Abstract
Myocardial ischemia–reperfusion injury (MIRI) induces life-threatening damages to the cardiac tissue, and pharmacological means to achieve cardioprotection are sorely needed. MIRI severity varies along the day–night cycle and is molecularly linked to components of the cellular clock, including the nuclear receptor REV-ERBα, a transcriptional repressor. Here we show that digoxin administration in mice is cardioprotective when timed to trigger REV-ERBα protein degradation. In cardiomyocytes, digoxin increases REV-ERBα ubiquitinylation and proteasomal degradation, which depend on REV-ERBα’s ability to bind its natural ligand, heme. Inhibition of the membrane-bound Src tyrosine-kinase partially alleviated digoxin-induced REV-ERBα degradation. In untreated cardiomyocytes, REV-ERBα proteolysis is controlled by several E3 ubiquitin ligases and the proteasome subunit PSMB5. Among these, only SIAH2 and PSMB5 contributed to digoxin-induced degradation of REV-ERBα. Thus, controlling REV-ERBα proteostasis through the ubiquitin–proteasome system is an appealing cardioprotective strategy. Our data support the timed use of clinically approved cardiotonic steroids in prophylactic cardioprotection.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
All data are available in the main text or the supplementary materials. Source data are available for this paper. Microarray data that support the finding of this study have been deposited in the National Center of Biotechnology Information’s Gene Expression Omnibus and are accessible through accession numbers GSE183659, GSE183660 and GSE183661. Other transcriptomic datasets have been described in ref. 10 and deposited under the accession number GSE62459. Whole mouse heart circadian gene expression was analyzed using the GSE180108 dataset47. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE77 partner repository with the dataset identifier PXD03660.
References
Patke, A., Young, M. W. & Axelrod, S. Molecular mechanisms and physiological importance of circadian rhythms. Nat. Rev. Mol. Cell Biol. 21, 67–84 (2020).
Martino, T. A. & Young, M. E. Influence of the cardiomyocyte circadian clock on cardiac physiology and pathophysiology. J. Biol. Rhythms 30, 183–205 (2015).
Zhang, J., Chatham, J. C. & Young, M. E. Circadian regulation of cardiac physiology: rhythms that keep the heart beating. Annu. Rev. Physiol. 82, 79–101 (2020).
Crnko, S., Du Pre, B. C., Sluijter, J. P. G. & Van Laake, L. W. Circadian rhythms and the molecular clock in cardiovascular biology and disease. Nat. Rev. Cardiol. 16, 437–447 (2019).
Hausenloy, D. J. & Yellon, D. M. Myocardial ischemia–reperfusion injury: a neglected therapeutic target. J. Clin. Invest. 123, 92–100 (2013).
Hausenloy, D. J. & Yellon, D. M. Ischaemic conditioning and reperfusion injury. Nat. Rev. Cardiol. 13, 193–209 (2016).
Rossello, X. & Yellon, D. M. The RISK pathway and beyond. Basic Res. Cardiol. 113, 2 (2018).
Durgan, D. J. et al. Short communication: ischemia/reperfusion tolerance is time-of-day-dependent: mediation by the cardiomyocyte circadian clock. Circ. Res. 106, 546–550 (2010).
Janszky, I. L. & Ljung, R. Shifts to and from daylight saving time and incidence of myocardial infarction. N. Engl. J. Med. 359, 1966–1968 (2008).
Montaigne, D. et al. Daytime variation of perioperative myocardial injury in cardiac surgery and its prevention by Rev-Erbα antagonism: a single-centre propensity-matched cohort study and a randomised study. Lancet 391, 59–69 (2018).
Davidson, S. M. et al. Multitarget strategies to reduce myocardial ischemia/reperfusion injury: JACC review topic of the week. J. Am. Coll. Cardiol. 73, 89–99 (2019).
Fellahi, J. L., Fischer, M. O., Daccache, G., Gerard, J. L. & Hanouz, J. L. Positive inotropic agents in myocardial ischemia–reperfusion injury. Anesthesiology 118, 1460–1472 (2013).
Matsui, H. & Schwartz, A. Mechanism of cardiac glycoside inhibition of the (Na+-K+)-dependent ATPase from cardiac tissue. Biochim. Biophys. Acta 151, 655–663 (1968).
Askari, A. The sodium pump and digitalis drugs: dogmas and fallacies. Pharmacol. Res. Perspect. 19, e00505 (2019).
Liang, M. et al. Identification of a pool of non-pumping Na/K-ATPase. J. Biol. Chem. 282, 10585–10593 (2007).
Marck, P. V. & Pierre, S. V. Na/K-ATPase signaling and cardiac pre/postconditioning with cardiotonic steroids. Int. J. Mol. Sci. 19, 2336 (2018).
Nelson, W., Kupferberg, H. & Halberg, F. Dose–response evaluations of a circadian rhythmic change in susceptibility of mice to ouabain. Toxicol. Appl. Pharmacol. 18, 335–339 (1971).
Patocka, J., Nepovimova, E., Wu, W. & Kuca, K. Digoxin: pharmacology and toxicology—a review. Environ. Toxicol. Pharmacol. 79, 103400 (2020).
Dostanic, I. et al. The alpha2 isoform of Na,K-ATPase mediates ouabain-induced cardiac inotropy in mice. J. Biol. Chem. 278, 53026–53034 (2003).
Iisalo, E. Clinical pharmacokinetics of digoxin. Clin. Pharmacokinet. 2, 1–16 (1977).
Davidson, M. M. et al. Novel cell lines derived from adult human ventricular cardiomyocytes. J. Mol. Cell. Cardiol. 39, 133–147 (2005).
Felippe Gonçalves-de-Albuquerque, C., Ribeiro Silva, A., Ignácio da Silva, C., Caire Castro-Faria-Neto, H. & Burth, P. Na/K pump and beyond: Na/K-ATPase as a modulator of apoptosis and autophagy. Molecules 22, 578 (2017).
Balsalobre, A., Marcacci, L. & Schibler, U. Multiple signaling pathways elicit circadian gene expression in cultured Rat-1 fibroblasts. Curr. Biol. 10, 1291–1294 (2000).
Katz, A. et al. Selectivity of digitalis glycosides for isoforms of human Na,K-ATPase. J. Biol. Chem. 285, 19582–19592 (2010).
Morgan, E. E. et al. Preconditioning by subinotropic doses of ouabain in the Langendorff perfused rabbit heart. J. Cardiovasc. Pharmacol. 55, 234–239 (2010).
Duan, Q. et al. Preconditioning and postconditioning by cardiac glycosides in the mouse heart. J. Cardiovasc. Pharmacol. 71, 95–103 (2018).
Zhang, Z. et al. Identification of hydroxyxanthones as Na/K-ATPase ligands. Mol. Pharmacol. 77, 961–967 (2010).
Orlov, S. N. et al. Na+i,K+i-dependent and -independent signaling triggered by cardiotonic steroids: facts and artifacts. Molecules 22, 635 (2017).
Huh, J. R. et al. Digoxin and its derivatives suppress TH17 cell differentiation by antagonizing RORγt activity. Nature 472, 486–490 (2011).
Ouyang, X. et al. Digoxin suppresses pyruvate kinase M2-promoted HIF-1α transactivation in steatohepatitis. Cell Metab 27, 339–350 (2018).
Zhang, H. et al. Digoxin and other cardiac glycosides inhibit HIF-1α synthesis and block tumor growth. Proc. Natl Acad. Sci. USA 105, 19579–19586 (2008).
Strickson, S. et al. The anti-inflammatory drug BAY 11-7082 suppresses the MyD88-dependent signalling network by targeting the ubiquitin system. Biochem. J. 451, 427–437 (2013).
Oikawa, D. et al. Molecular bases for HOIPINs-mediated inhibition of LUBAC and innate immune responses. Commun. Biol. 3, 163 (2020).
Xie, Z. et al. Gene set knowledge discovery with Enrichr. Curr. Protoc. 1, e90 (2021).
Brown, K., Gerstberger, S., Carlson, L., Franzoso, G. & Siebenlist, U. Control of IκB-α proteolysis by site-specific, signal-induced phosphorylation. Science 267, 1485–1488 (1995).
Wang, Y. et al. Cardiac glycosides induce autophagy in human non-small cell lung cancer cells through regulation of dual signaling pathways. Int. J. Biochem. Cell Biol. 44, 1813–1824 (2012).
Raghuram, S. et al. Identification of heme as the ligand for the orphan nuclear receptors REV-ERBα and REV-ERBβ. Nat. Struct. Mol. Biol. 14, 1207–1213 (2007).
Carter, E. L., Gupta, N. & Ragsdale, S. W. High affinity heme binding to a heme regulatory motif on the nuclear receptor Rev-erbβ leads to its degradation and indirectly regulates its interaction with nuclear receptor corepressor. J. Biol. Chem. 291, 2196–2222 (2016).
Mohammed, H. et al. Rapid immunoprecipitation mass spectrometry of endogenous proteins (RIME) for analysis of chromatin complexes. Nat. Protoc. 11, 316–326 (2016).
DeBruyne, J. P., Baggs, J. E., Sato, T. K. & Hogenesch, J. B. Ubiquitin ligase Siah2 regulates RevErbα degradation and the mammalian circadian clock. Proc. Natl Acad. Sci. USA 112, 12420–12425 (2015).
Zhao, X. et al. Circadian amplitude regulation via FBXW7-targeted REV-ERBα degradation. Cell 165, 1644–1657 (2016).
Yin, L., Joshi, S., Wu, N., Tong, X. & Lazar, M. A. E3 ligases Arf-bp1 and Pam mediate lithium-stimulated degradation of the circadian heme receptor Rev-erb α. Proc. Natl Acad. Sci. USA 107, 11614–11619 (2010).
Li, Y. et al. An integrated bioinformatics platform for investigating the human E3 ubiquitin ligase-substrate interaction network. Nat. Commun. 8, 347 (2017).
Yin, L., Wang, J., Klein, P. S. & Lazar, M. A. Nuclear receptor Rev-erbα is a critical lithium-sensitive component of the circadian clock. Science 311, 1002–1005 (2006).
Hill, R. J. W., Innominato, P. F., Levi, F. & Ballesta, A. Optimizing circadian drug infusion schedules towards personalized cancer chronotherapy. PLoS Comput. Biol. 16, e1007218 (2020).
Hermida, R. C. et al. Bedtime hypertension treatment improves cardiovascular risk reduction: the Hygia Chronotherapy Trial. Eur. Heart J. 41, 4565–4576 (2020).
Martino, T. et al. Day/night rhythms in gene expression of the normal murine heart. J. Mol. Med. 82, 256–264 (2004).
Podobed, P. et al. The day/night proteome in the murine heart. Am. J. Physiol. Regul. Integr. Comp. Physiol. 307, R121–R137 (2014).
Duan, Q. et al. Role of phosphoinositide 3-kinase IA (PI3K-IA) activation in cardioprotection induced by ouabain preconditioning. J. Mol. Cell. Cardiol. 80, 114–125 (2015).
Digitalis Investigation, G. The effect of digoxin on mortality and morbidity in patients with heart failure. N. Engl. J. Med. 336, 525–533 (1997).
Hallberg, P., Lindbäck, J., Lindahl, B., Stenestrand, U. & Melhus, H. Digoxin and mortality in atrial fibrillation: a prospective cohort study. Eur. J. Clin. Pharmacol. 63, 959–971 (2007).
Patel, N. J. et al. Digoxin significantly improves all-cause mortality in atrial fibrillation patients with severely reduced left ventricular systolic function. Int. J. Cardiol. 169, e84–e86 (2013).
Rana, S., Prabhu, S. D. & Young, M. E. Chronobiological influence over cardiovascular function: the good, the bad, and the ugly. Circ. Res. 126, 258–279 (2020).
Young, M. E. et al. Cardiomyocyte-specific BMAL1 plays critical roles in metabolism, signaling, and maintenance of contractile function of the heart. J. Biol. Rhythms 29, 257–276 (2014).
Tsimakouridze, E. V. et al. Chronomics of pressure overload-induced cardiac hypertrophy in mice reveals altered day/night gene expression and biomarkers of heart disease. Chronobiol. Int. 29, 810–821 (2012).
Carter, E. L., Ramirez, Y. & Ragsdale, S. W. The heme regulatory motif of nuclear receptor Rev-Erbβ is a key mediator of heme and redox signaling in circadian rhythm maintenance and metabolism. J. Biol. Chem. 292, 11280–11299 (2017).
Wang, J. & Lazar, M. A. Bifunctional role of Rev-erbα in adipocyte differentiation. Mol. Cell. Biol. 28, 2213–2220 (2008).
Kaasik, K. & Lee, C. C. Reciprocal regulation of haem biosynthesis and the circadian clock in mammals. Nature 430, 467–471 (2004).
Dioum, E. M. et al. NPAS2: a Gas-responsive transcription factor. Science 298, 2385–2387 (2002).
Yang, J. et al. A novel heme-regulatory motif mediates heme-dependent degradation of the circadian factor period 2. Mol. Cell. Biol. 28, 4697–4711 (2008).
Reitz, C. J. et al. SR9009 administered for one day after myocardial ischemia-reperfusion prevents heart failure in mice by targeting the cardiac inflammasome. Commun. Biol. 2, 353 (2019).
Mia, S. et al. Differential effects of REV-ERBα/β agonism on cardiac gene expression, metabolism, and contractile function in a mouse model of circadian disruption. Am. J. Physiol. Heart Circ. Physiol. 318, H1487–H1508 (2020).
Zhang, L. et al. REV-ERBα ameliorates heart failure through transcription repression. JCI Insight 2, e95177 (2017).
Stujanna, E. N. et al. Rev-erb agonist improves adverse cardiac remodeling and survival in myocardial infarction through an anti-inflammatory mechanism. PLoS ONE 12, e0189330 (2017).
Zhao, Y. et al. Disruption of circadian rhythms by shift work exacerbates reperfusion injury in myocardial infarction. J. Am. Coll. Cardiol. 79, 2097–2115 (2022).
Busonero, C. et al. Ouabain and digoxin activate the proteasome and the degradation of the eralpha in cells modeling primary and metastatic breast cancer. Cancers 12, 3840 (2020).
Wang, Y. et al. Bufalin is a potent small-molecule inhibitor of the steroid receptor coactivators SRC-3 and SRC-1. Cancer Res. 74, 1506–1517 (2014).
Woldt, E. et al. Rev-erb-α modulates skeletal muscle oxidative capacity by regulating mitochondrial biogenesis and autophagy. Nat. Med. 19, 1039–1046 (2013).
Bell, R. M., Mocanu, M. M. & Yellon, D. M. Retrograde heart perfusion: the Langendorff technique of isolated heart perfusion. J. Mol. Cell. Cardiol. 50, 940–950 (2011).
Schmittgen, T. D. & Livak, K. J. Analyzing real-time PCR data by the comparative CT method. Nat. Protoc. 3, 1101–1108 (2008).
Harris, V. M. Protein detection by Simple Western analysis. Methods Mol. Biol. 1312, 465–468 (2015).
Berthier, A. et al. Combinatorial regulation of hepatic cytoplasmic signaling and nuclear transcriptional events by the OGT/REV-ERBα complex. Proc. Natl Acad. Sci. USA 115, E11033–E11042 (2018).
Kondo, K., Klco, J., Nakamura, E., Lechpammer, M. & Kaelin, W. G. Jr. Inhibition of HIF is necessary for tumor suppression by the von Hippel-Lindau protein. Cancer Cell 1, 237–246 (2002).
Vandel, J. et al. GIANT: galaxy-based tool for interactive analysis of transcriptomic data. Sci. Rep. 10, 19835 (2020).
Hughes, M. E., Hogenesch, J. B. & Kornacker, K. JTK_CYCLE: an efficient nonparametric algorithm for detecting rhythmic components in genome-scale data sets. J. Biol. Rhythms 25, 372–380 (2010).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Perez-Riverol, Y. et al. The PRIDE database resources in 2022: a hub for mass spectrometry-based proteomics evidences. Nucleic Acids Res. 50, D543–d552 (2022).
Acknowledgements
We are grateful to J. Vandel for help with transcriptomic data visualization, S. Susen (CHRU Lille, France) for generously providing DigiFab, and F. Naji (PamGene, Netherlands) for help and advices for the PamGene data analysis. This work was supported by grants from INSERM (PL), Région Hauts-de-France and Université de Lille (REV-ERBalpha/SAS20215, P. L.), from ANR (LABX EGID (ANR-10-LABX-0046, B. S.); ANR VasCal (ANR46-CE14-0001-01, B. S.), ANR-CE14-0003-01 (D. M.), ANR-18-CE17-0003-02 (D. M.); PreciDiab ANR-18-IBHU-0001; 20001891/NP0025517; 2019_ESR_11 (D. M.), ANR-17-CE14-0034 (J.-S. A.), from the Leducq Foundation (LEAN network 16CVD01, B. S.), from Institut Pasteur de Lille (CTRL Melodie, J.-S. A.), from the Fondation pour la Recherche Médicale (EQU202103012732, J.-S. A.). B. S. is a recipient of an Advanced ERC Grant (694717)
Author information
Authors and Affiliations
Contributions
Conceptualization: M. V., P. L., B. S.; Methodology: M. V., J.-S. A., A. B., A. H., S. C., J. E., P. L.; Validation: M. V., C. G., A. H., S. C., A. B., P. L.; Formal analysis: M. V., P. L.; Investigation: M. V., C. G., X. M., A. B., A. H., S. C., R. B.; Resources: P. L., B. S., J. E., H. D., S. D., D. M., J.-S. A.; Data visualization: M. V., P. L.; Supervision: P. L., B. S.; Funding acquisition: P. L., B. S.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Cardiovascular Research thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Rhythmic gene and protein level in mouse heart whole tissue extracts.
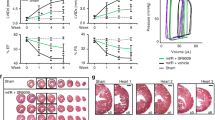
(a) WES analysis of REV-ERBα protein level in mouse hearts collected over a 24-hours period at rest phase (ZT0-ZT12) and active phase (ZT13-ZT24) (n = 4). HSP90α was used as a protein loading control. (b) WES analysis of REV-ERBα protein level in mouse hearts collected at ZT9 after vehicle or digoxin injection (0.1,0.5, and 0.1mpk at ZT5. (c) Cyclic Nr1d1, Nr1d2, Bmal1 and Cdkn1a mRNA expression in mouse hearts collected at rest (ZT0-ZT12) or active (ZT13-ZT24) phases Results are shown as mean + /−SEM (n = 3). (d) Nr1d1, Arntl (Bmal1) and Cdkn1a mRNA expression in mouse hearts collected at ZT9 after vehicle or digoxin (1mpk) injection at ZT5. Results are shown as mean + /−SEM (n = 8-9) which were compared using which were compared using a two-tailed Welch’s t-test. (e) P21 protein level in mouse hearts collected at ZT9 after vehicle or digoxin (1mpk) injection at ZT5. (f) REV-ERBα protein level in mouse hearts collected at ZT0 after vehicle or digoxin (1mpk) injection at ZT20. Basal REV-ERBα protein level at ZT9 is shown here as reference. Measurements were from distinct samples. Quantification of data in panels (a), (b) and (c) appears in Fig. 1.
Extended Data Fig. 2 NKA status in mouse heart and AC16 cells
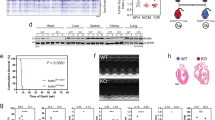
(a) Left panel: NKA isotype-encoding mRNA expression in mouse heart. Atp1a1, Atp1a2 and Atp1a3 expression levels are plotted as a function of time. Per1 mRNA expression is shown as a reference circadian gene. Middle panel: Relative expression of NKA isotype-encoding mRNA expression in mouse heart at ZT9. Results are shown as mean + /−SEM (n = 8-15) which were compared using Brown-Forsythe & Welch ANOVA test, followed by Dunnett’s multiple comparison test. Right panel: Effect of digoxin treatment on NKA isotype-encoding mRNA expression in mouse heart at ZT9. Results are shown as mean + /−SEM (n = 8-15) which were compared using Brown-Forsythe & Welch ANOVA test, followed by Dunnett’s multiple comparison test. (b) NKA isotype protein expression level in mouse heart as a function of time. Data quantification is shown (right panel) as mean + /−SEM (n = 3) which were compared using a one-way ANOVA test, followed by a Tukey’s multiple comparison test. *: P < 0.05, **: P < 0.01, ***: P < 0.001. (c) Left panel: Relative expression of NKA isotype-encoding mRNA expression in human AC16 cells (T24). Right panel: Effect of digoxin treatment on NKA isotype-encoding mRNA expression in human AC16 cells (T24). Results are shown as mean + /−SEM (n = 5-6) which were compared using a one-way ANOVA test, followed by a Tukey’s multiple comparison test. (d) NKA isotype protein expression level in human AC16 as a function of time. Data quantification is shown (right panel) as mean + /−SEM (n = 4) which were compared using a one-way ANOVA test, followed by a Tukey’s multiple comparison test. All measurements were from distinct samples. *: P < 0.05, **: P < 0.01
Extended Data Fig. 3 Effects of digoxin and of its structural analogs on REV-ERBα protein stability.
(a) AC16 cellular viability in the presence of increasing concentrations of digoxin. Data are plotted as mean + /−SEM (n = 4) which were compared using a one-way ANOVA test, followed by a Tukey’s multiple comparison test. (b) Protein array analysis of AC16 whole cell protein extract. AC16 cells were treated for 6 hours at T18 as indicated and extracts were probed on membrane spotted with antibodies specific for components of the apoptotic pathway. Fluorescence signals were quantified and were represented as a heatmap (n = 2). (c) Cyclic REV-ERBα protein levels in human AC16 cells. REV-ERBα levels were determined in synchronized human AC16 cells treated with vehicle or digoxin (0.5 µM) and harvested at T0, T6, T12, T18 and T24. Quantification of data is shown in Fig. 2a. (d) REV-ERBα protein level was assessed in AC16 cells treated with vehicle, digoxin (0.5 µM) and/or DigiFab (100 µg). HSP90α was used as a protein loading control. Right panel: normalized data are plotted as mean + /−SEM (n = 3) which were compared using a one-way ANOVA test, followed by a Tukey’s multiple comparison test. (e) REV-ERBα protein level was assessed in AC16 cells treated with vehicle or varying concentration of digoxin as indicated. HSP90α was used as a protein loading control Right panel: normalized data are plotted as mean + /−SEM (n = 3) and analyzed by nonlinear regression curve fitting. (f) REV-ERBα protein level in AC16 cells treated with vehicle or varying concentrations of bufalin (0.1, 1.0, 10 µM) (n = 2). (g) REV-ERBα protein level in AC16 cells treated with vehicle or ouabain (5 µM) (n = 2). HSP90α was used as a protein loading control. All measurements were from distinct samples. (h) Time course experiment in AC16 cells treated at T18 with vehicle or digoxin (5 µM). REV-ERBα protein level was assessed using the ProteinSimple Wes system (n = 2). HSP90α was used as a protein loading control. All measurements were from distinct samples. ***: P < 0.001.
Extended Data Fig. 4 Digoxin effect in U2OS cells.
(a) Rhythmic REV-ERBα and RORα protein level in synchronized U2OS cells treated with vehicle or digoxin (0.5 µM) and harvested over a 36-hour period as indicated. HSP90α was used as a protein loading control. (b) NR1D1 and BMAL1 mRNA expression in synchronized U2OS cells treated with vehicle or digoxin (0.5 µM) and harvested at T24. Normalized data are plotted as mean + /−SEM (n = 6) which were compared using a one-way ANOVA test, followed by a Tukey’s multiple comparison test. (c) Bioluminescent signal in U2OS cells transfected with the Bmal1-Luc plasmid and treated with vehicle or digoxin (0.5 µM). Detrended data are shown HSP90α was used as a protein loading control. All measurements were from distinct samples.
Extended Data Fig. 5 Effect of protein kinase inhibition or activation on digoxin-mediated REV-ERBα protein level decrease.
Representative WES analysis (related to quantified data in Fig. 4) are shown. REV-ERBα protein level in synchronized AC16 cells treated with REV-ERBα protein levels were quantified after a 6 hour-treatment with or without digoxin and with or without enzyme inhibitors: (a) PP2 (20 µM, Src kinase inhibitor), (b) ZSTK474 (10 µM, PI3 kinase inhibitor), (c) SCH772984 (10 µM, ERK 1&2 inhibitor), (d) MK2206 (1 µM, AKT1/2/3 kinase inhibitor), (e) KN93 (10 µM, CAMK 2&4 inhibitor), (f) PD98059 (10 µM, MEK 1&2 inhibitor), (g) RP-8-pCPT-cGMPS (20 µM, PKG1&2), (h) CRT0066101 (5 µM, PKD inhibitor), (i) PF-4708671 (10 µM, P70S6K1 inhibitor), (j) BAY11-7082 (10 µM, LUBAC and UPS inhibitor), (k) HOIPIN8 (10 µM, LUBAC-HOIP specific inhibitor), (l) TPCA-1 (20 nM, IKKβinhibitor), (m) TPCA-1 (5 µM, IKKβ/α inhibitor) (n) amlexanox (1 µM, IKKε & TBK1 inhibitor). HSP90α was used as a protein loading control.
Extended Data Fig. 6 Effect of protein kinase inhibition or activation on digoxin-mediated REV-ERBα protein level decrease.
Quantification of REV-ERBα protein levels in synchronized human AC16 cells (see Extended Data Fig. 7) (a) treated with vehicle, digoxin (0.5 µM) and/or SRC inhibitor (saracatinib), (b) treated with vehicle, digoxin (0.5 µM) and/or PI3K inhibitor (LY294002) (c) treated with vehicle, digoxin (0.5 µM) and/or PI3K/P70S6K inhibitor (dactolisib), (d) transfected with ERK1 expression vector, (e) treated with the GSK3β inhibitor lithium and/or digoxin, (f) transfected with scrambled siRNA (Scr siRNA) or CAMK4 siRNA and treated with vehicle or digoxin (0.5 µM), (g) transfected with scrambled siRNA (Scr siRNA) or ERK1&3 siRNA and treated with vehicle or digoxin (0.5 µM), (h) transfected with scrambled siRNA (Scr siRNA) or PKD1 siRNA and treated with vehicle or digoxin (0.5 µM), (i) transfected with scrambled siRNA (Scr siRNA) or PRKG1 siRNA and treated with vehicle or digoxin (0.5 µM), (j) transfected with scrambled siRNA (Scr siRNA) or HOIL siRNA and treated with vehicle or digoxin (0.5 µM) and (k) transfected with an empty (pcDNA3) or OTULIN-expressing (pcDNA3-OTULIN) vector and treated with vehicle or digoxin (0.5 µM). Normalized data are plotted as mean + /−SEM (n = 3-6) and compared using a one-way ANOVA test, followed by a Tukey’s multiple comparison test except for (c) for which groups were compared using an unpaired 2−tailed t-test. All measurements were from distinct samples.
Extended Data Fig. 7 Effect of protein kinase inhibition or activation on digoxin-mediated REV-ERBα protein level decrease.
WES analysis of REV-ERBα protein levels in synchronized human AC16 cells (a) treated with vehicle, digoxin (0.5 µM) and/or SRC inhibitor (saracatinib, 10 µM), (b) treated with vehicle, digoxin (0.5 µM) and/or PI3K inhibitor (LY294002, 20 µM) (c) treated with vehicle, digoxin (0.5 µM) and/or PI3K/P70S6K inhibitor (dactolisib, 0.1 µM), (d) transfected with ERK1 expression vector, (e) treated with the GSK3β inhibitor lithium (20 mM) and/or digoxin, (f) transfected with scrambled siRNA (Scr siRNA) or CAMK4 siRNA and treated with vehicle or digoxin (0.5 µM), (g) transfected with scrambled siRNA (Scr siRNA) or ERK1&3 siRNA and treated with vehicle or digoxin (0.5 µM), (h) transfected with scrambled siRNA (Scr siRNA) or PKD1 siRNA and treated with vehicle or digoxin (0.5 µM), (i) transfected with scrambled siRNA (Scr siRNA) or PKG1 siRNA and treated with vehicle or digoxin (0.5 µM), (j) transfected with scrambled siRNA (Scr siRNA) or HOIL siRNA and treated with vehicle or digoxin (0.5 µM) and (k) transfected with an empty (pcDNA3) or OTULIN-expressing (pcDNA3-OTULIN) vector and treated with vehicle or digoxin (0.5 µM).
Extended Data Fig. 8 Proteasome inhibition counteracts digoxin effect on REV-ERBα levels.
Quantification of data appears in Fig. 5. REV-ERBα protein level in synchronized AC16 cells (a) treated with either vehicle, digoxin (0.5 µM) and/or clasto-lactacystin β-lactone (10 µM, proteasomal Inhibitor), (b) treated with either vehicle, digoxin (0.5 µM) and/or bortezomib (200 nM, PSMB5 inhibitor), in synchronized U2OS cells treated (c) with either vehicle, digoxin (0.5 µM) and/or bortezomib (200 nM, PSMB5 inhibitor)in synchronized AC16 cells (d) in synchronized AC16 cells transfected with Flag-Rev-ERBα or Flag-Rev-ERBα-H602F then treated with vehicle or digoxin (0.5 µM). HSP90α was used as a protein loading control.
Extended Data Fig. 9 Screening E2/E3 ligases potentially involved in digoxin-induced REV-ERBα degradation.
(a) Quantification of REV-ERBα protein level in AC16 cells transfected either with scrambled siRNA (Scr siRNA) or SIAH2 siRNA and treated with vehicle or digoxin (0.5 µM). A strictly identical protocol was followed to assess the effect of FBXW7 (b), GSK3β (c), UBE2L3 (d), HUWE1 (e), PSME3 (f), TBL1XR1 (g), BUB3 (h), CBL, BRCA1, UBE4A (i), UBE4B, MDM2, STUB1 (j), and of TRIM21 and TRIM33 (k) knockdowns on REV-ERBα stability. Normalized data are plotted as mean + /−SEM (n = 3-6) and compared using a one-way ANOVA test, followed by a Tukey’s multiple comparison test. All measurements were from distinct samples.
Extended Data Fig. 10 Screening E2/E3 ligases potentially involved in digoxin-induced REV-ERBα degradation.
(a) REV-ERBα protein level in AC16 cells transfected either with scrambled siRNA (Scr siRNA) or SIAH2 siRNA and treated with vehicle or digoxin (0.5 µM). A strictly identical protocol was followed to assess the effect of FBXW7 (b), GSK3β (c), UBE2L3 (d), HUWE1 (e), PSME3 (f), TBL1XR1 (g), BUB3 (h), CBL, BRCA1, UBE4A (i), UBE4B, MDM2, STUB1 (j), and of TRIM21 and TRIM33 (k) knockdowns on REV-ERBα stability.
Supplementary information
Supplementary Information
Supplementary Information, Supplementary Figures 1–10, and Uncropped images for Supplementary Figures 1–10.
Supplementary Data 1
Gene expression profiling in ZT0 versus ZT12 mouse hearts. Mouse hearts were collected at ZT0 or ZT12 (n = 7 per group) and RNA was extracted and quantified by Affymetrix arrays. Results are expressed as fold change, and corrected P values are indicated (fold change > 1.2 with an FDR < 0.05, after a moderated t-test followed by a Benjamini–Hochberg multiple-testing correction). For the sake of clarity, only protein-encoding genes are indicated.
Supplementary Data 2
Gene expression profiling in Nr1d1+/+ vs Nr1d1−/− (ZT12) mouse hearts. Mouse hearts were collected at ZT12 (n = 7 per group) from Nr1d1+/+ versus Nr1d1−/− mice, and RNA was extracted and quantified by Affymetrix arrays. Results are expressed as fold change, and corrected P values are indicated (fold change > 1.2 with an FDR < 0.05, after a moderated t-test followed by a Benjamini–Hochberg multiple-testing correction). For the sake of clarity, only protein-encoding genes are indicated.
Supplementary Data 3
Transcriptomic alterations in digoxin-treated AC16 cells. AC16 cells were treated after synchronization by low (0.5 µM) or high (5 µM) digoxin concentrations (n = 3). mRNA levels were quantified by Affymetrix arrays. Results are expressed as fold change, and the corrected P values are indicated (fold change > 1.2 with an FDR < 0.05, after a moderated t-test followed by a Benjamini–Hochberg multiple-testing correction). For the sake of clarity, only protein-encoding genes are indicated.
Supplementary Data 4
Transcriptomic alterations in digoxin-treated mice. Mice were injected at ZT5 and hearts were collected at ZT9 (n = 3–5). After RNA extraction, gene expression was assayed and expressed as described in Supplementary Dataset 1 (fold change > 1.2 with an FDR < 0.05, after a moderated t-test followed by a Benjamini–Hochberg multiple-testing correction). For the sake of clarity, only protein-encoding genes are indicated.
Supplementary Data 5
Transcriptomic alterations common to mouse heart and human AC16 cells. Data from Supplementary Datasets 1 and 2 were compared and genes common to both lists are indicated.
Supplementary Data 6
REV-ERBα RIME data. REV-ERBα was pulled down from AC16 and HepG2 cells and associated proteins were identified by LC–MS/MS (n = 1 for HepG2 and n = 2 for U2OS). The list of common REV-ERBα interactants is shown, with selected interactants related to UPS indicated in bold. The threshold for significant interaction was arbitrarily set to 5 detected peptides to select a testable set of interactants.
Source data
Source Data Fig. 1
Statistical Source Data
Source Data Fig. 1
Unprocessed Western Blots and/or gels
Source Data Fig. 2
Statistical Source Data
Source Data Fig. 2
Unprocessed Western Blots and/or gels
Source Data Fig. 3
Statistical Source Data
Source Data Fig. 3
Unprocessed Western Blots and/or gels
Source Data Fig. 4
Statistical Source Data
Source Data Fig. 4
Unprocessed Western Blots and/or gels
Source Data Fig. 5
Statistical Source Data
Source Data Fig. 6
Statistical Source Data
Source Data Fig. 6
Unprocessed Western Blots and/or gels
Source Data Extended Data Fig. 1
Statistical Source Data
Source Data Extended Data Fig. 1
Unprocessed Western Blots and/or gels
Source Data Extended Data Fig. 2
Statistical Source Data
Source Data Extended Data Fig. 2
Unprocessed Western Blots and/or gels
Source Data Extended Data Fig. 3
Statistical Source Data
Source Data Extended Data Fig. 3
Unprocessed Western Blots and/or gels
Source Data Extended Data Fig. 4
Unprocessed Western Blots and/or gels
Source Data Extended Data Fig. 5
Source Data Extended Data Fig. 5Unprocessed Western Blots and/or gels
Source Data Extended Data Fig. 6
Statistical Source Data
Source Data Extended Data Fig. 7
Unprocessed Western Blots and/or gels
Source Data Extended Data Fig. 8
Unprocessed Western Blots and/or gels
Source Data Extended Data Fig. 9
Statistical Source Data
Source Data Extended Data Fig. 10
Unprocessed Western Blots and/or gels
Source data for Supplementary Information
Statistical Source Data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Vinod, M., Berthier, A., Maréchal, X. et al. Timed use of digoxin prevents heart ischemia–reperfusion injury through a REV-ERBα–UPS signaling pathway. Nat Cardiovasc Res 1, 990–1005 (2022). https://doi.org/10.1038/s44161-022-00148-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s44161-022-00148-z
This article is cited by
-
Timed use of cardiac glycoside protects the heart
Nature Cardiovascular Research (2022)