap junction plasticity as a mechanism to regulate network-wide oscillations. PLoS Comput. Biol. 14, e1006025 (2018).
Redolfi Riva, E. & Micera, S. Progress and challenges of implantable neural interfaces based on nature-derived materials. Bioelectron. Med. 7, 6 (2021).
Shim, J.-S., Rogers, J. A. & Kang, S.-K. Physically transient electronic materials and devices. Mater. Sci. Eng. R. Rep. 145, 100624 (2021).
Irimia-Vladu, M. “Green” electronics: biodegradable and biocompatible materials and devices for sustainable future. Chem. Soc. Rev. 43, 588–610 (2014).
Qin, Y. et al. Harnessing oxidative microenvironment for in vivo synthesis of subcellular conductive polymer microesicles enhances nerve reconstruction. Nano Lett. 22, 3825–3831 (2022).
Patel, S. R. & Lieber, C. M. Precision electronic medicine in the brain. Nat. Biotechnol. 37, 1007–1012 (2019).
Williams, D. C. et al. Rapid and permanent neuronal inactivation in vivo via subcellular generation of reactive oxygen with the use of KillerRed. Cell Rep. 5, 553–563 (2013).
Qi, Y. B., Garren, E. J., Shu, X., Tsien, R. Y. & Jin, Y. Photo-inducible cell ablation in Caenorhabditis elegans using the genetically encoded singlet oxygen generating protein miniSOG. Proc. Natl Acad. Sci. USA 109, 7499–7504 (2012).
Ng, J. et al. Genetically targeted 3D visualisation of Drosophila neurons under electron microscopy and X-ray microscopy using miniSOG. Sci. Rep. 6, 38863 (2016).
Liang, P., Kolodieznyi, D., Creeger, Y., Ballou, B. & Bruchez, M. P. Subcellular singlet oxygen and cell death: location matters. Front. Chem. 8, 1045 (2020).
Xie, W. et al. Chemoptogenetic ablation of neuronal mitochondria in vivo with spatiotemporal precision and controllable severity. eLife 9, e51845 (2020).
Acknowledgements
The Keck Foundation supported development of light-initiated polymerization and applications in living neural networks. The National Science Foundation Future Manufacturing Program grant (award no. 2037164) supported chemical method development for genetically targeted chemical assembly (GTCA). Both grants were awarded to K.D. and Z.B. based on the original GTCA concept and next-generation methods described previously15. A.Z. acknowledges support from the American Heart Association (AHA) (award no. 23POST1018301). K.Y.L. acknowledges support from the Stanford ChEM-H Chemistry/Biology Interface Predoctoral Training Program (NIH 5T32GM120007) and Bio-X Bowes Fellowship. Z.B. is a Chan Zuckerberg Biohub San Francisco investigator.
Author information
Authors and Affiliations
Contributions
A.Z., Z.B. and K.D. wrote the manuscript with edits from all authors. Z.B. and K.D. supervised all aspects of the work. All authors approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
All techniques and protocols are freely available to the academic community, and the authors provide free training in genetically targeted chemical assembly (GTCA) methods at Stanford in workshops that can be accessed for registration online (https://web.stanford.edu/group/dlab/optogenetics/oil.html). Z.B. and K.D. are co-inventors of the GTCA concept used here, in intellectual property filed and owned by Stanford University.
Peer review
Peer review information
Nature Reviews Bioengineering thanks Itaru Imayoshi, Brian Timko and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, A., Jiang, Y., Loh, K.Y. et al. Genetically targeted chemical assembly. Nat Rev Bioeng 2, 82–94 (2024). https://doi.org/10.1038/s44222-023-00110-z
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s44222-023-00110-z