Abstract
Genomic, transcriptomic and proteomic approaches have been used to gain insight into molecular underpinnings of aging in laboratory animals and in humans. However, protein function in biological systems is under complex regulation and includes factors besides abundance levels, such as modifications, localization, conformation and protein–protein interactions. By making use of quantitative chemical cross-linking technologies, we show that changes in the muscle mitochondrial interactome contribute to mitochondrial functional decline in aging in female mice. Specifically, we identify age-related changes in protein cross-links relating to assembly of electron transport system complexes I and IV, activity of glutamate dehydrogenase, and coenzyme-A binding in fatty acid β-oxidation and tricarboxylic acid cycle enzymes. These changes show a remarkable correlation with complex I respiration differences within the same young–old animal pairs. Each observed cross-link can serve as a protein conformational or protein–protein interaction probe in future studies, which will provide further molecular insights into commonly observed age-related phenotypic differences. Therefore, this data set could become a valuable resource for additional in-depth molecular studies that are needed to better understand complex age-related molecular changes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Mass spectrometry data have been deposited to the ProteomeXchange Consortium via the PRIDE repository under identifiers PXD031643 and PXD031644.
Cross-linking data have been uploaded to the XLinkdb online database (http://xlinkdb.gs.washington.edu/xlinkdb/Interactome_of_aged_muscle_mitochondria.php).
Code availability
R markdown used for statistical analysis and figure generation is available at https://github.com/brucelab/aging_mito_interactome_data_analysis.
References
López-Otín, C., Blasco, M. A., Partridge, L., Serrano, M. & Kroemer, G. The hallmarks of aging. Cell 153, 1194–1217 (2013).
Harman, D. Aging: a theory based on free radical and radiation chemistry. J. Gerontol. 11, 298–300 (1956).
Peterson, C. M., Johannsen, D. L. & Ravussin, E. Skeletal muscle mitochondria and aging: a review. J. Aging Res. 2012, 1–20 (2012).
Sun, N., Youle, R. J. & Finkel, T. The mitochondrial basis of Aging. Mol. Cell 61, 654–666 (2016).
Short, K. R. et al. Decline in skeletal muscle mitochondrial function with aging in humans. Proc. Natl Acad. Sci. 102, 5618–5623 (2005).
Hepple, R. T. Mitochondrial involvement and impact in aging skeletal muscle. Front. Aging Neurosci. 6, 211 (2014).
Ubaida-Mohien, C. et al. Discovery proteomics in aging human skeletal muscle finds change in spliceosome, immunity, proteostasis and mitochondria. eLife 8, e49874 (2019).
Tumasian, R. A. et al. Skeletal muscle transcriptome in healthy aging. Nat. Commun. 12, 2014 (2021).
Hwang, C. Y. et al. Quantitative proteome analysis of age-related changes in mouse gastrocnemius muscle using mTRAQ. Proteomics 14, 121–132 (2014).
Hunt, L. C. et al. Integrated genomic and proteomic analyses identify stimulus-dependent molecular changes associated with distinct modes of skeletal muscle atrophy. Cell Rep. 37, 109971 (2021).
Yu, Q. et al. Sample multiplexing for targeted pathway proteomics in aging mice. Proc. Natl Acad. Sci.117, 9723–9732 (2020).
Yeo, D., Kang, C. & Ji, L. L. Aging alters acetylation status in skeletal and cardiac muscles. GeroScience 42, 963–976 (2020).
Wagner, G. R. & Payne, R. M. Mitochondrial acetylation and diseases of aging. J. Aging Res. 2011, 234875 (2011).
Hofer, A. & Wenz, T. Post-translational modification of mitochondria as a novel mode of regulation. Exp. Gerontol. 56, 202–220 (2014).
Chavez, J. D., Keller, A., Mohr, J. P. & Bruce, J. E. Isobaric quantitative protein interaction reporter technology for comparative interactome studies. Anal. Chem. 92, 14094–14102 (2020).
Müller, F. & Rappsilber, J. A protocol for studying structural dynamics of proteins by quantitative crosslinking mass spectrometry and data-independent acquisition. J. Proteomics 218, 103721 (2020).
Chavez, J. D. et al. A general method for targeted quantitative cross-linking mass spectrometry. PLoS One 11, e0167547 (2016).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Rath, S. et al. MitoCarta3.0: an updated mitochondrial proteome now with sub-organelle localization and pathway annotations. Nucleic Acids Res. 49, D1541–D1547 (2021).
Figueiredo, P. A., Ferreira, R. M., Appell, H. J. & Duarte, J. A. Age-induced morphological, biochemical, and functional alterations in isolated mitochondria from murine skeletal muscle. J. Gerontol. Biol. Sci. Med. Sci. 63, 350–359 (2008).
Han, Y. et al. Transcriptome features of striated muscle aging and predictability of protein level changes. Mol. Omics 17, 796–808 (2021).
Wiley, C. D. & Campisi, J. The metabolic roots of senescence: mechanisms and opportunities for intervention. Nat. Metab. 3, 1290–1301 (2021).
Sánchez-Caballero, L., Guerrero-Castillo, S. & Nijtmans, L. Unraveling the complexity of mitochondrial complex I assembly: a dynamic process. Biochim. Biophys. Acta 1857, 980–990 (2016).
Kruse, S. E. et al. Age modifies respiratory complex I and protein homeostasis in a muscle type-specific manner. Aging Cell 15, 89–99 (2016).
Park, J. et al. SIRT5-mediated lysine desuccinylation impacts diverse metabolic pathways. Mol. Cell 50, 919–930 (2013).
Yang, W. et al. Mitochondrial sirtuin network reveals dynamic SIRT3-dependent deacetylation in response to membrane depolarization. Cell 167, 985–1000.e1021 (2016).
Guerrero-Castillo, S. et al. The assembly pathway of mitochondrial respiratory chain complex I. Cell Metab. 25, 128–139 (2017).
Hirst, J. & Roessler, M. M. Energy conversion, redox catalysis and generation of reactive oxygen species by respiratory complex I. Biochim. Biophys. Acta 1857, 872–883 (2016).
Szczepanowska, K. et al. A salvage pathway maintains highly functional respiratory complex I. Nat. Commun. 11, 1643 (2020).
Miwa, S. et al. Low abundance of the matrix arm of complex I in mitochondria predicts longevity in mice. Nat. Commun. 5, 3837 (2014).
Protasoni, M. et al. Respiratory supercomplexes act as a platform for complex III-mediated maturation of human mitochondrial complexes I and IV. EMBO J. 39, e102817 (2020).
Balsa, E. et al. NDUFA4 is a subunit of complex IV of the mammalian electron transport chain. Cell Metab. 16, 378–386 (2012).
Zong, S. et al. Structure of the intact 14-subunit human cytochrome c oxidase. Cell Res. 28, 1026–1034 (2018).
Caudal, A. et al. Mitochondrial interactome quantitation reveals structural changes in metabolic machinery in the failing murine heart. Nat. Cardiovasc. Res. 1, 855–866 (2022).
Clayton, S. A. et al. Inflammation causes remodeling of mitochondrial cytochrome. Sci. Adv. 7, eabl5182 (2021).
Pitceathly, R. D. S. et al. NDUFA4 mutations underlie dysfunction of a cytochrome c oxidase subunit linked to human neurological disease. Cell Rep. 3, 1795–1805 (2013).
Yuan, R. et al. Genetic coregulation of age of female sexual maturation and lifespan through circulating IGF1 among inbred mouse strains. Proc. Natl. Acad. Sci. USA 109, 8224–8229 (2012).
Ferrucci, L. & Kuchel, G. A. Heterogeneity of aging: individual risk factors, mechanisms, patient priorities, and outcomes. J. Am. Geriatr. Soc. 69, 610–612 (2021).
Smith, H. Q., Li, C., Stanley, C. A. & Smith, T. J. Glutamate dehydrogenase, a complex enzyme at a crucial metabolic branch point. Neurochem. Res. 44, 117–132 (2019).
Hamelin, M., Mary, J., Vostry, M., Friguet, B. & Bakala, H. Glycation damage targets glutamate dehydrogenase in the rat liver mitochondrial matrix during aging: glycation of glutamate dehydrogenase with aging. FEBS J. 274, 5949–5961 (2007).
Li, M., Li, C., Allen, A., Stanley, C. A. & Smith, T. J. The structure and allosteric regulation of mammalian glutamate dehydrogenase. Arch. Biochem. Biophys. 519, 69–80 (2012).
Banerjee, S., Schmidt, T., Fang, J., Stanley, C. A. & Smith, T. J. Structural studies on ADP activation of mammalian glutamate dehydrogenase and the evolution of regulation. Biochemistry 42, 3446–3456 (2003).
Hyyti, O. M., Ledee, D., Ning, X.-H., Ge, M. & Portman, M. A. Aging impairs myocardial fatty acid and ketone oxidation and modifies cardiac functional and metabolic responses to insulin in mice. Am. J. Physiol. Heart Circ. Physiol. 299, H868–H875 (2010).
Houtkooper, R. H. et al. The metabolic footprint of aging in mice. Sci. Rep. 1, 134 (2011).
Zhang, X. et al. Impaired mitochondrial energetics characterize poor early recovery of muscle mass following hind limb unloading in old mice. J. Gerontol. Ser. A 73, 1313–1322 (2018).
Fan, J. et al. Tetrameric acetyl-CoA acetyltransferase 1 is important for tumor growth. Mol. Cell 64, 859–874 (2016).
Wu, B. et al. Succinyl-CoA ligase deficiency in pro-inflammatory and tissue-invasive T cells. Cell Metab. 32, 967–980.e965 (2020).
Hayflick, L. The greatest risk factor for the leading cause of death is ignored. Biogerontology 22, 133–141 (2021).
Asadi Shahmirzadi, A. et al. Alpha-ketoglutarate, an endogenous metabolite, extends lifespan and compresses morbidity in aging mice. Cell Metab. 32, 447–456.e446 (2020).
Su, Y. et al. Alpha-ketoglutarate extends Drosophila lifespan by inhibiting mTOR and activating AMPK. Aging 11, 4183–4197 (2019).
Kaeberlein, M., Burtner, C. R. & Kennedy, B. K. Recent developments in yeast aging. PLoS Genet. 3, e84 (2007).
Chin, R. M. et al. The metabolite α-ketoglutarate extends lifespan by inhibiting ATP synthase and TOR. Nature 510, 397–401 (2014).
Hagopian, K. Caloric restriction increases gluconeogenic and transaminase enzyme activities in mouse liver. Exp. Gerontol. 38, 267–278 (2003).
Lyu, Y. et al. Drosophila serotonin 2A receptor signaling coordinates central metabolic processes to modulate aging in response to nutrient choice. eLife 10, e59399 (2021).
Lombardi, A. A. et al. Mitochondrial calcium exchange links metabolism with the epigenome to control cellular differentiation. Nat. Commun. 10, 4509 (2019).
Rardin, M. J. et al. Label-free quantitative proteomics of the lysine acetylome in mitochondria identifies substrates of SIRT3 in metabolic pathways. Proc. Natl Acad. Sci. USA 110, 6601–6606 (2013).
Mimaki, M., Wang, X., McKenzie, M., Thorburn, D. R. & Ryan, M. T. Understanding mitochondrial complex I assembly in health and disease. Biochim. Biophys. Acta 1817, 851–862 (2012).
Martínez-Reyes, I. & Chandel, N. S. Mitochondrial TCA cycle metabolites control physiology and disease. Nat. Commun. 11, 102 (2020).
Lee, S. H., Lee, S. K., Paik, D. & Min, K. J. Overexpression of fatty-acid-β-oxidation-related genes extends the lifespan of Drosophila melanogaster. Oxid. Med. Cell Longev. 2012, 854502 (2012).
Nguyen, D., Samson, S. L., Reddy, V. T., Gonzalez, E. V. & Sekhar, R. V. Impaired mitochondrial fatty acid oxidation and insulin resistance in aging: novel protective role of glutathione. Aging Cell 12, 415–425 (2013).
Keller, A., Chavez, J. D., Eng, J. K., Thornton, Z. & Bruce, J. E. Tools for 3D interactome visualization. J. Proteome Res. 18, 753–758 (2019).
Mänttäri, S. & Järvilehto, M. Comparative analysis of mouse skeletal muscle fibre type composition and contractile responses to calcium channel blocker. BMC Physiol. 5, 4 (2005).
Wallace, M. A. et al. The ketogenic diet preserves skeletal muscle with aging in mice. Aging Cell 20, e13322 (2021).
Hämäläinen, N. & Pette, D. The histochemical profiles of fast fiber types IIB, IID, and IIA in skeletal muscles of mouse, rat, and rabbit. J. Histochem. Cytochem. 41, 733–743 (1993).
Lark, D. S. et al. Direct real-time quantification of mitochondrial oxidative phosphorylation efficiency in permeabilized skeletal muscle myofibers. Am. J. Physiol. Cell Physiol. 311, C239–C245 (2016).
Figueiredo, P. A., Ferreira, R. M., Appell, H. J. & Duarte, J. A. Age-induced morphological, biochemical, and functional alterations in isolated mitochondria from murine skeletal muscle. J. Gerontol. Ser. Biol. Sci. Med. Sci. 63, 350–359 (2008).
Crouch, M.-L. et al. Cyclophosphamide leads to persistent deficits in physical performance and in vivo mitochondria function in a mouse model of chemotherapy late effects. PLoS One 12, e0181086 (2017).
Wittig, I., Karas, M. & Schägger, H. High resolution clear native electrophoresis for in-gel functional assays and fluorescence studies of membrane protein complexes. Mol. Cell Proteomics 6, 1215–1225 (2007).
Mohr, J. P., Perumalla, P., Chavez, J. D., Eng, J. K. & Bruce, J. E. Mango: a general tool for collision induced dissociation-cleavable cross-linked peptide identification. Anal. Chem. 90, 6028–6034 (2018).
Eng, J. K., Jahan, T. A. & Hoopmann, M. R. Comet: an open-source MS/MS sequence database search tool. Proteomics 13, 22–24 (2013).
Keller, A., Chavez, J. D. & Bruce, J. E. Increased sensitivity with automated validation of XL-MS cleavable peptide crosslinks. Bioinformatics 35, 895–897 (2019).
Wickham, H. et al. Welcome to the Tidyverse. J. Open Source Software 4, 1686 (2019).
Szklarczyk, D. et al. The STRING database in 2021: customizable protein–protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Res. 49, D605–D612 (2021).
Bullock, J. M. A., Thalassinos, K. & Topf, M. Jwalk and MNXL web server: model validation using restraints from crosslinking mass spectrometry. Bioinformatics 34, 3584–3585 (2018).
Acknowledgements
Figures were created with BioRender.com. We thank members of the Bruce laboratory for helpful discussion. This work was funded by National Institutes of Health grants P01-AG001751 (D.J.M. and M.D.C.), R56-AG070096 (D.J.M. and J.E.B.), T32 AG066574 (G.A.P.), R35GM136255 (J.E.B.) and R01HL144778 (J.E.B.).
Author information
Authors and Affiliations
Contributions
A.A.B., G.A.P., D.J.M. and J.E.B. designed the experiments. A.A.B., G.A.P., D.J.M. and J.E.B. wrote and edited the manuscript. G.A.P., M.D.C. and R.S.S. performed animal experiments. A.A.B. performed cross-linking experiments, mass spectrometry raw data acquisition and processing. A.K. developed computational tools to support structural protein analysis and cross-linking quantitation. D.J.M. and J.E.B. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Aging thanks Luca Scorrano, Nicolas Demaurex and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Global quantitative cross-linking measurements.
a. Histogram of calculated Euclidean distances for all intraprotein cross-links mapped to AlphaFold predicted structures. b. Boxplots of number of ions used for each log2 ratio in P1, P2, P3, and P4. X-axis is capped at 40 for readability but there are ratios in each sample with more than 40 ions. c. Histogram of differences between a mean log2 ratios for cross-linked residue pairs based on multiple cross-linked peptides (cross-links that connect the same lysines, but can be identified in differently cleaved or modified peptides) and each cross-linked peptide pair d. Submitochondrial localization of significantly changed interlinks (left) and intralinks (right). e. STRING network of proteins with significantly changed cross-links. f. Isolated gastrocnemius mitochondria oxphos capacity normalized by amount of protein from the 4 young (6 months) and 4 old (30 months) female mice. Mean ± SD.
Extended Data Fig. 2 Complex I and Complex IV cross-linking analysis.
a. Heatmap of log2 ratios of all NDUS1 and NDUV1 cross-links and interprotein cross-links to other CI subunits. b. Boxplots of all intralinks in CI subunits by a biological replicate. c. Boxplots of crosslinks downregulated in aging and non-changing intralinks based on all 4 biological replicates for CI (NDUV1XLs n = 34; other links n = 112; visualized as median and 25th and 75th percentiles, with whiskers indicating minima and maxima) and CIV (f) Interlinks n = 30; other links n = 14; visualized as median and 25th and 75th percentiles, with whiskers indicating minima and maxima). P values are from Welch two-sided t-test. d. Structure of supercomplex (PDB 5GUP) with cross-linked CI and CIII subunits highlighted (top) and specific CI–CIII cross-links mapped to the subunits (bottom); decreased cross-linked are in green. e. CIII cross-links mapped to a bovine structure. Subunits with decreased intralinks highlighted and zoomed in (right). g. Correlation plots of CI and CIV cross-links changing in aging mitochondria and CII OXPHOS with multiple R-squared displayed.
Extended Data Fig. 3 Complex I and Complex IV in gel activity measurements show decrease in aged muscle mitochondria.
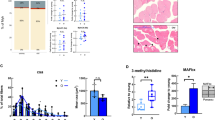
a. In-gel CI activity assay blot with identified bands. b. Quantification of total CI activity (left; statistical significance determined by unpaired two-tailed student’s t-test; p = 0.1726) and CI activity for each identified band (right; statistical significance determined by Ordinary Two-Way ANOVA with Sidak’s post hoc test; p = 0.0011 age main effect). c. In-gel CIV activity assay blot with identified bands. d. Quantification of total CIV activity (left; statistical significance determined by unpaired two-tailed student’s t-test; p = 0.1219) and CIV activity for each identified band (right; statistical significance determined by Ordinary Two-Way ANOVA with Sidak’s post hoc test; p = 0.0045 age main effect; p = 0.0059 for Band 1 Sidak’s post hoc test). Both activity assays were performed using isolated gastrocnemius mitochondria from young (4–6 month, n = 4) and old (27–29 month, n = 5) NIA C57BL/6 J female mice. Mean ± SD. ns, not significant, **p < 0.01 by Sidak’s post hoc test.
Extended Data Fig. 4 Antenna specific DHE3 cross-links decreased with aging.
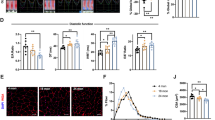
a. Decreased and non-changing cross-link levels in glutamate dehydrogenase highlighted on the volcano plot with Bonferroni corrected p value=0.05. b. Heatmap of all DHE3 cross-linked peptide pairs with each individual peptide sequence shown. Cross-linked lysine residues are in red. c. Correlation plots of DHE3 antenna cross-links changing in aged mitochondria and CII OXPHOS. d. Normalized glutamate stimulated respiration across a range of glutamate concentrations (left; statistical significance determined by Two-Way RM ANOVA with Sidak’s post hoc test; p = 0.0058 age main effect) and maximum respiration capacity with glutamate stimulation (right; statistical significance determined by unpaired two-tailed student’s t-test; p = 0.0009) in young (4–6 mo, n = 4) and old (27–29 mo, n = 4) female NIA C57BL/6 J isolated gastrocnemius mitochondria. e. The kinetics of glutamate stimulated respiration are altered with age (left; nonlinear regression determined by [Agonist] vs. normalized response–Variable slope) in young (4–6 mo, n = 4) and old (27–29 mo, n = 4) female NIA C57BL/6 J isolated gastrocnemius mitochondria. The amount of glutamate required to stimulate 50% respiration calculated from the nonlinear regression is increased in old (right; statistical significance determined by unpaired two-tailed student’s t-test; p = 0.0253). Mean ± SD. *p < 0.05, **p < 0.01.
Extended Data Fig. 5 TCA cycle and FAO enzymes show decrease in cross-links proximal to CoA binding sites.
a. Fumarate hydratase cross-links mapped to a E. Coli structure (PDB 4HGV). Decreased cross-link levels are shown in the zoomed in square. b. Heatmap of log2 ratios of fumarate hydratase cross-links. c. Boxplots for Acadvl cross-links based on all 4 biological replicates. CoA proximal XLs n = 21; other links n = 46; visualized as median and 25th and 75th percentiles, with whiskers indicating minima and maxima; P-values are from Welch’s two-sided t test.
Supplementary information
Supplementary Information
Supplementary results with Fig. 1 and discussion on limitations.
Supplementary Data 1
Calculations of log_2 changes for each young/old pair in mito yield, respiration and CS activity.
Source data
Source Data Fig. 2
Quantitative cross-linking data and functional measurements.
Source Data Fig. 3
Quantitative cross-linking data and functional measurements of complex I and complex IV.
Source Data Fig. 4
Quantitative cross-linking data and functional measurements of glutamate dehydrogenase.
Source Data Fig. 5
Quantitative cross-linking data for ACADVL, THIL and SUCA/SUCB.
Source Data Fig. 6
Correlation between quantitative cross-linking and functional measurements; XL clustering.
Source Data Extended Data Fig. 1
Global analysis of quantitative cross-linking measurements.
Source Data Extended Data Fig. 2
Quantitative cross-linking data and functional measurements of complex I and complex IV.
Source Data Extended Data Fig. 4a
Uncropped complex I activity gel.
Source Data Extended Data Fig. 4c
Uncropped complex IV activity gel.
Source Data Extended Data Fig. 3
Quantitation and statistical analysis of in gel CI and CIV activity for young and old mice.
Source Data Extended Data Fig. 4
Quantitative cross-linking data and functional measurements of DHE3.
Source Data Extended Data Fig. 5
Quantitative cross-linking data and functional measurements of ACADVL, THIL and SUCA/SUCB.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Bakhtina, A.A., Pharaoh, G.A., Campbell, M.D. et al. Skeletal muscle mitochondrial interactome remodeling is linked to functional decline in aged female mice. Nat Aging 3, 313–326 (2023). https://doi.org/10.1038/s43587-023-00366-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43587-023-00366-5