Abstract
Sea ice is a critical component of the Earth’s Climate System and a unique habitat. Sea-ice changes prior to the satellite era are poorly documented, and proxy methods are needed to constrain its past variability. Here, we demonstrate the potential of sedimentary DNA from Polarella glacialis, a sea-ice microalga, for tracing past sea-ice conditions. We quantified P. glacialis DNA (targeting the nuclear ribosomal ITS1 region) in Arctic marine and fjord surface sediments and a sediment core from northern Baffin Bay spanning 12,000 years. Sea ice and sediment trap samples confirmed that cysts of P. glacialis are common in first-year sea ice and sinking particulate matter following sea-ice melt. Its detection is more efficient with our molecular approach than standard micropaleontological methods. Given that the species inhabits coastal and marine environments in the Arctic and Antarctic, P. glacialis DNA has the potential to become a useful tool for circum-polar sea-ice reconstructions.
Similar content being viewed by others
Introduction
The polar regions are warming at a rate unprecedented in millennia, leading to profound changes in the cryosphere. First-year and multiyear sea ice in the Arctic are decreasing in extent, volume, and thickness, and multiyear sea ice is projected to disappear within this century under the current climate trajectory1. Sea ice is a key component of the cryosphere, regulating large-scale heat and gas exchange between the ocean and the atmosphere, and major oceanic currents2,3. To increase our confidence in sea-ice projections, it is important to have long-term data that can capture climate variability and contribute to model development and validation. However, long-term records of sea ice are scarce, and sea ice remains one of the most poorly understood elements of the Earth System. Sea ice leaves little evidence in the geological record, limiting our understanding of its natural variability and the magnitude of past changes.
The palaeoceanography community has for a long time explored ways to infer past sea-ice conditions from proxies preserved in geological records. These include records from marine sediments4 and ice cores (methanesulfonic acid and bromine)5,6, alongside other indicators such as driftwood7 and whale macrofossils8. The most commonly used sea-ice indicators in marine sediments include microfossils that preserve due to their silica frustules (e.g., diatoms), calcium carbonate tests (e.g., foraminifera) or organic-walled cysts (e.g., dinoflagellate cysts), and biogeochemical tracers such as highly-branched isoprenoid (HBI) biomarkers9 and alkenones from Isochrysidales10. Two microfossil groups have been extensively used to infer past sea-ice conditions: dinoflagellate cysts and diatoms. Past assemblages have been numerically compared to modern reference datasets, and quantitative reconstructions proposed based on transfer functions (diatoms) and the modern analogue technique (MAT, dinocysts)4. However, the reliability of these inference methods is limited by incomplete information on specific habitats and seasonal niches of sea-ice-associated species, limited knowledge of life cycles (e.g., unknown resting-vegetative stage relationships), and microfossil preservation biases4. In addition, the most widely used Arctic sea-ice indicators in microfossil records (such as the dinoflagellate cyst taxon Islandinium minutum and diatom Fragilariopsis oceanica) are not specifically sea ice-dependent and often bloom in cold and stratified waters that can be related to sea ice or glacial meltwater11. Frustules of sympagic diatoms are rarely preserved in the sedimentary record. However, some species synthesise characteristic HBI compounds that can be well preserved and stable in marine sediments12,13. The HBI compound IP25 (i.e., ice proxy with 25 carbon atoms) is considered a reliable proxy for seasonal sea ice in the Arctic, and semi-quantitative approaches combining IP25 with sterols or other HBIs are widely used9,14,15. However, there are still several challenges; the full range of biotic and abiotic factors that regulate HBI synthesis is poorly understood16, and IP25 concentrations in sediments appear to be reduced in fjord systems and coastal areas influenced by glacial/terrestrial runoff17,18,19,20,21. Even though IP25-producers have been found in sea ice from fjord environments, their abundance, growth and/or HBI synthesis appears to be inhibited, possibly due to reduced surface salinity and sea-ice microstructure affecting species composition or ecophysiology (e.g.18). These limitations highlight the need for exploration and development of additional sea-ice proxies.
Recent studies have shown that ancient DNA from diverse organisms can be preserved in marine sediments for at least up to one million years22 and that specific gene markers can be used to detect taxa beyond what is possible using traditional methods22,23,24,25,26. Two recent studies in the Greenland Sea and the Fram Strait investigated the potential of using metabarcoding of marine sedaDNA for sea-ice reconstructions. Both studies showed that DNA from a variety of organisms is preserved in marine sediment cores, and targeting a hypervariable region of the 18 S rRNA gene allowed the detection of high diversity across protists and other groups23,24. These included characteristic sea-ice groups such as diatom predators associated with brine communities27. Most sedimentary ancient DNA studies have focused on describing past environments and changes in community composition over time by using metabarcoding23,24 and shotgun sequencing approaches22,28. An alternative approach is to use droplet digital Polymerase Chain Reaction (ddPCR), which can provide sensitive quantification of DNA targets from complex environmental samples29. This approach has the potential for proxy development in the field of paleoclimate and paleoceanography, as it allows for the quantification of specific DNA markers that target taxa or genes indicative of particular environmental conditions.
The sea-ice dinoflagellate Polarella glacialis M. Montresor, G. Procaccini and D.K. Stoecker, is one of few microalgal species with a bi-polar distribution30,31. P. glacialis has two morphologically different life-cycle stages: a spiny, non-motile resting cyst (~14 µm long, 10 µm wide) and a slightly smaller motile vegetative cell stage (~11 µm long; 7 µm wide)32 (Fig. 1a, Supplementary Fig. 1, Supplementary Video 1); both stages have been found in first-year Arctic and Antarctic sea ice33,34. In Antarctica, encystment was reported to occur following nitrate depletion, shortly before sea-ice melt35. The Antarctic sea-ice isolates possess sufficient genomic plasticity to adjust to both hyposaline and hypersaline conditions36, and ecotypes of P. glacialis have colonised Antarctic saline lakes37. P. glacialis is also resilient to rapid temperature changes with a suite of coding genes38,39, consistent with survival in sea ice. In the Arctic, P. glacialis is found in the sea-ice matrix and sea-surface waters influenced by ice33,40,41,42,43. This small dinoflagellate is also reported from seasonally sea-ice-covered sub-Arctic seas44,45,46, but its presence has never been confirmed in areas without seasonal sea-ice occurrence. These observations suggest that P. glacialis is a widely distributed sea-ice microalgae. However, it is not often reported in microplankton surveys and its cysts have been only sporadically found in surface sediments47,48,49 and in sediment records50,51,52. The species may have been overlooked in both types of surveys due to sample processing protocols (e.g. sieve mesh-size, acid treatment for palynological analyses) and difficulties in separating P. glacialis from other small dinoflagellates using standard light microscopy. The species is not included in the dinocyst surface sample assemblage dataset that is used for (semi-) quantitative reconstructions of past sea-surface conditions using the MAT4,53.
a The two life-cycle stages of P. glacialis: vegetative cell and resting cysts. The scale bar is 10 µm. b Spatial distribution of different sample types where P. glacialis cells, cysts or DNA were detected: white circles represent sea-ice samples (cysts), blue circles represent water samples (vegetative cells and cysts), black circles represent sediment traps (cysts), green circles represent surface sediments (DNA), and orange circles sediment cores (DNA and cysts). High-resolution sampling was conducted around Greenland and in northern and central Baffin Bay. Sample sites include three Arctic fjords: Inglefield Bredning (IB), Independence Fjord (IF) and Young Sound (YS); two polynyas: North-East Water polynya (NEW) and North Water polynya (Pikialasorsuaq) (NOW); marine coastal and offshore sites near Qikiqtarjuaq (Q) and Qeqertarsuup Tunua (QT); central Baffin Bay and the Greenland Sea. The ice camp in Q and oceanographic campaign in QT include more than one sample type at the same location. Map created using General Bathymetric Chart of the Oceans (GEBCO)2019 grid86. Topographic data is derived from Esri Data and Maps for ArcGIS and Ice cap and glacier data from87,88.
P. glacialis was one of the first free-living extant taxa of the order Suessiales to be described and is phylogenetically distant from its closest relatives30,32,54. Compared to genome sizes of many dinoflagellates, with haploid genomes estimated to be up to 250 Gbp (Giga base pairs)55,56, P. glacialis has a relatively small diploid genome of just under 3 Gbp38 Despite its relatively small size and genome, P. glacialis DNA was detected in sediments dating back to ca. 33,700 calibrated years before present (cal yr BP) from the Greenland Sea23 and a late-Holocene sediment core record from an east Antarctic fjord26. This suggests that the species is a promising target for the development of a DNA-based paleo proxy. To further explore the potential of P. glacialis DNA as a sea-ice proxy, the following criteria and hypotheses were investigated: Specificity: P. glacialis is more common in sea-ice environments than previously reported, and its distribution in surface sediments reflects the presence of first-year sea-ice. Because it can tolerate a broad range of salinities, P. glacialis can be used to trace past sea-ice conditions in both marine systems and fjords and coastal areas influenced by freshwater runoff. Sensitivity and Stability: Species-specific gene markers are suitable to detect and quantify P. glacialis in marine sediments based on ddPCR, and P. glacialis DNA is sufficiently stable in sedimentary records to be quantifiable for past sea-ice reconstructions. We addressed these hypotheses by examining sea-ice cores, sea ice-loaded waters, sinking particulate matter collected using sediment traps, surface sediments and marine sediment cores from the Arctic. Targeting the nuclear ribosomal ITS1 gene region, we quantified P. glacialis DNA (Pgla-DNA) in surface sediments. The Pgla-DNA data were compared with sea-ice concentration data derived from remote sensing and in situ observations around Greenland. Furthermore, we examined the detection of Pgla-DNA compared to microscopy, following a standard micropaleontological approach, in a sediment core spanning the last ca. 12,000 years.
Results
We detected cysts, vegetative cells, and DNA of P. glacialis in a total of 181 samples from the Arctic region, including first-year sea ice, ice-loaded surface waters, sinking particulate matter captured by sediment traps, surface marine sediments, and sediment core samples (Fig. 1).
P. glacialis in Arctic sea ice
We observed cysts of P. glacialis in sea-ice samples from the Central Arctic Ocean, southwest and central Baffin Bay, as well as Qeqertarsuup Tunua (Disko Bay, central West Greenland) (Fig. 1b). While there was a large variation in the number of cysts detected and their relative abundance in relation to other species, P. glacialis was present in all analysed first-year sea-ice cores from the Central Arctic Ocean (relative abundances 6–82% of the total cell counts). The species was absent from the only multiyear sea-ice core that we analysed (Supplementary Table 1). At the ice camp south of Qikiqtarjuaq, southwest Baffin Bay (Fig. 1b), cysts were detected at low abundances by the Imaging Flow Cytobot system in sea-ice samples and ice-loaded surface waters from beneath the sea-ice coring sites (Supplementary Fig. 1), whereas in central Baffin Bay cysts were found in most sea-ice samples analysed (Supplementary Table 1). In Qeqertarsuup Tunua, P. glacialis cysts and vegetative cells (identified using light microscopy) were detected in every sea-ice sample collected in winter and spring. Cysts and vegetative cells were also identified using light microscopy in both ice-loaded surface waters and at the depth of the chlorophyll-a maximum in Qeqertarsuup Tunua. Notably, P. glacialis was not detected in water samples from Qeqertarsuup Tunua when the sea surface was ice-free, nor was it observed in the subsequent spring bloom. The 2011 spring bloom in Qeqertarsuup Tunua began in late April with increasing chlorophyll-a concentrations until late May57 (Supplementary Table 1).
Cysts of P. glacialis in sinking particulate matter
To assess the temporal and spatial occurrence of P. glacialis in sinking particulate matter originating from sea-ice environments, we investigated circum-Arctic sediment trap data from deployments over periods of a few weeks to 12 months and spanning about a decade (2005–2016); locations are given in Fig. 2a. Cysts were observed in annual sediment trap deployments at Arctic sites characterised by seasonal sea-ice cover. All traps captured cysts of P. glacialis during and after sea ice melt, typically from June to September (Fig. 2b). In the Laptev Sea and Amundsen Gulf, some cysts were also found during sea-ice formation (Fig. 2a). Additional deployments at high temporal resolution collected sinking particles using drifting sediment traps in central Qeqertarsuup Tunua (Fig. 3). The study areas had been covered by sea ice for ~7 months (central Baffin Bay) and ~3 months (Qeqertarsuup Tunua) prior to trap deployment. In central Baffin Bay, the sediment trap was deployed at 25 m depth, attached to a drifting sea-ice floe. Cysts of P. glacialis were absent at first but were continuously collected following the onset of sea-ice breakup (Fig. 3b–d). Cysts were also collected by drifting sediment traps simultaneously deployed at a depth of 50 m and 100 m in Qeqertarsuup Tunua when the sea ice started to melt but were absent both before (end-March) and after the ice melted (May) (Fig. 3e–g).
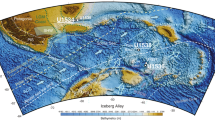
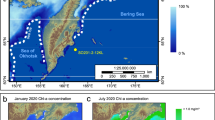
a The sediment trap location (black circles) in Rijpfjorden (Svalbard), Laptev Sea, Baffin Bay, and Beaufort Sea (Amundsen Gulf and Mackenzie Shelf), map created by ggOceanMaps89. b Timeline and characteristics of annual trap deployments (green quadrates indicate detection of P. glacialis, white quadrates no detection, double forward slash indicate the timing of sea-ice formation and backward slash sea-ice melt).
a Location of the drifting sediment traps in central Baffin Bay (CBB) and Qeqertarsuup Tunua (QT). Circles show the southward drift of the trap from 15 June to 7 July 2016 in CBB. b Fluxes of P. glacialis cysts at 25 m water-depth in CBB; the arrows point to the date of the sea-ice cover satellite image, and the empty circles denote samples counted without detection. c Satellite image retrieved on 17 June 2016 representing maximum sea-ice cover and d satellite image retrieved 5 July 2016 during sea-ice melt. e Fluxes of P. glacialis cysts at 50 m and 100 m water-depth in QT. f Satellite image retrieved on 28 March 2011 representing maximum sea-ice cover and g satellite image retrieved 24 April 2011 during sea-ice breakup. The scale bars on c–f represent 50 km. We acknowledge the use of imagery from the NASA Worldview Snapshots application (https://wvs.earthdata.nasa.gov), part of the Earth Observing System Data and Information System (EOSDIS).
Spatial and temporal distribution of Pgla-DNA in surface sediments
We successfully detected and quantified Pgla-DNA in 45 out of 48 surface sediment samples. The three sampling locations where P. glacialis was not detected include the ice-free waters of the eastern Greenland Sea (two sites) and one site near the permanently ice-covered area on the North-East Greenland shelf Greenland shelf bordering the North-East Water polynya.
The sampling locations included both marine and fjord sea-ice environments (Fig. 4a, Supplementary Fig. 2). The assay was carried out using ddPCR (see “Materials and methods” section), targeting a partial gene fragment from the nuclear ribosomal ITS1 region of P. glacialis. The concentrations of Pgla-DNA ranged from 3000 to 437,000 Pgla-DNA ITS1 gene copies g−1 (ITS1 gc g−1). Maximum concentrations of Pgla-DNA were found in Inglefield Bredning, adjacent to the North Water (Pikialasorsuaq), an area of intense sea-ice formation, with average annual sea-ice concentrations of 75–80%. There were no significant differences in the concentrations of Pgla-DNA between fjord and marine environments (marine locations n = 33, fjords n = 14, t-test p = 0.4). For the sites with up to 80% annual sea-ice cover, there was a positive linear relationship between the sea-ice concentration and Pgla-DNA (n = 29) with r2 of 0.4 and an improved fit with r2 of 0.6 using a polynomial cubic correlation (k = 2) (Fig. 4b, c). At locations with 80–100% sea-ice concentration (n = 19), the relationship was negative, with a linear regression r2 of 0.5 and polynomial cubic regression (k = 2) r2 of 0.7 (Fig. 4d, e).
a Bubble plot (green circles) of P. glacialis DNA concentration in surface sediments. The sites include three fjords—Inglefield Bredning (IB), Independence Fjord system (IF) and Young Sound (YS)—the North Water polynya (Pikialasorsuaq) (NOW) and the North-East Water polynya (NEW) (indicated by dashed white lines), and marine sites on the east coast of Greenland. ArcGIS was used to create the map. Topography and bathymetry are from IBCAO: https://www.gebco.net/about_us/committees_and_groups/scrum/ibcao/ibcao_v4.html. Ice coverage around Greenland is from90. Ice coverage elsewhere is based on the Randolph Glacier Inventory 6.0: https://www.glims.org/RGI/rgi60_dl.html. b–e The correlation between Pgla-DNA in surface sediment and sea-ice concentration (average of 6 years prior to sampling) in areas with up to 80% of sea-ice cover using b linear regression and c with improved fit using polynomial cubic regression (k = 2), and in areas with more than 80% sea-ice cover using d linear regression and e polynomial cubic regression (k = 2). The dashed lines denote the 95% confidence interval for the regression.
Detection of P. glacialis in Holocene sediment records (Pgla-DNA vs. cyst counts)
Pgla-DNA was detected and quantified in all samples analysed from a sediment core retrieved in northern Baffin Bay (Fig. 5a). The 599 cm-long core AMD16-117Q spans the last 12,000 ± 600 cal yr BP58. The total DNA yield (25–525 ng DNA g−1) was variable throughout the core and generally decreased with age (Fig. 5b). In core AMD16-117Q, the ddPCR assay detected between ~100 and 33,000 Pgla-DNA ITS1 gc g−1 (n = 16), with the highest quantities detected at 116.5 cm core depth, dated to ca. 2000 cal yr BP (Fig. 5c).
a Location of core AMD16-117Q (orange circle), map created by ggOceanMaps88. b Total DNA concentration given as loge. (black bars). c Concentration of ITS1 gene copies of P. glacialis given as loge. (green bars). d P. glacialis cyst concentrations determined by micropaleontological analyses (green bars, empty circles are samples for which standard assemblage counts detected no P. glacialis cysts. e Total dinoflagellate cyst concentrations (grey bars). f Relative abundance of Islandinium minutum, a cold-water taxon often associated with seasonal sea ice (brown bars). g Sea-ice concentration reconstruction based on dinocyst assemblages (excluding P. glacialis) following the Modern Analogue Technique (MAT) (black line median, red and blue lines minimum and maximum respectively). h Concentration of the sea-ice biomarker IP25 (black bars) as presented in58 normalised to total organic carbon (TOC). The unit for the y-axis is calibrated years before present (cal yr BP).
Dinoflagellate cyst assemblage analyses were performed for the sediment core AMD16-117Q at an average resolution of 8 cm (n = 69) following standard palynological procedures and using room-temperature HCl and HF acids. Cyst counts revealed that P. glacialis cysts were present but generally rare in the Holocene sediments examined, never exceeding 30 cysts g−1 (Fig. 5d) and never accounting for more than 1.2% of the total dinoflagellate cyst assemblages. In core AMD16-117Q, P. glacialis cysts were present in the top section dated to <1000 cal yr BP, an interval with mean dinocyst concentrations of ~10,000 ± 3800 cysts g−1 (Fig. 5e). Cysts were also present toward the bottom of the core dated to ca. 9000–11,000 cal yr BP, where the mean total cyst concentrations were ~1800 ± 900 cysts g−1. Notably, P. glacialis cysts were not found in the middle part of the core, where total cysts concentrations were the highest for the entire studied period (i.e., ~17,000 ± 5300 cysts g−1; Fig. 5e). A single cyst of P. glacialis was observed in two of the four samples for which counting efforts were increased tenfold (i.e., counting a minimum of 3000 cysts per sample). We estimated the detection limit in our samples to be on average 23 ± 17 cysts g−1 (n = 10) (meaning that if P. glacialis cyst concentrations were lower, the species would not have been detected following the standard micropaleontological approach, which typically involves counting 300 specimens per sample). In contrast, Pgla-DNA was successfully detected and quantified by the ddPCR assay in all samples. For comparison, the relative abundance of the cold water and seasonal sea-ice-associated dinoflagellate cyst Islandinium minutum is shown in Fig. 5f, alongside sea-ice concentration estimates based on the MAT using the modern dinocyst reference dataset n = 196853, which does not include P. glacialis cysts (Fig. 5g). The sea-ice biomarker IP25 is shown in Fig. 5h.
Throughout the core, the observed cysts showed no signs of cell wall degradation and cellular content was often visible (Fig. 6a). During tests conducted separately on marine and fjord sediments from the same region, we found that the cysts were prone to degradation if treated with warm HF during palynological preparation (Fig. 6b). This could partly explain the absence of P. glacialis cysts in previous studies that used heated acids during sediment processing including those that were integrated in existing modern reference databases53.
a Micrographs from core AMD16-117Q at: (1) 112–113 cm core depth, (2) 317–318 cm core depth, and (3) 494-495 cm core depth. Scale bars = 10 μm. Cysts were preserved throughout the core and some cysts had cell content (1 and 3) as far back as ca. 10,000 years. b Micrographs of P. glacialis cysts from arctic sediments (northern Baffin Bay and North-East Greenland) showing 1–3) intact cysts and 4–14) degradation of cysts following acid treatment using warm (50 °C) hydrofluoric acid (instead of room-temperature acid) during palynological processing.
Though the primers have been validated for species specificity on culture material23, we further tested if the primers would amplify off-target in eDNA samples from both surface sediments and the AMD16-117Q sediment core. In all samples, the sequence reads best matched references of P. glacialis in the National Center for Biotechnology Information (NCBI), with variants accounting for 0.02–0.04% of the total reads (Supplementary Fig. 3).
Discussion
We set out to explore the potential of P. glacialis as a sea-ice proxy. Its occurrence in Baffin Bay and around Greenland agrees with a specific first-year sea-ice distribution and adds to a recent compilation of >100 first-year and multiyear sea-ice cores from other Arctic locations42. Though its abundance was highly variable, we show that P. glacialis is common in Arctic first-year sea ice, similar to results previously reported in landfast ice off Antarctica59. In contrast to its wide distribution and common presence in first-year sea ice, P. glacialis was not detected in the multiyear sea ice core we analysed. Although sampling was limited, we speculate that the absence could be due to differences in ice properties such as brine channel structure, light limitation in the thicker multiyear ice, or due to life cycle requirements of P. glacialis, where cysts are released from melting ice, and vegetative cells recolonise new ice during freeze-up.
Though little is known about the life-cycle of sympagic microalgae in general, an important life-cycle transition has been reported for P. glacialis connected to first-year sea ice, where vegetative cells reproduce in early spring and cysts form in late spring following nitrate depletion35,60. Resting stages (cysts) are considered key life-cycle traits for sympagic microalgae, but characteristics of this life-cycle strategy in sea ice are generally poorly understood61. Observations of high concentrations of P. glacialis cysts in upper sections of first-year and second-year sea ice in late winter and early spring suggest that the species overwinters in the sea ice33,34,62. Our results indicate that, during and following sea-ice melt, cysts sink through the water column and towards the seafloor. This is coherent with the absence or rarity of P. glacialis in plankton surveys63,64,65. Marine metabarcoding studies using universal primers43 did not detect P. glacialis in water samples during spring/summer in northern Baffin Bay64 or Nares Strait63. These sites are close to where we found cysts in sediment traps, and both resting cysts and sedaDNA in sediments. Similarly, the large-scale marine sequencing effort of Tara Oceans retrieved no P. glacialis sequences38. However, we also noted that P. glacialis cysts were found in sediment traps at depths ranging from 75 to 200 m several weeks after sea ice melted. This could indicate slower vertical export of cysts under strong water-column stratification during summer, advection from surrounding regions, or resuspension. We found resting cysts of P. glacialis with intact cell content in surface sediments, indicating viability and potential for germination. While other studies have shown that newly-formed sea ice is seeded with vegetative cells of P. glacialis33, it remains to be verified to what extent these may originate from a benthic seed bank, a common strategy for coastal cyst-forming dinoflagellates66.
We hypothesised that because P. glacialis shows phenotypic plasticity and can adapt to changes in salinity and temperature36,39, it is able to thrive in both marine and fjord environments and thus may be a useful proxy for different sea-ice environments. Our results show that Pgla-DNA traces first-year sea ice over a range of settings, including high-Arctic fjord systems, coastal marine and polynya sites, as well as relatively low productivity sites on the North-East Greenland shelf. The positive correlation between sea-ice concentrations from 20 to 80% and Pgla-DNA gene copies in surface sediments reflects higher P. glacialis abundances with increasing sea-ice concentration (up to 80%). This is coherent with seasonal observations, indicating that P. glacialis cysts and vegetative cells are among the first microalgae to colonise young ice, and that their biomass increases as the sea-ice season progresses33,34,49,61,62. Though a linear relationship would not be expected, as there are multiple factors that control algal growth within the sea ice, the trend is significant (Fig. 4). At locations with more than 80% sea-ice cover, such as in North-East Greenland, Pgla-DNA was still detected but at relatively low concentrations. This might be related to these being generally very low productive environments, with limited availability of light (due to thicker sea ice) and generally low nutrient levels49.
To assess whether Pgla-DNA is stable and detectable on millennial timescales, we analysed sediment core samples from northern Baffin Bay spanning the Holocene epoch (~12,000 years). This area is known to have been seasonally covered by sea ice throughout the Holocene58,67,68. We detected Pgla-DNA in all the samples analysed from sediment core AMD16-117Q (Fig. 5). While a sea-ice reconstruction based on Pgla-DNA is beyond the scope of this study, we note that the number of P. glacialis ITS1 gene copies was highest in the most recent samples, reflecting the overall trend of increased sea-ice cover during the neoglacial period (ca. past 3000 years). This general trend is captured by the sea-ice biomarker IP25, the relative abundance of I. minutum cysts, and sea-ice concentration estimates based on the MAT on dinoflagellate cyst assemblage data. When comparing the detection of P. glacialis by Pgla-DNA analyses with classical micropaleontological methods (cyst counts), we conclude that sedaDNA improves detection of this species. The cysts we observed showed no signs of degradation (after palynological treatment using room temperature HF). Although P. glacialis cysts were found in sediments older than 9,000 cal yr BP, the species was rare and often below detection limit when using standard census procedures (counting 300 dinocysts per sample). Cysts were often below the detection limit at times of increased total dinocyst production (even when counting efforts were increased 10-fold), such as during the Mid- to Late Holocene, when the P. glacialis signal was diluted by the higher concentrations of other cyst-forming taxa in the sediments. As each cell of P. glacialis contains multiple gene copies of the ITS1 gene38, the molecular method would indeed be expected to be more sensitive. Furthermore, most dinoflagellate cysts are diploid, sexual stages. Although this has not been confirmed for P. glacialis, genomic data for this species strongly indicate the presence of diploidy38. This would mean that each cyst reaching the sediment carries double the amount of gene copies of the ITS1 gene targeted in this study compared to vegetative cells. The recent sequencing of P. glacialis genomes from both polar regions revealed some genetic dissimilarities between the two poles, such as the size of the genome, but not within the ITS gene region38. Cysts have been found in surface sediments in the Lützow-Holm Bay50(East-Antarctica), and P. glacialis inhabits land-fast ice in the Antarctic62 and has also been detected in inland saline lakes37. The apparent circumpolar distribution of this species at locations with first-year sea ice makes Pgla-DNA potentially the first biogenic sea-ice proxy that may be applicable to both polar regions.
The validation of specificity of the ITS1 primers used for ddPCR showed some variation in the 95 bp product in the ITS1 gene. The variation was equivalent to 1 or 2 single nucleotide variants (<2%), which is within the inter-strain divergence of P. glacialis sequences available in NCBI and similar to other Suessiales (e.g.69). We, therefore, conclude that the ITS 1 primers are specific for P. glacialis under these conditions and that the ddPCR results accurately reflect the copy number of amplicons for this species in an environmental sample.
Further studies are needed to explore how far back in time the Pgla-DNA proxy can be applied. Pgla-DNA has been reported from a single sediment sample dated to ca. 33,700 cal yr BP from the Greenland Sea23, but its full stratigraphic extent is currently unknown. We observed cysts with cell content in sediment traps and surface sediments. How long P. glacialis resting cysts can remain viable in sediments is also unknown, although 100-150 years longevity has been reported for other protist resting stages66. Cell content was also present in the cysts observed down-core, back to ca. 10,000 cal yr BP (Fig. 6). Even when resting stages lose their viability, DNA can remain inside structurally intact cyst walls, which likely reduces DNA degradation rates. It is thus reasonable to expect that the DNA of cyst-forming organisms such as P. glacialis will have a better preservation potential in marine sediments compared to non-cyst-forming taxa. Marine sedaDNA is a combination of extracellular DNA (adsorbed to minerals and other particles) and intracellular DNA (inside organisms either dead or alive). The differences in the preservation of extracellular vs. intracellular DNA, as well as DNA degradation rates in marine sediments, are poorly known. Hydrolysis is likely the main process contributing to DNA degradation in sedaDNA, whilst low temperatures such as those of the polar regions, high pH and high ionic strength appear to slow down degradation70,71,72,73. Furthermore, conditions at the water-sediment interface and bioturbation may also affect the long-term preservation of DNA. Ancient DNA sequences are often very short due to fragmentation, and a recent study showed the average base coverage for the 18 S rRNA gene V9 region in all eukaryotes and in diatoms to be around 100 base pairs (bp)72. The average fraction length for cyst-forming dinoflagellates is unknown, but we can assume the product length of the primers (95 bp) is suitable for detection in marine sediments further back in time.
The Pgla-DNA method we suggest here (Supplementary Protocol 1) is a relatively simple approach, permitting rapid preparation and analysis of hundreds of samples to facilitate high-resolution analysis. The laboratory procedures, data processing and costs involve only a fraction of the resources needed for other genomic approaches, such as metabarcoding and shotgun sequencing. Moreover, quantification of gene copies by ddPCR has the advantage of potentially producing an operational input for climate models. However, and as with all proxies, Pgla-DNA also has shortcomings. A known uncertainty, common to all proxies, is that they originate from sea ice and need to be transported to the sediments. Thus, there is a need to carefully consider how advection by currents, resuspension, bioturbation and other sedimentary processes may affect the fidelity of proxy records. Besides concerns with DNA degradation (discussed above), a limitation of the Pgla-DNA approach is that a null result can represent either permanently open waters or perennial sea-ice cover, as with the IP25 biomarker. Biomarker indices (e.g., PIP25) using related biomarkers presumably produced by open-water organisms have been proposed to distinguish multiyear sea-ice cover from open-water conditions74. We suggest that a similar approach can be developed by simultaneously targeting and quantifying DNA from a well-established sea-ice species with a robust genetic record (as demonstrated here for P. glacialis) with species exclusively associated with either permanent sea-ice cover or open water. We recommend carefully choosing sedaDNA markers based on sound knowledge of species ecologies or traits. Alongside new advances in the field of marine sedaDNA, improvement of genomic and taxonomic references, experimental studies on the ecophysiology and life cycles of sympagic species, as well as multi-proxy and proxy-model comparison studies, are needed to contribute to more accurate reconstructions and improve our understanding of past sea-ice changes.
Materials and methods
Sample collection
Sample collection was conducted over spring and summer during various oceanographic campaigns and ice camps that took place from 2005 to 2017 in the Arctic Ocean and adjacent seas. The collection of samples for sea ice and water near Qikiqtarjuaq and in Baffin Bay (between March and July of 2015 and 2016) was part of the GreenEdge project; details of the methods are given in75. Briefly: sea-ice cores were collected (minimum four each year) using a Kovacs Mark 14 cm diameter ice corer; water samples were collected through a hole in the ice using 10 L or 20 L Niskin bottles; samples were collected at the following water depths: 1.5, 5, 10, 20, 40, and 60 m. Sea ice and water sampling in Qeqertarsuup Tunua 2011, from March to May, is described in detail in41. Briefly, when sea ice was present, a block of sea ice was cut out with a chainsaw and removed with a lifting tong; adjacent water was sampled using 10 L Niskin bottles on a hand wire from a small motorboat. After the sea ice melted, Qeqertarsuup Tunua water was collected from the research vessel RV Porsild (Arctic Station, University of Copenhagen) using 10 L Niskin bottles mounted on a rosette system. Sea-ice cores from the Central Arctic Ocean were collected in August-September 2012 as described in76. Briefly, sea ice was sampled using a Kovacs Mark ice corer with a 9 cm diameter. All sea ice samples were processed following direct melting in the dark.
The drifting sediment trap in central Baffin Bay was deployed as part of the GreenEdge project, while sediment traps deployed in the North Water, Beaufort Sea, and Laptev Sea were deployed as part of the ArcticNet Long-Term Oceanic Observatories project. In central Baffin Bay, a sediment trap with 12 sampling cups (Technicap PPS4) was anchored 25 m below sea ice and programmed to rotate to a new sampling cup at 2-day intervals. All sampling cups were filled with filtered seawater and formalin buffered with sodium borate, and the salinity was adjusted to 38 with NaCl to avoid diffusion of content and ensure the preservation of samples during deployment and after recovery. In Qeqertarsuup Tunua, a set of drifting sediment traps with two cups was deployed simultaneously at 50 m and 100 m depth. The traps consisted of a double set of parallel acrylic cylinders mounted on a gimbaled frame (KC Denmark). The traps were filled with 0.2 µm filtered seawater from depths below 200 m with NaCl added to reach a salinity of ~38. The salinity of the surrounding water was ~33 and 34, respectively. No preservative was added as the deployment time of traps was short. Sediment traps deployed in the North Water, Beaufort Sea, and Laptev Sea are described in77. In Rijpfjorden (Svalbard), sediment traps (MCcLane Parflux) with a 21-bottle carousel and a 1 m2 funnel opening were deployed on a single-point mooring at a depth of 60 m (bottom depth of 235 m). The sample bottles were prefilled with filtered seawater adjusted with NaCl to salinity 38. To preserve deposited material, 4 % formalin buffered with sodium borate was added to the traps before deployment.
Surface sediment samples (0–1 cm) were collected during three cruises and one ice campaign (Supplementary Table 1). Sub-sampling for DNA was conducted immediately after collection. To avoid contamination, single-use sterile equipment was used, and the samples were placed into sterile WHIRL-PAK© sampling bags and frozen immediately at either −20 or −80 °C. The samples were freeze-dried in the same sterile sampling bags. During the ice2ice cruise GS15-198, surface sediments from the Greenland Sea were collected immediately from multicores, bagged and kept frozen at −20 °C until analysis.
Core AMD16-117Q (77°00.29’ N, 72°08.32’ W, 963 m water depth, 599 cm total sediment recovery) was retrieved from outside the Inglefield Bredning Fjord (Qaanaaq) during the CCGS Amundsen GreenEdge/ArcticNet 2016 Leg 1a expedition using a Calypso Square giant gravity coring device. Geochronology was determined by combining 15 210Pb measurements with 13 radiocarbon dates measured on biogenic carbonate (mixed benthic foraminifera, planktonic foraminifera Neogloboquadrina pachyderma sin. or bivalve shell fragments) and 4 radiocarbon dates measured on bulk organic carbon. Ages were modelled in the R (R Core Team, 2022) package bacon78. All radiocarbon dates were calibrated using the Marine13 calibration curve79 and an additional local reservoir correction (ΔR) of 140 ± 60 years; for more details on the geochronology see58. Discrete sub-samples were collected for biomarker analyses and sedaDNA on separate U-channels. The sediment at the surface was first scraped off and the freshly exposed sediment was then collected using single-use sterile syringes into sterile WHIRL-PAK© sampling bags and kept frozen until analysis.
Identification and abundance of P. glacialis cysts and vegetative cells
For cyst identification and abundance assessment in sea ice and surrounding water, an Imaging Flow Cytobot system (McLane labs) was used for observations near Qikiqtarjuaq and further north in Baffin Bay. Imaging Flow Cytobot system images are triggered by chlorophyll a fluorescence. For the samples collected in Qeqertarsuup Tunua, the abundance of cells was assessed by Utermöhl sedimentation chambers (Hydro-Bios) using an inverted microscope (Nikon) at 400× magnification. Cells were identified according to their morphology as vegetative cells or cysts, vegetative cells were 9–15 μm long and 5–9 μm wide (n = 19), and cysts were 12–19 μm long and 8–15 μm wide (n = 20). The identification of the vegetative stage was further confirmed by DNA from monocultures established by isolating single vegetative cells out of melted sea-ice samples from Qeqertarsuup Tunua. The cultures were grown at 3 °C in TL30 medium at a light intensity of ~25 µmol photons m−2 s−1 and an 18:6 h light:dark cycle until a density of approximately 4000 cells mL−1. Aliquots of the cultures (10 mL) were centrifuged down to a pellet, and the pellets were frozen for molecular analysis. The microscopic detection of P. glacialis in sea-ice samples from Qeqertarsuup Tunua was partly given in41 but for this study the identification by DNA was assessed by the ITS1 and 2 gene regions using the primers ITS-1F (5’-CTT GGT CAT TTA GAG GAA GTA A-3’) and ITS-4 (5’-TCC TCC GCT TAT TGA TAT GC-3’)80.
The bottom 5–10 cm of the sea-ice cores from the Central Arctic Ocean were left to melt overnight at 4 °C, and the cysts contained in the meltwater were identified using light microscopy (as above). The abundance of cells was assessed as described for the Qeqertarsuup Tunua samples.
To determine microalgal cyst abundance in the sediment traps from central Baffin Bay, each sample cup was gently homogenised before subsampling a volume ranging from 0.5 to 1.0 mL (depending on the quantity of material in the sample cup). A micropipette with a clipped tip was used to ensure the collection of large aggregates. The volume of each subsample was adjusted to 3 mL with filtered seawater in a Utermöhl chamber. After sedimentation, a minimum of 300 microalgal cells were counted and identified to the lowest taxonomic level possible by inverted light microscopy at 100×, 200×, or 400× magnification depending on cell size according to the Utermöhl method. The identification and abundance estimation of cysts in the sediment traps deployed in Qeqertarsuup Tunua was conducted for the sea ice and water at the same location.
DNA extractions
DNA extractions from the surface sediments took place at the Globe Institute, University of Copenhagen, Denmark and at the Norwegian Research Centre, NORCE, Bergen, Norway. Both facilities have compartmented laboratories, avoiding DNA and PCR contamination. Following the manufacturer’s instructions, depending on the amount of sediment (0.5–5 g), extractions were conducted using either DNeasy PowerSoil or DNeasy PowerSoil Max (Qiagen, Hilden, Germany). Lysing Matrix E (Thermo Fisher Scientific, Massachusetts, USA) was added to the PowerBead Max tubes. DNA from the two sediment cores was extracted at the ancient DNA laboratories at Globe Institute, University of Copenhagen. DNA from the AMD16-117Q core was extracted using DNeasy PowerSoil Max, with an addition of Lysing Matrix E (VWR) and the following modification: to the C1-solution provided in the kit, we added 50 mM Tris/HCl, 20 mM EDTA, 150 mM NaCl and 50 mM DTT. After the homogenisation step, 0.8 mg of proteinase K was added, and the sample was incubated on a rotating wheel overnight at 56 °C. The remaining extraction steps followed the manufacturer’s protocol.
Quantitative PCR
The ddPCR analysis was done at NORCE in Bergen (Norway) following an established protocol (23). We used 5.5 µl DNA templates in 20 µl ddPCR reactions volume containing 1X EvaGreen supermix, and 250 nM of each primer Polarella-ITS-44F (5’-CGA CTG GGT GGA GAT GGT TG-3’) and Polarella-ITS-138R (5’-CCC AGG TGT TTA AGC CAG GT-3’)23. Droplet generation and PCR enumeration were performed according to the manufacturer’s instructions (Bio-Rad). The results were normalised to gene copies per gram of wet sediment (gc g-1); see Supplementary Protocol 1. This primer set was previously tested on a pool of DNA extracts from nine different dinoflagellate species: Heterocapsa triquetra, Gyrodinium aureolum, Karlodinium armiger, Woloszynskia cincta, Pelagodinium beii, Alexandrium minutum, Scrippsiella donghaiensis, Symbiodinium sp., Biecheleria sp23. and confirmed to be species-specific under these conditions. To validate that the ITS1 primers did not amplify other taxa in the marine sediments, we sequenced reads from AMD16-117Q (n = 8) and surface sediments from Inglefield Bredning (n = 3). This was done by PCR amplification by the ddPCR primers, using Q5® High-Fidelity DNA Polymerase. The sequencing was conducted using the Illumina MiSeq v3 platform. Paired-end reads were merged and filtered for quality, and adapters were removed using AdaptorRemoval81 and Primers were removed using cutPrimers82. The resulting cleaned reads were then identified using the NCBIBLAST (September, 2020). All identified sequences matched P. glacialis as the best hit with 98-100 % identity; the variation is given in Supplementary Fig. 3, and the raw reads are available at the open data repository GEUS Dataverse83. Due to the potential risk of contamination during the freeze–drying step, blanks for this step were produced by placing artificial clay that was wetted with molecular-grade water into sterile WHIRL-PAK© bags and freeze-dried in parallel with the samples. Using primers that do amplify P. glacialis43, no P. glacialis reads were found in the blanks. In addition, negative controls were used at every stage: freeze–drying (2), weighing (1 blank per round of 6–8 samples of freeze-dried sediments), extraction blank (1 per round of 6–8 samples) and ddPCR (2 blanks per PCR run). All blanks were tested using ddPCR, and no contamination was detected.
Palynological analyses
Sediment core samples for dinoflagellate cyst counts were prepared at Ghent University, following the protocol described in84. Before treatment, a calibrated tablet of Lycopodium clavatum spores was added to each sample to allow later estimates of the absolute concentrations of dinoflagellate cysts. The sediment was treated with room-temperature hydrochloric acid (HCl, 2N) and room-temperature hydrofluoric acid (HF, 40%) to remove carbonates and silicates, respectively. Between each treatment, the sediment was rinsed with distilled water. To break up clusters of amorphous organic matter, the residue was sonicated for <30 s, and subsequently sieved through a 10 µm mesh. The final residues were mixed with glycerine jelly and mounted on microscope slides. Counts of cysts of P. glacialis and other dinoflagellate cyst taxa were done using an upright light microscope (Olympus BX53), at magnifications of 400× and 1000×.
In the present study, selected sediment samples (including surface sediments and sediment core samples) for which the presence of P. glacialis was verified were treated twice with 15–20 mL of HF (40%), at a temperature of 50 °C (i.e., warm acid treatment, standard palynological treatment in the past). After the first treatment, for around 20 minutes, the sediment was left in HF overnight. Following acid treatments and sieving, sediment residue used for palynological analyses was mounted onto permanent microscopy slides. All sediment samples treated with warm HF were treated using the same protocol (i.e., the same duration of acid treatment, acid concentration, and temperature).
Sea-ice concentration data
The number of days per year with sea ice-cover was retrieved from the National Snow and Ice Data Center (https://nsidc.org/data/g02135/versions/3), which uses passive microwave sea-ice concentration data with a spatial resolution of 25 × 25 km. The data were retrieved for each sampling station and converted into annual mean sea-ice concentrations. For the three Arctic fjord sites and the North-East Water polynya, we used NASA Worldview (https://worldview.earthdata.nasa.gov) imagery to estimate sea-ice concentrations. The sea-ice cover above the surface seabed sediment sampling sites was estimated by averaging annual sea-ice concentrations six years prior to the sampling year. The Polar Portal (http://polarportal.dk) was used to determine sea-ice cover of first-year or multiyear sea ice at the sampling locations. For observations in Qeqertarsuup Tunua (Disko Bay), data were provided by the Arctic Station, University of Copenhagen. Observations of sea-ice formation near Qikiqtarjuaq and further up central Baffin Bay in 2015 and 2016 are reported in75. Sea-ice concentrations above the sediment traps deployed in 2005-2006 in the North Water in Baffin Bay, in the Beaufort Sea, and in the northern Laptev Sea were previously reported in77. Sea-ice concentrations above the sediment traps in the Beaufort Sea (2008-2016) and Rijpfjorden (Svalbard) (2013-2014) were retrieved from the Polar Portal and NASA Worldview.
Numerical analysis
Measurements were taken from distinct samples, and the Shapiro–Wilks Normality test was used to verify that the data were normally distributed. To examine if there was a relationship between the two dynamic variables Pgla-DNA gene copy numbers and sea-ice concentration, a linear and polynomial regression was applied. To detect possible autocorrelation that could lead to misinterpretation in a small sample size (<50), the regressions were validated by a Durbin-Watson test. For the detection of outliers in the regression analysis, the Cook’s Distance test was performed. To obtain an improved fit by polynomial cubic regression, the optimal model was established by testing k = 1–5. Independent t-tests were applied to compare samples at different sites.
Regressions of Pgla-DNA indices and environmental parameters were performed via SigmaPlot Version 14.5 (2020) Systat Software, Inc., San Jose, California. A p-value lower than 0.05 was considered statistically significant.
The MAT was applied to the dinoflagellate cyst assemblage data down-core using the reference dataset n = 196853 with standard settings85.
Data availability
The datasets generated and analysed during the current study are available in the open data repository GEUS Dataverse: https://doi.org/10.22008/FK2/IPZG7V.
References
IPCC, 2022, “Climate Change 2022: Impacts, Adaptation, and Vulnerability” in Contribution of Working Group II to the Sixth Assessment Report of the Intergovernmental Panel on Climate Change. (Cambridge University Press, 2022).
Else, B. G. T. et al. Wintertime CO2 fluxes in an Arctic polynya using eddy covariance: evidence for enhanced air-sea gas transfer during ice formation. J. Geophys. Res. Oceans https://doi.org/10.1029/2010JC006760 (2011).
Yoshimori, M., Abe-Ouchi, A. & Laine, A. The role of atmospheric heat transport and regional feedbacks in the Arctic warming at equilibrium. Clim. Dynam. 49, 3457–3472 (2017).
de Vernal, A., Gersonde, R., Goosse, H., Seidenkrantz, M. S. & Wolff, E. W. Sea ice in the paleoclimate system: the challenge of reconstructing sea ice from proxies - an introduction. Q. Sci. Rev. 79, 1–8 (2013).
Curran, M. A. J., van Ommen, T. D., Morgan, V. I., Phillips, K. L. & Palmer, A. S. Ice core evidence for Antarctic sea ice decline since the 1950s. Science 302, 1203–1206 (2003).
Spolaor, A. et al. Canadian Arctic sea ice reconstructed from bromine in the Greenland NEEM ice core. Sci. Rep. 6, 33925 (2016).
Funder, S. et al. A 10,000-year record of arctic ocean sea-ice variability-view from the beach. Science 333, 747–750 (2011).
Dyke, A. S., Hooper, J. & Savelle, J. M. A history of sea ice in the Canadian Arctic Archipelago based on postglacial remains of the bowhead whale (Balaena mysticetus). Arctic 49, 235–255 (1996).
Kolling, H. M. et al. Biomarker distributions in (Sub)-arctic surface sediments and their potential for sea ice reconstructions. Geochem. Geophys. Geosyst. 21 e2019GC008629 (2020).
Wang, K. J. et al. Group 2i Isochrysidales produce characteristic alkenones reflecting sea ice distribution. Nat. Commun. https://doi.org/10.1038/s41467-020-20187-z (2021).
Weckström, K. et al. Improving the paleoceanographic proxy tool kit—on the biogeography and ecology of the sea ice-associated species Fragilariopsis oceanica, Fragilariopsis reginae-jahniae and Fossula arctica in the northern North Atlantic. Mar. Micropaleontol. https://doi.org/10.1016/j.marmicro.2020.101860 (2020).
Belt, S. T. et al. A novel chemical fossil of palaeo sea ice: IP25. Org. Geochem. 38, 16–27 (2007).
Belt, S. T. Source-specific biomarkers as proxies for Arctic and Antarctic sea ice. Org. Geochem. 125, 277–298 (2018).
Müller, J. et al. Towards quantitative sea ice reconstructions in the northern North Atlantic: a combined biomarker and numerical modelling approach. Earth Planet. Sci. Lett. 306, 137–148 (2011).
Pienkowski, A. J. et al. Seasonal sea ice persisted through the Holocene Thermal Maximum at 80 degrees N. Commun. Earth Environ. https://doi.org/10.1038/s43247-021-00191-x (2021).
Belt, S. T. What do IP25 and related biomarkers really reveal about sea ice change? Q. Sci. Rev. 204, 216–219 (2019).
Ribeiro, S. et al. Sea ice and primary production proxies in surface sediments from a High Arctic Greenland fjord: Spatial distribution and implications for palaeoenvironmental studies. Ambio 46, S106–S118 (2017).
Limoges, A. et al. Spring succession and vertical export of diatoms and IP25 in a seasonally ice-covered high arctic fjord. Front. Earth Sci. https://doi.org/10.3389/feart.2018.00226 (2018).
Horner, T., Stein, R., Fahl, K. & Birgel, D. Post-glacial variability of sea ice cover, river run-off and biological production in the western Laptev Sea (Arctic Ocean)—a high-resolution biomarker study. Q. Sci. Rev. 143, 133–149 (2016).
Su, L. et al. HBIs and sterols in surface sediments across the East Siberian Sea: implications for palaeo sea-ice reconstructions. Geochem. Geophys. Geosyst. https://doi.org/10.1029/2021GC009940 (2022).
Xiao, X. T., Fahl, K. & Stein, R. Biomarker distributions in surface sediments from the Kara and Laptev seas (Arctic Ocean): indicators for organic-carbon sources and sea-ice coverage. Q. Sci. Rev. 79, 40–52 (2013).
Armbrecht, L. et al. Ancient marine sediment DNA reveals diatom transition in Antarctica. Nat. Commun. https://doi.org/10.1038/s41467-022-33494-4 (2022).
De Schepper, S. et al. The potential of sedimentary ancient DNA for reconstructing past sea ice evolution. ISME J. 13, 2566–2577 (2019).
Zimmermann, H. H. et al. Changes in the composition of marine and sea-ice diatoms derived from sedimentary ancient DNA of the eastern Fram Strait over the past 30 000 years. Ocean Sci. 16, 1017–1032 (2020).
Armbrecht, L. H. The potential of sedimentary ancient DNA to reconstruct past ocean ecosystems. Oceanography 33, 116–123 (2020).
Boere, A. C. et al. Late-Holocene succession of dinoflagellates in an Antarctic fjord using a multi-proxy approach: paleoenvironmental genomics, lipid biomarkers and palynomorphs. Geobiology 7, 265–281 (2009).
Thaler, M. & Lovejoy, C. Distribution and diversity of a protist predator cryothecomonas (Cercozoa) in arctic marine waters. J. Eukaryot. Microbiol. 59, 291–299 (2012).
Zimmermann, H. H. et al. Marine ecosystem shifts with deglacial sea-ice loss inferred from ancient DNA shotgun sequencing. Nat. Commun. 14, 1650 (2023).
Hindson, B. J. et al. High-throughput droplet digital PCR system for absolute quantitation of DNA copy number. Anal. Chem. 83, 8604–8610 (2011).
Montresor, M., Procaccini, G. & Stoecker, D. K. Polarella glacialis, gen. nov., sp. nov. (Dinophyceae): suessiaceae are still alive! J. Phycol. 35, 186–197 (1999).
Marret, F. et al. From bi-polar to regional distribution of modern dinoflagellate cysts, an overview of their biogeography. Mar. Micropaleontol. https://doi.org/10.1016/j.marmicro.2019.101753 (2020).
Montresor, M., Lovejoy, C., Orsini, L., Procaccini, G. & Roy, S. Bipolar distribution of the cyst-forming dinoflagellate Polarella glacialis. Polar Biol. 26, 186–194 (2003).
Kauko, H. M. et al. Algal colonization of young arctic sea ice in spring. Front. Mar. Sci. https://doi.org/10.3389/fmars.2018.00199 (2018).
Stoecker, D. K., Gustafson, D. E., Black, M. M. D. & Baier, C. T. Population dynamics of microalgae in the upper land-fast sea ice at a snow-free location. J. Phycol. 34, 60–69 (1998).
Stoecker, D. K., Gustafson, D. E., Baier, C. T. & Black, M. M. D. Primary production in the upper sea ice. Aquat. Microb. Ecol. 21, 275–287 (2000).
Rengefors, K., Logares, R., Laybourn-Parry, J. & Gast, R. J. Evidence of concurrent local adaptation and high phenotypic plasticity in a polar microeukaryote. Environ. Microbiol. 17, 1510–1519 (2015).
Logares, R., Boltovskoy, A., Bensch, S., Laybourn-Parry, J. & Rengefors, K. Genetic diversity patterns in five protist species occurring in lakes. Protist 160, 301–317 (2009).
Stephens, T. G. et al. Genomes of the dinoflagellate Polarella glacialis encode tandemly repeated single-exon genes with adaptive functions. BMC Biol. https://doi.org/10.1186/s12915-020-00782-8 (2020).
Zheng, S. X., Wang, G. Z. & Lin, S. J. Heat shock effects and population survival in the polar dinoflagellate Polarella glacialis. J. Exp. Mar. Biol. Ecol. 438, 100–108 (2012).
Lovejoy, C., Legendre, L., Martineau, M. J., Bacle, J. & von Quillfeldt, C. H. Distribution of phytoplankton and other protists in the North Water. Deep Sea Res. Part II 49, 5027–5047 (2002).
Harðardóttir, S., Lundholm, N., Møestrup, O. & Nielsen, T. G. Description of Pyramimonas diskoicola sp. nov. and the importance of the flagellate Pyramimonas (Prasinophyceae) in Greenland sea ice during the winter-spring transition. Polar Biol. 37, 1479–1494 (2014).
Hop, H. et al. Changes in sea-ice protist diversity with declining sea ice in the arctic ocean from the 1980s to 2010s. Front. Mar. Sci. https://doi.org/10.3389/fmars.2020.00243 (2020).
Comeau, A. M. et al. Protists in Arctic drift and land-fast sea ice. J. Phycol. 49, 229–240 (2013).
Majaneva, M., Rintala, J. M., Piisila, M., Fewer, D. P. & Blomster, J. Comparison of wintertime eukaryotic community from sea ice and open water in the Baltic Sea, based on sequencing of the 18S rRNA gene. Polar Biol. 35, 875–889 (2012).
Heikkilä, M. et al. Dinoflagellate cyst production over an annual cycle in seasonally ice-covered Hudson Bay. Mar. Micropaleontol. 125, 1–24 (2016).
Luostarinen, T. et al. Seasonal and habitat-based variations in vertical export of biogenic sea-ice proxies in Hudson Bay. Commun. Earth Environ. https://doi.org/10.1038/s43247-023-00719-3 (2023).
Kunz-Pirrung, M. Rekonstruktion der Oberflächenmassen der östlichen Laptevsee im Holozän anhand aquatischen Palynomorphen (Aquatic palynomorphs: Reconstruction of Holocene sea-surface water masses in the eastern Laptev Sea). In Berichte zur Polarforschung (Reports on Polar Research) Vol. 281 (Alfred Wegener Institute for Polar and Marine Research, 1998) p. 1–117.
Heikkilä, M. et al. Surface sediment dinoflagellate cysts from the Hudson Bay system and their relation to freshwater and nutrient cycling. Mar. Micropaleontol. 106, 79–109 (2014).
Limoges, A. et al. Linking the modern distribution of biogenic proxies in high Arctic greenland shelf sediments to sea ice, primary production, and Arctic-Atlantic inflow. J. Geophys. Res. 123, 760–786 (2018).
Ichinomiya, M., Nakamachi, M., Fukuchi, M. & Taniguchi, A. Resting cells of microorganisms in the 20–100μm fraction of marine sediments in an Antarctic coastal area. Polar Sci. 2, 27–32 (2008).
Van Nieuwenhove, N., Limoges, A., Norgaard-Pedersen, N., Seidenkrantz, M. S. & Ribeiro, S. Episodic Atlantic water inflow into the independence fjord system (Eastern North Greenland) during the Holocene and Last Glacial Period. Front. Earth Sci. https://doi.org/10.3389/feart.2020.565670 (2020).
Koerner, K. A. et al. Late Holocene sea-surface changes in the North Water polynya reveal freshening of northern Baffin Bay in the 21st century. Glob. Planet Change https://doi.org/10.1016/j.gloplacha.2021.103642 (2021).
de Vernal, A. et al. Distribution of common modern dinoflagellate cyst taxa in surface sediments of the Northern Hemisphere in relation to environmental parameters: The new n=1968 database. Mar. Micropaleontol. https://doi.org/10.1016/j.marmicro.2019.101796 (2020).
Møestrup, O., Lindberg, K. & Daugbjerg, N. Studies on woloszynskioid dinoflagellates IV: The genus Biecheleria gen. nov. Phycol. Res. 57, 203–220 (2009).
Hidalgo, O. et al. Is there an upper limit to genome size? Trends Plant Sci. 22, 567–573 (2017).
Hou, Y. & Lin, S. Distinct gene number-genome size relationships for eukaryotes and non-eukaryotes: gene content estimation for dinoflagellate genomes. PloS ONE 4, e6978 (2009).
Riisgaard, K., Nielsen, T. G. & Hansen, P. J. Impact of elevated pH on succession in the Arctic spring bloom. Mar. Ecol. Progress Ser. 530, 63–75 (2015).
Jackson, R. et al. Holocene polynya dynamics and their interaction with oceanic heat transport in northernmost Baffin Bay. Sci. Rep. https://doi.org/10.1038/s41598-021-88517-9 (2021).
Thomson, P. G. et al. Antarctic distribution, pigment and lipid composition, and molecular identification of the brine dinoflagellate Polarella glacialis (Dinophyceae). J. Phycol. 40, 867–873 (2004).
van Leeuwe, M. A. et al. Microalgal community structure and primary production in Arctic and Antarctic sea ice: a synthesis. Elementa https://doi.org/10.1525/elementa.267’ (2018).
Stoecker, D. K., Gustafson, D. E., Merrell, J. R., Black, M. M. D. & Baier, C. T. Excystment and growth of chrysophytes and dinoflagellates at low temperatures and high salinities in Antarctic sea-ice. J. Phycol. 33, 585–595 (1997).
Werner, I., Ikavalko, J. & Schunemann, H. Sea-ice algae in Arctic pack ice during late winter. Polar Biol. 30, 1493–1504 (2007).
Kalenitchenko, D., Joli, N., Potvin, M., Tremblay, J. E. & Lovejoy, C. Biodiversity and species change in the arctic ocean: a view through the lens of Nares Strait. Front. Mar. Sci. https://doi.org/10.3389/fmars.2019.00479 (2019).
Joli, N. et al. Need for focus on microbial species following ice melt and changing freshwater regimes in a Janus Arctic Gateway. Sci. Rep. https://doi.org/10.1038/s41598-018-27705-6 (2018).
Elferink, S. et al. Molecular diversity patterns among various phytoplankton size-fractions in West Greenland in late summer. Deep Sea Res. Part I 121, 54–69 (2017).
Ellegaard, M. & Ribeiro, S. The long-term persistence of phytoplankton resting stages in aquatic ‘seed banks. Biol. Rev. 93, 166–183 (2018).
Georgiadis, E. et al. Local and regional controls on Holocene sea ice dynamics and oceanography in Nares Strait, Northwest Greenland. Mar. Geol. https://doi.org/10.1016/j.margeo.2020.106115 (2020).
Ribeiro, S. et al. Vulnerability of the North Water ecosystem to climate change. Nat. Commun. https://doi.org/10.1038/s41467-021-24742-0 (2021).
Pandeirada, M. S., Craveiro, S. C., Daugbjerg, N., Moestrup, Ø. & Calado, A. J. Fine-structural characterization and phylogeny of Sphaerodinium (Suessiales, Dinophyceae), with the description of an unusual type of freshwater dinoflagellate cyst. Eur. J. Protistol. 78, 125770 (2021).
Ellegaard, M. et al. Dead or alive: sediment DNA archives as tools for tracking aquatic evolution and adaptation. Commun. Biol. https://doi.org/10.1038/s42003-020-0899-z (2020).
Torti, A., Lever, M. A. & Jorgensen, B. B. Origin, dynamics, and implications of extracellular DNA pools in marine sediments. Mar. Genom. 24, 185–196 (2015).
Armbrecht, L. et al. Paleo-diatom composition from Santa Barbara Basin deep-sea sediments: a comparison of 18S-V9 and diat-rbcL metabarcoding vs shotgun metagenomics. ISME Commun. 1, 66 (2021).
Armbrecht, L., Hallegraeff, G., Bolch, C. J. S., Woodward, C. & Cooper, A. Hybridisation capture allows DNA damage analysis of ancient marine eukaryotes. Sci. Rep. https://doi.org/10.1038/s41598-021-82578-6 (2021).
Smik, L., Cabedo-Sanz, P. & Belt, S. T. Semi-quantitative estimates of paleo Arctic sea ice concentration based on source-specific highly branched isoprenoid alkenes: a further development of the PIP25 index. Org. Geochem. 92, 63–69 (2016).
Massicotte, P. et al. Green Edge ice camp campaigns: understanding the processes controlling the under-ice Arctic phytoplankton spring bloom. Earth Syst. Sci. Data 12, 151–176 (2020).
Fernandez-Mendez, M. et al. Photosynthetic production in the central Arctic Ocean during the record sea-ice minimum in 2012. Biogeosciences 12, 3525–3549 (2015).
Lalande, C., Forest, A., Barber, D. G., Gratton, Y. & Fortier, L. Variability in the annual cycle of vertical particulate organic carbon export on Arctic shelves: contrasting the Laptev Sea, Northern Baffin Bay and the Beaufort Sea. Cont. Shelf Res. 29, 2157–2165 (2009).
Blaauw, M. & Christen, J. A. Flexible paleoclimate age-depth models using an autoregressive gamma process. Bayesian Anal. 6, 457–474 (2011).
Reimer, P. J. et al. IntCal13 and Marine13 radiocarbon age calibration curves 0–50,000 years cal BP. Radiocarbon 55, 1869–1887 (2013).
White, T. J., Bruns, T., Lee, S. & Taylor, J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. PCR Protoc 18, 315–322 (1990).
Schubert, M., Lindgreen, S. & Orlando, L. AdapterRemoval v2: rapid adapter trimming, identification, and read merging. BMC Res. Notes 9, 1–7 (2016).
Kechin, A., Boyarskikh, U., Kel, A. & Filipenko, M. cutPrimers: a new tool for accurate cutting of primers from reads of targeted next generation sequencing. J. Comput. Biol. 24, 1138–1143 (2017).
Ribeiro, S., et al. Millennial-scale variations in Arctic sea ice are recorded in sedimentary ancient DNA of the microalga Polarella glacialis. GEUS Dataverse, V1 https://doi.org/10.22008/FK2/IPZG7V.
Quaijtaal, W., Donders, T. H., Persico, D. & Louwye, S. Characterising the middle Miocene Mi-events in the Eastern North Atlantic realm: a first high-resolution marine palynological record from the Porcupine Basin. Palaeogeogr. Palaeoclimatol. 399, 140–159 (2014).
de Vernal, A. et al. Reconstructing past sea ice cover of the Northern Hemisphere from dinocyst assemblages: status of the approach. Q. Sci. Rev. 79, 122–134 (2013).
General Bathymetric Chart of the Oceans (GEBCO) Blue Marble satellite mosaic, NASA’s Earth Observatory, www.nasa.gov/vision/earth/features/blue_marble.html World Vector Shoreline, National Geophysical Data Center, http://rimmer.ngdc.noaa.gov/mgg/coast/wvs.html.
GLIMS and NSIDC (2005, updated 2018): Global Land Ice Measurements from Space glacier database. Compiled and made available by the international GLIMS community and the National Snow and Ice Data Center, Boulder CO, USA. https://doi.org/10.7265/N5V98602.
RGI Consortium (2017). Randolph Glacier Inventory – A Dataset of Global Glacier Outlines: Version 6.0: Technical Report, Global Land Ice Measurements from Space, Colorado, USA. Digital Media. https://doi.org/10.7265/N5-RGI-60.
Vihtakari M. ggOceanMaps: Plot Data on Oceanographic Maps using ‘ggplot2’. R package version 2.1.12 (2023).
Citterio M., Ahlstrøm A. P., “Ice extent” GEUS Dataverse, V1 https://doi.org/10.22008/FK2/PRWITW (2022).
Acknowledgements
For assistance in the laboratory, we thank M. Potvin, C. Guilmette and, with map compilation K. K. Kjeldsen and C. Brogaard. We acknowledge the staff at the Arctic Station, University of Copenhagen and the help of Abel Brand and other local hunters from Qeqertarsuaq. The GreenEdge project was carried out with the support of the Hamlet of Qikiqtarjuaq, members of the community, including the Inuksuit School and Jacqueline Arsenault. We thank the crew and staff of CCGS Amundsen and Amundsen Science. We acknowledge the contributions of individuals from Takuvik, CNRS, the University of Manitoba and the Vagabond, who carried out sampling and logistic coordination for both the ice camp and the Amundsen mission. This study received funding from the European Union’s Horizon 2020 research and innovation programme under the Marie Skłodowska-Curie grant agreement No. 846860 (grant to S.H., hosts S.Ri and C.Lo), the VILLUM Foundation grant nr: VKR023454 to Sri, the Danish Council for Independent Research (DFF Sapere Aude grant no. 9064-00039B to S.Ri, and grants No. 7014-00113B (G-Ice) and 0135-798 00165B (GreenShelf) to MSS, with additional funding from the EU H2020 Grant Agreement No. 869383 (ECOTIP). The NorthGreen17 expedition was funded by the Danish Centre for Marine Research and the Natural Sciences and Engineering Research Council of Canada (NSERC). The GreenEdge project was conducted under the scientific coordination of the Canada Excellence Research Chair on Remote sensing of Canada’s new Arctic frontier (CERC-new frontier) and the CNRS & Université Laval Takuvik Joint International Laboratory (UMI3376). The GreenEdge work was funded by the following: France, ANR (Contract #111112), CNES (project #131425), French Arctic Initiative, Foundation Total, CSA, LEFE and IPEV (project #1164) and in Canada ArcticNet (A network of Centers for excellence, Canada), CERC-new frontier and the CNRS & Université Laval Takuvik Joint International Laboratory (UMI3376). The Canadian research icebreaker CCGS Amundsen received support from the Amundsen Science programme funded by the Canada Foundation for Innovation (CFI) Major Science Initiatives (MSI) Fund. The Long-Term Oceanic Observatories project of ArcticNet, to C.La for sediment trap deployments. Natural Science and Engineering Research Council (NSERC-Canada) discovery grants to C.Lo and A.L. (discovery grant RGPIN-2018-03984). The ice2ice project was funded by the European Research Council under the European Community’s Seventh Framework Programme (grant agreement 610055), and S.D.S, J.L.R and K.S.S are funded via the European Research Council under the European Union’s Horizon 2020 research and innovation programme (grant agreement 818449, AGENSI). The Qeqertarsuup Tunua (Spring in Disko) project was financed by grant DFF–1323-00258 from the Danish Research Council to N.L.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Earth & Environment thanks Cheong Xin Chan and Timothy Stephens for their contribution to the peer review of this work. Primary Handling Editor: Aliénor Lavergne. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Harðardóttir, S., Haile, J.S., Ray, J.L. et al. Millennial-scale variations in Arctic sea ice are recorded in sedimentary ancient DNA of the microalga Polarella glacialis. Commun Earth Environ 5, 25 (2024). https://doi.org/10.1038/s43247-023-01179-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s43247-023-01179-5
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.