Abstract
Gene clusters rich in carbohydrate-active enzymes within Flavobacteriia genera provide a competitiveness for their hosts to degrade diatom-derived polysaccharides. One such widely distributed polysaccharide is glucuronomannan, a main cell wall component of diatoms. A conserved gene cluster putatively degrading glucuronomannan was found previously among various flavobacterial taxa in marine metagenomes. Here, we aimed to visualize two glycoside hydrolase family 92 genes coding for α-mannosidases with fluorescently-labeled polynucleotide probes using direct-geneFISH. Reliable in situ localization of single-copy genes was achieved with an efficiency up to 74% not only in the flavobacterial strains Polaribacter Hel1_33_49 and Formosa Hel1_33_131 but also in planktonic samples from the North Sea. In combination with high-resolution microscopy, direct-geneFISH gave visual evidence of the contrasting lifestyles of closely related Polaribacter species in those samples and allowed for the determination of gene distribution among attached and free-living cells. We also detected highly similar GH92 genes in yet unidentified taxa by broadening probe specificities, enabling a visualization of the functional trait in subpopulations across the borders of species and genera. Such a quantitative insight into the niche separation of flavobacterial taxa complements our understanding of the ecology of polysaccharide-degrading bacteria beyond omics-based techniques on a single-cell level.
Similar content being viewed by others
Introduction
Glucuronomannan is a universal polysaccharide found in diatom cell walls. Its backbone consists of α-1,3-mannose alternating with β-glucuronic acid residues and is often branched and sulphated [1,2,3,4]. Since diatoms are a major polysaccharide source and contribute substantially to the marine food web by attributing to 40% of primary production [5,6,7], it can be assumed that glucuronomannan is widely distributed wherever diatoms are found. Hence, marine bacteria are adapted to its degradation and, especially Bacteroidetes, can digest substrates rich in sulphated α-mannose using complementary carbohydrate active enzymes (CAZymes), such as glycoside hydrolases (GHs) of family 92 [8]. The GH92 family codes for exo-acting α-mannosidases and thus far, catalytic activity has been detected on α-1,2-, α-1,3-, α-1,4- and α-1,6-linked mannose [9]. GH92 genes are common in many flavobacterial species. Therefore, a co-evolution along with diverse cell wall components of algae seems plausible and would explain the many GH92 variants. Extensive analysis of metagenome and metaproteome data retrieved from bacterioplankton seawater samples revealed conserved GH92-rich polysaccharide utilization loci (PULs) that contain sulfatases and cluster for their SusC-like TonB-dependent transporters and SusD-like substrate-binding proteins (SusCD-like tandem pairs) [10]. Their specific combination of CAZymes strongly suggests a degradation potential for sulphated α-glucomannans such as glucuronomannan [11]. Hence, GH92s from these particular PULs may act in a concerted manner on the different linkage types of the glucuronomannan, maximizing cell wall degradation of diatoms. The highly similar sequences of these PULs were retrieved from species of the Formosa and Polaribacter genera, which recurrently dominated the bacteroidetal response to algae blooms in 2010–2012 in the North Sea [10]. It was also shown from bacterioplankton-derived (meta-) proteomic data, that the expression levels of GH92s were consistently high during phytoplankton spring blooms in the North Sea [12]. As well as being recurrent in bacterioplankton-derived metagenomic data from the North Sea [13], GH92 genes were also identified in samples from the North Atlantic Ocean [14], which underlines their ecological importance.
In this study, we investigated the in situ distribution of these conserved putative glucuronomannan PULs among various taxa in complex planktonic samples from a time series taken in the North Sea to visualize the hypothesis raised by the metagenomic analyses. To this end, we targeted the two most common variants of GH92 genes, GH92_a and GH92_b, in two representative strains of recurrently blooming flavobacterial clades isolated from the North Sea: Formosa Hel1_33_131 [15, 16] and Polaribacter Hel1_33_49 [15, 17, 18]. We applied direct-geneFISH [19], a molecular tool that uses fluorescence in situ hybridization (FISH), to directly link a taxonomic identity to a putative glucuronomannan degradation potential. The method identifies cells via 16S ribosomal RNA (rRNA)-targeted oligonucleotides and simultaneously detects the gene(s) of interest with double-stranded DNA polynucleotide probes (typical length of 300–500 bp), allowing the visualization of key genes directly within the bacterial host. Furthermore, we evaluated whether the specificity of gene-targeting polynucleotide probes can be broadened by relaxing the hybridization conditions. In contrast to oligonucleotide probes, which need to be applied under stringent conditions during hybridization to gain specific signals, polynucleotide probes tolerate a certain amount of mismatches without compromising specificity. This property can be used to detect genes within one species but also highly similar genes in closely related taxa. Consequently, applying direct-geneFISH at relaxed conditions would allow to analyze an environmental sample based on a functional trait rather than on taxonomic affiliation alone.
In combination with high-resolution microscopy, direct-geneFISH has the additional advantage of providing direct, visual evidence of individual lifestyles and a quantitative insight into niche separation, which extends beyond what can be predicted from metagenome studies. Investigating flavobacterial genetic repertoires and their potential to degrade polysaccharides in combination with their visualization can help to define the niche spaces of these recurring clades and to understand their contribution to the marine carbon cycle.
Material and methods
Plankton samples and representative pure cultures
Plankton samples were collected in the German Bight at Helgoland Roads, station ‘Kabeltonne’ (54° 11.3′ N, 07° 54.0′ E) in the framework of a time series as described in [12, 13]. We used subsamples from the fixed filters taken on 3rd April 2009, 20th and 26th April 2010, 26th April and 3rd May 2012. Pure cultures of Formosa Hel1_33_131 [15, 16] (UBA3537 according to Genome Taxonomy Database (GTDB) release 83 [20]) and Polaribacter Hel1_33_49 were previously obtained by dilution cultivation from the same sampling site on 20th April 2010 [15, 17]. Cells were grown in HaHa_100 medium [15], fixed in exponential phase with formaldehyde (2%) and filtered onto polycarbonate filters (0.2 μm pore size).
Synteny of putative glucuronomannan PULs
For analysis of PUL synteny, we retrieved all putative glucuronomannan PULs from 38 metagenomes taken at station ‘Kabeltonne’ in the years 2010–2012 during spring phytoplankton blooms, published in [10]. They were annotated as described therein and defined by the existence of at least one susC- or susD-like gene and at least two degradative CAZymes from the glycoside hydrolase or polysaccharide lyase families and additionally by a combination of predicted GH92 and sulfatase genes. We included representatives even if they occurred only in a single metagenome. All remaining PUL contigs were annotated using Prokka v 1.12 [21] (with adaptions for the –c and –m options within Prodigal as described in [22]) with manual annotation of the predicted genes and shortened to the genetic region of the PULs. Their taxonomic affiliation was determined using the metagenome-assembled genome (MAG) affiliation from [10] and the respective taxonomic classification using the GTDB-Tk genome-based taxonomy (GTDB-Tk v0.0.8 with GTDB release 83 [20]). PULs from Formosa Hel1_33_131 and Polaribacter Hel1_33_49 genomes were extracted accordingly. Respective GenBank files for all shortened contigs and genomes are available as Supplementary Data 1. The synteny map of the PULs in Fig. 1 was calculated with clinker to directly visualize the structural similarity on the protein level using GenBank files as input [23].
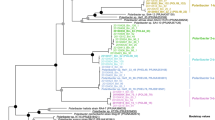
Synteny map of PULs likely encoding glucuronomannan degradation, retrieved from 2010–2012 Helgoland MAGs and isolates and a PUL found on a fosmid retrieved from the North Atlantic Ocean in 2006 [14]. The synteny map is aligned on the SusCD-like tandem pairs (SusC-like: orange, SusD-like: green). GH92_a are highlighted in yellow and GH92_b in magenta (coherent with the color-coding in following figures), all other GH92s in light blue. Sulfatases are displayed in gray. The synteny map was calculated with clinker [23] based on the amino acid sequences, but shown here are the corresponding nucleotide sequence identities. These numbers are only true for the immediate neighbors and are meant to showcase their remarkable similarity. PULs were sorted by complexity except for the PUL from the fosmid that was added for comparison at the top. Continuing contigs are indicated by “…” and end of contigs by “II”. The three GH92 genes for which polynucleotide probe sets were designed are marked with asterisks. NA = contig not assigned (binned) to a MAG, thus taxonomy unassigned.
Phylogenetic tree of Flavobacteriaceae in Helgoland metagenomes
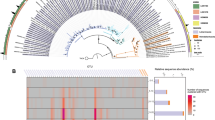
To visualize the distribution of glucuronomannan PULs within the family of Flavobacteriaceae, we constructed a reduced tree (Supplementary Fig. 1) based on the phylogenetic tree published in [10] using only sequences classified as Flavobacteriaceae. The tree was computed as in [10] with the exception of using RAxML v8.2.11 [24]. It was visualized using iTOL [25].
GH92 identities and taxonomic affiliation of non-PUL GH92s
Nucleotide and amino acid identity values for all GH92 genes on representative glucuronomannan PULs were calculated using multiple sequence alignments computed with MAFFT v7.017 [26] with E-INS-I algorithm. Pairwise identities and stretches of mismatches or gaps of each polynucleotide probe (FORM-GH92_a_1-5, FORM-GH92_b_1-5 and POL-GH92_a_1-5) to the corresponding GH92_a or GH92_b in PULs of Fig. 1 were calculated using Nucleotide-Nucleotide BLAST [27] within Geneious 8.1.8 (https://www.geneious.com) (Supplementary Table 1). To check for the presence of GH92 genes in the 38 bacterioplankton metagenomes that were not located within one of the putative glucuronomannan PULs but maintained a nucleotide sequence similarity of 80% or higher, we compared all sequences of the designed gene-targeted polynucleotide probes to all predicted GH92s in the metagenomes using Nucleotide-Nucleotide BLAST v2.8.1 [27]. A hit was accepted if at least four of five nucleotide probes from a set had a hit with ≧80%. Taxonomic affiliation of these hits was again determined using the MAG affiliation from [10] and the respective taxonomic classification using the GTDB-Tk genome-based taxonomy (Supplementary Table 2).
Oligonucleotide 16S rRNA probe design
All 16S rRNA oligonucleotide probes are summarized in Supplementary Table 3 and were ordered directly labeled with four fluorescent dyes from biomers.net GmbH (Ulm, Germany) that are specified in Supplementary Table 4. In this study, new helper oligonucleotides that flank the 16S rRNA probe FORM181B on either side were designed with ARB software [28] on the Silva SSURef NR 99 v128 16S rRNA sequence database [29], to improve accessibility for the probe to its target sequence.
Polynucleotide probes
For the design of the GH92-targeted polynucleotide probe sets, the published genomes of Formosa Hel1_33_131 (accession number in NCBI’s Genbank: CP017260.1) and Polaribacter Hel1_33_49 (JPDI00000000) were used. The three most widely distributed GH92 genes in our metagenome analyses served as targets for direct-geneFISH: FORM-GH92_a (gene locus tag: FORMB_02890, protein_id: AOR27350.1), FORM-GH92_b (gene locus tag: FORMB_02900, protein_id: AOR27351.1) and POL-GH92_a (gene locus tag: PHEL49_1329, protein_id: KGL60439.1). For each gene, a set of five consecutive polynucleotide probes of 442-456 bp length and their respective primer sets were designed with SnapGene® software (from GSL Biotech; available at snapgene.com) (Supplementary Table 5, Supplementary Data 2). The polynucleotide probes within each set were designed to have a similar melting temperature (Tm) and GC content to enable simultaneous hybridization. To investigate the stringency boundaries that determine the probe specificities in direct-geneFISH, we used the Wetmur formula for DNA:DNA hybrids, where [Na+] = molar Na+ concentration, GC = GC content [%], N = probe length [bp], M = mismatch [%] and FA = formamide concentration [%], [30]:
For each polynucleotide probe, the corresponding melting temperatures were calculated across varying parameters, such as formamide (FA) concentration and the respective allowed percentage of mismatch for the probe to the target DNA.
The polynucleotide probes were synthesized as described in [19] and either individually labeled or pooled in equi-molar probe mixes and then labeled with the Thermo Fisher Ulysis™ Alexa Fluor™ Nucleic Acid Labeling Kit (catalog numbers: Alexa Fluor® 488: U21650, Alexa Fluor® 594: U21654, Alexa Fluor® 647: U21660), according to the manufacturer’s instructions with the following modifications: The amount of dye added to 1 μg DNA was increased to 3 μl for Alexa Fluor® 488, 15 μl for Alexa Fluor® 594 and 10 μl for Alexa Fluor® 647, which successfully enhanced the degree of labeling (number of dyes per 100 bp) (Supplementary Table 6). The time for the labeling reagent to react with the probe DNA was increased to 30 min. Unbound dyes were removed with Bio-Spin™ P-30 Gel Columns (Tris Buffer, catalog number: 7326223, Bio-Rad, Hercules, California, USA).
As negative controls, we used polynucleotide probes that target a gene of unknown function (unk) originating from a Pseudoalteromonas phage PSA-HP1 (synthesized as described in [31], displayed as negative control (NC) in the figures), as well as hybridizations without any added gene probes (data not shown). To confirm that the unk-targeted polynucleotide probes used as NC did not hybridize to any DNA in the plankton samples, their probe sequences were blasted against all predicted genes within the analyzed 38 metagenomes and no hits were detected.
Direct-geneFISH
Direct-geneFISH was performed as described in [19] with a few modifications. The permeabilization step was omitted for Formosa Hel1_33_131 to preserve cell morphology, even though it decreased the detection efficiency for the gene slightly. Permeabilization of Polaribacter cells was performed for 5–10 min at room temperature with 0.5 mg ml−1 lysozyme in buffer (1× PBS, 0.05 M ethylenediaminetetraacetic acid (EDTA), 0.1 M Tris-HCl). Hybridized samples were counterstained with 4′,6-diamidino-2-phenylindole (DAPI) and mounted with ProLong™ Gold Antifade Mountant (catalog number: P36930, Thermo Fisher Scientific) on a microscope slide, covered with a #1.5 coverslip and cured for 24 h at room temperature.
Microscopy
The Airyscan detector upgrade on a confocal laser scanning microscope Zeiss LSM 780 was run in super-resolution mode (63× plan apochromatic oil immersion objective, 32 GaAsP detectors) [32]. The excitation lasers and emission detection windows are listed in Supplementary Table 7. For each field of view, a z-stack was taken and after post-processing with default Airyscan filtering, a 3D reconstruction was performed with Zen Black 2.1 software (Version 13.0.0.0, Carl Zeiss, Germany) with automatic Airyscan filter strength. The final images were obtained by a maximum intensity projection of the z-stack. The histogram of signal intensities was adjusted to maximize visibility of gene probe-conferred positive signals and the same values were used as thresholds to evaluate negative controls of the respective experiment to estimate eventual false positives. For better representation, fluorescent signals were consistently pseudocoloured, not resembling their original labeling dye color.
Gene detection efficiencies and relative gene abundances
For each direct-geneFISH polynucleotide probe set, we determined the gene detection efficiency under stringent hybridization conditions on pure cultures. Per experiment, we analyzed 3–8 fields of view in which 721–2747 cells were counted. The efficiencies are given as the mean of the percentages of gene-positive cells ((number of cells with gene signal/total number of cells) * 100) of each evaluated field of view and the corresponding standard deviation. In plankton samples, we report the fraction of 16S rRNA-targeted cells carrying a gene signal as the relative gene abundances (RGAs) in [%] and the corresponding standard deviation. Only cells with both a 16S rRNA-FISH and DAPI-DNA signal were considered for counting. Per experiment on plankton samples, we analyzed 6–20 fields of view in which 41–387 cells were counted. More details, such as the total number of cells counted per experiment, are given in Supplementary Table 4. P-values for evaluating differences in efficiencies were calculated using a two-tailed t-test assuming equal variances (confirmed by two-sample F-test for variances) in Excel.
Results
Synteny of glucuronomannan PULs
We adapted direct-geneFISH to target genes from conserved, putative glucuronomannan-degrading PULs in the flavobacterial groups Polaribacter and Formosa. A preceding analysis of (meta-) genomic data obtained from sampling campaigns in the German Bight in 2010–2012 indicated that the gene sequences of susCD-like tandem pairs retrieved from GH92- and sulfatase-rich PULs, clustered together based on their amino acid sequence similarity [10]. A more detailed analysis revealed that these PULs were not only taxonomically linked to representative strains and MAGs of the genera Formosa and Polaribacter [10], but also to MAGs affiliated to Aurantivirga and Algibacter (Fig. 1, Supplementary Fig. 1). We examined the synteny of these PULs and found that they share a high sequence identity for their SusCD-like proteins, GH92s and sulfatases, within and across genera (Fig. 1). Furthermore, we could confirm that the PULs share similarly high sequence identity with a PUL found on a fosmid affiliated to Polaribacter sp. (fosmid S3-860) retrieved from bacterioplankton samples taken in the North Atlantic Ocean in 2006 [14]. GH92 genes at the same positions within the PULs are homologous across strains, sharing a high percentage of sequence identity (Fig. 1). For example, FORM-GH92_b and POL-GH92_b share a nucleotide sequence identity of 86% (Fig. 1, Supplementary Table 8). However, the sequence comparison of the different GH92 genes within the same PUL showed they have less than 50% nucleotide sequence identity, most likely reflecting differences in substrate specificity of these exo-α-mannosidases. When compared to GH92 protein sequences within the CAZy database [33], no discrete specificity could be determined for GH92_a, while the closest characterized proteins to GH92_b and GH92_c were α-1,3-mannosidases. GH92_d and GH92_e clustered with α-1,2-mannosidases. Interestingly, while a homolog of GH92_b is present in all and of GH92_a in most of the putative glucuronomannan PULs in Fig. 1, other GH92 variants were found only sporadically, which could also be due to incomplete assemblies of the respective MAGs.
Gene detection efficiency of each gene-targeting probe set
For each gene-targeting polynucleotide probe set, we determined the gene detection efficiencies under stringent conditions in pure cultures of Polaribacter and Formosa. We aimed to target the most widely distributed GH92s in our metagenome analyses. The polynucleotide probe sets were designed based on the genomes of two representative strains to target the genes POL-GH92_a in Polaribacter Hel1_33_49 and FORM-GH92_a and FORM-GH92_b in Formosa Hel1_33_131. Each set consists of five adjacent probes between 442–456 bp in length (Supplementary Table 5). When blasted against each other, the longest stretch of mismatches between single probes and the target DNA from respective GH92 genes within examined PULs was 12 bp (Supplementary Table 1). The pairwise identity of each probe compared to the equivalent GH92 variant in Fig. 1 ranged between 73 and 100%, with an average of 88% (Supplementary Table 1). Per probe set, we could incorporate between 38 and 77 dyes (details in Supplementary Table 6). We performed direct-geneFISH successfully on pure cultures of Polaribacter Hel1_33_49 and for the probe set POL-GH92_a, we achieved a detection efficiency of 68.4% ± 7.9% (negative control (NC): 2.9% ± 0.4%) (Table 1, Fig. 2A). Additionally, we targeted the genes FORM-GH92_a and FORM-GH92_b individually in Formosa Hel1_33_131, which resulted in similar efficiencies: 74.3% ± 5.0% (NC: 1.8% ± 0.7%) and 54.5% ± 5.2% (NC: 0.0% ± 0.0%), respectively. Detecting both genes simultaneously in a double gene hybridization, the efficiencies reached 72.8% ± 6.5% for FORM-GH92_a and 72.7% ± 6.9% for FORM-GH92_b (NC: 1.8% ± 0.4%) (Table 1, Fig. 2B). For targeting the 16S rRNA, we used either species-specific probes (POL183a or FORM181B) or the general bacterial probe EUB338I-III.
Detection efficiencies of three probe sets targeting GH92 genes with direct-geneFISH in pure cultures of Polaribacter Hel1_33_49 and Formosa Hel1_33_131. A POL-GH92_a in Polaribacter was targeted with Alexa 488-labeled polynucleotide probes (yellow) and the 16S rRNA with the species-specific probe POL183a (cyan). B FORM-GH92_a in Formosa was targeted with Alexa 594-labeled polynucleotide probes (yellow), FORM-GH92_b with Alexa 647-labeled polynucleotide probes (magenta) and the 16S rRNA with a species-specific probe FORM181B or general bacterial probe EUB338I-III (cyan). The DNA was counterstained with DAPI (appears blue in “all channels”). Shown micrographs are maximum intensity projections of processed images achieved with Airyscan microscopy in super-resolution mode. NC = negative control, ns = not significant, scale bar: 1 µm, error bars in histograms represent standard deviations of the mean efficiencies. Experimental details are given in Table 1 and Supplementary Table 4.
Detection of relative gene abundance in plankton samples
In bacterioplankton samples collected during the phytoplankton spring bloom in the German Bight near Helgoland in 2010 relative abundances of GH92 genes were determined with direct-geneFISH. Based on catalyzed reporter deposition (CARD)-FISH counts, the relative abundance of cells affiliated to Polaribacter spp. “cluster 3a”, with the 16S rRNA-targeted probe POL183a, was 5.1% on 20th April 2010 (Fig. 3A, [18]). “Cluster 3a” was only detected in metagenomic reads from 2010 and includes the strain Polaribacter Hel1_33_49. The RGA of gene POL-GH92_a detected in DAPI-stained cells with a positive 16S rRNA probe signal (POL183a), was comparable to the gene detection efficiency in cultured representatives: 72.5% ± 9.6% (NC: 3.0% ± 3.8%) (Table 1, Fig. 3B). The relative abundance of Formosa cells (FORM181B) was 3.5% on 26th April 2010 (Fig. 3A) [13]. The RGAs for genes FORM-GH92_a and FORM-GH92_b were 55.7% ± 13.0% for FORM-GH92_a and 29.2% ± 11% for FORM-GH92_b (NC: 6.9% ± 2.1%) (Table 1, Fig. 3B). In plankton samples taken on 3rd April 2009, we obtained RGAs of 42.1% ± 29.5% for FORM-GH92_a and 31.9% ± 24.8% for FORM-GH92_b (NC: 4.2% ± 12.5%). More detailed information and additional experiments can be found in the Supplementary Text and Supplementary Table 4.
Detection of GH92 genes with direct-geneFISH in plankton samples taken during the spring bloom in the German Bight. A Relative cell abundances of Polaribacter spp. of “cluster 3a” targeted with the 16S rRNA probe POL183a and Formosa spp. targeted with the 16S rRNA probe FORM181B based on CARD-FISH counts [13]. Chlorophyll a concentration was taken as a proxy for algae biomass (gray area). Black arrows indicate dates from which samples were used for experiments (left: 20th April 2010, right: 26th April 2010). B Relative gene abundance (RGA), NC = negative control, error bars in histograms represent standard deviations of mean RGAs. Significance is indicated by asterisks (p ≤ 0.001). C Overview image to the magnified regions of interest to the right of Polaribacter spp. of “cluster 3a” targeted with the 16S rRNA probe POL183a and POL-GH92_a in a plankton sample. D Overview image to the magnified regions of interest to the right of Formosa spp. targeted with the 16S rRNA probe FORM181B and FORM-GH92_a and _b in a plankton sample. DNA was counterstained with DAPI (appears blue in ‘all channels’). Arrows in magenta and yellow show the respective gene signals and white arrows indicate gene-like signals in non-target organisms. Arrow in cyan points to a cell with a low ribosome content. All micrographs are maximum intensity projections of processed images achieved with Airyscan microscopy in super-resolution mode. Scale bar: 1 μm.
Broadening polynucleotide probe specificity to include close relatives
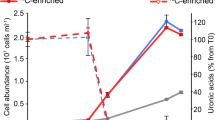
For an application on environmental samples, we aimed to broaden polynucleotide probe specificities to allow for the detection of similar genes beyond the borders of species and across genera. This was achieved by carefully relaxing the stringency during hybridization. The homologous genes FORM-GH92_b from Formosa and POL-GH92_b from Polaribacter share a nucleotide sequence identity of 86% (Supplementary Table 8). Theoretically, the gene POL-GH92_b can be detected with the probe set designed for FORM-GH92_b by relaxing the hybridization conditions and adjusting the FA concentration within the hybridization buffer from 35% to 15% (Supplementary Table 9). To test this hypothesis, we performed direct-geneFISH with the FORM-GH92_b gene probe set on pure cultures of Polaribacter Hel1_33_49 with hybridization buffers containing 35% and 15% FA. The GH92_b genes in Polaribacter were detected with an efficiency of 43.6% ± 1.9% (NC: 9.3% ± 5.5%) at relaxed conditions (15% FA) and with a significantly lower efficiency of 24.8% ± 11.6% (NC: 5.4% ± 1.0%) at stringent conditions (35% FA) (p < 0.01, Fig. 4A, Table 1). These results confirmed that the target specificity of the polynucleotide probe sets can be broadened by relaxing the stringency of hybridization conditions.
A Detection of GH92_b with FORM-GH92_b gene probe set in pure culture of Polaribacter Hel1_33_49 targeted with general bacterial 16 S rRNA probe EUB338I-III. B Detection of GH92_b with FORM-GH92_b gene probe set in Polaribacter spp. of “cluster 1a” (targeted with 16S rRNA probe POL405) in plankton samples taken in the German Bight on the 26th April 2012 and 3rd May 2012. Significance levels are indicated by asterisks (***: p ≤ 0.001, **: p ≤ 0.01), ns = not significant. Arrows in magenta show the detected gene signals, white arrows display gene-like signals in other non-target organisms, arrows in cyan point to cells that have a low ribosomal content. Micrographs display experiments marked with a cross in the bar plots. Shown micrographs are maximum intensity projections of processed images achieved with Airyscan microscopy in super-resolution mode. DNA was counterstained with DAPI (appears blue in ‘all channels’). RGA = relative gene abundance, NC = negative control, scale bar: 1 μm, error bars represent the standard deviations.
From the study of Avcı et al. [18], we know that Polaribacter spp. of “cluster 3a” were only abundant in the year 2010, whereas Polaribacter spp. of “cluster 1a” were abundant in 2012. “Cluster 1a” consists of four MAGs that are targeted by the 16S rRNA probe POL405, but only POL1a_84 harbors a putatively glucuronomannan degrading PUL [18] that clusters with the other PULs shown in Fig. 1. The GH92_b gene of MAG POL1a_84 has a nucleotide percentage identity of 87.7% with POL-GH92_b from Polaribacter Hel1_33_49 and 90.3% with FORM-GH92_b from Formosa Hel1_33_181 (blast analysis in Supplementary Table 8). Therefore, we combined the FORM-GH92_b probe set and the 16S rRNA probe POL405 to target “cluster 1a” and the respective GH92_b gene in two samples taken during the spring phytoplankton bloom in the German Bight in 2012 (Table 1, Fig. 4B). Indeed, we were able to detect the GH92_b gene from MAG POL1a_84 on 26th April 2012 with a relative abundance of 28.0% ± 5.0% (NC: 4.6% ± 1.0%) at relaxed conditions and with 9.1% ± 3.2% (NC: 2.5% ± 0.6%) at stringent conditions (p < 0.001). On 3rd May 2012, we could detect the gene with an RGA of 26.2% ± 8.4% (NC: 3.8% ± 1.3%) at relaxed conditions and with 21.9% ± 4.6% (NC: 4.7% ± 4.2%) at stringent conditions (p > 0.05). This is particularly interesting, since the relative abundance based on CARD-FISH counts was much lower on the second date (1.8%) than on the first date (12.7%) and yet, successful gene detection was possible.
In situ localization
By combining direct-geneFISH with high-resolution microscopy, we determined whether cells carrying a GH92 gene are rather particle-associated or free-living. On plankton samples from the German Bight of two consecutive years, only 18.6% ± 18.6% (3rd April 2009) and 19.4% ± 10.2% (26th April 2010) of cells identified with 16S rRNA probe FORM181B were attached to algal debris (Fig. 3D). In contrast, 58% ± 21.9% of cells of Polaribacter “cluster 3a” were attached to algal-like remnants on samples from 20th April 2010 (Fig. 3C, Supplementary Figs. 2, 3). Cells of Polaribacter “cluster 1a” however, were rather found free-living. In samples from two dates in 2012, only 13% ± 5.7% (26th April 2012) and 29% ± 7.1% (3rd May 2012) of the cells were attached to algae debris (Fig. 4B, Supplementary Fig. 4). The microscopic analysis also provided the opportunity to determine whether GH92 genes were over- or underrepresented in either the attached or the free-living fraction of cells, but both genes were nearly evenly distributed among both fractions (data not shown).
Discussion
In this study, we targeted the two most common GH92 variants from conserved and putatively glucuronomannan degrading PULs with direct-geneFISH. With an efficiency of up to 74%, we report, so far, the highest numbers for the detection of single-copy genes in pure cultures and environmental samples with (direct-) geneFISH (compare for example [19, 34, 35]).
After successful tests on two flavobacterial pure cultures, Polaribacter Hel1_33_49 and Formosa Hel1_33_131, we examined plankton samples from the German Bight taken in 2010. We could show that almost three quarter of Polaribacter individuals of “cluster 3a” carried the POL-GH92_b gene, reaching the same numbers as in pure culture experiments. More than half of all cells in plankton samples identified as Formosa sp. carried the FORM-GH92_a gene but only one third the FORM-GH92_b gene. These differences, however, were not a result of strain variation within Formosa spp., evidenced by the even distribution of read mapping coverage for both genes and the switching of the dyes for the gene-targeting probes that resulted in RGAs not differing significantly in gene proportions (see Supplementary Text for details). Therefore, we do not think that the different gene hybridization pattern represents a biological pattern, but suggest it is rather based on the delicate cells of Formosa. During direct-geneFISH, Formosa cells tend to break and leak cell content, which could lead to some loss of geneFISH signals (some cell disruption is also visible in Fig. 2B).
Even though it extends beyond the scope of this study, we hypothesize that the high similarity of our conserved glucuronomannan PULs that are unevenly distributed among four genera (Supplementary Fig. 1) suggests a horizontal gene transfer between them. Other studies already showed that CAZymes and entire syntenic PULs are commonly transferred between both closely and distantly related bacteroidetal species [36,37,38,39]. To detect possible gene variants across the borders of species and genera with direct-geneFISH, it is necessary to relax the hybridization conditions due to the less conserved nature of the targeted genes compared to the ribosomal RNA (see also [40]). Indeed, we could show that by relaxing the stringency of hybridization, the probe sets can be used to detect genes with up to 20% mismatches. Just recently, it was similarly shown in pure cultures that the detection of genes with mismatches of up to 20% is possible [34] using the original CARD-geneFISH protocol [35] and a single digoxigenin-labeled polynucleotide probe with antibody amplification, proving the robustness of this approach. Taking this to the next step, we successfully tested our hypothesis on North Sea plankton samples taken in 2012: Almost 30% of cells affiliated to Polaribacter sp. from “cluster 1a” carried the GH92_b gene detected with the Formosa-specific probe set FORM-GH92_b (Fig. 4B). Our experiments prove that by modulating the stringency via the FA-concentration, direct-geneFISH can provide an in situ link of a functional trait even to a subpopulation of yet uncultured Polaribacter spp. in plankton samples of which we only have evidence from metagenomic data. Our findings open up new perspectives for the detection and visualization of functional guilds that share a similar trait and degradation potential in their genomes but belong to different taxa [41]. To narrow down the identity of the host carrying the functional genes, the use of multiple polynucleotide probes with high signal intensities in combination with mild cell-fixation can be sufficient for fluorescence-activated flow cytometric cell sorting based on target genes. Sorted cells carrying the functional gene of interest can then be subjected to genome amplification and sequencing for taxonomic identification [42].
Micrographs of our plankton samples also displayed cells that showed gene-like signals without a corresponding 16S rRNA signal. Such signals could derive from bacterioplankton species from the four genera of Formosa, Polaribacter, Aurantivirga, and Algibacter that harbor that particular gene (or a highly similar version of it) but are not identified by the applied 16S rRNA probe (Supplementary Fig. 1). From the relative abundances of Formosa and Polaribacter spp. over the course of the spring bloom in 2010, we know that they were present on both sampling dates (Fig. 3A) and that their GH92s can be detected across genera with a smaller but yet pronounced detection rate (Fig. 4A). Therefore, the white arrows in Fig. 3D might point to cells which are affiliated to Polaribacter spp. that also carry the target gene. Another example can be seen in Fig. 4B, where GH92_b genes were detected in samples from 26th April 2012. The relative cell abundances were 12.7% for the 16S rRNA targeted probe POL405 (“cluster 1a”), 0.5% for POL183a (“cluster 3a”) and 0% for FORM181B [13, 18]. Thus, the white arrows in Fig. 4B most likely point to cells of “cluster 3a” that were also present on that date. However, blast analysis of all polynucleotide gene probe sequences against predicted GH92 genes in the spring bloom metagenomes yielded additional GH92 variants which are not encoded for in conserved glucuronomannan PULs, like those defined for Fig. 1. Still, they share a sequence similarity ≧80% and could be targeted by one of our probes sets (Supplementary Fig. 1, Supplementary Table 2). The GH92-containing contigs that were identified, taxonomically belonged exclusively to the four genera of Formosa, Polaribacter, Aurantivirga, and Algibacter and most of them even to the corresponding species or clusters shown in Fig. 1.
Combining direct-geneFISH with high-resolution microscopy also provides indications on the lifestyles of the targeted bacterioplankton clades. Earlier studies suggested a free-living lifestyle for Formosa clade A and B (Hel1_33_131), due to their rather small genome size [15], in contrast to the related Formosa agariphila for which an attached lifestyle was suggested [43]. Indeed, after thorough microscopic analysis of plankton samples taken from the German Bight in 2010, we confirmed that only 20% of the individual cells identified with 16S rRNA probe FORM181B were attached to algal debris (Fig. 3D). The evaluation of micrographs from the same spring bloom showed a strikingly different scheme for cells from Polaribacter “cluster 3a” (POL183a): 60% were, on average, attached to algal-like remnants (Fig. 3C, Supplementary Figs. 2, 3), even though previous studies suggested a free-living lifestyle due to their genome size and limited enzyme repertoire [17]. In contrast to that, the MAGs of Polaribacter “cluster 1a” harbor an extensive repertoire of CAZymes and PULs associated with the utilization of nine different polysaccharides, suggesting a specialization for growth on microalgae and brown algae with a potential attached lifestyle [18]. “Cluster 1a” includes several species-representative MAGs which is reflected in the different morphologies of the FISH-stained cells, ranging from coccoid to rod shaped (Supplementary Fig. 4). From the micrographs, we could further infer that, while being high in relative abundance on 26th April 2012 with 12.7%, only 13% of “cluster 1a” cells were attached to algae debris (Fig. 4). This was in stark contrast to their distribution only days later, when the relative abundance dropped to 1.8% on 3rd May 2012 and proportions of cells attached to algae and their remnants more than doubled. Our microscopy analysis of plankton samples suggests that even though the examined Formosa and Polaribacter spp. share the glucuronomannan-degradation potential encoded in conserved PULs, they appear to occupy different niche spaces in the same habitat.
In future experiments, it would be promising to also microscopically co-localize the polysaccharide, which is putatively degraded, by staining with specific fluorescently-labeled antibodies or carbohydrate binding modules [44] in combination with direct-geneFISH to visualize the CAZyme-encoding gene. We can envision a reliable application on sediment or soil samples if critical steps, such as cell permeabilization, are adapted beforehand. Also, the background autofluorescence of such samples has to be considered and either bleached or tackled with a careful choice of fluorophores and corresponding embedding medium. For example, Rahlff et al. [45] already performed direct-geneFISH on biofilms without any major alterations of the protocol.
We have shown that we can reliably detect GH92 genes with direct-geneFISH in pure cultures and plankton samples, visually linking function and phylogeny even across the borders of genera. Our results emphasize the importance to not only analyze metagenomic data but also to integrate FISH-based methods and (high-resolution) microscopy for visual evidence of individual lifestyles of defined bacterial clades.
References
McConville MJ, Wetherbee R, Bacic A. Subcellular location and composition of the wall and secreted extracellular sulphated polysaccharides/proteoglycans of the diatom Stauroneis amphioxys Gregory. Protoplasma. 1999;206:188–200. https://doi.org/10.1007/bf01279266
Chiovitti A, Harper RE, Willis A, Bacic A, Mulvaney P, Wetherbee R. Variations in the substituted 3-linked mannans closely associated with the silicified walls of diatoms. J Phycol. 2005;41:1154–61. https://doi.org/10.1111/j.1529-8817.2005.00140.x
Gügi B, Le Costaouëc T, Burel C, Lerouge P, Helbert W, Bardor M. Diatom-specific oligosaccharide and polysaccharide structures help to unravel biosynthetic capabilities in diatoms. Mar Drugs. 2015;13:5993–6018. https://doi.org/10.3390/md13095993
Le Costaouëc T, Unamunzaga C, Mantecon L, Helbert W. New structural insights into the cell-wall polysaccharide of the diatom Phaeodactylum tricornutum. Algal Res. 2017;26:172–9. https://doi.org/10.1016/j.algal.2017.07.021
Field CB. Primary production of the biosphere: integrating terrestrial and oceanic components. Science. 1998;281:237–40. https://doi.org/10.1126/science.281.5374.237
Tréguer P, Bowler C, Moriceau B, Dutkiewicz S, Gehlen M, Aumont O, et al. Influence of diatom diversity on the ocean biological carbon pump. Nat Geosci. 2017;11:27–37. https://doi.org/10.1038/s41561-017-0028-x
Arnosti C, Wietz M, Brinkhoff T, Hehemann JH, Probandt D, Zeugner L, et al. The biogeochemistry of marine polysaccharides: sources, inventories, and bacterial drivers of the carbohydrate cycle. Annu Rev Mar Sci. 2021;13:81–108. https://doi.org/10.1146/annurev-marine-032020-012810
Chen J, Robb CS, Unfried F, Kappelmann L, Markert S, Song T, et al. Alpha- and beta-mannan utilization by marine Bacteroidetes. Environ Microbiol. 2018;20:4127–40. https://doi.org/10.1111/1462-2920.14414
Zhu Y, Suits MDL, Thompson AJ, Chavan S, Dinev Z, Dumon C, et al. Mechanistic insights into a Ca2+-dependent family of α-mannosidases in a human gut symbiont. Nat Chem Biol. 2009;6:125–32. https://doi.org/10.1038/nchembio.278
Krüger K, Chafee M, Francis TB, del Rio TG, Becher D, Schweder T, et al. In marine Bacteroidetes the bulk of glycan degradation during algae blooms is mediated by few clades using a restricted set of genes. ISME J. 2019;13:2800–16. https://doi.org/10.1038/s41396-019-0476-y
Kappelmann L, Krüger K, Hehemann JH, Harder J, Markert S, Unfried F, et al. Polysaccharide utilization loci of North Sea Flavobacteriia as basis for using SusC/D-protein expression for predicting major phytoplankton glycans. ISME J. 2018;13:76–91. https://doi.org/10.1038/s41396-018-0242-6
Teeling H, Fuchs BM, Becher D, Klockow C, Gardebrecht A, Bennke CM, et al. Substrate-controlled succession of marine bacterioplankton populations induced by a phytoplankton bloom. Science. 2012;336:608–11. https://doi.org/10.1126/science.1218344
Teeling H, Fuchs BM, Bennke CM, Krüger K, Chafee M, Kappelmann L, et al. Recurring patterns in bacterioplankton dynamics during coastal spring algae blooms. eLife, 5, 2016. https://doi.org/10.7554/eLife.11888
Gómez-Pereira PR, Schüler M, Fuchs BM, Bennke C, Teeling H, Waldmann J, et al. Genomic content of uncultured Bacteroidetes from contrasting oceanic provinces in the North Atlantic Ocean. Environ Microbiol. 2012;14:52–66. https://doi.org/10.1111/j.1462-2920.2011.02555.x
Hahnke RL, Bennke CM, Fuchs BM, Mann AJ, Rhiel E, Teeling H, et al. Dilution cultivation of marine heterotrophic bacteria abundant after a spring phytoplankton bloom in the North Sea. Environ Microbiol. 2014;17:3515–26. https://doi.org/10.1111/1462-2920.12479
Unfried F, Becker S, Robb CS, Hehemann JH, Markert S, Heiden SE, et al. Adaptive mechanisms that provide competitive advantages to marine Bacteroidetes during microalgal blooms. ISME J. 2018;12:2894–906. https://doi.org/10.1038/s41396-018-0243-5
Xing P, Hahnke RL, Unfried F, Markert S, Huang S, Barbeyron T, et al. Niches of two polysaccharide-degrading Polaribacter isolates from the North Sea during a spring diatom bloom. ISME J. 2015;9:1410–22. https://doi.org/10.1038/ismej.2014.225
Avcı B, Krüger K, Fuchs BM, Teeling H, Amann RI. Polysaccharide niche partitioning of distinct Polaribacter clades during North Sea spring algal blooms. ISME J. 2020;14:1369–83. https://doi.org/10.1038/s41396-020-0601-y
Barrero-Canosa J, Moraru C, Zeugner L, Fuchs BM, Amann RI. Direct-geneFISH: a simplified protocol for the simultaneous detection and quantification of genes and rRNA in microorganisms. Environ Microbiol. 2017;19:70–82. https://doi.org/10.1111/1462-2920.13432
Parks DH, Chuvochina M, Waite DW, Rinke C, Skarshewski A, Chaumeil PA, et al. A standardized bacterial taxonomy based on genome phylogeny substantially revises the tree of life. Nat Biotechnol. 2018;36:996–1004. https://doi.org/10.1038/nbt.4229
Seemann T. Prokka: rapid prokaryotic genome annotation. Bioinformatics. 2014;30:2068–9. https://doi.org/10.1093/bioinformatics/btu153
Francis TB, Krüger K, Fuchs BM, Teeling H, Amann RI. Candidatus Prosiliicoccus vernus, a spring phytoplankton bloom associated member of the Flavobacteriaceae. Syst Appl Microbiol. 2019;42:41–53. https://doi.org/10.1016/j.syapm.2018.08.007
Gilchrist CLM, Chooi YH. clinker & clustermap.js: automatic generation of gene cluster comparison figures. Bioinformatics. 2021. https://doi.org/10.1093/bioinformatics/btab007
Stamatakis A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics. 2014;30:1312–3. https://doi.org/10.1093/bioinformatics/btu033
Letunic I, Bork P. Interactive tree of life (iTOL) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 2021;49:W293–W296. https://doi.org/10.1093/nar/gkab301
Katoh K, Standley DM. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol. 2013;30:772–80. https://doi.org/10.1093/molbev/mst010
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215:403–10. https://doi.org/10.1016/s0022-2836(05)80360-2
Ludwig W, Strunk O, Westram R, Richter L, Meier H, Yadhukumar, et al. ARB: a software environment for sequence data. Nucleic Acids Res. 2004;32:1363–71. https://doi.org/10.1093/nar/gkh293
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 2012;41:D590–D596. https://doi.org/10.1093/nar/gks1219
Wetmur JG. DNA probes: applications of the principles of nucleic acid hybridization. Crit Rev Biochem Mol Biol. 1991;26:227–59. https://doi.org/10.3109/10409239109114069
Allers E, Moraru C, Duhaime MB, Beneze E, Solonenko N, Barrero-Canosa J, et al. Single-cell and population level viral infection dynamics revealed by phageFISH, a method to visualize intracellular and free viruses. Environ Microbiol. 2013;15:2306–18. https://doi.org/10.1111/1462-2920.12100
Huff J. The Airyscan detector from ZEISS: confocal imaging with improved signal-to-noise ratio and super-resolution. Nat Methods. 2015;12:i–ii. https://doi.org/10.1038/nmeth.f.388
Lombard V, Ramulu HG, Drula E, Coutinho PM, Henrissat B. The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res. 2013;42:D490–D495. https://doi.org/10.1093/nar/gkt1178
Richards PM, Mattes TE. Detection of an alkene monooxygenase in vinyl chloride-oxidizing bacteria with GeneFISH. J Microbiol Methods. 2021;181:106147 https://doi.org/10.1016/j.mimet.2021.106147
Moraru C, Lam P, Fuchs BM, Kuypers MMM, Amann RI. GeneFISH—an in situ technique for linking gene presence and cell identity in environmental microorganisms. Environ Microbiol. 2010;12:3057–73. https://doi.org/10.1111/j.1462-2920.2010.02281.x
Ficko-Blean E, Préchoux A, Thomas F, Rochat T, Larocque R, Zhu Y, et al. Carrageenan catabolism is encoded by a complex regulon in marine heterotrophic bacteria. Nat Commun. 2017; 8. https://doi.org/10.1038/s41467-017-01832-6
Hehemann JH, Kelly AG, Pudlo NA, Martens EC, Boraston AB. Bacteria of the human gut microbiome catabolize red seaweed glycans with carbohydrate-active enzyme updates from extrinsic microbes. Proc Natl Acad Sci USA. 2012;109:19786–91. https://doi.org/10.1073/pnas.1211002109
Larsbrink J, Rogers TE, Hemsworth GR, McKee LS, Tauzin AS, Spadiut O, et al. A discrete genetic locus confers xyloglucan metabolism in select human gut Bacteroidetes. Nature. 2014;506:498–502. https://doi.org/10.1038/nature12907
Déjean G, Tamura K, Cabrera A, Jain N, Pudlo NA, Pereira G, et al. Synergy between cell surface glycosidases and glycan-binding proteins dictates the utilization of specific beta(1,3)-glucans by human gut Bacteroides. mBio. 2020;11. https://doi.org/10.1128/mbio.00095-20
Moraru C, Moraru G, Fuchs BM, Amann RI. Concepts and software for a rational design of polynucleotide probes. Environ Microbiol Rep. 2011;3:69–78. https://doi.org/10.1111/j.1758-2229.2010.00189.x
Barton AD, Pershing AJ, Litchman E, Record NR, Edwards KF, Finkel ZV, et al. The biogeography of marine plankton traits. Ecol Lett. 2013;16:522–34. https://doi.org/10.1111/ele.12063
Grieb A, Bowers RM, Oggerin M, Goudeau D, Lee J, Malmstrom RR, et al. A pipeline for targeted metagenomics of environmental bacteria. Microbiome. 2020;8. https://doi.org/10.1186/s40168-020-0790-7
Mann AJ, Hahnke RL, Huang S, Werner J, Xing P, Barbeyron T, et al. The genome of the alga-associated marine flavobacterium Formosa agariphila KMM 3901T reveals a broad potential for degradation of algal polysaccharides. Appl Environ Microbiol. 2013;79:6813–22. https://doi.org/10.1128/aem.01937-13
Vidal-Melgosa S, Pedersen HL, Schückel J, Arnal G, Dumon C, Amby DB, et al. A new versatile microarray-based method for high throughput screening of carbohydrate-active enzymes. J Biol Chem. 2015;290:9020–36. https://doi.org/10.1074/jbc.m114.630673
Rahlff J, Turzynski V, Esser SP, Monsees I, Bornemann TLV, Figueroa-Gonzalez PA, et al. Lytic archaeal viruses infect abundant primary producers in Earth’s crust. Nat Commun. 2021;12. https://doi.org/10.1038/s41467-021-24803-4
Acknowledgements
We thank Hanno Teeling and Thomas Ben Francis for in-depth discussions. Special thanks to Andreas Ellrott for excellent technical support with the microscopes and to Miriam Philippi for her support in the laboratory, for many fruitful discussions and critical reading of the manuscript. We are grateful to Greta Giljan and Taylor Priest for critical reading of the manuscript. This study was funded by the Max Planck Society and Laura E. Zeugner is a member of the International Max Planck Research School of Marine Microbiology (MarMic).
Author information
Authors and Affiliations
Contributions
RIA, BMF, LEZ, and JBC conceived and designed the analysis. LEZ performed the experiments, microscopy and corresponding data analysis. KK contributed the bioinformatic data analysis of MAGs, PULs and GH92 gene sequences. All authors contributed to the interpretation of the retrieved data. LZE, BFU, RIA, and KK drafted the manuscript and wrote the paper. All authors revised the article critically and approved the version to be published.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zeugner, L.E., Krüger, K., Barrero-Canosa, J. et al. In situ visualization of glycoside hydrolase family 92 genes in marine flavobacteria. ISME COMMUN. 1, 81 (2021). https://doi.org/10.1038/s43705-021-00082-4
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s43705-021-00082-4