Abstract
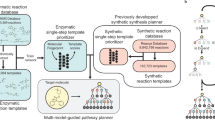
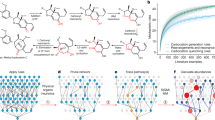
Efficient syntheses of complex small molecules, such as bioactive natural products, often involve detailed retrosynthetic planning and experimental evaluation of speculative synthetic routes. The central challenge of such an approach is that experimental evaluation of high-risk strategies is resource intensive because it requires iterative attempts at unsuccessful strategies. Along with the rapid development of cheminformatics and artificial intelligence, computer-aided synthetic planning has emerged to address this challenge. Herein, we report a complementary strategy that combines human-generated synthetic plans with computational prediction of the feasibility of key steps in the proposed synthesis. A neural network model (NNET) was trained on a literature-based dataset (from Reaxys) to predict the outcome of a generally disfavoured transformation, 6-endo-trig radical cyclization. The model performance was rigorously tested by experimental validation. On the basis of the virtual screening of potential substrates with our NNET model, optimal disconnections and structural modifications were chosen, resulting in five- to eight-step syntheses of three clovane sesquiterpenoids. This work establishes how a machine learning model informs human design and guides multistep syntheses of complex small molecules.

This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The data supporting the findings of this study are available within the paper and its Supplementary Information.
Code availability
All code used to support the findings of this work is supplied as Supplementary Information. The code is also available on GitHub (https://github.com/Newhouse-Group/6-Endo-Radical-Cyclization). Source data are provided with this paper.
References
Corey, E. J. Robert Robinson Lecture. Retrosynthetic thinking—essentials and examples. Chem. Soc. Rev. 17, 111–133 (1988).
Shen, Y. et al. Automation and computer-assisted planning for chemical synthesis. Nat. Rev. Methods Primers 1, 23 (2021).
Coley, C. W., Eyke, N. S. & Jensen, K. F. Autonomous discovery in the chemical sciences part I: progress. Angew. Chem. Int. Ed. 59, 22858–22893 (2020).
Segler, M. H. S., Preuss, M. & Waller, M. Planning chemical syntheses with deep neural networks and symbolic AI. Nature 555, 604–610 (2018).
Szymkuc, S. et al. Computer-assisted synthetic planning: the end of the beginning. Angew. Chem. Int. Ed. 55, 5904–5937 (2016).
Marth, C. J. et al. Network-analysis-guided synthesis of weisaconitine D and liljestrandinine. Nature 528, 493–498 (2015).
Mo, Y. et al. Evaluating and clustering retrosynthesis pathways with learned strategy. Chem. Sci. 12, 1469–1478 (2021).
Mikulak-Klucznik, B. et al. Computational planning of the synthesis of complex natural products. Nature 588, 83–88 (2020).
Bideau, F. L. et al. Tricyclic sesquiterpenes from marine origin. Chem. Rev. 117, 6110–6159 (2017).
Chung, H.-M. et al. Rumphellclovane B, a novel clovane analogue from the gorgonian coral rumphella antipathies. Bull. Chem. Soc. Jpn 84, 119–121 (2011).
Chung, H.-M. et al. Rumphellclovanes C–E, new clovane-type sesquiterpenoids from the gorgonian coral Rumphella antipathies. Tetrahedron Lett. 69, 2740–2744 (2013).
Cheng, X. et al. Neuronal growth promoting sesquiterpene–neolignans; syntheses and biological studies. Org. Biomol. Chem. 10, 383–393 (2012).
Yang, J. et al. Asymmetric total synthesis of (−)-clovan-2,9-dione using Rh(I)-catalyzed [3+2+1] cycloaddition of 1-yne-vinylcyclopropane and CO. Org. Lett. 19, 6040–6043 (2017).
Liu, G., Zhang, Z., Fu, S. & Liu, B. Asymmetric total synthesis of rumphellclovane E. Org. Lett. 23, 290–295 (2021).
de Souza, G. G. et al. Biotransformation of clovane derivatives. Whole cell fungi mediated domino synthesis of rumphellclovane A. Org. Biomol. Chem. 10, 3315–3320 (2012).
Wright, P. M., Seiple, I. B. & Myers, A. G. The evolving role of chemical synthesis in antibacterial drug discovery. Angew. Chem. Int. Ed. 53, 8840–8869 (2014).
Romero, K. J., Galliher, M. S., Pratt, D. A. & Stephenson, C. R. J. Radicals in natural product synthesis. Chem. Soc. Rev. 47, 7851–7866 (2018).
Sha, C.-K., Jean, T.-S. & Wang, D.-C. Intramolecular radical cyclization of silylacetylenic or olefinic α-iodo ketones: application to the total synthesis of (±)-modhephene. Tetrahedron Lett. 31, 3745–3748 (1990).
Baldwin, J. E. Rules for ring closure. J. Chem. Soc., Chem. Commun. 18, 734–736 (1976).
Jorner, K. et al. Organic reactivity from mechanism to machine learning. Nat. Rev. Chem. 5, 240–255 (2021).
Reid, J. P. & Sigman, M. S. Holistic prediction of enantioselectivity in asymmetric catalysis. Nature 571, 343–348 (2019).
Ahneman, D. T. et al. Predicting reaction performance in C-N cross-coupling using machine learning. Science 360, 186–190 (2018).
Shields, B. J. et al. Bayesian reaction optimization as a tool for chemical synthesis. Nature 590, 89–96 (2021).
Zahrt, A. F. et al. Prediction of higher-selectivity catalysts by computer-driven workflow and machine learning. Science 363, eaau5631 (2019).
Schwaller, P., Vaucher, A., Laino, T. & Reymond, J.-L. Prediction of chemical reaction yields using deep learning. Mach. Learn.: Sci. Technol. 2, 015016 (2021).
Kim, D. E., Zweig, J. E. & Newhouse, T. R. Total synthesis of paspaline A and emindole PB enabled by computational augmentation of a transform-guided retrosynthetic strategy. J. Am. Chem. Soc. 141, 1479–1483 (2019).
Beckwith, A. L. J. & Schiesser, C. H. Regio- and stereo-selectivity of alkenyl radical ring closure: a theoretical study. Tetrahedron 41, 3925–3941 (1985).
Wenderski, T. A. et al. Principal component analysis as a tool for library design: a case study investigating natural products, brand-name drugs, natural product-like libraries, and drug-like libraries. Methods Mol. Biol. 1263, 225–242 (2015).
Estrada, J. G. et al. Response to comment on ‘Predicting reaction performance in C–N cross-coupling using machine learning’. Science 362, eaat8763 (2018).
Valerio, V., Mostinski, Y., Kotikalapudi, R. & Tsvelikhovsky, D. Stereo- and regioselective synthesis of tricyclic spirolactones by diastereoisomeric differentiation of a collective key precursor. Chem. Eur. J. 22, 2640–2647 (2016).
Siewert, J., Textor, A., Grond, S. & von Zezschwitz, P. Spirodionic acid, a novel metabolite from Streptomyces sp., part 2: total synthesis through a twofold Michael addition as a selective spiroannelation strategy. Chem. Eur. J. 13, 7424–7431 (2007).
Benohoud, M., Tuokko, S. & Pihko, P. M. Stereoselective hydrosilylation of enals and enones catalysed by palladium nanoparticles. Chem. Eur. J. 17, 8404–8413 (2011).
Chung, H.-M. et al. Rumphellclovane A, a novel clovane-related sesquiterpenoid from the gorgonian coral Rumphella antipathies. Tetrahedron Lett. 51, 2734–2736 (2010).
Matsumoto, T. et al. Lignan dicarboxylates and terpenoids from the flower buds of cananga odorata and their inhibitory effects on melanogenesis. J. Nat. Prod. 77, 990–999 (2014).
Acknowledgements
We are grateful for financial support from Yale University, the Sloan Foundation, Boehringer Ingelheim, Genentech and the National Institutes of Health (grant no. GR100045). Student support included Chemical Biology Training grant (no. T32 GM067543 to R.L.C.) and an Anderson Postdoctoral Fellowship (P.Z.). We gratefully acknowledge Yale University’s High-Performance Computing Center for providing resources for this work. F. Menges of Yale Chemical and Biophysical Instrumentation Center is gratefully acknowledged for obtaining the high-resolution mass spectrometry data.
Author information
Authors and Affiliations
Contributions
M.E., P.Z. and T.R.N. initiated the project. Y.Z. and J.E. synthesized clovan-2,9-dione. P.Z. and R.L.C. synthesized rumphellclovane A and canangaterpene II. J.E. and R.L.C. carried out DFT and nuclear magnetic resonance spectroscopy calculations. P.Z., M.E. and Y.Z. performed the ML modelling. P.Z. and J.E. carried out experimental validation. All co-authors wrote and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Synthesis thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editor: Peter Seavill, in collaboration with the Nature Synthesis team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Experimental Details, Supplementary Sections I–X, Figs. 1–15 and Tables 1–7.
Source data
Source Data Fig. 2
Reported 6-endo-trig radical cyclization yields with calculated ΔG of reactions.
Source Data Fig. 4
ML-predicted yields and experimental yields.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, P., Eun, J., Elkin, M. et al. A neural network model informs the total synthesis of clovane sesquiterpenoids. Nat. Synth 2, 527–534 (2023). https://doi.org/10.1038/s44160-023-00271-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s44160-023-00271-0
This article is cited by
-
Valence-isomer selective cycloaddition reaction of cycloheptatrienes-norcaradienes
Nature Communications (2024)
-
Predicting synthetic viability
Nature Synthesis (2023)