Abstract
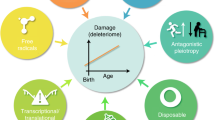
Information storage and retrieval is essential for all life. In biology, information is primarily stored in two distinct ways: the genome, comprising nucleic acids, acts as a foundational blueprint and the epigenome, consisting of chemical modifications to DNA and histone proteins, regulates gene expression patterns and endows cells with specific identities and functions. Unlike the stable, digital nature of genetic information, epigenetic information is stored in a digital–analog format, susceptible to alterations induced by diverse environmental signals and cellular damage. The Information Theory of Aging (ITOA) states that the aging process is driven by the progressive loss of youthful epigenetic information, the retrieval of which via epigenetic reprogramming can improve the function of damaged and aged tissues by catalyzing age reversal.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Johnson, F. B., Sinclair, D. A. & Guarente, L. Molecular biology of aging. Cell 96, 291–302 (1999).
Lopez-Otin, C. et al. Hallmarks of aging: an expanding universe. Cell 186, 243–278 (2023).
Munsky, B. & Neuert, G. From analog to digital models of gene regulation. Phys. Biol. 12, 045004 (2015).
Bernstein, B. E., Meissner, A. & Lander, E. S. The mammalian epigenome. Cell 128, 669–681 (2007).
Shannon, C. E. A mathematical theory of communication. Bell Syst. Tech. J. 27, 379–423 (1948).
Oberdoerffer, P. et al. SIRT1 redistribution on chromatin promotes genomic stability but alters gene expression during aging. Cell 135, 907–918 (2008).
Sinclair, D. A. & LaPlante, M. D. Lifespan: Why We Age—and Why We Don’t Have To (Atria Books, Simon and Schuster, 2019).
Sinclair, D. A., Mills, K. & Guarente, L. Accelerated aging and nucleolar fragmentation in yeast sgs1 mutants. Science 277, 1313–1316 (1997).
Mills, K. D., Sinclair, D. A. & Guarente, L. MEC1-dependent redistribution of the Sir3 silencing protein from telomeres to DNA double-strand breaks. Cell 97, 609–620 (1999).
Yang, J. H. et al. Loss of epigenetic information as a cause of mammalian aging. Cell 186, 305–326 (2023).
Lu, Y. et al. Reprogramming to recover youthful epigenetic information and restore vision. Nature 588, 124–129 (2020).
Szilard, L. On the nature of the aging process. Proc. Natl Acad. Sci. USA 45, 30–45 (1959).
Cagan, A. et al. Somatic mutation rates scale with lifespan across mammals. Nature 604, 517–524 (2022).
Schumacher, B. et al. The central role of DNA damage in the ageing process. Nature 592, 695–703 (2021).
White, R. R. et al. Controlled induction of DNA double-strand breaks in the mouse liver induces features of tissue ageing. Nat. Commun. 6, 6790 (2015).
Sinclair, D. A. & Oberdoerffer, P. The ageing epigenome: damaged beyond repair? Ageing Res. Rev. 8, 189–198 (2009).
Kaya, A., Lobanov, A. V. & Gladyshev, V. N. Evidence that mutation accumulation does not cause aging in Saccharomyces cerevisiae. Aging Cell 14, 366–371 (2015).
Robinson, P. S. et al. Increased somatic mutation burdens in normal human cells due to defective DNA polymerases. Nat. Genet. 53, 1434–1442 (2021).
Kato, T. et al. Dynamic stem cell selection safeguards the genomic integrity of the epidermis. Dev. Cell 56, 3309–3320 (2021).
Kim, J. et al. Controlled DNA double-strand break induction in mice reveals post-damage transcriptome stability. Nucleic Acids Res. 44, e64 (2016).
Horvath, S. DNA methylation age of human tissues and cell types. Genome Biol. 14, R115 (2013).
Lu, A. T. et al. Universal DNA methylation age across mammalian tissues. Nat. Aging 3, 1144–1166 (2023).
Bertucci-Richter, E. M. & Parrott, B. B. The rate of epigenetic drift scales with maximum lifespan across mammals. Nat. Commun. 14, 7731 (2023).
Sinclair, K. D. et al. Healthy ageing of cloned sheep. Nat. Commun. 7, 12359 (2016).
Burgstaller, J. P. & Brem, G. Aging of cloned animals: a mini-review. Gerontology 63, 417–425 (2017).
Macip, C. C. et al. Gene therapy mediated partial reprogramming extends lifespan and reverses age-related changes in aged mice. Preprint at bioRxiv https://doi.org/10.1101/2023.01.04.522507 (2023).
Alle, Q. et al. A single short reprogramming early in life initiates and propagates an epigenetically related mechanism improving fitness and promoting an increased healthy lifespan. Aging Cell 21, e13714 (2022).
Kennedy, B. K. et al. Mutation in the silencing gene SIR4 can delay aging in S. cerevisiae. Cell 80, 485–496 (1995).
Kaeberlein, M., McVey, M. & Guarente, L. The SIR2/3/4 complex and SIR2 alone promote longevity in Saccharomyces cerevisiae by two different mechanisms. Genes Dev. 13, 2570–2580 (1999).
Imai, S. et al. Transcriptional silencing and longevity protein Sir2 is an NAD-dependent histone deacetylase. Nature 403, 795–800 (2000).
Sinclair, D. A. & Guarente, L. Extrachromosomal rDNA circles—a cause of aging in yeast. Cell 91, 1033–1042 (1997).
Xu, C. et al. SIRT1 is downregulated by autophagy in senescence and ageing. Nat. Cell Biol. 22, 1170–1179 (2020).
Kennedy, B. K. et al. Redistribution of silencing proteins from telomeres to the nucleolus is associated with extension of life span in S. cerevisiae. Cell 89, 381–391 (1997).
Ono, T. & Cutler, R. G. Age-dependent relaxation of gene repression: increase of endogenous murine leukemia virus-related and globin-related RNA in brain and liver of mice. Proc. Natl Acad. Sci. USA 75, 4431–4435 (1978).
Oberdoerffer, P. & Sinclair, D. A. The role of nuclear architecture in genomic instability and ageing. Nat. Rev. Mol. Cell Biol. 8, 692–702 (2007).
Lu, T. et al. REST and stress resistance in ageing and Alzheimer’s disease. Nature 507, 448–454 (2014).
Nalapareddy, K. et al. Canonical Wnt signaling ameliorates aging of intestinal stem cells. Cell Rep. 18, 2608–2621 (2017).
Mortusewicz, O. et al. Recruitment of DNA methyltransferase I to DNA repair sites. Proc. Natl Acad. Sci. USA 102, 8905–8909 (2005).
Chou, D. M. et al. A chromatin localization screen reveals poly (ADP ribose)-regulated recruitment of the repressive polycomb and NuRD complexes to sites of DNA damage. Proc. Natl Acad. Sci. USA 107, 18475–18480 (2010).
Lu, J. Y. et al. Comparative transcriptomics reveals circadian and pluripotency networks as two pillars of longevity regulation. Cell Metab. 34, 836–856 (2022).
Tian, X. et al. SIRT6 is responsible for more efficient DNA double-strand break repair in long-lived species. Cell 177, 622–638 (2019).
Tan, L. et al. Naked mole rat cells have a stable epigenome that resists iPSC reprogramming. Stem Cell Rep. 9, 1721–1734 (2017).
Kanfi, Y. et al. The sirtuin SIRT6 regulates lifespan in male mice. Nature 483, 218–221 (2012).
Satoh, A. et al. Sirt1 extends life span and delays aging in mice through the regulation of Nk2 homeobox 1 in the DMH and LH. Cell Metab. 18, 416–430 (2013).
Taylor, J. R. et al. Sirt6 regulates lifespan in Drosophila melanogaster. Proc. Natl Acad. Sci. USA 119, e2111176119 (2022).
Williams, G. C. Pleiotropy, natural selection, and the evolution of senescence. Evolution 11, 398–411 (1957).
Dobbin, M. M. et al. SIRT1 collaborates with ATM and HDAC1 to maintain genomic stability in neurons. Nat. Neurosci. 16, 1008–1015 (2013).
Kerepesi, C. et al. Epigenetic clocks reveal a rejuvenation event during embryogenesis followed by aging. Sci. Adv. 7, eabg6082 (2021).
Gurdon, J. B., Elsdale, T. R. & Fischberg, M. Sexually mature individuals of Xenopus laevis from the transplantation of single somatic nuclei. Nature 182, 64–65 (1958).
Wilmut, I. et al. Viable offspring derived from fetal and adult mammalian cells. Nature 385, 810–813 (1997).
Yu, J. et al. Induced pluripotent stem cell lines derived from human somatic cells. Science 318, 1917–1920 (2007).
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006).
Lapasset, L. et al. Rejuvenating senescent and centenarian human cells by reprogramming through the pluripotent state. Genes Dev. 25, 2248–2253 (2011).
Wahlestedt, M. et al. An epigenetic component of hematopoietic stem cell aging amenable to reprogramming into a young state. Blood 121, 4257–4264 (2013).
Mertens, J. et al. Directly reprogrammed human neurons retain aging-associated transcriptomic signatures and reveal age-related nucleocytoplasmic defects. Cell Stem Cell 17, 705–718 (2015).
Lee, H. Y. et al. Identifying molecular targets for reverse aging using integrated network analysis of transcriptomic and epigenomic changes during aging. Sci. Rep. 11, 12317 (2021).
Birnbaum, A. et al. Age-dependent changes in transcription factor FOXO targeting in female Drosophila. Front. Genet. 10, 312 (2019).
Schaum, N. et al. Ageing hallmarks exhibit organ-specific temporal signatures. Nature 583, 596–602 (2020).
Statello, L. et al. Gene regulation by long non-coding RNAs and its biological functions. Nat. Rev. Mol. Cell Biol. 22, 96–118 (2021).
Peng, Y. & Croce, C. M. The role of MicroRNAs in human cancer. Signal Transduct. Target Ther. 1, 15004 (2016).
Kinser, H. E. & Pincus, Z. MicroRNAs as modulators of longevity and the aging process. Hum. Genet. 139, 291–308 (2020).
Du, W. W. et al. miR-17 extends mouse lifespan by inhibiting senescence signaling mediated by MKP7. Cell Death Dis. 5, e1355 (2014).
Kumar, S. et al. MicroRNA-455-3p improves synaptic, cognitive functions and extends lifespan: relevance to Alzheimer’s disease. Redox Biol. 48, 102182 (2021).
Aguilera, A. & Garcia-Muse, T. R loops: from transcription byproducts to threats to genome stability. Mol. Cell 46, 115–24 (2012).
Ellis, D. A. et al. R-loops and regulatory changes in chronologically ageing fission yeast cells drive non-random patterns of genome rearrangements. PLoS Genet. 17, e1009784 (2021).
Jauregui-Lozano, J. et al. Proper control of R-loop homeostasis is required for maintenance of gene expression and neuronal function during aging. Aging Cell 21, e13554 (2022).
Strom, A. R. et al. Phase separation drives heterochromatin domain formation. Nature 547, 241–245 (2017).
Villeponteau, B. The heterochromatin loss model of aging. Exp. Gerontol. 32, 383–394 (1997).
Imai, S. & Kitano, H. Heterochromatin islands and their dynamic reorganization: a hypothesis for three distinctive features of cellular aging. Exp. Gerontol. 33, 555–570 (1998).
Ni, Z. et al. Two SET domain containing genes link epigenetic changes and aging in Caenorhabditis elegans. Aging Cell 11, 315–325 (2012).
Larson, K. et al. Heterochromatin formation promotes longevity and represses ribosomal RNA synthesis. PLoS Genet. 8, e1002473 (2012).
Wood, J. G. et al. Chromatin remodeling in the aging genome of Drosophila. Aging Cell 9, 971–978 (2010).
De Cecco, M. et al. Transposable elements become active and mobile in the genomes of aging mammalian somatic tissues. Aging 5, 867–883 (2013).
Zhang, W. et al. Aging stem cells. A Werner syndrome stem cell model unveils heterochromatin alterations as a driver of human aging. Science 348, 1160–1163 (2015).
De Cecco, M. et al. L1 drives IFN in senescent cells and promotes age-associated inflammation. Nature 566, 73–78 (2019).
Simon, M. et al. LINE1 derepression in aged wild-type and SIRT6-deficient mice drives inflammation. Cell Metab. 29, 871–885 (2019).
Benayoun, B. A. et al. Remodeling of epigenome and transcriptome landscapes with aging in mice reveals widespread induction of inflammatory responses. Genome Res. 29, 697–709 (2019).
Freund, A. et al. Lamin B1 loss is a senescence-associated biomarker. Mol. Biol. Cell 23, 2066–75 (2012).
Scaffidi, P. & Misteli, T. Lamin A-dependent nuclear defects in human aging. Science 312, 1059–1063 (2006).
van Steensel, B. & Belmont, A. S. Lamina-associated domains: links with chromosome architecture, heterochromatin, and gene repression. Cell 169, 780–791 (2017).
Criscione, S. W. et al. Reorganization of chromosome architecture in replicative cellular senescence. Sci. Adv. 2, e1500882 (2016).
Dixon, J. R. et al. Chromatin architecture reorganization during stem cell differentiation. Nature 518, 331–336 (2015).
Fu, V. X. et al. A loss of insulin-like growth factor-2 imprinting is modulated by CCCTC-binding factor down-regulation at senescence in human epithelial cells. J. Biol. Chem. 279, 52218–52226 (2004).
Pal, S. et al. Impaired cohesion and homologous recombination during replicative aging in budding yeast. Sci. Adv. 4, eaaq0236 (2018).
Pouikli, A. et al. Chromatin remodeling due to degradation of citrate carrier impairs osteogenesis of aged mesenchymal stem cells. Nat. Aging 1, 810–825 (2021).
Tan, M. et al. Identification of 67 histone marks and histone lysine crotonylation as a new type of histone modification. Cell 146, 1016–1028 (2011).
Dang, W. et al. Histone H4 lysine 16 acetylation regulates cellular lifespan. Nature 459, 802–807 (2009).
Liu, L. et al. Chromatin modifications as determinants of muscle stem cell quiescence and chronological aging. Cell Rep. 4, 189–204 (2013).
O’Sullivan, R. J. et al. Reduced histone biosynthesis and chromatin changes arising from a damage signal at telomeres. Nat. Struct. Mol. Biol. 17, 1218–1225 (2010).
Sidler, C. et al. A role for SUV39H1-mediated H3K9 trimethylation in the control of genome stability and senescence in WI38 human diploid lung fibroblasts. Aging 6, 545–563 (2014).
Kane, A. E. & Sinclair, D. A. Epigenetic changes during aging and their reprogramming potential. Crit. Rev. Biochem Mol. Biol. 54, 61–83 (2019).
Benayoun, B. A., Pollina, E. A. & Brunet, A. Epigenetic regulation of ageing: linking environmental inputs to genomic stability. Nat. Rev. Mol. Cell Biol. 16, 593–610 (2015).
Feser, J. et al. Elevated histone expression promotes life span extension. Mol. Cell 39, 724–35 (2010).
Pal, S. & Tyler, J. K. Epigenetics and aging. Sci. Adv. 2, e1600584 (2016).
Bollati, V. et al. Decline in genomic DNA methylation through aging in a cohort of elderly subjects. Mech. Ageing Dev. 130, 234–239 (2009).
Rakyan, V. K. et al. Human aging-associated DNA hypermethylation occurs preferentially at bivalent chromatin domains. Genome Res 20, 434–439 (2010).
Bocklandt, S. et al. Epigenetic predictor of age. PLoS ONE 6, e14821 (2011).
Hannum, G. et al. Genome-wide methylation profiles reveal quantitative views of human aging rates. Mol. Cell 49, 359–367 (2013).
Bell, C. G. et al. DNA methylation aging clocks: challenges and recommendations. Genome Biol. 20, 249 (2019).
Meer, M. V. et al. A whole lifespan mouse multi-tissue DNA methylation clock. Elife 7, e40675 (2018).
Thompson, M. J. et al. An epigenetic aging clock for dogs and wolves. Aging 9, 1055–1068 (2017).
Lowe, R. et al. DNA methylation clocks as a predictor for ageing and age estimation in naked mole-rats, Heterocephalus glaber. Aging 12, 4394–4406 (2020).
de Magalhaes, J. P. Ageing as a software design flaw. Genome Biol. 24, 51 (2023).
Abad, M. et al. Reprogramming in vivo produces teratomas and iPS cells with totipotency features. Nature 502, 340–345 (2013).
Ocampo, A. et al. In vivo amelioration of age-associated hallmarks by partial reprogramming. Cell 167, 1719–1733 (2016).
Parras, A. et al. In vivo reprogramming leads to premature death linked to hepatic and intestinal failure. Nat. Aging, https://doi.org/10.1038/s43587-023-00528-5 (2023).
Karg, M. M. et al. Sustained vision recovery by OSK gene therapy in a mouse model of glaucoma. Cell. Reprogram. 25, https://doi.org/10.1089/cell.2023.0074 (2023).
Drake, S. S. et al. Cellular rejuvenation protects neurons from inflammation mediated cell death. Preprint at bioRxiv, https://doi.org/10.1101/2023.09.30.560301 (2023).
Ksander, B. R. et al. Epigenetic reprogramming - a novel gene therapy that restores vision loss in a nonhuman primate model of NAION. Invest. Ophthalmol. Vis. Sci. 64, 474 (2023).
Roux, A. E. et al. Diverse partial reprogramming strategies restore youthful gene expression and transiently suppress cell identity. Cell Syst. 13, 574–587 (2022).
Browder, K. C. et al. In vivo partial reprogramming alters age-associated molecular changes during physiological aging in mice. Nat. Aging 2, 243–253 (2022).
Chondronasiou, D. et al. Multi-omic rejuvenation of naturally aged tissues by a single cycle of transient reprogramming. Aging Cell 21, e13578 (2022).
Hishida, T. et al. In vivo partial cellular reprogramming enhances liver plasticity and regeneration. Cell Rep. 39, 110730 (2022).
Chen, Y. et al. Reversible reprogramming of cardiomyocytes to a fetal state drives heart regeneration in mice. Science 373, 1537–1540 (2021).
Rodriguez-Matellan, A. et al. In vivo reprogramming ameliorates aging features in dentate gyrus cells and improves memory in mice. Stem Cell Rep. 15, 1056–1066 (2020).
Sarkar, T. J. et al. Transient non-integrative expression of nuclear reprogramming factors promotes multifaceted amelioration of aging in human cells. Nat. Commun. 11, 1545 (2020).
de Lazaro, I. et al. Non-viral, tumor-free induction of transient cell reprogramming in mouse skeletal muscle to enhance tissue regeneration. Mol. Ther. 27, 59–75 (2019).
Humphreys, T. et al. Ancestral stem cell reprogramming genes active in hemichordate regeneration. Front. Ecol. Evol. 10, 769433 (2022).
Ingles, M. et al. Centenarians overexpress pluripotency-related genes. J. Gerontol. A Biol. Sci. Med. Sci. 74, 1391–1395 (2019).
McLaughlin, K. A. & Levin, M. Bioelectric signaling in regeneration: mechanisms of ionic controls of growth and form. Dev. Biol. 433, 177–189 (2018).
Tai, W. L. et al. Regulation of retinal ganglion cell axon growth and optic nerve regeneration by DNA methyltransferase. Invest. Ophthalmol. Vis. Sci. 64, 2840 (2023).
Zheng, Z. et al. The DNA methylation inhibitor RG108 protects against noise-induced hearing loss. Cell Biol. Toxicol. 37, 751–771 (2021).
Zhang, B. et al. Multi-omic rejuvenation and lifespan extension on exposure to youthful circulation. Nat. Aging 3, 948–964 (2023).
Horvath, S. et al. Reversal of biological age in multiple rat organs by young porcine plasma fraction. GeroScience, https://doi.org/10.1007/s11357-023-00980-6 (2023).
Tavenier, J. et al. Association of GDF15 with inflammation and physical function during aging and recovery after acute hospitalization: a longitudinal study of older patients and age-matched controls. J. Gerontol. A Biol. Sci. Med. Sci. 76, 964–974 (2021).
Yoshida, M. et al. Extracellular vesicle-contained eNAMPT delays aging and extends lifespan in mice. Cell Metab. 30, 329–342 (2019).
Sahu, A. et al. Regulation of aged skeletal muscle regeneration by circulating extracellular vesicles. Nat. Aging 1, 1148–1161 (2021).
De Miguel, Z. et al. Exercise plasma boosts memory and dampens brain inflammation via clusterin. Nature 600, 494–499 (2021).
Blanchard, J. W. et al. Replacing reprogramming factors with antibodies selected from combinatorial antibody libraries. Nat. Biotechnol. 35, 960–968 (2017).
Hou, P. et al. Pluripotent stem cells induced from mouse somatic cells by small-molecule compounds. Science 341, 651–654 (2013).
Guan, J. et al. Chemical reprogramming of human somatic cells to pluripotent stem cells. Nature 605, 325–331 (2022).
Yang, J. H. et al. Chemically induced reprogramming to reverse cellular aging. Aging 15, 5966–5989 (2023).
Mitchell, W. et al. Multi-omics characterization of partial chemical reprogramming reveals evidence of cell rejuvenation. eLife 12, RP90579 (2023).
Schoenfeldt, L. et al. Chemical reprogramming ameliorates cellular hallmarks of aging and extends lifespan. Preprint at bioRxiv https://doi.org/10.1101/2022.08.29.505222 (2022).
Chin, R. M. et al. The metabolite alpha-ketoglutarate extends lifespan by inhibiting ATP synthase and TOR. Nature 510, 397–401 (2014).
Asadi Shahmirzadi, A. et al. Alpha-ketoglutarate, an endogenous metabolite, extends lifespan and compresses morbidity in aging mice. Cell Metab. 32, 447–456 (2020).
Demidenko, O. et al. Rejuvant, a potential life-extending compound formulation with alpha-ketoglutarate and vitamins, conferred an average 8 year reduction in biological aging, after an average of 7 months of use, in the TruAge DNA methylation test. Aging 13, 24485–24499 (2021).
Huangfu, D. et al. Induction of pluripotent stem cells by defined factors is greatly improved by small-molecule compounds. Nat. Biotechnol. 26, 795–797 (2008).
Simonsson, S. & Gurdon, J. DNA demethylation is necessary for the epigenetic reprogramming of somatic cell nuclei. Nat. Cell Biol. 6, 984–990 (2004).
He, S. et al. Passive DNA demethylation preferentially up-regulates pluripotency-related genes and facilitates the generation of induced pluripotent stem cells. J. Biol. Chem. 292, 18542–18555 (2017).
Wu, X. & Zhang, Y. TET-mediated active DNA demethylation: mechanism, function and beyond. Nat. Rev. Genet 18, 517–534 (2017).
Gontier, G. et al. Tet2 rescues age-related regenerative decline and enhances cognitive function in the adult mouse brain. Cell Rep. 22, 1974–1981 (2018).
Wang, D. et al. Active DNA demethylation promotes cell fate specification and the DNA damage response. Science 378, 983–989 (2022).
Gill, D. et al. Multi-omic rejuvenation of human cells by maturation phase transient reprogramming. Elife 11, e71624 (2022).
Tharakan, R. et al. Whole-genome methylation analysis of aging human tissues identifies age-related changes in developmental and neurological pathways. Aging Cell 22, e13847 (2023).
Rizzo, J. F. III et al. The role of epigenetics in accelerated aging: a reconsideration of later-life visual loss after early optic neuropathy. J. Neuroophthalmol., https://doi.org/10.1097/WNO.0000000000002041 (2023)
Poganik, J. R. et al. Biological age is increased by stress and restored upon recovery. Cell Metab. 35, 807–820 (2023).
Koblan, L. W. et al. In vivo base editing rescues Hutchinson–Gilford progeria syndrome in mice. Nature 589, 608–614 (2021).
Zeng, J. et al. Therapeutic base editing of human hematopoietic stem cells. Nat. Med. 26, 535–541 (2020).
Waddington, C. H. The Strategy of the Genes; A Discussion of Some Aspects of Theoretical Biology (George Allen & Unwin, 1957).
Becker, J. S., Nicetto, D. & Zaret, K. S. H3K9me3-dependent heterochromatin: barrier to cell fate changes. Trends Genet. 32, 29–41 (2016).
Cutler, R. G. The dysdifferentiative hypothesis of mammalian aging and longevity. Aging Brain 20, 1–18 (1982).
Xu, Q. et al. Stress induced aging in mouse eye. Aging Cell 21, e13737 (2022).
Kuo, P. L. et al. Epigenetic age acceleration and hearing: observations from the Baltimore Longitudinal Study of Aging. Front. Aging Neurosci. 13, 790926 (2021).
Schaible, R., Sussman, M. & Kramer, B. H. Aging and potential for self-renewal: hydra living in the age of aging—a mini-review. Gerontology 60, 548–556 (2014).
Reddien, P. W. & Sanchez Alvarado, A. Fundamentals of planarian regeneration. Annu. Rev. Cell Dev. Biol. 20, 725–757 (2004).
Beffagna, G. Zebrafish as a smart model to understand regeneration after heart injury: how fish could help humans. Front. Cardiovasc. Med. 6, 107 (2019).
Nowoshilow, S. et al. The axolotl genome and the evolution of key tissue formation regulators. Nature 554, 50–55 (2018).
Takeo, M. et al. Wnt activation in nail epithelium couples nail growth to digit regeneration. Nature 499, 228–232 (2013).
Maden, M. & Varholick, J. A. Model systems for regeneration: the spiny mouse, Acomys cahirinus. Development 147, dev167718 (2020).
Seifert, A. W. et al. Skin shedding and tissue regeneration in African spiny mice (Acomys). Nature 489, 561–565 (2012).
Wong, W. et al. Spiny mice (Acomys) exhibit attenuated hallmarks of aging and rapid cell turnover after UV exposure in the skin epidermis. PLoS ONE 15, e0241617 (2020).
Saxena, S. et al. Connective tissue fibroblasts from highly regenerative mammals are refractory to ROS-induced cellular senescence. Nat. Commun. 10, 4400 (2019).
Michalopoulos, G. K. & DeFrances, M. C. Liver regeneration. Science 276, 60–66 (1997).
Onal, P. et al. Gene expression of pluripotency determinants is conserved between mammalian and planarian stem cells. EMBO J. 31, 2755–2769 (2012).
Sarkar, A. et al. STAT3 promotes a youthful epigenetic state in articular chondrocytes. Aging Cell 22, e13773 (2023).
Lee, J. Y. et al. Misexpression of genes lacking CpG islands drives degenerative changes during aging. Sci. Adv. 7, eabj9111 (2021).
Dixon, G. et al. QSER1 protects DNA methylation valleys from de novo methylation. Science 372, eabd0875 (2021).
Mozhui, K. & Pandey, A. K. Conserved effect of aging on DNA methylation and association with EZH2 polycomb protein in mice and humans. Mech. Ageing Dev. 162, 27–37 (2017).
Moqri, M. et al. PRC2 clock: a universal epigenetic biomarker of aging and rejuvenation. Preprint at bioRxiv https://doi.org/10.1101/2022.06.03.494609 (2022).
Acknowledgements
We thank E. Smith for figure illustrations and R. Rogers-Hammond and M. LaPlante for manuscript advice. Y.R.L. was supported by postdoctoral fellowships from Glenn/AFAR (by Michael Shen, MIT'13) and the Life Sciences Research Foundation (by Lei-Luo Life Science Fund). X.T. was supported by a NIH/NIA award K99AG068303. This work was also funded by NIH/NIA R01AG019719, Harvard Medical School Epigenetics Grants, gifts from M. Chambers, R. Rosenkrantz, T. Robbins, P. Diamandis, S. Aoki, D. and S. Hoff, and the Glenn Foundation for Medical Research to D.A.S.
Author information
Authors and Affiliations
Contributions
Y.R.L., X.T. and D.A.S. wrote this Perspective.
Corresponding author
Ethics declarations
Competing interests
Y.R.L., D.A.S. and X.T. are inventors on patent applications licensed to Life Biosciences, a company developing epigenetic reprogramming-based therapies, in which Y.R.L. and D.A.S. have equity. Complete details of all relationships for profit and not-for-profit for D.A.S. can be found in the Supplementary Information.
Peer review
Peer review information
Nature Aging thanks Vera Gorbunova and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lu, Y.R., Tian, X. & Sinclair, D.A. The Information Theory of Aging. Nat Aging 3, 1486–1499 (2023). https://doi.org/10.1038/s43587-023-00527-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43587-023-00527-6