Abstract
Deserts represent an extreme challenge for photosynthetic life. Despite their aridity, they are often inhabited by diverse microscopic communities of cyanobacteria. These organisms are commonly found in lithic habitats, where they are partially sheltered from extremes of temperature and UV radiation. However, living under the rock surface imposes additional constraints, such as limited light availability, and enrichment of longer wavelengths than are typically usable for oxygenic photosynthesis. Some cyanobacteria from the genus Chroococcidiopsis can use this light to photosynthesize, in a process known as far-red light photoacclimation, or FaRLiP. This genus has commonly been reported from both hot and cold deserts. However, not all Chroococcidiopsis strains carry FaRLiP genes, thus motivating our study into the interplay between FaRLiP and extreme lithic environments. The abundance of sequence data and strains provided the necessary material for an in-depth phylogenetic study, involving spectroscopy, microscopy, and determination of pigment composition, as well as gene and genome analyses. Pigment analyses revealed the presence of red-shifted chlorophylls d and f in all FaRLiP strains tested. In addition, eight genus-level taxa were defined within the encompassing Chroococcidiopsidales, clarifying the phylogeny of this long-standing polyphyletic order. FaRLiP is near universally present in a generalist genus identified in a wide variety of environments, Chroococcidiopsis sensu stricto, while it is rare or absent in closely related, extremophile taxa, including those preferentially inhabiting deserts. This likely reflects the evolutionary process of gene loss in specialist lineages.
Similar content being viewed by others
Introduction
Environment shapes evolution, in both the macro- and microcosmos. Among bacteria, the oxygenic photosynthetic cyanobacteria have diversified to occupy numerous niches by synthesizing specific pigments, forming biofilms, or fixing atmospheric nitrogen [1, 2]. Early colonizing cyanobacteria are often the most significant carbon inputs in extreme, nutrient-poor environments, including deserts [3, 4]. Defined in this work as (hyper-)arid environments (Aridity Index <0.2, or Potential Evapotranspiration <400 mm for cold deserts) [5], they encompass places as varied as the Atacama Desert and University Valley in Antarctica, and are often subject to additional stresses, such as high or low temperatures, large daily temperature variations, ice crystal nucleation, and intense solar radiation [4]. For this reason, many desert cyanobacteria colonize the subsurface or the interior of rocks [4, 6, 7].
A microorganism commonly inhabiting deserts is the cyanobacterium Chroococcidiopsis. This genus is characterized by non-motile solitary spherical cells forming non-polarized agglomerations, and the ability to reproduce by non-motile baeocytes [8, 9]. It has been isolated from cold and hot deserts, hot springs, and spacecraft assembly clean rooms [7, 10,11,12,13,14,15,16], with some isolates even surviving space [17, 18]. Previous studies have highlighted extreme temperature tolerance as a lineage-specific trait, with different lineages characteristic of hot or cold deserts, respectively [15, 19]. Nevertheless, strains have also been found in seemingly moderate environments [12, 20, 21]. This environmental diversity, combined with the large number of strains available from culture collections, make Chroococcidiopsis an ideal test subject for the study of adaptations to extreme environments.
It has become clear that at least some members of the Chroococcidiopsis genus can also thrive in the absence of visible light, for instance in soil, biofilms, or within sedimentary rocks, by using far-red light / near-infrared radiation (~700–750 nm) [16, 22,23,24,25,26]. In order to do so, they undergo a process called far-red light photoacclimation (FaRLiP). This involves the extensive remodeling of the photosynthetic apparatus and the synthesis of new pigments, namely chlorophyll f, as well as traces (<1%) of chlorophyll d [23, 27, 28]. The significance of chlorophyll d in particular is an unsolved puzzle, as some studies locate it in the reaction center of far-red Photosystem II [23, 29], yet many strains appear to lack it altogether [21, 30,31,32,33,34]. In addition, FaRLiP relies on a 20-gene cluster encoding paralogues of Photosystem I, Photosystem II and phycobilisome components, as well as a phytochrome regulatory cascade [24]. It is present in a small but diverse range of cyanobacteria, and is mainly associated with shading by other photosynthetic organisms [22, 24]. However, certain types of rock such as quartz may create similar microenvironments through the preferential transmission of longer wavelengths [6, 16]. As desert Chroococcidiopsis strains have been frequently isolated from lithic habitats, they might provide models for understanding not only the evolution of far-red light photoacclimation, but also its role in the environment.
Some Chroococcidiopsis lineages are either hot or cold desert specialists [15, 19]. Other strains from moderate environments might be generalists, based on the wide spread of this genus [12, 20]. Comparing their FaRLiP capacities could highlight selective pressures and constraints for the habitat. Nevertheless, recent research casts doubt on taxonomically assigning certain extremophilic lineages to the same genus. For example, the lineage known as “Cold Desert Chroococcidiopsis” has recently been typified as the separate genus Aliterella [35]. Still, there is strong evidence of FaRLiP being present in the last common ancestor of Chroococcidiopsidales (the order containing Chroococcidiopsis), thereby motivating our research [22, 36]. Traditionally, many related strains have been intermixed and simplified as “Chroococcidiopsis”. Recent research has been addressing this topic, but an overarching view that combines fine-scale 16S rRNA gene, genomic and ecological data is still missing [37,38,39,40].
The availability of cultured strains sampled from all over the world, together with the recent expansion in sequencing data, including metadata and metagenome-assembled genomes (MAGs), enabled this investigation. Therefore, this work presents a fine-grained evolutionary history of far-red light photoacclimation within the genus Chroococcidiopsis, and reveals a stark contrast in FaRLiP maintenance in generalist versus specialist taxa. In the process, this work also clarifies phylogenetic relationships within this genus and its associated order.
Materials and methods
Cyanobacterial strains and cultivation
The 43 cyanobacterial strains were obtained from the culture collections PCC, SAG, CCALA, CCMEE, and BCCM/ULC (Table S1). The last two provided the extremophilic strains. All strains were grown in BG11 [9] at room temperature, apart from Antarctic BCCM/ULC strains (10 °C). Far-red light incubators used LEDs centered at 750 nm (LED750-03AU, Roithner). Cultures were grown in parallel under white light. The photon flux ranged between 10 and 30 μmol photons m–2 s–1.
Fluorescence microscopy and spectra
In order to investigate photosynthetic acclimation, fluorescence images and spectral scans of cyanobacterial samples were acquired with an inverted confocal laser scanning microscope (CLSM; Leica SP8) using a HC PL APO x63/1.40 oil CS2 objective. Strains were grown for more than 20 days under FRL, immobilized onto BG11 medium containing 1% agar (w/v) and excited at 488 nm. Fluorescence images were taken by detecting the emission from phycobilisomes and chlorophyll a at 660–700 nm and from chlorophyll f at 720–750 nm. CLSM lambda-scans were obtained by collecting the emission from 550 to 800 nm with 3 nm steps and 5 nm bandwidth. Images were analyzed with Leica LAS X (version 3.5.6) and FIJI [41]. Spectral analysis was performed using Jupyter Notebook (version 6) running Python 3.
Pigment analysis
Pigments were extracted and analyzed by HPLC using an Agilent 1100 HPLC system. Samples were run on a Supelco Discovery HS C18 column (5 μm particle size, 120 Å pore size, 250 × 4.6 mm) at 1 ml·min–1 and 40 ˚C. Solvent A was 64:16:20 (v/v/v) methanol:acetone:H2O, while solvent B was 80:20 (v/v) methanol:acetone. Solvent B was held at 50% for the first 2 min, increased linearly to 100% over 10 min, and was held there for 25 min. Elution of chlorophylls a, d, and f was detected by monitoring absorbance at 665, 696, and 705 nm, respectively.
DNA extraction and quantification
Genomic DNA (gDNA) was extracted using Quick-DNA Fungal/Bacterial Miniprep Kit (Zymo Research), with a longer bead-beating time (20 min). DNA concentration was measured with a NanoDrop spectrometer or a Qubit fluorometer (Thermo Fisher Scientific).
Amplification of marker genes
The ability of strains for FaRLiP was assessed by amplifying the far-red specific marker gene apcE2 by PCR from genomic DNA. This has been previously shown to be a simple, reliable assay in a wide variety of species [22, 42]. Two additional primer sets were used for the amplification of 16S rRNA gene and for the hypervariable intergenic sequence (ITS) between the 16S and 23 S rRNA genes. The PCR and extraction protocol used was the same as published before, with 25 ng gDNA per 25 µl reaction [22]. The amplicons obtained were sequenced (Microsynth Seqlab). For apcE2 amplicons in particular, 5ʹ tags were used as sequencing primers. All primers are listed in the Supplementary Material (Table S2).
Single-gene phylogenies and rooting
In addition to the amplicons obtained above, 16S rRNA and apcE2 gene sequences were also recovered from NCBI [43, 44]. Search settings for apcE2: BLASTp (nr database) and tBLASTn (WGS database, “cyanobacteria”). Search settings for 16S rRNA genes: BLASTn, nr, 1000 results. Query: 16S rRNA of Chroococcidiopsis thermalis PCC 7203. Sequences were aligned with MAFFT using Jalview [45, 46]. Phylogenies were built with RaxML on the CIPRES webserver [47, 48]. Settings: RAxML-HPC2 on XSEDE, model GTR CAT, bootstrap iterations 100. Trees were edited with iTOL [49] and Inkscape 0.92. For the graphical abstract, BioRender was used. Networks were built with Splitstree for alternative visualization [50]. Large branches with uncertain labels (e.g. “uncultured bacterium”), or outside the scope of this study, were removed. So were 16 rRNA genes showing high similarity to Chroococcidiopsis sequences, and assumed to be close relatives in previous work [38], but revealed by genome phylogenies to be distantly related [36]. “Chroococcidiopsis” sequences belonging to different orders were also largely filtered out (Table S8). Additional divergent sequences, including partial sequences (700 nt) were recovered through BLAST searches, for a total alignment length of 1284 ± 169 bp. For the apcE2 phylogeny, the root point was confirmed with Minimal Ancestral Deviation (MAD) [51] and previous work [22].
For the 16S rRNA gene phylogeny, rooting was done with Pleurocapsa sp. PCC 7327 as an outgroup. This strain is capable of FaRLiP, and branches out from just outside the group containing the Chroococcidiopsidales together with their closest relatives, the Nostocales [24, 36]. Except for the position of the Nostocales as an artifact in 16S rRNA gene trees, the branching pattern closely mirrored the genome tree and was therefore regarded as overall more accurate [36]. Using either Nostocales or Gloeobacter as an outgroup significantly changed this branching pattern, and was therefore considered less accurate. Previous work also shows this branching instability [15, 35, 37, 38, 52].
Sequence comparison of 16S rRNA genes
Simple sequence similarity can be a useful phylogenetic proxy. The 16S rRNA genes were aligned with MUSCLE in a ClustalW format [53]. The output was submitted to Clustal Omega in order to generate a percentage identity matrix (PIM) [54]. The matrix was parsed and statistics calculated with Python 3.7 using PyCharm (modules: pandas, statistics) [55].
RNA structure prediction
D1-D1’ and Box B loop structures from the ITS were predicted by running SPOT-RNA on a Linux server [56]. This included non-canonical base pairs/pseudoknots stabilized by tertiary interactions. Results were cross-checked with Mfold and RNA-fold [57, 58].
Genome data sourcing: sequencing and assembly
Based on their phylogenetic position, Chroococcidiopsis sp. SAG 39.79, SAG 2023, SAG 2025, and Chroococcopsis gigantea SAG 12.99 were sequenced using Illumina and Oxford Nanopore Technologies (Table S3). Samples were sequenced according to vendor protocols on a MiSeq and MinION (MinKNOW version 1.10.23), the former using Nextera XT v2 and the latter a R9.4.1 flowcell with base-calling via Albacore (version 2.1.7). In addition to these samples, a metagenomic dataset from the Atacama Desert (NCBI codes SRR2394720 and SRR2396013), associated with ignimbrite rock, was reassembled. Its sourcing is described in the following section.
Trimming and quality control was performed with BBDuk (settings: qtrim=r trimq=10 minlen=30 ktrim=r k = 23 mink=11 hdist=1 tpe tbo) [59]. Assembly was executed for each individual sample with MEGAHIT (settings: meta-sensitive, paired) [60] for the metagenome, and the hybrid assembler Unicycler for the cultured strains [61]. Samples were co-binned with vamb [62]. CheckM, Prokka, and the BBMap statswrapper module were used to estimate bin quality after refinement with anvi’o [59, 63,64,65]. The SAG 39.79 assembly was further improved with LongStitch (settings: tigmint-ntLink-arks) [66].
Genome-level analyses
Besides the locally sequenced samples, genomes were recovered by genus name, as well as by BLAST searches [43]. For clades with little to no representation, four additional metagenome bins were obtained: “ignimbrite12”, “ignimbrite01”, “Atacama+Negev” and “mojave”. The first two are associated with samples from the Atacama Desert (Fig. S3) [67]. Their raw data was shown to contain apcE2 fragments corresponding to Chroococcidiopsidales in a previous study using SearchSRA for data mining [22, 68]. Bin “Atacama+Negev” was identified from the literature as Chroococcidiopsis [16], and confirmed with a genome tree. It is representative for three near-identical FaRLiP endolithic bins from the Atacama and Negev deserts (IMG/MER bin used 3300037877_1; similar bins 3300039401_1 and 3300039404_1) [16]. Bin ‘mojave’ was originally sampled from the Mojave desert, and recovered with IMG/MER’s 16S rRNA-based BLAST search (original IMG/MER bin ID 3300034134_4; permission granted by Kirsten Fisher from California State University, USA) [69].
The genome phylogeny was built with OrthoFinder, and included 13 strains which belong to the Chroococcidiopsidales [70]. Orthofinder uses proteome inputs; where not already available, these were obtained from genomic sequences with Prokka [65]. Despite the different genus name, both the 16S rRNA gene and the genome labeled Scytonema millei VB511283 (JTJC00000000.3) clustered with Chroococcidiopsis strains.
To identify clade-specific genes, the genomes and metagenome bins were submitted to OrthoVenn2 [71]. Datasets chosen as representatives for their respective clades: Chroococcidiopsis thermalis PCC 7203, Gloeocapsa sp. PCC 7428, Aliterella atlantica CENA595, the original ignimbrite12 metagenome, the mislabeled Scytonema millei VB511283, Chroococcidiopsis sp. CCALA 051, Chroococcales cyanobacterium IPPAS B-1203, Mojave metagenome. As ignimbrite12 was the only representative of its clade, clade-specific genes were approximated by dataset-specific genes. Default settings. The orthologue sets recovered were submitted to BlastKOALA and KofamKOALA on the KEGG webserver [72, 73], with KEGG Mapper enabling a better understanding of genus-specific functional pathways [74].
Environment classification
Desert environments were classified into hot and cold depending on the main reason for their aridity: evaporation or low temperatures, respectively [5]. This has been typically used in cyanobacterial work [15]. Equivalents in the Köppen–Geiger climatic scheme: Bwh for hot deserts, and ET (polar or alpine) for cold deserts [75]. For the purposes of this study, the mild Atacama Desert was considered ‘hot’. In addition, a subset of hot deserts with large seasonal variations, where average monthly temperatures might drop below 0° C, were additionally labeled “hot-and-cold” [76]. This category also included sampling sites from unspecified deserts. For 16S rRNA genes, all NCBI metadata was considered, together with citing literature when necessary / available.
Results and discussion
Red-shifted chlorophylls d and f are extensively present in the genus Chroococcidiopsis
A broad range of 43 samples labeled as Chroococcidiopsis were obtained from culture collections. These strains are representative of either non-extreme environments (19) or deserts (20 from hot deserts, 4 from cold deserts) (Table S1). The strains were tested for their ability the perform FaRLiP by using a combination of molecular biology, biochemical and biophysical methods. A first screening approach was performed by PCR for the FaRLiP marker gene apcE2 (Table S2). This gene has always and exclusively been found co-occurring with chlF (psbA4), the gene encoding the chlorophyll f synthase [22]. Out of the 43 tested strains, 22 were positive for the presence of apcE2.
A phylogenetic tree was built with the amino acid sequences encoded by these 22 apcE2 genes, together with homologs recovered from the NCBI. The ApcE2 phylogeny is split into two main branches (Fig. 1A: I and II). The majority of sequences fall in group I, and are associated with a narrow definition of Chroococcidiopsis around the type strain Chroococcidiopsis thermalis PCC 7203 (hereafter referred to as Chroococcidiopsis sensu stricto). These strains have been largely sampled from non-extreme environments (Fig. 1B). However, sequences for “Chroococcidiopsis’ CCMEE 10, CCMEE 130, and ignimbrite12 (a metagenomics sequence from ignimbrite rock in the Atacama Desert) are clearly divergent from the others (Fig. 1A: II). These strains have been sampled from hot deserts and appear to belong to a different genus, yet to be defined.
A Phylogeny of ApcE2 marker sequences from Chroococcidopsis strains, or strains previously classified as such. Bullet points mark full-length proteins. There is a clear split between two branches, I and II, with the largest one (I) associated with Chroococcidiopsis sensu stricto. Type species Chroococcidiopsis thermalis PCC 7203 is listed in bold. B Distribution of FaRLiP in the genus Chroococcidiopsis sensu stricto (I). The 16S rRNA gene phylogeny includes strains which tested positive for apcE2 and/or strains with a known FaRLiP cluster (red). Most of them also survived under far-red light, except for a minority of non-axenic strains (CCALA 44, 47, 51, 52, 927). One strain tested negative for apcE2 and did not survive under FRL (black square). HPLC data showed the presence of chlorophylls d and f (circles) or only chlorophyll f, as reported in previous studies (triangles). Trees built with RaxML. Bootstrap values < 30 not shown.
The ApcE2 phylogeny (Fig. 1A) matches the 16S rRNA gene phylogeny (Fig. 1B), the genome tree discussed later, as well as a higher-resolution Chroococcidiopsis tree based on MALDI-TOF data [20]. These results are consistent with vertical descent driving the distribution of FaRLiP within this genus, as opposed to horizontal gene transfer as has previously been suggested for this phenotype [24]. As the apcE2 gene is an indirect, though fast marker for FaRLiP [22], additional methods were used to assess the cellular response. These included monitoring the long-term survival of the strains under far-red light (as judged by pigmentation), recording fluorescence emission spectra and determining pigment composition by HPLC. Out of the 22 apcE2-positive strains, 18 strains were selected for further analyses as representative of phylogenetic and environmental variation. Out of these, 13 strains survived in far-red light (Table S1). They included 11 strains from group I and 2 from group II. The remaining 5 cultures (group I) showed abundant non-cyanobacterial growth, which likely outcompeted the cyanobacteria under the test conditions. FaRLiP is a slow process, taking 12-14 days for full acclimation, during which cyanobacteria do not grow [77]. None of the 8 apcE2-negative strains tested survived these conditions.
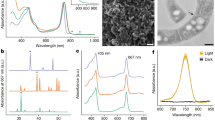
To accurately distinguish the FaRLiP cyanobacterial cells from contaminants on a single-cell level, we recorded fluorescence emission spectra using confocal microscopy. All strains surviving in far-red light showed emission peaks that indicated the presence of red-shifted chlorophylls in the photosystems (Fig. 2, or S1 for full results). The chlorophylls involved extend the emission range to 720–750 nm, in addition to the commonly found fluorescence attributed to chlorophyll a and phycobilisomes in the range of 640–700 nm at room temperature. This response is specific to far-red light and it does not occur under standard, white-light conditions [24]. As confocal microscopy is unable to identify the specific pigments, HPLC analysis was performed on a subset of 9 strains from groups I and II that were phylogenetically representative and exhibited low contamination levels.
The synthesis of red-shifted chlorophylls and their incorporation into photosystems can be seen in confocal micrographs (left). In addition to florescence emission from chlorophyll a and phycobilisomes (magenta, 660–700 nm), there is emission from chlorophyll f (yellow, 720–750 nm). Rightmost micrographs mark the overlay of both channels. Excitation, 488 nm. Scale bar, 10 µm. Spectral characteristics can be also observed through a fluorescence emission scan, with the aforementioned channels highlighted (second column from the right). Y-axis: Intensity (a.u.). Composite graph for a minimum of 3 cells. Pigments were extracted and analyzed by HPLC (right). Absorbance traces at 696 nm are shown, indicating the presence of chlorophyll d (16.3 min), f (16.7 min) and a (19.0 min).
All of the HPLC-tested strains contained chlorophylls a, d, and f when cultured under far-red light (Fig. 2, Table S1). Chromatogram peaks were visually consistent with the published percentages of approx. 90% chl a, 9% chl f and 1% chl d of the chlorophyll content [24]. Chlorophylls d and f were always observed in HPLC data for samples that showed a red-shifted fluorescence peak under far-red light, and these peaks were always observed for samples containing these red-shifted chlorophylls (Fig. 2). It has been shown that chlorophyll f is essential for FaRLiP [24], and the trace pigment chlorophyll d has been reported from the majority of strains undergoing this process [27]. However, the literature is divided on the significance of the latter. Some strains appear to lack chlorophyll d [21, 30,31,32,33,34], including Chroococcidiopsis strains from a subtropical forest [21]. These strains are 98.2-100% similar in 16S rRNA sequence to ours [21, 23]. All of our phylogenetically diverse samples contained chlorophyll d (Fig. 1B). This is consistent with previous studies indicating that Photosystem II (PSII) from far-red grown FaRLiP cyanobacteria contain a single chlorophyll d pigment in the reaction center, including in the type strain Chroococcidiopsis thermalis PCC 7203 [23, 29]. Since the amount of PSII per cell is known to vary depending on growth conditions [78], the detection of chlorophyll d might thus be more difficult under certain conditions and explain its apparent absence in some cases. Therefore, we suggest that having one chlorophyll d per PSII may be a conserved attribute of all FaRLiP strains. Future studies could consider growth phase in their chlorophyll d pigment analyses.
Overall, it is clear that the capacity for FaRLiP is correlated with the phylogenetic lineage. Specifically, within Chroococcidiopsis sensu stricto (I), all but one of 24 strains (96%), tested positive for chlorophyll f synthesis via multiple methods (marker gene presence, confocal microscopy, HPLC) (Fig. 1). In contrast, in lineage II, the majority of strains (18, or 73%) tested negative for chlorophyll f (Fig. S2). Out of 24 desert strains tested, only 3 were positive, belonging to lineage II. This seemingly contradicted our initial expectation that extreme endolithic environments would be enriched in FaRLiP cyanobacteria. Therefore, in order to investigate the connection between FaRLiP and habitat, and whether this is influenced by phylogenetic history, we set out to understand phylogenetic groupings within the Chroococcidiopsidales, and the adaptive niches they may be associated with.
Improved phylogeny of the order Chroococcidiopsidales
Triggered by the observed divergence in the ApcE2 tree (Fig. 1A), a 16S rRNA gene phylogeny (Fig. 3) and a sequence similarity network (Fig. S3) were built to better understand the evolutionary history of the order Chroococcidiopsidales. Trees based on 16S rRNA genes carry limited phylogenetic information and are therefore prone to issues such as unreliable branching patterns, polyphyly, and/or mislabeling. In contrast to them, genome-level phylogenies are less sensitive to noise, but full genomes are available for far fewer strains [36, 40]. This study took advantage of both methods, by building a genome phylogeny (Fig. 3B) and using it to correct and interpret the 16S rRNA gene phylogeny (Fig. 3A).
Phylogenetic trees of the Chroococcidiopsidales order based on 16S rRNA genes (A) and genome information (B). The two trees mirror and complement each other. In (A), it can be seen that the genus ‘Chroococcidiopsis’ is currently polyphyletic. Strains identified as such actually belong to multiple clades, some of which may be highly supported, but whose relationships with each other is sometimes unclear. Large, highly supported clades and/or clades with type species have been collapsed. Each color represents a genus-level taxon, some with associated genome information (B). These taxa include well-defined genera such as Chroococcidiopsis sensu stricto (I), Sinocapsa (IV), Aliterella (VI), Haliplanktos (VII) and Pseudocyanosarcina (VIII). They also encompass yet-undefined genera lacking a type species—Hot desert Chroococcidiopsidales (II) and ‘Additional desert clade’ (V). Moreover, ‘Gloeocapsa’ (III) represents a cluster of taxonomically ambiguous, closely related strains (Gloeocapsa, Gloeocapsopsis, Chroogloeocystis and Speleotes). Although showing low support in this tree, the Sinocapsa (IV) branch is stable across multiple tree-building methods. The related Nostocales are indicated by a gray rectangle (see “Materials and Methods”: Single-gene phylogenies and rooting). The tree was rooted with Pleurocapsa sp. PCC 7327, another FaRLiP strain, as an outgroup.
The resulting Chroococcidiopsidales genome tree covered a diversity of lineages, by combining publicly available NCBI data (12 genomes), metagenome bins (3) and locally sequenced strains (2). Of the latter, Chroococcidiopsis sp. SAG 2025 was sequenced to the level of a complete, circularized genome (Table S3). Combining our trees with other recent work, showed six highly supported clades (III-VIII) besides the two already mentioned (I-II) (Fig. 3; See Figs. S2, S4–S8 for individual trees) [35, 37,38,39]. These include five genera, such as Chroococcidiopsis sensu stricto (I), Sinocapsa (IV), Aliterella (previously ‘Cold desert Chroococcidiopsis’) (VI), Haliplanktos (VII) Pseudocyanosarcina (VIII), and the two additional taxa ‘Hot desert Chroococcidiopsidales’ (previously ‘Hot desert Chroococcidiopsis’) (II) and ‘Additional desert clade’ (V). One extra clade is represented by a monophyletic cluster of genera related to Gloeocapsa (III), with >95% similarity between 16S rRNA gene sequences. Because the taxonomy of Gloeocapsa-related strains requires revision (Fig. S4), for the purposes of this paper they will be discussed together.
All of these clades, including those yet to be described, are likely equivalent to genera. This can be seen from 16S rRNA gene similarities (>95% within clades) (Table S4) [79], from the shape of D1-D1’ (Fig. S9) and Box-B loops of the rRNA operon’s hypervariable region (Fig. S10), from shared genes (Fig. S11), as well as metadata. For ease of cross-referencing, names used in previous publications were collated (Table S5).
Although it was noted as a possibility from an early landmark paper [80], polyphyly within Chroococcidiopsis has been insufficiently acknowledged in the literature [15, 19]. This can lead to biased assumptions in comparative microbiological studies, which can propagate through later research. For example, a model organism used for cryo-EM studies of Photosystem I is Chroococcidiopsis sp. TS-821 [81], which we show actually belongs to ‘Gloeocapsa’ (III) (Fig. 3). Awareness of ‘Chroococcidiopsis’ diversity grew in recent years through the introduction of genera such as Aliterella and Sinocapsa [35, 37,38,39]. Our work highlights two additional genus-level clades (II and V) which have been commonly mislabeled as ‘Chroococcidiopsis’. The present phylogeny considers all the data available (16S rRNA genes, genomes, MAGs) to produce a state-of-the-art understanding of evolutionary relationships within the Chroococcidiopsidales, and assist further research on this order.
FaRLiP within the Chroococcidiopsidales is common to generalists, not specialists
By recovering sampling location data, many clades could be associated with particular (micro)environments, and were therefore defined as ‘specialists’ (Fig. 4). The majority of strains in specialist clades do not appear to have FaRLiP, indicative of gene loss. FaRLiP-lacking clades include ‘Gloeocapsa’ (III), which is strongly associated with hot spring or saline environments (Supplementary Text 1), and Aliterella (VI) which appears linked to cold deserts, such as polar and alpine regions (Supplementary Text 2).
This figure considers the temperature/humidity of the original sampling sites (inside row), as well as the capacity to utilize far-red light (outside row). Lineages are listed on the tree branches. Lineage II appears to be highly specialized to living in hot deserts. In contrast, Aliterella (VI) sequences have often found in cold deserts. Gloeocapsa and related strains (III) are characteristic of hot springs, while Chroococcidiopsis sensu stricto (I) appears to have a wider distribution. Chroococcidiopsis sensu stricto is the largest of the clades recovered, and it is strongly associated with the ability to perform FaRLiP. FaRLiP capacity was assessed by combining experimental and genome data. Strains listed as positive survived under far-red light and / or carried the entire FaRLiP gene cluster. Negative strains did not survive under far-red light, failing the apcE2 PCR assay, and / or lacked the gene within a complete genome. Tree built with RaxML. An extended version with labels can be accessed at https://itol.embl.de/tree/1301336668200071662129117.
In specialist lineage II, only 3 out of the 18 tested strains proved capable of FaRLiP. We defined this clade as ‘Hot desert Chroococcidiopsidales’. Out of 35 available 16S rRNA gene sequences, 33 were recovered from hypo-/endolithic niches in hot deserts (Fig. 4, S2). Adaptations to extremophilic life have been documented by considerable laboratory work (Supplementary Text 3) [15, 19]. The three FaRLiP strains are CCMEE 10, 12 and 130, with the first two identical in 16S rRNA gene sequences. Out of them, CCMEE 10 has been recently shown to only carry a partial, though functional, FaRLiP cluster, lacking Photosystem I genes [42]. The partial or total loss of FaRLiP genes seems surprising given that these strains are found in far-red light enriched habitats. It is possible that living in stable, extreme environments may result in less competition for light, leading to gene loss. Although gene loss in this phylum has been mainly studied in the highly reduced genomes of marine picocyanobacteria, the environmental stability hypothesized to underpin this genome optimization [36] might also apply to deserts.
The majority of FaRLiP-positive strains in this study fall under the classification of Chroococcidiopsis sensu stricto (I) (Fig. 4, Table S6). All but one of the tested strains in this genus were FaRLiP-positive. Considerable support exists for FaRLiP in this group, as the genus included most of the genomes, sequences and strains recovered. Strains within Chroococcidiopsis sensu stricto can be defined as generalists, as they appear to inhabit highly diverse environments. They have been found in freshwater, saltwater, as well as in dry soil and on rocks. However, they have only rarely been identified in hyperarid areas, unlike the closely related clade II, Hot desert Chroococcidiopsidales. Therefore, in contrast to the generally held view, they appear to be generalists with limited extremotolerance potential, supported by field and laboratory studies [16, 19]. Genome comparisons suggest that, in contrast to related genera, they may carry a conserved group of genes associated with hypoxic conditions (Supplementary Text 4). We hypothesize that these genes could have been co-selected, together with FaRLiP, for living in shaded environments such as soil or microbial mats.
These widespread generalist strains have mostly been underrepresented in discussions of Chroococcidiopsis phylogenetic history. Yet, not only do they form their own clade, but it is the largest among the known Chroococcidiopsidales. While this might be partially explained by sampling bias (non-extreme environments may be easier to sample), these strains appear to be extremely successful. They live in highly diverse environments, unlike those in extremophile genera. This might be due to higher competition, predation and many additional challenges in these moderate environments that are difficult to discern but shape evolution as much as the more obvious abiotic factors [82].
Not all lineages observed in this study were as straightforward to investigate as Chroococcidiopsis sensu stricto. No genomes, and very few strains, were available for clades IV-V and VII-VIII. Therefore, we were unable to assess their FaRLiP capacity. Regarding environments, for Sinocapsa (IV) and ‘Additional desert clade’ (V), there were at least sufficient 16S rRNA gene sequences available with associated metadata to suggest that these groups may be extremotolerant (Fig. 4). The latter group included sequences sampled a wide variety of desert environments (Utah, Tibet, Atacama). For groups VII-VIII, there were few sequences available, and no inferences could be made.
This study relied on recovering sampling locations from 16S rRNA datasets to balance out the limited strain availability. It proved particularly valuable for taxonomic groups for which cultured strains are rare, difficult to obtain/grow (as is the case with many polar microbes) or labeled with varying accuracy. It also allowed for correction of previous assumptions about prokaryote dispersal. Likely due to limited data, Chroococcidiopsis-related lineages have long been considered to be largely constrained to specific continents [15]. The present study, supported by additional work [12], contradicts this, showing that gene flow occurs frequently. Although nearby sampling locations may share closely related strains (sequences with >99% identity from similar locations were trimmed in our study) [15], this does not scale at higher taxonomic or geographic levels. For example, within a small branch of the Aliterella 16S rRNA gene tree, there are sequences from four continents, in no particular pattern. This is especially remarkable for extremophile clades, as deserts have been described functionally as “islands” [15]. Yet widespread geographic dispersal undoubtedly also plays a significant role for generalists, as they encounter diverse environments where survival may involve relying on various acclimation processes, such as FaRLiP.
Conclusion
Chroococcidiopsis has been previously defined as an extremophile cyanobacterium. However, many of the strains known under this name form separate genera. Some of them are specialists adapted to extreme environments such as hot or cold deserts; others are generalists. By combining bioinformatics data mining with large-scale laboratory assays, we have been able to connect genes, genomes, environments and adaptations. FaRLiP has been inherited from a common ancestor, and it remains near-universal in Chroococcidiopsis sensu stricto, a generalist genus. However, it has been commonly lost in specialist extremophile genera. It may have been co-selected with genes for hypoxic environments. Chlorophyll d was present in all our FaRLiP samples, supporting its predicted significant role in FaRLiP.
Data availability
The expanded version of the large 16S rRNA gene phylogeny can be accessed at iTOL (https://itol.embl.de/tree/1301336668200071662129117) (Legend in Figure S12). Genomic bins for Chroococcidiopsis sp. SAG 2025 and Chroococcopsis gigantea SAG 12.99 have been uploaded to the WGS database (JAOCNC and JAODIG, respectively). Refined ignimbrite bins and the Chroococcidiopsis sp. SAG 2023 genome can be found at figshare (https://figshare.com/projects/Chroococcidiopsis-related_metagenomic_data/149038). Accession numbers for additional sequences are listed in Table S7.
References
Quesada A, Vincent WF. Strategies of adaptation by antarctic cyanobacteria to ultraviolet radiation. Eur J Phycol. 1997;32:335–42.
Genuário DB, Corrêa DM, Komárek J, Fiore MF. Characterization of freshwater benthic biofilm-forming Hydrocoryne (Cyanobacteria) isolates from Antarctica. J Phycol. 2013;49:1142–53.
Makhalanyane TP, Valverde A, Gunnigle E, Frossard A, Ramond JB, Cowan DA. Microbial ecology of hot desert edaphic systems. FEMS Microbiol Rev. 2015;39:203–21.
Pointing SB, Belnap J. Microbial colonization and controls in dryland systems. Nat Rev Microbiol. 2012;10:551–62.
Cherlet M, Hutchinson C, Reynolds J, Hill J, Sommer S, von Maltitz G. World atlas of desertification. In: Middleton N, Thomas D, (eds). World atlas of desertification. 3rd ed. Luxembourg: Publications Office of the European Union; 2018. p. 72–74.
Schlesinger WH, Pippen JS, Wallenstein MD, Hofmockel KS, Klepeis DM, Mahall BE. Community composition and photosynthesis by photoautotrophs under quartz pebbles, southern Mojave Desert. Ecology. 2003;84:3222–31.
Friedmann EI, Ocampo R. Endolithic blue-green algae in the Dry Valleys: primary producers in the Antarctic Desert ecosystem. Science. 1976;193:1247–9.
Waterbury JB, Stanier RY. Patterns of growth and development in Pleurocapsalean cyanobacteria. Microbiol Rev. 1978;42:2–44.
Rippka R, Deruelles J, Waterbury JB. Generic assignments, strain histories and properties of pure cultures of cyanobacteria. J Gen Microbiol. 1979;111:1–61.
Billi D. Anhydrobiotic rock-inhabiting cyanobacteria: Potential for astrobiology and biotechnology. In: Stan-Lotter H, Fendrihan S, (eds). Adaption of Microbial Life to Environmental Extremes: Novel Research Results and Application. Vienna: Springer; 2012. p. 119–32.
Moissl-Eichinger C, Auerbach AK, Probst AJ, Mahnert A, Tom L, Piceno Y, et al. Quo vadis? Microbial profiling revealed strong effects of cleanroom maintenance and routes of contamination in indoor environments. Sci Rep. 2015;5:1–13.
Donner A. The case of Chroococcidiopsis: New phylogenetic and morphological insights into ecological important Cyanobacteria. Kaiserslautern, Germany: Doctoral dissertation, Technische Universität Kaiserslautern; 2013.
Chrismas NAM, Anesio AM, Sánchez-Baracaldo P. Multiple adaptations to polar and alpine environments within cyanobacteria: A phylogenomic and Bayesian approach. Front Microbiol. 2015;6:1–10.
Büdel B, Becker U, Follmann G. Sterflinger K. Algae, fungi, and lichens on inselbergs. In: Porembski S, Barthlott W, (eds). Inselbergs. Berlin, Heidelberg: Springer; 2000. p. 69–90.
Bahl J, Lau MCY, Smith GJD, Vijaykrishna D, Cary SC, Lacap DC, et al. Ancient origins determine global biogeography of hot and cold desert cyanobacteria. Nat Commun. 2011;2:1–6.
Murray B, Ertekin E, Dailey M, Soulier NT, Shen G, Bryant DA, et al. Adaptation of Cyanobacteria to the endolithic light spectrum in hyper-arid deserts. Microorganisms. 2022;10:1–12.
Cockell CS, Rettberg P, Rabbow E, Olsson-Francis K. Exposure of phototrophs to 548 days in low Earth orbit: Microbial selection pressures in outer space and on early earth. ISME J. 2011;5:1671–82.
Billi D, Staibano C, Verseux C, Fagliarone C, Mosca C, Baqué M, et al. Dried biofilms of desert strains of Chroococcidiopsis survived prolonged exposure to space and Mars-like conditions in Low Earth Orbit. Astrobiology. 2019;19:1008–17.
Fagliarone C, Mosca C, Ubaldi I, Verseux C, Baqué M, Wilmotte A, et al. Avoidance of protein oxidation correlates with the desiccation and radiation resistance of hot and cold desert strains of the cyanobacterium Chroococcidiopsis. Extremophiles. 2017;21:981–91.
Šebela M, Jahodářová E, Raus M, Lenobel R, Hašler P. Intact cell MALDI-TOF mass spectrometric analysis of Chroococcidiopsis cyanobacteria for classification purposes and identification of possible marker proteins. PLoS One. 2018;13:1–21.
Zhang ZC, Li ZK, Yin YC, Li Y, Jia Y, Chen M, et al. Widespread occurrence and unexpected diversity of red-shifted chlorophyll producing cyanobacteria in humid subtropical forest ecosystems. Environ Microbiol. 2019;21:1497–510.
Antonaru LA, Cardona T, Larkum AWD, Nürnberg DJ. Global distribution of a chlorophyll f cyanobacterial marker. ISME J. 2020;14:2275–87.
Nürnberg DJ, Morton J, Santabarbara S, Telfer A, Joliot P, Antonaru LA, et al. Photochemistry beyond the red limit in chlorophyll f–containing photosystems. Science. 2018;360:1210–3.
Gan F, Shen G, Bryant DA. Occurrence of far-red light photoacclimation (FaRLiP) in diverse cyanobacteria. Life. 2015;5:4–24.
Trampe E, Kühl M. Chlorophyll f distribution and dynamics in cyanobacterial beachrock biofilms. J Phycol. 2016;52:990–6.
Kühl M, Trampe E, Mosshammer M, Johnson M, Larkum AWD, Koren K. Substantial near-infrared radiation-driven photosynthesis of chlorophyll f-containing cyanobacteria in a natural habitat. Elife. 2020;1:1–15.
Gan F, Bryant DA. Adaptive and acclimative responses of cyanobacteria to far-red light. Environ Microbiol. 2015;17:3450–65.
Gan F, Zhang S, Rockwell NC, Martin SS, Lagarias JC, Bryant DA. Extensive remodeling of a cyanobacterial photosynthetic apparatus in far-red light. Science. 2014;345:1312–7.
Gisriel CJ, Shen G, Ho M-YY, Kurashov V, Flesher DA, Wang J, et al. Structure of a monomeric photosystem II core complex from a cyanobacterium acclimated to far-red light reveals the functions of chlorophylls d and f. J Biol Chem. 2022;298:1–15.
Behrendt L, Brejnrod A, Schliep M, Sørensen SJ, Larkum AWD, Kühl M. Chlorophyll f-driven photosynthesis in a cavernous cyanobacterium. ISME J. 2015;9:2108–11.
Akutsu S, Fujinuma D, Furukawa H, Watanabe T, Ohnishi-Kameyama M, Ono H, et al. Pigment analysis of a chlorophyll f-containing cyanobacterium strain KC1 isolated from Lake Biwa. Photomed Photobiol. 2011;33:35–40.
Averina S, Velichko N, Senatskaya E, Pinevich A. Far-red light photoadaptations in aquatic cyanobacteria. Hydrobiologia. 2018;813:1–17.
Chen M, Li Y, Birch D, Willows RD. A cyanobacterium that contains chlorophyll f - A red-absorbing photopigment. FEBS Lett. 2012;586:3249–54.
Ohkubo S, Miyashita H. A niche for cyanobacteria producing chlorophyll f within a microbial mat. ISME J. 2017;11:2368–78.
Rigonato J, Gama WA, Alvarenga DO, Branco LHZ, Brandini FP, Genuário DB, et al. Aliterella atlantica gen. nov., sp. nov., and Aliterella antarctica sp. nov., novel members of coccoid Cyanobacteria. Int J Syst Evol Microbiol. 2016;66:2853–61.
Chen MY, Teng WK, Zhao L, Hu CX, Zhou YK, Han BP, et al. Comparative genomics reveals insights into cyanobacterial evolution and habitat adaptation. ISME J. 2020;15:211–27.
Wang Y, Cai F, Jia N, Li R. Description of a novel coccoid cyanobacterial genus and species Sinocapsa zengkensis gen. nov. sp. nov. (Sinocapsaceae, incertae sedis), with taxonomic notes on genera in Chroococcidiopsidales. Phytotaxa. 2019;409:146–60.
Jung P, Brust K, Schultz M, Büdel B, Donner A, Lakatos M. Opening the gap: rare lichens with rare cyanobionts—unexpected cyanobiont diversity in cyanobacterial lichens of the order Lichinales. Front Microbiol. 2021;12:1–24.
Panou M, Gkelis S. Unravelling unknown cyanobacteria diversity linked with HCN production. Mol Phylogenet Evol. 2022;166:107322.
Strunecký O, Ivanova AP, Mareš J. An updated classification of cyanobacterial orders and families based on phylogenomic and polyphasic analysis. J Phycol. 2022;51:12–51.
Schindelin J, Arganda-Carreras I, Frise E, Kaynig V, Longair M, Pietzsch T, et al. Fiji: An open-source platform for biological-image analysis. Nat Methods. 2012;9:676–82.
Billi D, Napoli A, Mosca C, Fagliarone C, De Carolis R, Balbi A, et al. Identification of far-red light acclimation in an endolithic Chroococcidiopsis strain and associated genomic features: Implications for oxygenic photosynthesis on exoplanets. Front Microbiol. 2022;13:1–12.
Sayers EW, Cavanaugh M, Clark K, Ostell J, Pruitt KD, Karsch-Mizrachi I. GenBank. Nucleic Acids Res. 2020;48:D84–86.
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215:403–10.
Katoh K, Kuma KI, Toh H, Miyata T. MAFFT version 5: Improvement in accuracy of multiple sequence alignment. Nucleic Acids Res. 2005;33:511–8.
Waterhouse AM, Procter JB, Martin DMA, Clamp M, Barton GJ. Jalview Version 2-A multiple sequence alignment editor and analysis workbench. Bioinformatics. 2009;25:1189–91.
Stamatakis A. RAxML version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics. 2014;30:1312–13.
Miller MA, Pfeiffer W, Schwartz T. Creating the CIPRES Science Gateway for inference of large phylogenetic trees. 2010 Gatew. Comput. Environ. Work. New Orleans, LA, USA: IEEE; 2010. p. 1–8.
Letunic I, Bork P. Interactive tree of life (iTOL) v3: an online tool for the display and annotation of phylogenetic and other trees. Nucleic Acids Res. 2016;44:W242–45.
Huson DH, Bryant D. Application of phylogenetic networks in evolutionary studies. Mol Biol Evol. 2006;23:254–67.
Tria FDK, Landan G, Dagan T. Phylogenetic rooting using minimal ancestor deviation. Nat Ecol Evol. 2017;1:1–7.
Jung P, Mikhailyuk T, Emrich D, Baumann K, Dultz S, Büdel B. Shifting boundaries: ecological and geographical range extension based on three new species in the cyanobacterial genera Cyanocohniella, Oculatella, and Aliterella. J Phycol. 2020;56:1216–31.
Edgar RC. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004;32:1792–7.
Sievers F, Higgins DG. Clustal Omega for making accurate alignments of many protein sequences. Protein Sci. 2018;27:135–45.
McKinney W. Data structures for statistical computing in Python. Proc 9th Python Sci Conf. 2010;1697900:51–56.
Singh J, Hanson J, Paliwal K, Zhou Y. RNA secondary structure prediction using an ensemble of two-dimensional deep neural networks and transfer learning. Nat Commun. 2019;10:1–13.
Zuker M. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 2003;31:3406–15.
Lorenz R, Bernhart SH, Höner zu Siederdissen C, Tafer H, Flamm C, Stadler PF, et al. ViennaRNA Package 2.0. Algorithms Mol Biol. 2011;6:1–14.
Bushnell B. BBMap. BBMap short read aligner, and other bioinformatic tools. https://sourceforge.net/projects/bbmap/. Accessed 15 Oct 2020.
Li D, Liu CM, Luo R, Sadakane K, Lam TW. MEGAHIT: An ultra-fast single-node solution for large and complex metagenomics assembly via succinct de Bruijn graph. Bioinformatics. 2015;31:1674–6.
Wick RR, Judd LM, Gorrie CL, Holt KE. Unicycler: Resolving bacterial genome assemblies from short and long sequencing reads. PLoS Comput Biol. 2017;13:1–22.
Nissen JN, Johansen J, Allesøe RL, Sønderby CK, Armenteros JJA, Grønbech CH, et al. Improved metagenome binning and assembly using deep variational autoencoders. Nat Biotechnol. 2021;39:555–60.
Parks DH, Imelfort M, Skennerton CT, Hugenholtz P, Tyson GW. CheckM: Assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 2015;25:1043–55.
Eren AM, Kiefl E, Shaiber A, Veseli I, Miller SE, Schechter MS, et al. Community-led, integrated, reproducible multi-omics with anvi’o. Nat Microbiol. 2021;6:3–6.
Seemann T. Prokka: Rapid prokaryotic genome annotation. Bioinformatics. 2014;30:2068–9.
Coombe L, Li JX, Lo T, Wong J, Nikolic V, Warren RL, et al. LongStitch: high-quality genome assembly correction and scaffolding using long reads. BMC Bioinformatics. 2021;22:1–13.
Crits-Christoph A, Robinson CK, Ma B, Ravel J, Wierzchos J, Ascaso C, et al. Phylogenetic and functional substrate specificity for endolithic microbial communities in hyper-arid environments. Front Microbiol. 2016;7:1–15.
Levi K, Abeysinghe E, Rynge M, Edwards RA. Searching the sequence read archive using Jetstream and Wrangler. Pract. Exp. Adv. Res. Comput. ‘18 (PEARC ’18). 2018. Pittsburgh, PA, USA, pp 1–7.
Chen IMA, Chu K, Palaniappan K, Pillay M, Ratner A, Huang J, et al. IMG/M v.5.0: an integrated data management and comparative analysis system for microbial genomes and microbiomes. Nucleic Acids Res. 2019;47:D666–77.
Emms DM, Kelly S. OrthoFinder: phylogenetic orthology inference for comparative genomics. Genome Biol. 2019;20:1–14.
Xu L, Dong Z, Fang L, Luo Y, Wei Z, Guo H, et al. OrthoVenn2: a web server for whole-genome comparison and annotation of orthologous clusters across multiple species. Nucleic Acids Res. 2019;47:W52–58.
Kanehisa M, Sato Y, Morishima K. BlastKOALA and GhostKOALA: KEGG tools for functional characterization of genome and metagenome sequences. J Mol Biol. 2016;428:726–31.
Aramaki T, Blanc-Mathieu R, Endo H, Ohkubo K, Kanehisa M, Goto S, et al. KofamKOALA: KEGG Ortholog assignment based on profile HMM and adaptive score threshold. Bioinformatics. 2020;36:2251–2.
Kanehisa M, Sato Y. KEGG Mapper for inferring cellular functions from protein sequences. Protein Sci. 2020;29:28–35.
Beck HE, Zimmermann NE, McVicar TR, Vergopolan N, Berg A, Wood EF. Present and future Köppen-Geiger climate classification maps at 1-km resolution. Sci Data. 2018;5:1–12.
Feddema JJ. A revised Thornthwaite-type global climate classification. Phys Geogr. 2005;26:442–66.
Ho MY, Gan F, Shen G, Zhao C, Bryant DA. Far-red light photoacclimation (FaRLiP) in Synechococcus sp. PCC 7335: I. Regulation of FaRLiP gene expression. Photosynth Res. 2017;131:173–86.
Luimstra VM, Schuurmans JM, de Carvalho CFM, Matthijs HCP, Hellingwerf KJ, Huisman J. Exploring the low photosynthetic efficiency of cyanobacteria in blue light using a mutant lacking phycobilisomes. Photosynth Res. 2019;141:291–301.
Yarza P, Yilmaz P, Pruesse E, Glöckner FO, Ludwig W, Schleifer KH, et al. Uniting the classification of cultured and uncultured bacteria and archaea using 16S rRNA gene sequences. Nat Rev Microbiol. 2014;12:635–45.
Fewer D, Friedl T, Büdel B. Chroococcidiopsis and heterocyst-differentiating cyanobacteria are each other’s closest living relatives. Mol Phylogenet Evol. 2002;23:82–90.
Semchonok DA, Mondal J, Cooper CJ, Schlum K, Li M, Amin M, et al. Cryo-EM structure of a tetrameric photosystem I from Chroococcidiopsis TS-821, a thermophilic, unicellular, non-heterocyst-forming cyanobacterium. Plant Commun. 2022;3:1–18.
Feng W, Zhang Y, Lai Z, Qin S, Yan R, Sun Y, et al. Soil bacterial and eukaryotic co-occurrence networks across a desert climate gradient in northern China. L Degrad Dev. 2021;32:1938–50.
Acknowledgements
This research was supported by the Deutsche Forschungsgemeinschaft (DFG; Emmy Noether project award no. NU 421/1 to DJN), the Biotechnology and Biological Sciences Research Council (BBSRC; Grant ref. BB/W008076/1 to DPC), the National Institute for Health Research Health Protection Research Unit (NIHR HPRU) in Healthcare Associated Infections and Antimicrobial Resistance at the University of Oxford in partnership with Public Health England (PHE) (HPRU-2012-10041 to DJW) and the Italian Space Agency (ASI, DC-VUM-2017-034, Grant no. 2019-3 U.0 Life in Space to DB). LAA was supported by a postdoctoral fellowship from the Freie Universität Berlin and a Schrödinger fellowship from Imperial College London. DJW by a Big Data Institute Robertson Fellowship and a Sir Henry Dale Fellowship, jointly funded by the Wellcome Trust and the Royal Society (Grant ref. 101237/Z/13/B). We would also like to acknowledge the assistance of the Core Facility BioSupraMol supported by the DFG for confocal microscopy. The HPC facilities at Imperial College London and Freie Universität Berlin maintain the Linux clusters used for the computations. The BCCM/ULC collection of cyanobacteria at University of Liège (BE) is acknowledged for the gift of three strains. We thank Kirsten Fisher for allowing the use of the Mojave metagenome, Annick Wilmotte for the helpful discussions on the manuscript and David Griffiths for assistance with DNA sequencing. The views expressed are those of the author(s) and not necessarily those of the funders.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
DJN and LAA designed the study. DJN, DB and DJW acquired the research funds. DB and BB contributed strains and strain expertise. Cyanobacterial cultivation, the survival assay, as well as the PCR assay, were done by DJN, LAA, VMS, GDS and DB. Microscopy was performed by VMS and DJN. DPC designed the HPLC protocol; DPC and LAA performed the HPLC experiments. NDS, LB and DJW were responsible for genome sequencing. LAA performed the bioinformatics analyses. LAA and PJ collaborated on the phylogenetic trees. LAA and DJN wrote the manuscript with contributions from all co-authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Antonaru, L.A., Selinger, V.M., Jung, P. et al. Common loss of far-red light photoacclimation in cyanobacteria from hot and cold deserts: a case study in the Chroococcidiopsidales. ISME COMMUN. 3, 113 (2023). https://doi.org/10.1038/s43705-023-00319-4
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s43705-023-00319-4