Abstract
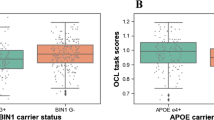
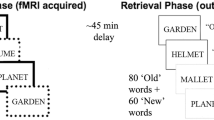
Although APOE ε4 carriers are at substantially higher risk of developing Alzheimer’s disease than noncarriers1, controversial evidence suggests that APOE ε4 might confer some advantages, explaining the survival of this gene (antagonistic pleiotropy)2,3. In a population-based cohort born in one week in 1946 (assessed aged 69–71 years), we assessed differential effects of APOE ε4 and β-amyloid pathology (quantified using 18F-Florbetapir-PET) on visual working memory (object–location binding). In 398 cognitively normal participants, APOE ε4 and β-amyloid had opposing effects on object identification, predicting better and poorer recall, respectively. ε4 carriers also recalled locations more precisely, with a greater advantage at higher β-amyloid burden. These results provide evidence of superior visual working memory in ε4 carriers, showing that some benefits of this genotype are demonstrable in older age, even in the preclinical stages of Alzheimer’s disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
All data from NSHD are curated and stored by the Lifelong Health and Aging Unit at UCL. Anonymized data will be shared by request from qualified investigators (https://skylark.ucl.ac.uk/NSHD/doku.php).
Code availability
Code for the 2D-mixture model (MATLAB) is freely available at https://doi.org/10.5281/zenodo.3752705. Code for statistical analyses conducted in Stata is provided in Supplementary Information.
References
Liu, C.-C., Kanekiyo, T., Xu, H. & Bu, G. Apolipoprotein E and Alzheimer disease: risk, mechanisms and therapy. Nat. Rev. Neurol. 9, 106–118 (2013).
Smith, C. J., Ashford, J. W. & Perfetti, T. A. Putative survival advantages in young apolipoprotein ɛ4 carriers are associated with increased neural stress. J. Alzheimers Dis. 68, 885–923 (2019).
Tuminello, E. R. & Duke Han, S. The apolipoprotein E antagonistic pleiotropy hypothesis: review and recommendations. Int. J. Alzheimers Dis. 2011, 726197 (2011).
Safieh, M., Korczyn, A. D. & Michaelson, D. M. ApoE4: an emerging therapeutic target for Alzheimer’s disease. BMC Med. 17, 64 (2019).
Jack, C. R. et al. Tracking pathophysiological processes in Alzheimer’s disease: an updated hypothetical model of dynamic biomarkers. Lancet Neurol. 12, 207–216 (2013).
Villemagne, V. L. et al. Amyloid β deposition, neurodegeneration, and cognitive decline in sporadic Alzheimer’s disease: a prospective cohort study. Lancet Neurol. 12, 357–367 (2013).
Baker, J. E. et al. Cognitive impairment and decline in cognitively normal older adults with high amyloid-β: a meta-analysis. Alzheimers Dement. 6, 108–121 (2017).
Brookmeyer, R. & Abdalla, N. Estimation of lifetime risks of Alzheimer’s disease dementia using biomarkers for preclinical disease. Alzheimers Dement. 14, 981–988 (2018).
Byars, S. G. & Voskarides, K. Antagonistic pleiotropy in human disease. J. Mol. Evol. 88, 12–25 (2020).
Jasienska, G. et al. Apolipoprotein E (ApoE) polymorphism is related to differences in potential fertility in women: a case of antagonistic pleiotropy? Proc. R. Soc. B Biol. Sci. 282, 20142395 (2015).
Duke Han, S. & Bondi, M. W. Revision of the apolipoprotein E compensatory mechanism recruitment hypothesis. Alzheimers Dement. 4, 251–254 (2008).
Rusted, J. M. et al. APOE e4 polymorphism in young adults is associated with improved attention and indexed by distinct neural signatures. Neuroimage 65, 364–373 (2013).
O’Donoghue, M. C., Murphy, S. E., Zamboni, G., Nobre, A. C. & Mackay, C. E. APOE genotype and cognition in healthy individuals at risk of Alzheimer’s disease: a review. Cortex 104, 103–123 (2018).
Iacono, D. & Feltis, G. C. Impact of apolipoprotein E gene polymorphism during normal and pathological conditions of the brain across the lifespan. Aging 11, 787–816 (2019).
Zink, N., Bensmann, W., Arning, L., Beste, C. & Stock, A. K. Apolipoprotein ε4 is associated with better cognitive control allocation in healthy young adults. Neuroimage 185, 274–285 (2019).
Marchant, N. L., King, S. L., Tabet, N. & Rusted, J. M. Positive effects of cholinergic stimulation favor young APOE e4 carriers. Neuropsychopharmacology 35, 1090–1096 (2010).
D’Souza, H. et al. Differential associations of apolipoprotein E ε4 genotype with attentional abilities across the life span of individuals with Down syndrome. JAMA Netw. Open 3, e2018221 (2020).
Austad, S. N. & Hoffman, J. M. Is antagonistic pleiotropy ubiquitous in aging biology? Evol. Med. Public Health 2018, 287–294 (2018).
Abondio, P. et al. The genetic variability of APOE in different human populations and its implications for longevity. Genes (Basel) 10, 222 (2019).
Raichlen, D. A. & Alexander, G. E. Exercise, APOE genotype, and the evolution of the human lifespan. Trends Neurosci. 37, 247–255 (2014).
Weissberger, G. H., Nation, D. A., Nguyen, C. P., Bondi, M. W. & Han, S. D. Meta-analysis of cognitive ability differences by apolipoprotein e genotype in young humans. Neurosci. Biobehav. Rev. 94, 49–58 (2018).
Pertzov, Y., Dong, M. Y., Peich, M.-C. & Husain, M. Forgetting what was where: the fragility of object-location binding. PLoS ONE 7, e48214 (2012).
Zokaei, N. et al. Sex and APOE: a memory advantage in male APOE ε4 carriers in midlife. Cortex 88, 98–105 (2017).
Zokaei, N. et al. Dissociable effects of the apolipoprotein-E (APOE) gene on short- and long-term memories. Neurobiol. Aging 73, 115–122 (2019).
Zokaei, N. et al. Short-term memory advantage for brief durations in human APOE ε4 carriers. Sci. Rep. 10, 9503 (2020).
Ma, W. J., Husain, M. & Bays, P. M. Changing concepts of working memory. Nat. Neurosci. 17, 347–356 (2014).
Zokaei, N., Burnett Heyes, S., Gorgoraptis, N., Budhdeo, S. & Husain, M. Working memory recall precision is a more sensitive index than span. J. Neuropsychol. 9, 319–329 (2015).
Lane, C. A. et al. Study protocol: Insight 46 – a neuroscience sub-study of the MRC National Survey of Health and Development. BMC Neurol. 17, dec2017 (2017).
Kuh, D. et al. The MRC National Survey of Health and Development reaches age 70: maintaining participation at older ages in a birth cohort study. Eur. J. Epidemiol. 31, 1135–1147 (2016).
Roberts, R. O. et al. Prevalence and outcomes of amyloid positivity among persons without dementia in a longitudinal, population-based setting. JAMA Neurol. 75, 970–979 (2018).
Kern, S. et al. Prevalence of preclinical Alzheimer disease: comparison of current classification systems. Neurology 90, e1682–e1691 (2018).
Prins, N. D. & Scheltens, P. White matter hyperintensities, cognitive impairment and dementia: an update. Nat. Rev. Neurol. 11, 157–165 (2015).
Pertzov, Y., Heider, M., Liang, Y. & Husain, M. Effects of healthy ageing on precision and binding of object location in visual short term memory. Psychol. Aging 30, 26–35 (2015).
Grogan, J. P. et al. A new toolbox to distinguish the sources of spatial memory error. J. Vis. 20, 6 (2020).
Di Battista, A. M., Heinsinger, N. M. & Rebeck, G. W. Alzheimer’s disease genetic risk factor APOE-ε4 also affects normal brain function. Curr. Alzheimer Res. 13, 1200–1207 (2016).
Scheller, E. et al. APOE moderates compensatory recruitment of neuronal resources during working memory processing in healthy older adults. Neurobiol. Aging 56, 127–137 (2017).
Rawle, M. J. et al. Apolipoprotein-E (Apoe) ε4 and cognitive decline over the adult life course. Transl. Psychiatry 8, 18 (2018).
Salvato, G. Does apolipoprotein e genotype influence cognition in middle-aged individuals? Curr. Opin. Neurol. 28, 612–617 (2015).
Small, B. J., Rosnick, C. B., Fratiglioni, L. & Bäckman, L. Apolipoprotein E and cognitive performance: a meta-analysis. Psychol. Aging 19, 592–600 (2004).
Vermunt, L. et al. Duration of preclinical, prodromal, and dementia stages of Alzheimer’s disease in relation to age, sex, and APOE genotype. Alzheimers Dement. 15, 888–898 (2019).
Lim, Y. Y. et al. Aβ-related memory decline in APOE ε4 noncarriers: implications for Alzheimer disease. Neurology 86, 1635–1642 (2016).
Liang, Y. et al. Visual short-term memory binding deficit in familial Alzheimer’s disease. Cortex 78, 150–164 (2016).
Lu, K. et al. Cognition at age 70: life course predictors and associations with brain pathologies. Neurology 93, e2144–e2156 (2019).
Lu, K. et al. Increased variability in reaction time is associated with amyloid beta pathology at age 70. Alzheimers Dement. (Amst.) 12, e12076 (2020).
James, S.-N. et al. Using a birth cohort to study brain health and preclinical dementia: recruitment and participation rates in Insight 46. BMC Res. Notes 11, 885 (2018).
Livingston, G. et al. Dementia prevention, intervention, and care: 2020 report of the Lancet Commission. Lancet 396, 413–446 (2020).
Lane, C. A. et al. Associations between blood pressure across adulthood and late-life brain structure and pathology in the neuroscience substudy of the 1946 British birth cohort (Insight 46): an epidemiological study. Lancet Neurol. 18, 942–952 (2019).
Lu, K. et al. Visuomotor integration deficits are common to both familial and sporadic preclinical Alzheimer’s disease. Brain Commun. 3, fcab003 (2021).
Pertzov, Y. et al. Binding deficits in memory following medial temporal lobe damage in patients with voltage-gated potassium channel complex antibody-associated limbic encephalitis. Brain 136, 2474–2485 (2013).
Grogan, J. P. johnPGrogan/MemToolbox2D: updated release. Zenodo https://zenodo.org/record/3752705/export/hx#.YSi5UPlKjIU (2020).
Parker, T. D. et al. Hippocampal subfield volumes and pre-clinical Alzheimer’s disease in 408 cognitively normal adults born in 1946. PLoS ONE 14, e0224030 (2019).
Sudre, C. H. et al. Bayesian model selection for pathological neuroimaging data applied to white matter lesion segmentation. IEEE Trans. Med. Imaging 34, 2079–2102 (2015).
Cardoso, J. et al. STEPS: similarity and truth estimation for propagated segmentations and its application to hippocampal segmentation and brain parcelation. Med. Image Anal. 17, 671–684 (2013).
Malone, I. B. et al. Accurate automatic estimation of total intracranial volume: a nuisance variable with less nuisance. Neuroimage 104, 366–372 (2015).
Richards, M. & Sacker, A. Lifetime antecedents of cognitive reserve. J. Clin. Exp. Neuropsychol. 25, 614–624 (2003).
Richards, M. et al. Identifying the lifetime cognitive and socioeconomic antecedents of cognitive state: seven decades of follow-up in a British birth cohort study. BMJ Open 9, e024404 (2019).
Armstrong, R. A. When to use the Bonferroni correction. Ophthalmic Physiol. Opt. 34, 502–508 (2014).
Althouse, A. D. Adjust for multiple comparisons? It’s not that simple. Ann. Thorac. Surg. 101, 1644–1645 (2016).
Acknowledgements
This study was principally funded by grants from Alzheimer’s Research UK (nos. ARUK-PG2014-1946 and ARUK-PG2017-1946), the Medical Research Council Dementias Platform UK (no. CSUB19166) and the Selfridges Group Foundation (no. PR/ylr/18575). Genetic analyses were funded by the Brain Research Trust (no. UCC14191). The Florbetapir amyloid tracer was kindly provided by AVID Radiopharmaceuticals (a wholly owned subsidiary of Eli Lilly), who had no part in the design of the study. NSHD is funded by the Medical Research Council (nos. MC_UU_12019/06 and MC_UU_12019/08). The funders of the study had no role in study design, data collection, analysis, interpretation, report writing or the decision to submit the article for publication. T.D.P. was supported by a Wellcome Trust Clinical Research Fellowship (no. 200109/Z/15/Z). A.K. was supported by a Wolfson Clinical Research Fellowship. C.H.S. is supported by an Alzheimer’s Society Junior Fellowship (no. AS-JF-17-011). N.C.F. acknowledges support from the UK Dementia Research Institute at University College London, the National Institute for Health Research (Senior Investigator award) and University College London Hospitals Biomedical Research Centre. J.M.S. is supported by University College London Hospitals Biomedical Research Centre, Engineering and Physical Sciences Research Council (no. EP/J020990/1), British Heart Foundation (no. PG/17/90/33415) and EU’s Horizon 2020 research and innovation program (no. 666992). We thank participants both for their contributions to Insight 46 and for their commitments to research over the past seven decades. We are grateful to the radiographers and nuclear medicine physicians (A. Groves, J. Bomanji and I. Kayani) at the UCL Institute of Nuclear Medicine, and to the staff at the Leonard Wolfson Experimental Neurology Centre at UCL. We thank D. Marcus and R. Herrick for assistance with XNAT, P. Curran for assistance with data sharing with the MRC Unit for Lifelong Health and Ageing, the DRC trials team for assistance with imaging quality control, M. White for his work on data connectivity and J. Dickson, A. Barnes and D. Thomas for help with imaging.
Author information
Authors and Affiliations
Contributions
J.M.S., S.J.C., M.R. and N.C.F. conceptualized and led the Insight 46 study. Y.P. and M.H. designed the visual working memory experiment. K.L., I.M.P. and S.-N.J. collected data for the visual working memory test. T.D.P., C.A.L., A.K., S.E.K. and S.M.B. collected clinical and neuroimaging data. H.M.-S. and A.W. were responsible for study coordination and data management. K.L., S.M.D.H., J.M.S. and S.J.C. conceived the manuscript. J.G. and M.H. developed the 2D-mixture model. K.L. analyzed data and drafted the initial manuscript. J.M.N. provided statistical support. D.M.C., I.B.M., C.H.S. and W.C. generated neuroimaging outcomes. K.L., S.M.D.H., J.G., M.H., J.M.S. and S.J.C. aided in manuscript preparation and interpretation. All authors revised and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Aging thanks Duke Han, Miranka Wirth and the other, anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
43587_2021_117_MOESM1_ESM.pdf
Supplementary analyses and discussion in nine subsections, incorporating Tables 1–5 and Figs. 1–6; code for statistical analyses reported in the main manuscript and supplementary analyses; and visual representation of the raw response locations for each trial.
Rights and permissions
About this article
Cite this article
Lu, K., Nicholas, J.M., Pertzov, Y. et al. Dissociable effects of APOE ε4 and β-amyloid pathology on visual working memory. Nat Aging 1, 1002–1009 (2021). https://doi.org/10.1038/s43587-021-00117-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43587-021-00117-4
This article is cited by
-
Updating the study protocol: Insight 46 – a longitudinal neuroscience sub-study of the MRC National Survey of Health and Development – phases 2 and 3
BMC Neurology (2024)
-
Genetic association between the APOE ε4 allele, toxicant exposures and Gulf war illness diagnosis
Environmental Health (2023)
-
Inter- and intra-chromosomal modulators of the APOE ɛ2 and ɛ4 effects on the Alzheimer’s disease risk
GeroScience (2023)
-
Genetic polymorphism in BIN1 rather than APOE is associated with poor recognition memory among men without dementia
Scientific Reports (2022)
-
The Power of Birth Cohorts to Study Risk Factors for Cognitive Impairment
Current Neurology and Neuroscience Reports (2022)