Abstract
Metabolite exchange within marine microbial communities transfers carbon and other major elements through global cycles and forms the basis of microbial interactions. Yet lack of gene annotations and concern about the quality of existing ones remain major impediments to revealing currencies of carbon flux. We employed an arrayed mutant library of the marine bacterium Ruegeria pomeroyi DSS-3 to experimentally annotate substrates of organic compound transporter systems, using mutant growth and compound drawdown analyses to link transporters to their cognate substrates. Mutant experiments verified substrates for thirteen R. pomeroyi transporters. Four were previously hypothesized based on gene expression data (taurine, glucose/xylose, isethionate, and cadaverine/putrescine/spermidine); five were previously hypothesized based on homology to experimentally annotated transporters in other bacteria (citrate, glycerol, N-acetylglucosamine, fumarate/malate/succinate, and dimethylsulfoniopropionate); and four had no previous annotations (thymidine, carnitine, cysteate, and 3-hydroxybutyrate). These bring the total number of experimentally-verified organic carbon influx transporters to 18 of 126 in the R. pomeroyi genome. In a longitudinal study of a coastal phytoplankton bloom, expression patterns of the experimentally annotated transporters linked them to different stages of the bloom, and also led to the hypothesis that citrate and 3-hydroxybutyrate were among the most highly available bacterial substrates. Improved functional annotation of the gatekeepers of organic carbon uptake is critical for deciphering carbon flux and fate in microbial ecosystems.
Similar content being viewed by others
Introduction
The ocean microbiome plays a central role in mediating carbon and element cycles through its unique ability to process organic carbon dissolved in seawater [1,2,3]. Ultimately, bacteria take up and assimilate as much as half of marine net primary production (NPP) in the form of exometabolites derived from excretion and death of phytoplankton and other microbes [3, 4]. Given that current and future controls over this globally significant carbon flux are poorly understood, identification of the metabolites produced and consumed by ocean microbes is vitally important [5].
One approach to unraveling marine metabolite flux is through the application of transcriptomic and proteomic tools by which dynamics of the chemical environment can be gleaned from changes in the expression of microbial genes. Such approaches are easy to scale with advancements in sequencing and data sharing [6, 7] and have successfully addressed metabolite dynamics in various microbial systems such as model communities [8], phytoplankton blooms [9, 10], oligotrophic ocean regions [11, 12], and global-scale ocean surveys [13,14,15]. Transporter genes in particular are of value in such approaches because they are a cell’s interface with its environment and their expression can reveal the identity of available metabolites [16]. A key limitation to their use, however, is a dependence on accurate gene annotation to identify protein function. For most microbial transporters, the cognate substrate is still unknown. Others are annotated computationally based on homology [17,18,19], yet this is error prone when relationships to experimentally annotated genes are distant [20]. Indeed, transporters have a lower rate of successful annotation based on homology than catabolic enzymes [19].
Experimental confirmation of gene function is the gold standard for annotation, but is both time and resource intensive. Moreover, it is largely limited to cultured species for which genetic systems are available, leaving out much of the diversity represented in environmental bacteria. An alternate approach uses pooled transposon mutants whose fitness under defined selection pressure provides hypotheses of gene function [21,22,23,24]. This method requires only a minimal genetic system to introduce small DNA fragments (transposons) and a protein that catalyzes insertion of the transposons (transposase) into bacterial cells. A recent high-throughput advancement of this method, termed RB-TnSeq [25, 26], uses unique barcodes that link each transposon insertion site to the specific gene it disrupts, thereby allowing mutant pools to be analyzed for fitness through cost-effective amplicon sequencing. A wide taxonomic range of bacteria have been found amenable to RB-TnSeq library construction, resulting in hypothesis generation for gene functions that include stress response, metabolism, phage resistance, and transport [25, 27,28,29]. Hypotheses can be confirmed experimentally if targeted single-gene mutants are subsequently constructed, as for those predicting membrane proteins [28] and catabolic enzymes [30].
For a small number of well-studied model bacterial species, genome-wide mutant arrays have been constructed through painstaking targeted gene deletions to produce libraries of single-gene knockouts across the genome. Excellent tools for gene annotation, these arrayed libraries are currently available for species such as Escherichia coli [31], Acinetobacter baylyi [32], Bacillus subtilis [33, 34], and Salmonella enterica [35]. Pooled transposon mutant libraries have been used successfully as the starting material for such arrayed libraries [24, 36, 37], but require individual sequencing of tens of thousands of colonies to determine transposon insertion location.
Recently, a modification of the RB-TnSeq approach was used to create an inexpensive arrayed mutant library of the model marine bacterium Ruegeria pomeroyi DSS-3 [38]. The method took advantage of the ease of insertion site identification in RB-TnSeq libraries to map 270,000 barcodes to transposon insertions which were condensed into a library of 4991 mutants, similar to the approach used recently for the anaerobic gut microbe Bacteroides thetaiotaomicron [39]. R. pomeroyi is known for its ecological association with marine phytoplankton and ability to grow on plankton-derived metabolites [16, 40, 41], but to this point substrates of only four of the 126 putative organic compound influx transporters have been experimentally verified via knockout mutants: choline [42], dihydroxypropanesulfonate (DHPS) [41, 43], ectoine [44], and trimethylamine N-oxide [13]. Here we leveraged a set of 156 influx transporter mutants from the R. pomeroyi arrayed RB-TnSeq (arrayed-RB-TnSeq) library in high-throughput screens against 70 possible substrates to increase knowledge of transporter function. Resulting gene annotations were then applied to a longitudinal study of R. pomeroyi transcriptomes following introduction of the bacterium into Monterey Bay, CA, USA seawater on 14 dates during a dinoflagellate bloom [9] in the manner of an ecological invasion study [45]. The 13 newly verified transporter annotations provided insights into the metabolites serving roles as substrates to heterotrophic bacteria during a coastal bloom.
Methods
RB-TnSeq library generation and mapping
Full methods for generating and arraying the R. pomeroyi barcoded mutant library are provided in Mejia et al. [38]. Briefly, a pool of randomly barcoded transposon mutants was constructed according to Wetmore et al. [26] by conjugating R. pomeroyi DSS-3 with E.coli WM3064 containing the pKMW7 Tn5 barcoded library (strain APA766). The transposon insertion sites were subsequently linked to the unique barcodes through random barcode transposon-site sequencing (RB-TnSeq) [26]. RB-TnSeq was achieved by sequencing on an Illumina NovaSeq6000 with 150-bp paired-end reads (Novogene Co., Sacramento, CA). Bioinformatic analysis of the reads used custom scripts described in Wetmore et al. [26]. To construct the arrayed libraries, individual mutants were isolated on ½ YTSS solid medium amended with 100 µg ml−1 kanamycin. Colonies were picked after 2 d (Qpix2 automated colony picker; Molecular Devices, San Jose, CA) and arrayed into 384 well plates containing 80 µl of ½ YTSS + kanamycin broth. Plates were incubated at 30 oC for 2–5 d until visible growth appeared and then replicated. Glycerol was added to a final concentration of 20% and plates were frozen at −80 oC. To identify the mutant locations, barcodes were amplified in each well using a unique combination of indexed 16 forward and 24 reverse location primers, allowing indexing of the well position. These barcode amplicons were re-amplified with dual indexed Illumina adapters, and amplicons from this second round of PCR were pooled within a plate for sequencing [38]. Of the 270,510 unique barcodes originally present in the pooled RB-TnSeq library, the condensed RB-TnSeq arrayed library contained 4991 mutants covering 3048 of 4284 protein encoding genes with either 1 or 2 mutants, accounting for 71% of R. pomeroyi’s genes. From these, the first 156 putative organic compound influx transporter mutants identified in the arraying procedure were re-arrayed into two plates for subsequent screening (Table S1).
Growth screens
Mutant cultures were pre-grown overnight in ½ YTSS broth with 50 μg ml−1 kanamycin. Screens were performed in L1 minimal medium [46] (https://doi.org/10.17504/protocols.io.jvccn2w) modified to a salinity of 20 ppt and amended with ammonium (3 mM), kanamycin (50 μg ml−1), and phosphorus as PO43- at 36 μM. For the initial screen, overnight cultures of individual mutants (2 μl) were inoculated into 198 μl of modified L1 with a single substrate as the sole carbon source at 8 mM carbon. Plates were incubated at 25 oC with shaking, and optical density (OD600) was read at intervals of 6–12 h using a SpectraMax M3 (Molecular Devices, San Jose, CA) until cultures entered stationary phase at ~24–48 h. Mutants exhibiting phenotypes in the initial screen were moved to the targeted screen in which 4 replicate 200 µl mutant cultures were prepared by inoculating 2 μl of washed (3x) overnight culture into 96 well plates containing 198 μl modified L1 medium and a single substrate at 8 mM carbon. As a positive control, four wells with the same medium were inoculated with washed overnight cultures of the pooled-RB-TnSeq library, used as a proxy for wild-type R. pomeroyi growth but harboring a transposon/kanamycin resistance gene insertion. Cultures were grown at 25 oC in a Synergy H1 plate reader (BioTek, Winooski, VT, USA) shaking at 425 rpm for 68–72 h. OD600 readings were collected once each hour and corrected to a pathlength of 1 cm assuming a volume of 200 µl.
Mutant defect was identified by comparison to the OD600 achieved by the pooled-RB-TnSeq library (n = 4; ANOVA and TukeyHSD; p ≤ 0.05) (Table 1). Mutants with significantly lower OD600 on multiple substrates were regrown on rich medium to check for viability, and removed from further consideration if they broadly demonstrated poor growth; one mutant was removed after this viability check (SPO2952).
Metabolite drawdown screen
For each mutant-substrate pair identified from the growth screens, 3 replicate 220 µl cultures were prepared in 96 well plates by inoculating 3 μl of washed (3x) overnight mutant cultures into minimal medium containing the candidate substrate at 8 mM carbon. Cultures were grown shaking at 25 oC for 24 h or 36 h, depending on the growth rate supported by the carbon source. At termination, 200 µl of medium were collected and centrifuged at 3700 rpm for 10 min, and the supernatant was stored at −80 oC. Metabolite analysis was performed using a Bruker Avance lll 600 MHz spectrometer (Bruker, Billerica, MA, USA) equipped with a 5-mm TCI cryoprobe. Samples were prepared with addition of a deuterated phosphate buffer (30 mmol L−1, pH 7.4) and the internal standard 2,2-dimethyl-2-silapentane-5-sulfonate-d6 (DSS, 1 mmol L−1) (10:1 vol:vol) and transferred to 3 mm NMR tubes (Bruker). Data were acquired by a one dimensional 1H experiment with water suppression (noesypr1d, Bruker) at 298 K using TopSpin 3.6.4 (Bruker). For glycerol, a 1H J-resolved experiment (jresgpprqf) was used to avoid overlapping background peaks. Spectra were processed using NMRPipe on NMRbox [47, 48], and the processed data were analyzed using Metabolomics Toolbox (https://github.com/artedison/Edison_Lab_Shared_Metabolomics_UGA) and MATLAB R2022a (MathWorks). For quantification of metabolites, spectra were normalized to the DSS standard and peak area for representative peaks was calculated. TopSpin experiment settings, NMRpipe spectra processing parameters, and MATLAB data analysis scripts are available in Metabolomics Workbench (see Data Availability).
Pooled-RB-TnSeq experiment
Minimal medium was prepared for 10 substrates at 8 mM carbon in a 96 well plate, 180 μl per well (n = 4). Each well was inoculated with 20 μl of washed (3x) overnight culture of the R. pomeroyi pooled-RB-TnSeq library. After growth with shaking at 25 oC for 72 h, cultures were serially transferred into fresh media three times. After the third transfer, the full culture volume of each well (200 μl) was transferred to 800 μl of minimal medium with substrate for a final 72 h grow out at 25 oC. The libraries averaged 15.5 generations under selection (ranging from 13.5 for xylose to 18.5 for N-acetylglucosamine). These 1 ml cultures were then transferred to 1.5 ml tubes, pelleted by centrifugation at 8000 × g for 3 min, and stored at −80 oC until further processing. Genomic DNA was extracted from the cell pellets using the DNEasy blood and tissue kit (Qiagen, Hilden, Germany). PCR amplification of RB-TnSeq barcodes was performed using primers modified from Wetmore et al. [26] with PhusionHF master mix (Fisher, Pittsburgh, PA). An aliquot of 8 ng of product from each sample was pooled, purified using HiPrep beads (MagBio, Gaithersburg, MD, USA), and sequenced on a NextSeq SE150 Mid Output flow cell (SE150) at the Georgia Genomics and Bioinformatics Core Facility (Athens, Georgia, USA). The initial processing and demultiplexing of sequence data was performed using Perl scripts (MapTnSeq.pl, DesignRandomPool.pl, and MultiCodes.pl) provided in Wetmore et al. [26]. Following quality control, an average of 35,090 unique barcodes mapped to insertions that fell within the interior 10 to 90% of R. pomeroyi coding sequences. In total, 55 million reads were mapped to insertions in 3048 genes (out of 4284 protein-encoding genes in the R. pomeroyi genome) with a median of 404,513 mapped reads per sample. Further processing of demultiplexed reads was performed in R v3.6.1. Reads mapping to different insertion sites within the same coding sequence were pooled for subsequent analyses. Two sample T tests with multiple comparison adjustment (FDR) were used to identify enrichment or depletion of normalized reads for each transporter on a given substrate relative to all other substrates tested, a modification of the reference medium method for RB-TnSeq analysis described in Borchert et al. [49]. To display enrichment or depletion of multiple transporters and substrates on the same scale, we calculated the fold change of each transporter’s normalized abundance for a given substrate relative to the normalized abundance of that transporter on all other substrates.
Transporter expression during a Monterey Bay bloom
Processed R. pomeroyi transcriptome data (transcripts per million and Z-scores), metadata, and complete experimental methods are available elsewhere [9]. Briefly, on 14 days over 5 weeks, R. pomeroyi cells were added to 350 ml of unfiltered Monterey Bay surface water (n = 3). R. pomeroyi was inoculated at cell numbers equivalent to that of the natural heterotrophic bacteria. Subsequent sequencing analysis indicated that R. pomeroyi transcripts averaged 38% of the bacterial reads in the metatranscriptome datasets [50]. Cells were collected by filtration after 90 min and processed for RNAseq analysis.
Homologs in the Roseobacter group
Roseobacter strains with complete genomes were selected based on Simon et al. [51]. Phylogenic analysis of the 14 selected strains was carried out with a set of 117 single copy genes using GToTree v1.6.37 [52]. R. pomeroyi transporter genes with homologs in the other strains were identified by BLASTp using Diamond v2.0.14.152 [53], threshold: E ≤ 10−5 and identity ≥ 60% across the full sequence, and checked with a reciprocal best hits analysis. Data analysis and figure generation was performed with R v3.6.1. Manual checks of gene neighborhoods were performed when BLASTp results showed that multicomponent transporters were missing one or more component genes.
Results and discussion
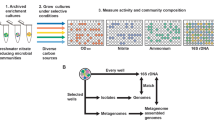
From a pooled-RB-TnSeq transposon mutant library of R. pomeroyi prepared according to Wetmore et al. [26], colonies were individually arrayed into 384 well plates (Fig. 1). The gene disrupted in each arrayed mutant was determined by sequencing the transposon barcode in conjunction with indexed primers that indicated plate column and row [38], creating a library that covers 3048 of 4284 protein-encoded genes in the R. pomeroyi genome (71%). From the genome annotation [54, 55], the first 156 mutants that were predicted to encode for 104 organic compound influx transporter proteins (Table S1) were re-arrayed into multi-well plates to facilitate functional screens using 70 compounds (Table S2) known to be produced by marine phytoplankton [56].
Growth screens identify 13 transporters of 18 substrates
Initial screens of the 156 mutants identified candidate substrates of transporter genes based on OD600 deficits after 24–72 h (n = 2). These mutants were transferred to a second round of screening in which each candidate substrate/mutant pair was monitored for growth with hourly OD readings and higher replication (n = 4). A positive control treatment consisting of the pooled-RB-TnSeq library approximated wild-type growth (Fig. 2A, S1). We used mutants of three previously confirmed transporters as positive controls for the screening protocols; these were the betT (SPO1087) for choline uptake [42], hpsKLM (SPO0591-0593) for DHPS uptake [41, 43], and uehABC (SPO1145-1147) for ectoine uptake [44] (Table 1, Fig. S1). One mutant (SPO2952) was excluded from consideration because it exhibited growth defects on multiple structurally dissimilar substrates (alanine, arginine, glycolate, malate, spermidine) in initial growth screens and pooled-RB-TnSeq screens, suggesting a general fitness defect not necessarily related to transport function.
a Growth of transporter mutants compared to growth of the pooled-RB-TnSeq library (an analog for wild-type growth but carrying transposon and resistance gene insertions) on selected marine plankton metabolites. Shaded regions indicate 95% confidence intervals (n = 4). Numbers refer to Ruegeria pomeroyi DSS-3 locus tags. b Substrate concentrations (1H-NMR peak area) after growth of mutants (brown symbols, n = 3) or the pooled-RB-TnSeq library (green symbols, n = 3), and at inoculation (gray symbols, n = 2). Letters that differ indicate that peak areas for the mutants were significantly higher than for the pooled-RB-TnSeq library (ANOVA, n = 3, p ≤ 0.05), with a TukeyHSD test carried out when multiple mutants for the same substrate were tested (p ≤ 0.05). For full results, see Table 1.
The growth-based screening resulted in substrate predictions for 13 R. pomeroyi transporters (Fig. 2A, S1, Table 1). Four of these were consistent with target metabolites hypothesized based on previous gene expression data: xylFGH (SPO0861-0863) (glucose/xylose) [57], iseKLM (SPO2356-2358) (isethionate) [58], potFGHI (SPO3466-3469) (polyamines: cadaverine, spermidine, and putrescine) [59], and tauABC (SPO0674-0676) (taurine) [60]. Four were consistent with target metabolites hypothesized based on in silico analysis by the GapMind tool for carbon sources, which combines pathway annotations and gap-filling using mutant fitness data:[19] tctABC (SPO0184-00186) (citrate), dctMPQ (SPO2626-2628) (C4 organic acids succinate, fumarate, and malate), nagTUVW (SPO1835, SPO1837-1839) (N-acetylglucosamine), and glpVSTPQ (SPO0608-0612) (glycerol). One was consistent with a target metabolite based on homology to an experimentally verified transporter in the closely related species Roseovarius nubinhibens:[61] dmdT (SPO3186) [dimethylsulfoniopropionate (DMSP)] (Table 1). Four were novel annotations with no previous substrate predictions: cntTUVW (SPO2995-2998) (carnitine), cuyTUVW (SPO3040-3043) (cysteate), hbtABC (SPO2571-2573) (3-hydroxybutyrate), and nupABC (SPO0376-0379) (thymidine) (Table 1). Previous studies had identified all hypothesized substrates as endometabolites in cultured phytoplankton or natural plankton communities or as exometabolites in phytoplankton cultures or seawater [56, 62,63,64,65,66,67,68,69].
Metabolite drawdown confirms gene knockout results
Substrate identifications emerging from the growth screens were further tested in metabolite drawdown experiments. Similar to the design of the growth screens, isolated mutants were inoculated into minimal medium with a single substrate as the sole carbon source (n = 3), alongside positive control treatments inoculated with the pooled-RB-TnSeq library as an analog for wild type. Spent media samples were collected at 24 h or, for substrates that supported slower growth, at 36-48 h. Substrate concentration was measured by 1H-NMR and a mutant drawdown defect was defined as significantly higher substrate concentration in the mutant cultures compared to the pooled-RB-TnSeq library (ANOVA and TukeyHSD, p ≤ 0.05, Table 1). All transporter annotations that had emerged from the growth screens were subsequently upheld in these drawdown screens (Fig. 2B, S2), consistent with gene disruption reducing or eliminating substrate uptake (Fig. 2A, S1).
Some transporter mutants, such as betT, were completely unable to grow on or draw down the substrate (Figs. S1, S2). This is the expected pattern if the disrupted transporter is the only system for uptake by R. pomeroyi. Alternatively, some mutants, such as dmdT, were capable of partial growth and drawdown, but significantly less than the mutant pool (Fig. 2, S1, S2). This pattern suggests that more than one transporter in the R. pomeroyi genome can take up the compound. For example, dmdT belongs to the BCCT-type family whose members frequently have low substrate affinity [70], suggesting to us that a second, high-affinity transporter was available when substrates became depleted. Indeed, a recent paper identified a high affinity DMSP transporter in R. pomeroyi (SPO2441-2443; dmpXWV) that, like dmdT, only partially explained observed DMSP uptake;[71] this second DMSP transporter now brings the number of experimentally verified transporters to 18. In a mixed result, complete loss of growth and drawdown for fumarate yet partial losses for succinate and malate suggests that dctMPQ is the only fumarate transporter system in the R. pomeroyi genome while the other C4 organic acids have a second transporter (Fig. 2B, S2, Table 1). We note that while we identified three transporters with multiple substrates (C4 organic acids, glucose/xylose, and multiple polyamines), additional multi-substrate transporters would be missed in our analysis if the other target substrate(s) was not among the 70 screened compounds. Further, the efficacy of growth-based screens for identifying transporter substrates is hampered by poor knowledge of the diversity of metabolites that support heterotrophic growth;[56] by the inability to test substrates that don’t support growth as a sole carbon source; by substrates that can be taken up by more than one transporter; and by the limited availability of transporter mutant collections that can facilitate matching transporters with their cognate substrates.
Pooled-RB-TnSeq studies are consistent with mutant screens
Another approach to identify substrates of bacterial transporters is to place a pooled-RB-TnSeq library under selection on a single carbon source [25]. In this case, transporter mutants that exhibit poor growth are identified as candidate uptake systems. We asked whether the pooled-RB-TnSeq approach would have been sufficient to recognize the R. pomeroyi transporters identified here, saving the effort of arraying the RB-TnSeq library while also providing additional information on catabolic and regulatory genes that may support metabolite utilization.
Mutant abundance was calculated for members of the pooled-RB-TnSeq library following selection for growth on ten substrates used in the growth screens (Fig. 3). Selection occurred over four growth dilution cycles of 72 h each. Amplicon sequencing of the pooled library at the beginning and end of selection [26] was used to calculate relative enrichment/depletion for each mutant in the pool as a proxy for fitness. For five substrates, the pooled RB-TnSeq results agreed with results from the arrayed mutant screens, identifying the same transporter systems for DHPS, ectoine, glucose, 3-hydroxybutyrate, and spermidine (n = 4; T test, p ≤ 0.05) (Fig. 3). For five other substrates, the known transporter mutant was either not significantly depleted from the mutant pool or significantly enriched, and thus transporters were not correctly identified for DMSP, malate, xylose, cysteate, and N-acetylglucosamine. Hypotheses for why these were not identified include the presence of a second transporter in the genome (DMSP, malate), poorer growth leading to weaker selection (xylose), and high substrate concentrations decreasing the need for the transporter substrate binding component (cysteate). In a counterintuitive finding, the N-acetylglucosamine transporter mutants (nagTUVW) were the most enriched populations in the pool, indicating a fitness gain for cells unable to take up the only provided substrate. We hypothesize that this was due to cross-feeding of an N-acetylglucosamine degradation product released by the other mutants and initially used only by the N-acetylglucosamine transporter mutant. While these results demonstrate that pooled-RB-TnSeq mutant libraries are excellent tools for low-cost, high-throughput hypothesis generation, predicted transporter annotations nonetheless require experimental follow-up [28, 30].
Green symbols indicate significant mutant depletion (t test, n = 4, Benjamini-Hochberg adjusted p ≤ 0.05) and gray symbols indicate non-significant changes. The larger filled symbols indicate the identified transporter for that metabolite as determined from growth and drawdown assays of individual mutants, and is colored green if it was correctly identified, and colored gray if not. Mutant enrichment/depletion for multi-gene transporter systems is plotted as the average of all components.
Transporter expression reveals the metabolite landscape of a coastal phytoplankton bloom
We used an R. pomeroyi gene expression dataset from a natural phytoplankton bloom in Fall 2016 in Monterey Bay, CA, USA [50] to assess the ecological relevance of the verified transporters. On 14 dates over 5 weeks during the decline of a bloom dominated by the dinoflagellate Akashiwo sanguinea, R. pomeroyi cells were introduced into samples from the natural community for 90 min [9]. Metatranscriptomic data from each sample were subsequently mapped to the R. pomeroyi genome to identify environmental conditions eliciting transcriptional responses on each sample date. We reanalyzed this dataset in light of the new information on transporter function, with the goal of generating insights into bloom-associated metabolites supporting heterotrophic bacterial growth.
To first evaluate the internal consistency of the expression data, pairwise correlation coefficients were calculated for the individual components of the 14 multi-component transporters across the sample dates. Nine systems had within-transporter correlation coefficients above 0.84 (Pearson correlation, p ≤ 0.05), confirming coherence in the expression patterns for genes in the same transporter system (Fig. 4a). The remaining four had within-transporter correlation coefficients ranging from 0.10 to 0.60; three of these, however, had particularly low expression in Monterey Bay (Fig. 4b) that may have affected analytical accuracy.
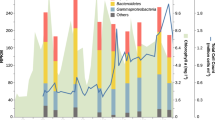
a Mean correlation coefficients of relative expression levels of genes within multi-component transporter systems in Monterey Bay, in Fall, 2016. b Mean relative expression levels for the 126 R. pomeroyi transporter systems when introduced into Monterey Bay seawater, averaged across 14 dates. c Relative expression of selected R. pomeroyi DSS-3 transporters in Monterey Bay seawater on each of 14 dates over 35 days, normalized as Z-scores. Expression of all verified transporters is shown in Fig. S3. Transporters have 1–5 component genes (each colored in different shades of brown) and each component gene has three replicates plotted individually. Lines connect the component mean expression through time. Total phytoplankton biomass (μg C L–1) during the 5-week sampling period is also shown.
Expression patterns of the carnitine, choline, taurine, and glycerol transporters were positively related to phytoplankton biomass through the bloom (Pearson correlation, p ≤ 0.05) (Fig. 4c, S3), and we hypothesize that these compounds were consistently present in the exometabolite pool. Expression of the C4 organic acid transporter and polyamine transporter had peaks coinciding with the largest drop in phytoplankton biomass (Fig. 4c), and we hypothesize that these compounds leaked from senescing or dead phytoplankton. Transcripts from R. pomeroyi’s 126 transporter systems were ranked by their abundance in the transcriptomes [mean transcripts per million (TPM), averaged across components for multi-gene transporters]. Making the assumption that heterotrophic bacterial transporter expression is regulated by substrate detection [admittedly an oversimplification [72]], citrate, 3-hydroxybutyrate, taurine, and DMSP, were among the most important sources of organic carbon to R. pomeroyi in this dinoflagellate-dominated bloom (expression ranked in the top 25% of transporters). Conversely, DHPS and cysteate were among the least important (ranked in the bottom 25%) (Fig. 4b, S3), both of which are found in endometabolites of diatoms and coccolithophores but not dinoflagellates [64]. The transporter for 3-hydroxybutyrate was of particular interest for two reasons. First, this newly annotated hbtABC is the only confirmed bacterial uptake system for 3-hydroxybutyrate [73]. Second, hbtABC was the third-most highly expressed R. pomeroyi transporter in Monterey Bay. While 3-hydroxybutyrate is well studied as the monomer of the bacterial storage compound polyhydroxybutanoate [74], it is not recognized as an ecologically-important component of the marine dissolved organic carbon pool. This is also the case for citrate, whose transporter was the most highly expressed of all the R. pomeroyi transporters (~5-fold higher than the second highest; Fig. 4b). Previous research showed that this central metabolite of the tricarboxylic acid (TCA) cycle and precursor for amino acid and cofactor biosynthesis was among the more abundant metabolites measured in phytoplankton cells in North Pacific surface seawater [62]. Thus its source in Monterey Bay is likely to be the ongoing phytoplankton bloom.
Orthologous transporters are present in other Roseobacter group members
R. pomeroyi and its relatives in the Roseobacter group are recognized for their high abundance in coastal marine environments [75, 76]. The cultured members of this group typically have large, well-regulated genomes capable of diverse metabolisms [77] and are often associated with phytoplankton blooms [78, 79]. To determine the distribution of the 17 verified transporters in Roseobacter genomes, 13 other strains having closed genomes and representing a broad sampling of the group’s phylogenetic diversity [51] were selected for analysis. Transporters for N-acetylglucosamine and carnitine are present only in close relatives of R. pomeroyi, consistent with vertical transmission (Fig. 5). Transporters for the organic sulfur compounds DHPS, taurine, and isethionate are common in deeply branching strains but retained in few lineages. The transporters for cysteate and ectoine are unique or nearly so to R. pomeroyi, suggestive of specialized niche dimensions. Finally, transporters for thymidine, citrate, glycerol, and 3-hydroxybutyrate are well conserved throughout Roseobacter genomes (Fig. 5), indicating broad importance of these substrates to the ecology of this group. Patchy distribution of transporter orthologs relative to the Roseobacter phylogeny has been reported previously [80].
Each row indicates a single gene and shading groups the genes that make up multi-component transporters. Black circles denote orthologs identified by BLASTp using e ≤ 10-5, identity thresholds ≥60%, and reciprocal best hit analysis. Black triangles denote orthologs of multicomponent transporters that did not meet the BLAST thresholds but were co-located in a transporter operon with components that did. Yellow circles denote potential orthologs in the outgroup A. macleodii ATCC27126 (identities <40%). Species phylogeny is based on analysis of 117 single copy genes.
Conclusions
Thirteen R. pomeroyi transporter annotations were confirmed in a screen of 70 metabolites against 156 transporter mutants representing 104 of the bacterium’s 126 organic carbon influx transporter systems. The verified gene functions provided new insights into in a longitudinal dataset of R. pomeroyi transcription through a natural phytoplankton bloom, revealing details of the metabolite landscape and generating hypotheses that citrate, 3-hydroxybutyrate, taurine, and DMSP were highly available metabolites during the dinoflagellate-dominated Monterey Bay bloom. Comparative analysis of the verified transporters across Roseobacter genomes revealed, on the one hand, narrow niche dimensions restricted to subgroups (R. pomeroyi and its closest relatives), and on the other, broad ecological characteristics common across the group and reflecting core ecological roles. As is the case for many marine bacterial taxa [81], the streamlined Roseobacter species that are more numerous in ocean microbial communities are poorly represented in culture collections [82]. As such, experimental gene annotation is central for analyzing, or re-analyzing, microbial gene, transcript, and protein data that harbor extensive untapped knowledge. For model organism R. pomeroyi, this study brings the percent of organic compound influx transporters with identified substrates to 14% of the 126 gene systems that acquire metabolites from the ocean’s organic carbon pools.
Data availability
All growth and RB-TnSeq data are available through BCO-DMO project 884792. All raw NMR data, processing scripts, and processed files for the metabolite drawdown experiment are available in Metabolomics Workbench with Study ID ST002381 (https://doi.org/10.21228/M8ST4T).
References
Azam F, Fenchel T, Field J, Grey J, Meyer-Reil L, Thingstad F. The ecological role of water-column microbes. Mar Ecol Prog Ser. 1983;10:257–63.
Anderson TR, Ducklow HW. Microbial loop carbon cycling in ocean environments studied using a simple steady-state model. Aquat Microb Ecol. 2001;26:37–49.
Moran MA, Ferrer‐González FX, Fu H, Nowinski B, Olofsson M, Powers MA, et al. The ocean’s labile DOC supply chain. Limnol Oceanogr. 2022;67:1007–21.
Cole JJ, Likens GE, Strayer DL. Photosynthetically produced dissolved organic carbon: An important carbon source for planktonic bacteria. Limnol Oceanogr. 1982;27:1080–90.
Cavicchioli R, Ripple WJ, Timmis KN, Azam F, Bakken LR, Baylis M, et al. Scientists’ warning to humanity: microorganisms and climate change. Nat Rev Microbiol. 2019;17:569–86.
Nayfach S, Roux S, Seshadri R, Udwary D, Varghese N, Schulz F, et al. A genomic catalog of Earth’s microbiomes. Nat Biotechnol. 2021;39:499–509.
Wang M, Carver JJ, Phelan VV, Sanchez LM, Garg N, Peng Y, et al. Sharing and community curation of mass spectrometry data with Global Natural Products Social Molecular Networking. Nat Biotechnol. 2016;34:828–37.
Amin S, Hmelo L, Van Tol H, Durham B, Carlson L, Heal K, et al. Interaction and signalling between a cosmopolitan phytoplankton and associated bacteria. Nature. 2015;522:98–101.
Nowinski B, Moran MA. Niche dimensions of a marine bacterium are identified using invasion studies in coastal seawater. Nat Microbiol. 2021;6:524–32.
Teeling H, Fuchs BM, Becher D, Klockow C, Gardebrecht A, Bennke CM, et al. Substrate-controlled succession of marine bacterioplankton populations induced by a phytoplankton bloom. Science. 2012;336:608–11.
Poretsky RS, Hewson I, Sun SL, Allen AE, Zehr JP, Moran MA. Comparative day/night metatranscriptomic analysis of microbial communities in the North Pacific subtropical gyre. Environ Microbiol. 2009;11:1358–75.
Sowell SM, Wilhelm LJ, Norbeck AD, Lipton MS, Nicora CD, Barofsky DF, et al. Transport functions dominate the SAR11 metaproteome at low-nutrient extremes in the Sargasso Sea. ISME J. 2009;3:93–105.
Lidbury I, Murrell JC, Chen Y. Trimethylamine N-oxide metabolism by abundant marine heterotrophic bacteria. Proc Natl Acad Sci USA. 2014;111:2710–5.
Tang K, Jiao N, Liu K, Zhang Y, Li S. Distribution and functions of TonB-dependent transporters in marine bacteria and environments: implications for dissolved organic matter utilization. PloS One. 2012;7:e41204.
Salazar G, Paoli L, Alberti A, Huerta-Cepas J, Ruscheweyh H-J, Cuenca M, et al. Gene expression changes and community turnover differentially shape the global ocean metatranscriptome. Cell. 2019;179:1068–83.
Ferrer-González FX, Widner B, Holderman NR, Glushka J, Edison AS, Kujawinski EB, et al. Resource partitioning of phytoplankton metabolites that support bacterial heterotrophy. ISME J. 2021;15:762–73.
Chang Y-C, Hu Z, Rachlin J, Anton BP, Kasif S, Roberts RJ, et al. COMBREX-DB: an experiment centered database of protein function: knowledge, predictions and knowledge gaps. Nucleic Acids Res. 2016;44:D330–D5.
Erbilgin O, Rübel O, Louie KB, Trinh M, Raad MD, Wildish T, et al. MAGI: A method for metabolite annotation and gene integration. ACS Chem Biol. 2019;14:704–14.
Price MN, Deutschbauer AM, Arkin AP. Filling gaps in bacterial catabolic pathways with computation and high-throughput genetics. PLoS Genet. 2022;18:e1010156.
Schnoes AM, Brown SD, Dodevski I, Babbitt PC. Annotation error in public databases: misannotation of molecular function in enzyme superfamilies. PLoS Comput Biol. 2009;5:e1000605.
Griffin JE, Gawronski JD, DeJesus MA, Ioerger TR, Akerley BJ, Sassetti CM. High-resolution phenotypic profiling defines genes essential for mycobacterial growth and cholesterol catabolism. PLoS Pathog. 2011;7:e1002251.
Van Opijnen T, Camilli A. A fine scale phenotype–genotype virulence map of a bacterial pathogen. Genome Res. 2012;22:2541–51.
Chao MC, Abel S, Davis BM, Waldor MK. The design and analysis of transposon insertion sequencing experiments. Nat Rev Microbiol. 2016;14:119–28.
Cameron DE, Urbach JM, Mekalanos JJ. A defined transposon mutant library and its use in identifying motility genes in Vibrio cholerae. Proc Natl Acad Sci USA. 2008;105:8736–41.
Price MN, Wetmore KM, Waters RJ, Callaghan M, Ray J, Liu H, et al. Mutant phenotypes for thousands of bacterial genes of unknown function. Nature. 2018;557:503–9.
Wetmore KM, Price MN, Waters RJ, Lamson JS, He J, Hoover CA, et al. Rapid quantification of mutant fitness in diverse bacteria by sequencing randomly bar-coded transposons. MBio. 2015;6:e00306–15.
Adler BA, Kazakov AE, Zhong C, Liu H, Kutter E, Lui LM, et al. The genetic basis of phage susceptibility, cross-resistance and host-range in Salmonella. Microbiology. 2021;167:001126.
Jensen HM, Eng T, Chubukov V, Herbert RA, Mukhopadhyay A. Improving membrane protein expression and function using genomic edits. Sci Rep. 2017;7:1–14.
Wehrs M, Thompson MG, Banerjee D, Prahl J-P, Morella NM, Barcelos CA, et al. Investigation of Bar-seq as a method to study population dynamics of Saccharomyces cerevisiae deletion library during bioreactor cultivation. Microb Cell Fact. 2020;19:1–15.
Clark IC, Youngblut M, Jacobsen G, Wetmore KM, Deutschbauer A, Lucas L, et al. Genetic dissection of chlorate respiration in Pseudomonas stutzeri PDA reveals syntrophic (per) chlorate reduction. Environ Microbiol. 2016;18:3342–54.
Baba T, Ara T, Hasegawa M, Takai Y, Okumura Y, Baba M, et al. Construction of Escherichia coli K‐12 in‐frame, single‐gene knockout mutants: the Keio collection. Mol Syst Biol. 2006;2:2006.0008.
De Berardinis V, Vallenet D, Castelli V, Besnard M, Pinet A, Cruaud C, et al. A complete collection of single‐gene deletion mutants of Acinetobacter baylyi ADP1. Mol Syst Biol. 2008;4:174.
Koo B-M, Kritikos G, Farelli JD, Todor H, Tong K, Kimsey H, et al. Construction and analysis of two genome-scale deletion libraries for Bacillus subtilis. Cell Syst. 2017;4:291–305.e7.
Kobayashi K, Ehrlich SD, Albertini A, Amati G, Andersen K, Arnaud M, et al. Essential Bacillus subtilis genes. Proc Natl Acad Sci USA. 2003;100:4678–83.
Porwollik S, Santiviago CA, Cheng P, Long F, Desai P, Fredlund J, et al. Defined single-gene and multi-gene deletion mutant collections in Salmonella enterica sv Typhimurium. PloS One. 2014;9:e99820.
Liberati NT, Urbach JM, Miyata S, Lee DG, Drenkard E, Wu G, et al. An ordered, nonredundant library of Pseudomonas aeruginosa strain PA14 transposon insertion mutants. Proc Natl Acad Sci USA. 2006;103:2833–8.
Ramage B, Erolin R, Held K, Gasper J, Weiss E, Brittnacher M, et al. Comprehensive arrayed transposon mutant library of Klebsiella pneumoniae outbreak strain KPNIH1. J Bacteriol. 2017;199:e00352–17.
Mejia C, Trujillo Rodriguez L, Poudel R, Ellington A, Rivers A, Reisch C An arrayed transposon library of Ruegeria pomeroyi DSS-3. bioRxiv. 2022:2022.09.11.507510.
Shiver AL, Culver R, Deutschbauer AM, Huang KC. Rapid ordering of barcoded transposon insertion libraries of anaerobic bacteria. Nat Protoc. 2021;16:3049–71.
Uchimiya M, Schroer W, Olofsson M, Edison AS, Moran MA. Diel investments in metabolite production and consumption in a model microbial system. ISME J. 2022;16:1–12.
Landa M, Burns AS, Roth SJ, Moran MA. Bacterial transcriptome remodeling during sequential co-culture with a marine dinoflagellate and diatom. ISME J. 2017;11:2677–90.
Lidbury I, Kimberley G, Scanlan DJ, Murrell JC, Chen Y. Comparative genomics and mutagenesis analyses of choline metabolism in the marine Roseobacter clade. Environ Microbiol. 2015;17:5048–62.
Mayer J, Huhn T, Habeck M, Denger K, Hollemeyer K, Cook AM. 2, 3-Dihydroxypropane-1-sulfonate degraded by Cupriavidus pinatubonensis JMP134: purification of dihydroxypropanesulfonate 3-dehydrogenase. Microbiology. 2010;156:1556–64.
Schulz A, Stöveken N, Binzen IM, Hoffmann T, Heider J, Bremer E. Feeding on compatible solutes: a substrate‐induced pathway for uptake and catabolism of ectoines and its genetic control by EnuR. Environ Microbiol. 2017;19:926–46.
Mallon CA, Van Elsas JD, Salles JF. Microbial invasions: the process, patterns, and mechanisms. Trends Microbiol. 2015;23:719–29.
Guillard R, Hargraves P. Stichochrysis immobilis is a diatom, not a chrysophyte. Phycologia. 1993;32:234–6.
Maciejewski MW, Schuyler AD, Gryk MR, Moraru II, Romero PR, Ulrich EL, et al. NMRbox: a resource for biomolecular NMR computation. Biophys J. 2017;112:1529–34.
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A. NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR. 1995;6:277–93.
Borchert AJ, Bleem A, Beckham GT. Experimental and analytical approaches for improving the resolution of randomly barcoded transposon insertion sequencing (RB-TnSeq) studies. ACS Synth Biol. 2022;11:2015–21.
Nowinski B, Smith CB, Thomas CM, Esson K, Marin R, Preston CM, et al. Microbial metagenomes and metatranscriptomes during a coastal phytoplankton bloom. Sci Data. 2019;6:1–7.
Simon M, Scheuner C, Meier-Kolthoff JP, Brinkhoff T, Wagner-Döbler I, Ulbrich M, et al. Phylogenomics of Rhodobacteraceae reveals evolutionary adaptation to marine and non-marine habitats. ISME J. 2017;11:1483–99.
Lee MD. GToTree: a user-friendly workflow for phylogenomics. Bioinformatics. 2019;35:4162–4.
Buchfink B, Reuter K, Drost H-G. Sensitive protein alignments at tree-of-life scale using DIAMOND. Nat Methods. 2021;18:366–8.
Rivers AR, Burns AS, Chan L-K, Moran MA. Experimental identification of small non-coding RNAs in the model marine bacterium Ruegeria pomeroyi DSS-3. Front Microbiol. 2016;7:380.
Moran MA, Buchan A, González JM, Heidelberg JF, Whitman WB, Kiene RP, et al. Genome sequence of Silicibacter pomeroyi reveals adaptations to the marine environment. Nature. 2004;432:910–3.
Moran MA, Kujawinski EB, Schroer WF, Amin SA, Bates NR, Bertrand EM, et al. Microbial metabolites in the marine carbon cycle. Nat Microbiol. 2022;7:508–23.
Wiegmann K, Hensler M, Wöhlbrand L, Ulbrich M, Schomburg D, Rabus R. Carbohydrate catabolism in Phaeobacter inhibens DSM 17395, a member of the marine Roseobacter clade. Appl Environ Microbiol. 2014;80:4725–37.
Weinitschke S, Sharma PI, Stingl U, Cook AM, Smits TH. Gene clusters involved in isethionate degradation by terrestrial and marine bacteria. Appl Environ Microbiol. 2010;76:618–21.
Mou X, Sun S, Rayapati P, Moran MA. Genes for transport and metabolism of spermidine in Ruegeria pomeroyi DSS-3 and other marine bacteria. Aquat Microb Ecol. 2010;58:311–21.
Gorzynska AK, Denger K, Cook AM, Smits TH. Inducible transcription of genes involved in taurine uptake and dissimilation by Silicibacter pomeroyi DSS-3. Arch Microbiol. 2006;185:402–6.
Sun L, Curson AR, Todd JD, Johnston AW. Diversity of DMSP transport in marine bacteria, revealed by genetic analyses. Biogeochemistry. 2012;110:121–30.
Boysen AK, Carlson LT, Durham BP, Groussman RD, Aylward FO, Ribalet F, et al. Particulate metabolites and transcripts reflect diel oscillations of microbial activity in the surface ocean. mSystems. 2021;6:e00896–20.
Dawson H, Heal K, Torstensson A, Carlson L, Ingalls A, Young J. Large diversity in nitrogen-and sulfur-containing compatible solute profiles in polar and temperate diatoms. Integr Comp Biol. 2020;60:1401–13.
Durham BP, Boysen AK, Carlson LT, Groussman RD, Heal KR, Cain KR, et al. Sulfonate-based networks between eukaryotic phytoplankton and heterotrophic bacteria in the surface ocean. Nat Microbiol. 2019;4:1706–15.
Durham BP, Boysen AK, Heal KR, Carlson LT, Boccamazzo R, Deodato CR, et al. Chemotaxonomic patterns in intracellular metabolites of marine microbial plankton. Front Mar Sci. 2022;9:1749.
Heal KR, Kellogg NA, Carlson LT, Lionheart RM, Ingalls AE. Metabolic consequences of cobalamin scarcity in the diatom Thalassiosira pseudonana as revealed through metabolomics. Protist. 2019;170:328–48.
Sacks JS, Heal KR, Boysen AK, Carlson LT, Ingalls AE. Quantification of dissolved metabolites in environmental samples through cation‐exchange solid‐phase extraction paired with liquid chromatography–mass spectrometry. Limnol Oceanogr Methods. 2022;20:683–700.
Shibl AA, Isaac A, Ochsenkühn MA, Cárdenas A, Fei C, Behringer G, et al. Diatom modulation of select bacteria through use of two unique secondary metabolites. Proc Natl Acad Sci USA. 2020;117:27445–55.
Weber L, Armenteros M, Soule MK, Longnecker K, Kujawinski EB, Apprill A. Extracellular reef metabolites across the protected Jardines de la Reina, Cuba reef system. Front Mar Sci. 2020;7:582161.
Chen C, Beattie GA. Pseudomonas syringae BetT is a low-affinity choline transporter that is responsible for superior osmoprotection by choline over glycine betaine. J Bacteriol. 2008;190:2717–25.
Li C-Y, Mausz MA, Murphy A, Zhang N, Chen X-L, Wang S-Y, et al. Ubiquitous occurrence of a dimethylsulfoniopropionate ABC transporter in abundant marine bacteria. ISME J. 2023;17:1–9.
Moran MA, Satinsky B, Gifford SM, Luo H, Rivers A, Chan L-K, et al. Sizing up metatranscriptomics. ISME J. 2013;7:237–43.
Mierziak J, Burgberger M, Wojtasik W. 3-hydroxybutyrate as a metabolite and a signal molecule regulating processes of living organisms. Biomolecules. 2021;11:402.
Findlay RH, White DC. Polymeric beta-hydroxyalkanoates from environmental samples and Bacillus megaterium. Appl Environ Microbiol. 1983;45:71–8.
González JM, Moran MA. Numerical dominance of a group of marine bacteria in the alpha-subclass of the class Proteobacteria in coastal seawater. Appl Environ Microbiol. 1997;63:4237–42.
Giebel H-A, Kalhoefer D, Lemke A, Thole S, Gahl-Janssen R, Simon M, et al. Distribution of Roseobacter RCA and SAR11 lineages in the North Sea and characteristics of an abundant RCA isolate. ISME J. 2011;5:8–19.
Luo H, Moran MA. Evolutionary ecology of the marine Roseobacter clade. Microbiol Mol Biol Rev. 2014;78:573–87.
González JM, Simó R, Massana R, Covert JS, Casamayor EO, Pedrós-Alió C, et al. Bacterial community structure associated with a dimethylsulfoniopropionate-producing North Atlantic algal bloom. Appl Environ Microbiol. 2000;66:4237–46.
Buchan A, LeCleir GR, Gulvik CA, González JM. Master recyclers: features and functions of bacteria associated with phytoplankton blooms. Nat Rev Microbiol. 2014;12:686–98.
Newton RJ, Griffin LE, Bowles KM, Meile C, Gifford S, Givens CE, et al. Genome characteristics of a generalist marine bacterial lineage. ISME J. 2010;4:784–98.
Stewart EJ. Growing unculturable bacteria. J Bacteriol. 2012;194:4151–60.
Buchan A, González JM, Moran MA. Overview of the marine Roseobacter lineage. Appl Environ Microbiol. 2005;71:5665–77.
Acknowledgements
We thank C. Smith, C. Sanlatte, and J. Schreier for advice and assistance. This work was supported by Simons Foundation grant 542391 to MAM within the Principles of Microbial Ecosystems Collaborative, and NSF award OCE-2019589 to MAM. This is the Center for Chemical Currencies of a Microbial Planet (C-CoMP) publication #022.
Author information
Authors and Affiliations
Contributions
WFS, MAM, and CRR conceived of and designed the research; WFS, HEK, CM, and LTR performed the experiments; MU performed NMR analysis; CM, LTR, and CRR generated and arrayed the mutant library; WFS and MAM conducted statistical analysis, generated figures, and wrote the manuscript with significant input from all co-authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Schroer, W.F., Kepner, H.E., Uchimiya, M. et al. Functional annotation and importance of marine bacterial transporters of plankton exometabolites. ISME COMMUN. 3, 37 (2023). https://doi.org/10.1038/s43705-023-00244-6
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s43705-023-00244-6