Abstract
A repressive chromatin state featuring trimethylated lysine 36 on histone H3 (H3K36me3) and DNA methylation suppresses cryptic transcription in embryonic stem cells. Cryptic transcription is elevated with age in yeast and nematodes and reducing it extends yeast lifespan, though whether this occurs in mammals is unknown. We show that cryptic transcription is elevated in aged mammalian stem cells, including murine hematopoietic and neural stem cells and human mesenchymal stem cells. Precise mapping allowed quantification of age-associated cryptic transcription in human mesenchymal stem cells aged in vitro. Regions with significant age-associated cryptic transcription have a unique chromatin signature: decreased H3K36me3 and increased H3K4me1, H3K4me3 and H3K27ac with age. Genomic regions undergoing such changes resemble known promoter sequences and are bound by TATA-binding protein, even in young cells. Hence, the more permissive chromatin state at intragenic cryptic promoters likely underlies increased cryptic transcription in aged mammalian stem cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All RNA-seq, ChIP-seq, WGBS and CMS-IP-seq data have been deposited in the GEO database at NCBI under accession code GSE156409.
Code availability
All code for implementing the analyses described in this paper is available at GitHub at https://github.com/NyxSLY/ASCT.
References
López-Otín, C., Blasco, M. A., Partridge, L., Serrano, M. & Kroemer, G. The hallmarks of aging. Cell 153, 1194 (2013).
Booth, L. N. & Brunet, A. The aging epigenome. Mol. Cell 62, 728–744 (2016).
Zhan, M. et al. Temporal and spatial transcriptional profiles of aging in Drosophila melanogaster. Genome Res. 17, 1236–1243 (2007).
Lai, R. W. et al. Multi-level remodeling of transcriptional landscapes in aging and longevity. BMB Rep. 52, 86–108 (2019).
Son, H. G., Altintas, O., Kim, E. J. E., Kwon, S. & Lee, S. J. V. Age-dependent changes and biomarkers of aging in Caenorhabditis elegans. Aging Cell 18, 1–11 (2019).
Sen, P. et al. H3K36 methylation promotes longevity by enhancing transcriptional fidelity. Genes Dev. 29, 1362–1376 (2015).
McCauley, B. S. & Dang, W. Histone methylation and aging: lessons learned from model systems. Biochim. Biophys. Acta Gene Regul. Mech. 1839, 1454–1462 (2014).
Sen, P., Shah, P. P., Nativio, R. & Berger, S. L. Epigenetic mechanisms of longevity and aging. Cell 166, 822–839 (2016).
Hennig, B. P. & Fischer, T. Chromatin and cryptic transcription. Transcription 4, 97–101 (2013).
Carvalho, S. et al. Histone methyltransferase SETD2 coordinates FACT recruitment with nucleosome dynamics during transcription. Nucleic Acids Res. 41, 2881–2893 (2013).
Neri, F. et al. Intragenic DNA methylation prevents spurious transcription initiation. Nature 543, 72–77 (2017).
Xie, L. et al. KDM5B regulates embryonic stem cell self-renewal and represses cryptic intragenic transcription. EMBO J. 30, 1473–1484 (2011).
Venkatesh, S. & Workman, J. L. Histone exchange, chromatin structure and the regulation of transcription. Nat. Rev. Mol. Cell Biol. 16, 178–189 (2015).
Belotserkovskaya, R. et al. FACT facilitates transcription-dependent nucleosome alteration. Science 301, 1090–1093 (2003).
Kaplan, C. D., Laprade, L. & Winston, F. Transcription elongation factors repress transcription initiation from cryptic sites. Science 301, 1096–1099 (2003).
Carrozza, M. J. et al. Histone H3 methylation by Set2 directs deacetylation of coding regions by Rpd3S to suppress spurious intragenic transcription. Cell 123, 581–592 (2005).
Pu, M. et al. Trimethylation of Lys36 on H3 restricts gene expression change during aging and impacts life span. Genes Dev. 29, 718–731 (2015).
Ni, Z., Ebata, A., Alipanahiramandi, E. & Lee, S. S. Two SET domain containing genes link epigenetic changes and aging in Caenorhabditis elegans. Aging Cell 11, 315–325 (2012).
Goodell, M. A. & Rando, T. A. Stem cells and healthy aging. Science 350, 1199–1204 (2015).
Sun, D. et al. Epigenomic profiling of young and aged HSCs reveals concerted changes during aging that reinforce self-renewal. Cell Stem Cell 14, 673–688 (2014).
Wagner, W. et al. Aging and replicative senescence have related effects on human stem and progenitor cells. PLoS ONE https://doi.org/10.1371/journal.pone.0005846 (2009).
Ferrari, K. J. et al. Polycomb-dependent H3K27me1 and H3K27me2 regulate active transcription and enhancer fidelity. Mol. Cell 53, 49–62 (2014).
Zhang, Y. et al. H3K36 histone methyltransferase Setd2 is required for murine embryonic stem cell differentiation toward endoderm. Cell Rep. 8, 1989–2002 (2014).
Xu, Q. et al. SETD2 regulates the maternal epigenome, genomic imprinting and embryonic development. Nat. Genet. 51, 844–856 (2019).
Urbán, N., Blomfield, I. M. & Guillemot, F. Quiescence of adult mammalian neural stem cells: a highly regulated rest. Neuron 104, 834–848 (2019).
Adelman, E. R. et al. Aging human hematopoietic stem cells manifest profound epigenetic reprogramming of enhancers that may predispose to leukemia. Cancer Discov. 9, 1080–1101 (2019).
Boisvert, M. M., Erikson, G. A., Shokhirev, M. N. & Allen, N. J. The aging astrocyte transcriptome from multiple regions of the mouse brain. Cell Rep. 22, 269–285 (2018).
Clarke, L. E. et al. Normal aging induces A1-like astrocyte reactivity. Proc. Natl Acad. Sci. USA 115, E1896–E1905 (2018).
Fleischer, J. G. et al. Predicting age from the transcriptome of human dermal fibroblasts. Genome Biol. 19, 1–8 (2018).
Kaisers, W. et al. Age, gender and UV-exposition related effects on gene expression in in vivo aged short term cultivated human dermal fibroblasts. PLoS ONE 12, 1–21 (2017).
MacRae, S. L. et al. DNA repair in species with extreme lifespan differences. Aging 7, 1171–1184 (2015).
Marthandan, S. et al. Conserved senescence associated genes and pathways in primary human fibroblasts detected by RNA-seq. PLoS ONE 11, 1–31 (2016).
Marthandan, S. et al. Similarities in gene expression profiles during in vitro aging of primary human embryonic lung and foreskin fibroblasts. Biomed Res. Int. https://doi.org/10.1155/2015/731938 (2015).
Rai, T. S. et al. HIRA orchestrates a dynamic chromatin landscape in senescence and is required for suppression of Neoplasia. Genes Dev. 28, 2712–2725 (2014).
Stilling, R. M. et al. De-regulation of gene expression and alternative splicing affects distinct cellular pathways in the aging hippocampus. Front. Cell Neurosci. 8, 1–15 (2014).
Johnson, B. S. et al. Biotin tagging of MeCP2 in mice reveals contextual insights into the Rett syndrome transcriptome. Nat. Med. 23, 1203–1214 (2017).
Zhang, W. et al. A Werner syndrome stem cell model unveils heterochromatin alterations as a driver of human aging. Science 348, 1160–1163 (2015).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
McDaniel, S. L. et al. H3K36 methylation regulates nutrient stress response in Saccharomyces cerevisiae by enforcing transcriptional fidelity. Cell Rep. 19, 2371–2382 (2017).
Haupt, S., Söntgerath, V. S. A., Leipe, J., Schulze-Koops, H. & Skapenko, A. Methylation of an intragenic alternative promoter regulates transcription of GARP. Biochim. Biophys. Acta Gene Regul. Mech. 1859, 223–234 (2016).
Cheung, V. et al. Chromatin- and transcription-related factors repress transcription from within coding regions throughout the Saccharomyces cerevisiae genome. PLoS Biol. 6, 2550–2562 (2008).
Ernst, J. & Kellis, M. ChromHMM: automating chromatin-state discovery and characterization. Nat. Methods 9, 215–216 (2012).
Knudsen, S. Promoter2.0: For the recognition of PolII promoter sequences. Bioinformatics 15, 356–361 (1999).
Yagi, S. & Galea, L. A. M. Sex differences in hippocampal cognition and neurogenesis. Neuropsychopharmacology 44, 200–213 (2019).
Challen, G. A. et al. Dnmt3a and Dnmt3b have overlapping and distinct functions in hematopoietic stem cells. Cell Stem Cell 15, 350–364 (2014).
Ziller, M. J. et al. Dissecting the functional consequences of de novo DNA methylation dynamics in human motor neuron differentiation and physiology. Cell Stem Cell 22, 559–574.e9 (2018).
Stewart, M. H. et al. The histone demethylase Jarid1b is required for hematopoietic stem cell self-renewal in mice. Blood 125, 2075–2078 (2015).
Dimri, G. P. et al. A biomarker that identifies senescent human cells in culture and in aging skin in vivo. Proc. Natl Acad. Sci. USA 92, 9363–9367 (1995).
Mori, E. et al. Impaired adipogenic capacity in induced pluripotent stem cells from lipodystrophic patients with BSCL2 mutations. Metabolism 65, 543–556 (2016).
Liu, B. et al. A protocol for isolation and identification and comparative characterization of primary osteoblasts from mouse and rat calvaria. Cell Tissue Bank 20, 173–182 (2019).
Dang, W. et al. Histone H4 lysine 16 acetylation regulates cellular lifespan. Nature 459, 802–807 (2009).
Lister, R. et al. Human DNA methylomes at base resolution show widespread epigenomic differences. Nature 462, 315–322 (2009).
Pastor, W. A. et al. Genome-wide mapping of 5-hydroxymethylcytosine in embryonic stem cells. Nature 473, 394–397 (2011).
Huang, Y., Pastor, W. A., Zepeda-Martínez, J. A. & Rao, A. The anti-CMS technique for genome-wide mapping of 5-hydroxymethylcytosine. Nat. Protoc. 7, 1897–1908 (2012).
Codega, P. et al. Prospective identification and purification of quiescent adult neural stem cells from their in vivo niche. Neuron 82, 545–559 (2014).
Leeman, D. S. et al. Lysosome activation clears aggregates and enhances quiescent neural stem cell activation during aging. Science 359, 1277–1283 (2018).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Krueger, F. & Andrews, S. R. Bismark: a flexible aligner and methylation caller for bisulfite-seq applications. Bioinformatics 27, 1571–1572 (2011).
Amemiya, H. M., Kundaje, A. & Boyle, A. P. The ENCODE blacklist: identification of problematic regions of the genome. Sci. Rep. 9, 1–5 (2019).
Dunham, I. et al. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017).
Liao, Y., Smyth, G. K. & Shi, W. FeatureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Lovén, J. et al. Selective inhibition of tumor oncogenes by disruption of super-enhancers. Cell 153, 320–334 (2013).
Whyte, W. A. et al. Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell 153, 307–319 (2013).
Lun, A. T. L. & Smyth, G. K. Csaw: a Bioconductor package for differential binding analysis of ChIP-seq data using sliding windows. Nucleic Acids Res. 44, e45 (2015).
Robinson, M. D. & Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. https://doi.org/10.1186/gb-2010-11-3-r25 (2010).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B 57, 289–300 (2016).
Dennis, G. et al. DAVID: Database for Annotation, Visualization, and Integrated Discovery. Genome Biol. https://doi.org/10.1186/gb-2003-4-9-r60 (2003).
Yu, G., Wang, L. G. & He, Q. Y. ChIP seeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics 31, 2382–2383 (2015).
Yu, G., Wang, L.-G., Han, Y. & He, Q.-Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Johnson, N. L., Kotz, S. & Kemp, A. W. Univariate Discrete Distrubtions 2nd Edn (John Wiley and Sons, 1992).
Ramírez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Akalin, A. et al. MethylKit: a comprehensive R package for the analysis of genome-wide DNA methylation profiles. Genome Biol. https://doi.org/10.1186/gb-2012-13-10-r87 (2012).
Zhang, Y. et al. Model-based analysis of ChIP-seq (MACS). Genome Biol. https://doi.org/10.1186/gb-2008-9-9-r137 (2008).
Gel, B. et al. RegioneR: an R/Bioconductor package for the association analysis of genomic regions based on permutation tests. Bioinformatics 32, 289–291 (2016).
Harmanci, A., Rozowsky, J. & Gerstein, M. MUSIC: identification of enriched regions in ChIP-seq experiments using a mappability-corrected multiscale signal processing framework. Genome Biol. 15, 474 (2014).
Acknowledgements
We thank R. Chen and the Human Genome Sequence Center at Baylor College of Medicine for performing the Illumina sequencing reported here. This work was funded by National Institutes of Health grants R01AG052507 to W.D. and R01AG053268 to A.E.W.; R01HL134780 and R01HL146852 to Y.H.; CPRIT award R1306 to W.D.; and a Ted Nash Long Life Foundation research grant to W.D. B.S.M. was supported by National Institutes of Health training grant T32AG000183.
Author information
Authors and Affiliations
Contributions
Author contributions were as follows: conceptualization, W.D., B.S.M. and L.S.; methodology, W.D., B.S.M., L.S. and Y.H.; investigation, B.S.M., L.S., R.Y., M.L., H.L., D.S.L., Y.H., A.E.W. and W.D.; writing of original draft, B.S.M., L.Y. and W.D.; writing, review and editing, B.S.M., L.S., R.Y., D.S.L., Y.H., A.E.W., M.K. and W.D.; funding acquisition, W.D., Y.H. and A.E.W.; and supervision, W.D.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Aging thanks Bérénice Benayoun, Aaron Viny and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Additional analysis of aging RNA-seq from mHSCs and hMSCs.
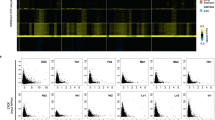
a, Growth curve of culture-expanded hMSCs; PD: population doubling. b, Proportion of senescence-associated β-galatosidase stained hMSCs at the indicated PDs, showing standard error of the mean. In total, 1,629, 1,641, and 293 cells were analyzed in PD 12, PD 32 and PD 42, respectively. c, Adipogenic and osteogenic differentiation of hMSCs is shown by Oil Red O (ORO) and Alizarin Red S (ARS) staining. Experiments were performed 3 times. d, Boxplots of the log2-transformed ratio of reads mapping to the indicated exon vs. reads mapping to the first or second exon (dark and light orange, respectively) in Setd2-perturbed vs. control samples (ratio in Setd2-perturbed divided by ratio in control, or ratio of ratios). Samples used: Setd2 knockout (n = 6,869, GSE72855; n = 7,821, E-GEOD-54932)11,23 or knockdown (n = 6,606, E-GEOD-51006)22 in murine embryonic stem cells and knockout in murine oocytes (n = 7,143, GSE112832)24. e,f, Boxplots showing the log2-transformed ratio of RNA-seq reads mapping to the indicated exons vs. the first exon of genes in mHSCs in e and hMSCs in f. Young samples are in blue and old in red; Y: young, O: old. g,h, Boxplots of the log2-transformed ratio of reads mapping to the indicated exons vs. reads mapping to the first exon in old vs. young samples (ratio in old divided by ratio in young, or ratio of ratios) divided by expression quartile in mHSCs in g and hMSCs in h. i, j, Bar charts of transcripts in which the indicated exon has a 2-fold increase (red) or decrease (blue) in TPM vs. the first exon; mHSCs in i and hMSCs in j. k,l, Histograms of the CT scores of major transcripts; mHSCs in k and hMSCs in l. m,n, Scatterplots showing the log2-transformed CT scores in old vs. young samples; mHSCs in m and hMSCs in n. Blue indicates an age-associated increase in cryptic transcription. o,p, Length distribution of transcripts with increased cryptic transcription (n = 210 for mHSCs and n = 305 for hMSCs) with age vs. expressed major transcripts with at least 3 exons. mHSCs are in o and hMSCs in p. For boxplots, bounds of box show the 25th and 75th percentiles; the central lines in the box plots represent the median value; and the whiskers show 1.5-fold of the interquartile range. P values were calculated using a two-sided Wilcoxon signed-rank test vs. the null hypothesis that the samples have the same average value or the log2-transformed ratio of ratios equals 1. Exact P values for panels D, G, and H are provided in Supplementary Table 1. Expressed major transcripts with at least 3 exons were included in the cryptic transcription analyses for mHSCs (n = 10,068, panels e, g, and o) and hMSCs (n = 9,230, panels f, h, and p).
Extended Data Fig. 2 Analysis of aging RNA-seq in NSCs and other tissues.
a,b, Boxplots showing the log2-transformed ratio of ratios (indicated exon vs. first exon) for transcripts in aNSCs, separated into quartiles by expression levels. aNSCs isolated from female mice are shown in a and from males in b; Y indicates young and O old. Expressed major transcripts with at least 3 exons were included in the analyses for females (n = 6,110) and males (n = 46,54). P values were calculated using a two-sided Wilcoxon signed-rank test with the null hypothesis that the calculated log2 ratios are equal to 0; exact P values are provided in Supplementary Table 1. c, Comparison of the distribution of the length of transcripts with an increase in cryptic transcription with age vs. expressed major transcripts with at least 3 exons in aNSCs, shown as a histogram and boxplot. aNSCs isolated from females on top (n = 266 for genes with age-increased cryptic transcription and n = 6,110 for all major transcripts) and from males on the bottom (n = 237 for genes with age-increased cryptic transcription and n = 4,654 for all major transcripts). P values were calculated using a two-sided Wilcoxon signed-rank test. d, Heatmap depicting the log2-transformed ratio of ratios (indicated exon vs. the first exon) from a variety of mammalian aging or senescence RNA-seq datasets, identified in the figure (E-GEOD-59966; E-GEOD-46486; GSE53330; E-MTAB-4879; and refs. 26,27,28,29,30,31,32,33,34,35). e,f, Boxplots showing the log2-transformed ratio of ratios (indicated exon vs. first or second exon) for transcripts in fibroblasts from Rett syndrome patients vs. controls36 in e and cells engineered to carry a mutation in LMNA that causes Werner syndrome37 in f. Expressed major transcripts with at least 3 exons were included in the analysis (Rett syndrome, n = 7,302; Werner syndrome MSCs, n = 8,934, Werner syndrome ESCs, n = 10,185). No significant result founds were in e and f using a two-sided Wilcoxon signed-rank test. For boxplots, bounds of box show the 25th and 75th percentiles; the central lines in the box plots represent the median value; and whiskers show 1.5-fold of the interquartile range.
Extended Data Fig. 3 Additional analysis of cryptic transcription in aging hMSCs.
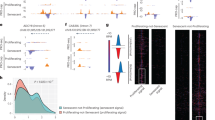
a, DECAP-seq read pile ups around cTSSs that were identified as having higher DECAP-seq peaks in the young hMSC sample vs. the old, that is, sites where cryptic transcription decreases with age. b, Genes were ranked by the ratio of their FPKM in young cells vs. FPKM in old. Histograms showing the ranked distribution of genes in the following categories: all genes (top); genes with sites that have an age-associated increase in cryptic transcription (middle); and genes with sites that have a decrease in cryptic transcription with age (bottom). FC indicates fold change. c, RT-qPCR results showing a mild decrease (~30%) in SETD2 RNA levels upon SETD2 knockdown in hMSCs. d, Growth curve showing growth of hMSCs expressing a control, non-targeting (NT) shRNA vs. those expressing SETD2 shRNA. e, Complete HOMER de novo motif results of the significant motifs found from age-increased cTSS flanking regions (±200 bp). Known promoter elements are highlighted in green. The at-AC splice acceptor sequence is shown in blue. TFs that bind motifs similar to the ones identified by HOMER are shown in grey if they are not expressed in hMSCs (FPKM < 1); ones listed in black are expressed in hMSCs. TFs highlighted in bold and indicated with an asterisk show the highest age-associated changes in expression and were included in a GO analysis. The P value was directly calculated by HOMER Motif Analysis38. f, GO analysis of putative targets of the indicated transcription factors in ENCODE datasets. Gene ratio indicates the proportion of genes in the dataset that fall in the GO cluster. In all panels, cTSS: cryptic transcription start site. Enrichment P values were generated by a one-sided hypergeometric test to determine if the list contains more genes for the GO cluster than expected by chance.
Extended Data Fig. 4 Genome-wide analysis of chromatin states.
a, Emission parameters of the ten state ChromHMM model in hMSCs. b, Enrichment of the ChromHMM states in the indicated genomic regions in young hMSCs. c, As b, except in old hMSCs. d, Enrichment of the ChromHMM states around annotated TSSs and TESs in young and old hMSCs. e, Transition map of ChromHMM states in old vs. young hMSCs. State in old is along the x-axis and state in young along the y-axis. f, Two examples of a decline in H3K9me3 (ChromHMM state 1) enrichment at LADs with age. Normalized mapped reads are shown in blue for young hMSCs and in red for old. g, Distribution of the chromatin states of age-decreased cTSSs determined by DECAP-seq in young and old hMSCs. h, Methylated CpG distributions in young (left) and old (right) hMSCs. i, Number of CMS-IP-seq (5-hydroxymethylcytosine) peaks in young and old hMSCs. j, Graphical representation of a one-sided permutation test with the null hypothesis that the number of CMS-IP-seq peaks that overlap with age-increased DECAP-seq peaks is equal to the background level of CMS-IP-peaks. This shows a significant overlap of CMS-IP-seq peaks with age-increased cTSSes. In all panels, Y: young; O: old; TSS: transcription start site; TES: transcription end site; LAD: lamin-associated domain.
Extended Data Fig. 5 Chromatin state changes around age-increased cTSSes.
a, Read pile ups of H3K4me3 around annotated TSSs (left), age-increased cTSSs (middle), and age-decreased cTSSs (right), independently clustered into 3 groups. b, As in a, except H3K4me1 enrichment is shown. c, As in a, except depicting H3K27ac enrichment. d, Boxplots showing H3K36me3 enrichment around age-increased cTSSs (n = 1,373) and endogenous TSSs (n = 13,802) in young and old hMSCs. P values were calculated using a two-sided Wilcoxon signed-rank test with the null hypothesis that enrichment was equal in the young and old samples. e, Bar chart showing the proportion of TBP ChIP-seq peaks around endogenous TSSs in young and old hMSCs. f, Metagene plot of TBP enrichment around annotated TSSs in hMSCs. g, DECAP-seq signal around putative age-associated cTSSs predicted in hMSCs by the chromatin state model. Averaged read depth of putative age-associated promoter regions (±1 kb of the midpoint of the identified region) in young (blue) and old (red) is shown on the left at 100 bp resolution; a boxplot of the log2-transformed ratio of signal in old vs. signal in young shown on the right (n = 166). Distinct random genic non-promoter regions (length = 2 kb) were used as control (n = 2,774). P values were calculated using a two-sided Wilcoxon signed-rank test vs. the hypothesis that the RNA-seq ratios were equal in the putative age-increased cTSSs vs. control regions, as appropriate. Regions without DECAP-seq signal were excluded from analysis. In all panels, Y: young; O: old; TSS: transcription start site; cTSS: cryptic transcription start site. For boxplots, bounds of box show the 25th and 75th percentiles; the central lines in the box plots represent the median value; and whiskers show 1.5-fold of the interquartile range.
Supplementary information
Supplementary Information
Supplementary Fig. 1 and associated legend.
Supplementary Tables
Contains Supplementary Table 1 (exact P values for Figs. 1 and 2 and Extended Data Figs. 1 and 2); Supplementary Table 2 (DECAP-seq peak and gene lists for DAVID analysis); Supplementary Table 3 (DAVID analysis of DECAP-seq peaks in young hMSCs); Supplementary Table 4 (DAVID analysis of DECAP-seq peaks in old hMSCs); Supplementary Table 5 (DAVID analysis of age-increased DECAP-seq peaks); and Supplementary Table 6 (GO analysis of target genes of cryptic transcription-associated TFs in ENCODE datasets).
Rights and permissions
About this article
Cite this article
McCauley, B.S., Sun, L., Yu, R. et al. Altered chromatin states drive cryptic transcription in aging mammalian stem cells. Nat Aging 1, 684–697 (2021). https://doi.org/10.1038/s43587-021-00091-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43587-021-00091-x
This article is cited by
-
Vitamin B12 is a limiting factor for induced cellular plasticity and tissue repair
Nature Metabolism (2023)
-
5-hydroxymethylcytosine stabilizes transcription by preventing aberrant initiation in gene bodies
Nature Genetics (2023)
-
Spurious intragenic transcription is a feature of mammalian cellular senescence and tissue aging
Nature Aging (2023)
-
Methylation across the central dogma in health and diseases: new therapeutic strategies
Signal Transduction and Targeted Therapy (2023)
-
Structural basis of nucleosome deacetylation and DNA linker tightening by Rpd3S histone deacetylase complex
Cell Research (2023)