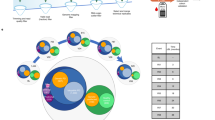
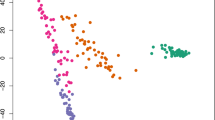
Abstract
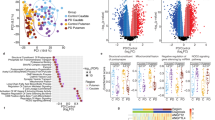
Changes in the blood-based RNA transcriptome have the potential to inform biomarkers of Parkinson’s disease (PD) progression. Here we sequenced a discovery set of whole-blood RNA species in 4,871 longitudinally collected samples from 1,570 clinically phenotyped individuals from the Parkinson’s Progression Marker Initiative (PPMI) cohort. Samples were sequenced to an average of 100 million read pairs to create a high-quality transcriptome. Participants with PD in the PPMI had significantly altered RNA expression (>2,000 differentially expressed genes), including an early and persistent increase in neutrophil gene expression, with a concomitant decrease in lymphocyte cell counts. This was validated in a cohort from the Parkinson’s Disease Biomarkers Program (PDBP) consisting of 1,599 participants and by alterations in immune cell subtypes. This publicly available transcriptomic dataset, coupled with available detailed clinical data, provides new insights into PD biological processes impacting whole blood and new paths for developing diagnostic and prognostic PD biomarkers.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Raw sequencing data (FASTQ files), alignment files (BAM files), TPM data and counts for each sample are available at the LONI IDA. (https://fairsharing.org/, IDA; LONI IDA, https://doi.org/10.25504/FAIRsharing.r4ph5f). Data are also available through the AMP PD (https://amp-pd.org/). These are the requirements for downloading from the AMP PD: (1) personal and institutional or company details; (2) description of intended data use, for example, proposed analyses; (3) institutional signature on the AMP PD Data Use Agreement (for researchers requesting access to individual-level, ’omics data). Additional data, including but not limited to study arm, motor assessments, DaTscan and MRI imaging, genetic testing results, whole-exome and genome sequencing data, patient history and standardized techniques and protocols for data collection are also available through the IDA. To access complete data, researchers need to fill out a data-use agreement. Data are available in a public (institutional, general or participant-specific) repository that does not issue datasets with DOIs (non-mandated deposition).
Code availability
All code used for data analysis is available at GitHub: https://github.com/tgen/ppmi-qc-wt-paper and https://github.com/tgen/ppmi-rnaseq-wb-paper.
References
Dorsey, E. R., Sherer, T., Okun, M. S. & Bloem, B. R. The emerging evidence of the Parkinson pandemic. J. Parkinsons Dis. 8, S3–S8 (2018).
Yang, W. et al. Current and projected future economic burden of Parkinson’s disease in the U.S. NPJ Parkinsons Dis. 6, 15 (2020).
Latourelle, J. C. et al. Large-scale identification of clinical and genetic predictors of motor progression in patients with newly diagnosed Parkinson’s disease: a longitudinal cohort study and validation. Lancet Neurol. 16, 908–916 (2017).
Marek, K. et al. The Parkinson’s Progression Markers Initiative (PPMI)—establishing a PD biomarker cohort. Ann. Clin. Transl. Neurol. 5, 1460–1477 (2018).
Nalls, M. A. et al. Baseline genetic associations in the Parkinson’s Progression Markers Initiative (PPMI). Mov. Disord. 31, 79–85 (2016).
Nalls, M. A. et al. Diagnosis of Parkinson’s disease on the basis of clinical and genetic classification: a population-based modelling study. Lancet Neurol. 14, 1002–1009 (2015).
Parkinson Progression Marker Initiative. The Parkinson Progression Marker Initiative (PPMI). Prog. Neurobiol. 95, 629–635 (2011).
Simuni, T. et al. Longitudinal change of clinical and biological measures in early Parkinson’s disease: Parkinson’s Progression Markers Initiative cohort. Mov. Disord. 33, 771–782 (2018).
Caspell-Garcia, C. et al. Multiple modality biomarker prediction of cognitive impairment in prospectively followed de novo Parkinson disease. PLoS ONE 12, e0175674 (2017).
Kang, J. H. et al. Association of cerebrospinal fluid β-amyloid 1-42, T-tau, P-tau181, and α-synuclein levels with clinical features of drug-naive patients with early Parkinson disease. JAMA Neurol. 70, 1277–1287 (2013).
Simuni, T. et al. Clinical and dopamine transporter imaging characteristics of non-manifest LRRK2 and GBA mutation carriers in the Parkinson’s Progression Markers Initiative (PPMI): a cross-sectional study. Lancet Neurol. 19, 71–80 (2020).
Polymeropoulos, M. H. et al. Mutation in the α-synuclein gene identified in families with Parkinson’s disease. Science 276, 2045–2047 (1997).
Hernandez, D. G., Reed, X. & Singleton, A. B. Genetics in Parkinson disease: Mendelian versus non-Mendelian inheritance. J. Neurochem. 139, 59–74 (2016).
Bandres-Ciga, S., Diez-Fairen, M., Kim, J. J. & Singleton, A. B. Genetics of Parkinson’s disease: an introspection of its journey towards precision medicine. Neurobiol. Dis. 137, 104782 (2020).
Funayama, M. et al. An LRRK2 mutation as a cause for the parkinsonism in the original PARK8 family. Ann. Neurol. 57, 918–921 (2005).
Paisan-Ruiz, C. et al. Cloning of the gene containing mutations that cause PARK8-linked Parkinson’s disease. Neuron 44, 595–600 (2004).
Zimprich, A. et al. Mutations in LRRK2 cause autosomal-dominant parkinsonism with pleomorphic pathology. Neuron 44, 601–607 (2004).
Sardi, S. P., Cedarbaum, J. M. & Brundin, P. Targeted therapies for Parkinson’s disease: from genetics to the clinic. Mov. Disord. 33, 684–696 (2018).
Hirsch, E. C. & Hunot, S. Neuroinflammation in Parkinson’s disease: a target for neuroprotection? Lancet Neurol. 8, 382–397 (2009).
McGeer, P. L., Itagaki, S., Boyes, B. E. & McGeer, E. G. Reactive microglia are positive for HLA-DR in the substantia nigra of Parkinson’s and Alzheimer’s disease brains. Neurology 38, 1285–1291 (1988).
Damier, P., Hirsch, E. C., Zhang, P., Agid, Y. & Javoy-Agid, F. Glutathione peroxidase, glial cells and Parkinson’s disease. Neuroscience 52, 1–6 (1993).
Brochard, V. et al. Infiltration of CD4+ lymphocytes into the brain contributes to neurodegeneration in a mouse model of Parkinson disease. J. Clin. Invest. 119, 182–192 (2009).
Garretti, F., Agalliu, D., Lindestam Arlehamn, C. S., Sette, A. & Sulzer, D. Autoimmunity in Parkinson’s disease: the role of α-synuclein-specific T cells. Front. Immunol. 10, 303 (2019).
Akil, E. et al. The increase of carcinoembryonic antigen (CEA), high-sensitivity C-reactive protein, and neutrophil/lymphocyte ratio in Parkinson’s disease. Neurol. Sci. 36, 423–428 (2015).
Jin, H., Gu, H. Y., Mao, C. J., Chen, J. & Liu, C. F. Association of inflammatory factors and aging in Parkinson’s disease. Neurosci. Lett. 736, 135259 (2020).
Byron, S. A., Van Keuren-Jensen, K. R., Engelthaler, D. M., Carpten, J. D. & Craig, D. W. Translating RNA sequencing into clinical diagnostics: opportunities and challenges. Nat. Rev. Genet. 17, 257–271 (2016).
Burgos, K. et al. Profiles of extracellular miRNA in cerebrospinal fluid and serum from patients with Alzheimer’s and Parkinson’s diseases correlate with disease status and features of pathology. PLoS ONE 9, e94839 (2014).
Borrageiro, G., Haylett, W., Seedat, S., Kuivaniemi, H. & Bardien, S. A review of genome-wide transcriptomics studies in Parkinson’s disease. Eur. J. Neurosci. 47, 1–16 (2018).
Nalls, M. A. et al. Identification of novel risk loci, causal insights, and heritable risk for Parkinson’s disease: a meta-analysis of genome-wide association studies. Lancet Neurol. 18, 1091–1102 (2019).
Uhlen, M. et al. A genome-wide transcriptomic analysis of protein-coding genes in human blood cells. Science 366, eaax9198 (2019).
Scherzer, C. R. et al. Molecular markers of early Parkinson’s disease based on gene expression in blood. Proc. Natl Acad. Sci. USA 104, 955–960 (2007).
Soreq, L. et al. Long non-coding RNA and alternative splicing modulations in Parkinson’s leukocytes identified by RNA sequencing. PLoS Comput. Biol. 10, e1003517 (2014).
Valentine, M. N. Z. et al. Multi-year whole-blood transcriptome data for the study of onset and progression of Parkinson’s disease. Sci. Data 6, 20 (2019).
Parnetti, L. et al. CSF and blood biomarkers for Parkinson’s disease. Lancet Neurol. 18, 573–586 (2019).
Wang, Y. & Wang, Z. An integrated network analysis of mRNA and gene expression profiles in Parkinson’s disease. Med. Sci. Monit. 26, e920846 (2020).
Amor, S. et al. Inflammation in neurodegenerative diseases—an update. Immunology 142, 151–166 (2014).
Prinz, M. & Priller, J. The role of peripheral immune cells in the CNS in steady state and disease. Nat. Neurosci. 20, 136–144 (2017).
Haghshomar, M. et al. White matter changes correlates of peripheral neuroinflammation in patients with Parkinson’s disease. Neuroscience 403, 70–78 (2019).
Gao, A. Identification of blood-based biomarkers for early stage Parkinson’s disease. Preprint at medRxiv https://doi.org/10.1101/2020.10.22.20217893 (2020).
Courtney, E., Kornfeld, S., Janitz, K. & Janitz, M. Transcriptome profiling in neurodegenerative disease. J. Neurosci. Methods 193, 189–202 (2010).
Grunblatt, E. et al. Gene expression profiling of parkinsonian substantia nigra pars compacta; alterations in ubiquitin–proteasome, heat shock protein, iron and oxidative stress regulated proteins, cell adhesion/cellular matrix and vesicle trafficking genes. J. Neural Transm. 111, 1543–1573 (2004).
Calligaris, R. et al. Blood transcriptomics of drug-naive sporadic Parkinson’s disease patients. BMC Genomics 16, 876 (2015).
Kedmi, M., Bar-Shira, A., Gurevich, T., Giladi, N. & Orr-Urtreger, A. Decreased expression of B cell related genes in leukocytes of women with Parkinson’s disease. Mol. Neurodegener. 6, 66 (2011).
Locascio, J. J. et al. Association between α-synuclein blood transcripts and early, neuroimaging-supported Parkinson’s disease. Brain 138, 2659–2671 (2015).
Mollenhauer, B. et al. α-synuclein and tau concentrations in cerebrospinal fluid of patients presenting with parkinsonism: a cohort study. Lancet Neurol. 10, 230–240 (2011).
Jellinger, K. A. Synuclein deposition and non-motor symptoms in Parkinson disease. J. Neurol. Sci. 310, 107–111 (2011).
Cai, C. et al. Is human blood a good surrogate for brain tissue in transcriptional studies? BMC Genomics 11, 589 (2010).
Sullivan, P. F., Fan, C. & Perou, C. M. Evaluating the comparability of gene expression in blood and brain. Am. J. Med. Genet. B Neuropsychiatr. Genet. 141B, 261–268 (2006).
Kern, F. et al. Deep sequencing of sncRNAs reveals hallmarks and regulatory modules of the transcriptome during Parkinson’s disease progression. Nat. Aging 1, 309–322 (2021).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Frankish, A. et al. GENCODE reference annotation for the human and mouse genomes. Nucleic Acids Res. 47, D766–D773 (2019).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Li, H. A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 27, 2987–2993 (2011).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Hoffman, G. E. & Schadt, E. E. variancePartition: interpreting drivers of variation in complex gene expression studies. BMC Bioinformatics 17, 483 (2016).
Law, C. W. et al. RNA-seq analysis is easy as 1-2-3 with limma, Glimma and edgeR. F1000Res 5, 1408 (2016).
Law, C. W., Chen, Y., Shi, W. & Smyth, G. K. voom: precision weights unlock linear model analysis tools for RNA-seq read counts. Genome Biol. 15, R29 (2014).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Prufer, K. et al. FUNC: a package for detecting significant associations between gene sets and ontological annotations. BMC Bioinformatics 8, 41 (2007).
Manjang, K., Tripathi, S., Yli-Harja, O., Dehmer, M. & Emmert-Streib, F. Graph-based exploitation of gene ontology using GOxploreR for scrutinizing biological significance. Sci. Rep. 10, 16672 (2020).
Gentleman, R. C. et al. Bioconductor: open software development for computational biology and bioinformatics. Genome Biol. 5, R80 (2004).
Acknowledgements
This research was supported and funded by the MJFF under grant numbers 12749 (K.V.K.-J.), 12749.01 (K.V.K.-J., M.R.C.) and 14696 (K.V.K.-J., A.K., D.W.C.). Many MJFF staff assisted in harmonizing and transferring data. This research was supported in part by the Intramural Research Program of the National Institutes of Health, National Institute on Aging. Data used in the preparation of this article were obtained from the PPMI database (https://www.ppmi-info.org/data). For up-to-date information on the study, visit https://www.ppmi-info.org/. We also acknowledge industry partners of the PPMI: AbbVie, Allergan, Amathus Therapeutics, Avid Radiopharmaceuticals, Biogen, BioLegend, Bristol Myers Squibb, Celgene, Denali, GE Healthcare, Genentech, GlaxoSmithKline, Golub Capital, Handl Therapeutics, Insitro, Janssen Neuroscience, Lilly, Lundbeck, Merck, Meso Scale Discovery, Neurocrine Biosciences, Pfizer, Piramal, Prevail Therapeutics, Roche, Sanofi Genzyme, Servier, Takeda, Teva, UCB, Verily and Voyager Therapeutics. We also thank the AMP PD for allowing us access to PDBP data. The PDBP consortium is supported by the National Institute of Neurological Disorders and Stroke at the National Institutes of Health. A full list of PDBP investigators can be found at https://pdbp.ninds.nih.gov/policy. PDBP investigators have not participated in reviewing the data analysis or content of the text. Data used in the preparation of this article were obtained from the AMP PD Knowledge Platform. For up-to-date information on the study, visit https://www.amp-pd.org. The AMP PD, a public–private partnership, is managed by the FNIH and funded by Celgene, GlaxoSmithKline, the MJFF, the National Institute of Neurological Disorders and Stroke, Pfizer, Sanofi and Verily. We thank all people with PD and families for participating in the study and donating their samples and time. We would like to thank the group at IU for RNA isolation, QC metrics and safe shipment of samples. We thank the group at the HAIB for running sequencing and sample tracking.
Author information
Authors and Affiliations
Consortia
Contributions
D.W.C., E.H., I.V., E.A., J.R.G., T.G.W., B.C., A.R., S.H., M.F., A.K., M.R.C. and K.V.K.-J. each contributed to the overall study design. D.W.C., E.H., I.V., E.A., S.K., F.K., T. Fehlman, A.K., M.R.C. and K.V.K.-J. each contributed to primary data analysis, including statistical evaluation. D.W.C., E.H., I.V., E.A., K.L.C. and A.W.T. contributed to data organization and data storage. T. Foroud, S.L. and M.R. each contributed to nucleic acid isolation and sequencing. D.W.C., E.H., T.G.W., M.R.C. and K.V.K.-J. each substantially contributed to writing the manuscript. All authors contributed to data interpretation and review and editing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
B.C., S.H., A.R. and M.F. are employees of the MJFF. E.H., D.W.C., I.V., E.A., K.L.C., A.W.T. and K.V.K.-J. are funded by the MJFF. The other authors declare no competing interests.
Additional information
Peer review information Nature Aging thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
PPMI author list and Supplementary Methods, Figs. 1–10 and Table 2.
Supplementary Tables
Supplementary Tables 1 and 3–11.
Rights and permissions
About this article
Cite this article
Craig, D.W., Hutchins, E., Violich, I. et al. RNA sequencing of whole blood reveals early alterations in immune cells and gene expression in Parkinson’s disease. Nat Aging 1, 734–747 (2021). https://doi.org/10.1038/s43587-021-00088-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43587-021-00088-6
This article is cited by
-
Prodromal Parkinson disease signs are predicted by a whole-blood inflammatory transcriptional signature in young Pink1−/− rats
BMC Neuroscience (2024)
-
Systematic analysis of multi-omics data reveals component-specific blood-based biomarkers for Parkinson’s disease
Translational Medicine Communications (2024)
-
Experimental capture of miRNA targetomes: disease-specific 3′UTR library-based miRNA targetomics for Parkinson’s disease
Experimental & Molecular Medicine (2024)
-
Early-stage idiopathic Parkinson’s disease is associated with reduced circular RNA expression
npj Parkinson's Disease (2024)
-
Extraction of high quality and high yield RNA from frozen EDTA blood
Scientific Reports (2024)