Abstract
Venetoclax with azacitidine (ven/aza) has emerged as a promising treatment regimen for acute myeloid leukemia (AML), with a high percentage of clinical remissions in newly diagnosed patients. However, approximately 30% of newly diagnosed patients and the majority of patients who have relapsed do not achieve remission with ven/aza. We previously reported that ven/aza efficacy is based on eradication of AML stem cells through a mechanism involving inhibition of amino acid metabolism, a process required in primitive AML cells to drive oxidative phosphorylation. Herein we demonstrate that resistance to ven/aza occurs via upregulation of fatty acid oxidation (FAO), which occurs either due to RAS pathway mutations or as a compensatory adaptation in relapsed disease. Utilization of FAO obviates the need for amino acid metabolism, thereby rendering ven/aza ineffective. Pharmacological inhibition of FAO restores sensitivity to ven/aza in drug-resistant AML cells. We propose inhibition of FAO as a therapeutic strategy to address ven/aza resistance.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Patient-related clinical data not included in the paper were generated as part of a multicenter clinical trial (NCT02203773). A detailed description of the dose-escalation portion of the study has been published (Dinardo et al.)2. All DNA and RNA raw and analyzed sequencing data can be found at the GEO database and are available via accession numbers GSE156041 and GSE143363 (single-cell RNA-seq) and accession numbers GSE156008 and GSE155431 (bulk RNA-seq). The Beat AML dataset can be downloaded from the cBIOPortal (data listed in dataset denoted as OHSU, Nature 2018). Source data are provided with this paper. Other data supporting the findings of this study are available from the corresponding author on reasonable request.
References
Pollyea, D. A. & Jordan, C. T. Therapeutic targeting of acute myeloid leukemia stem cells. Blood 129, 1627–1635 (2017).
DiNardo, C. D. et al. Venetoclax combined with decitabine or azacitidine in treatment-naive, elderly patients with acute myeloid leukemia. Blood 133, 7–17 (2019).
Jones, C. L. et al. Inhibition of amino acid metabolism selectively targets human leukemia stem cells. Cancer Cell 34, 724–740 (2018).
Pollyea, D. A. et al. Venetoclax with azacitidine disrupts energy metabolism and targets leukemia stem cells in patients with acute myeloid leukemia. Nat. Med. 24, 1859–1866 (2018).
Lagadinou, E. D. et al. BCL-2 inhibition targets oxidative phosphorylation and selectively eradicates quiescent human leukemia stem cells. Cell Stem Cell 12, 329–341 (2013).
Sriskanthadevan, S. et al. AML cells have low spare reserve capacity in their respiratory chain that renders them susceptible to oxidative metabolic stress. Blood 125, 2120–2130 (2015).
Skrtic, M. et al. Inhibition of mitochondrial translation as a therapeutic strategy for human acute myeloid leukemia. Cancer Cell 20, 674–688 (2011).
Cole, A. et al. Inhibition of the mitochondrial protease ClpP as a therapeutic strategy for human acute myeloid leukemia. Cancer Cell 27, 864–876 (2015).
Chan, S. M. et al. Isocitrate dehydrogenase 1 and 2 mutations induce BCL-2 dependence in acute myeloid leukemia. Nat. Med. 21, 178–184 (2015).
Jones, C. L. et al. Cysteine depletion targets leukemia stem cells through inhibition of electron transport complex II. Blood 134, 389–394 (2019).
DiNardo, C. D. et al. Safety and preliminary efficacy of venetoclax with decitabine or azacitidine in elderly patients with previously untreated acute myeloid leukaemia: a non-randomised, open-label, phase 1b study. Lancet Oncol. 19, 216–228 (2018).
DiNardo, C. D. et al. Azacitidine and venetoclax in previously untreated acute myeloid leukemia. N. Engl. J. Med. 383, 617–629 (2020).
DiNardo, C. D. et al. Clinical experience with the BCL2-inhibitor venetoclax in combination therapy for relapsed and refractory acute myeloid leukemia and related myeloid malignancies. Am. J. Hematol. 93, 401–407 (2018).
Nechiporuk, T. et al. The TP53 apoptotic network is a primary mediator of resistance to BCL2 inhibition in AML cells. Cancer Discov. 9, 910–925 (2019).
Chen, X. et al. Targeting mitochondrial structure sensitizes acute myeloid leukemia to venetoclax treatment. Cancer Discov. 9, 890–909 (2019).
Karjalainen, R. et al. Elevated expression of S100A8 and S100A9 correlates with resistance to the BCL-2 inhibitor venetoclax in AML. Leukemia 33, 2548–2553 (2019).
Kuusanmaki, H. et al. Phenotype-based drug screening reveals association between venetoclax response and differentiation stage in acute myeloid leukemia. Haematologica https://doi.org/10.3324/haematol.2018.214882 (2020).
Pei, S. et al. Monocytic subclones confer resistance to venetoclax-based therapy in acute myeloid leukemia patients. Cancer Discov. https://doi.org/10.1158/2159-8290.Cd-19-0710 (2020).
Jones, C. L. et al. Nicotinamide metabolism mediates resistance to venetoclax in relapsed acute myeloid leukemia stem cells. Cell Stem Cell https://doi.org/10.1016/j.stem.2020.07.021 (2020).
Pei, S. et al. AMPK/FIS1-mediated mitophagy is required for self-renewal of human AML stem cells. Cell Stem Cell 23, 86–100 (2018).
Reisz, J. A., Zheng, C., D’Alessandro, A. & Nemkov, T. Untargeted and semi-targeted lipid analysis of biological samples using mass spectrometry-based metabolomics. Methods Mol. Biol. 1978, 121–135 (2019).
Escudero, S. et al. Dynamic regulation of long-chain fatty acid oxidation by a noncanonical interaction between the MCL-1 BH3 helix and VLCAD. Mol. Cell 69, 729–743 (2018).
Perciavalle, R. M. et al. Anti-apoptotic MCL-1 localizes to the mitochondrial matrix and couples mitochondrial fusion to respiration. Nat. Cell Biol. 14, 575–583 (2012).
Lee, K.-M. et al. MYC and MCL1 cooperatively promote chemotherapy-resistant breast cancer stem cells via regulation of mitochondrial oxidative phosphorylation. Cell Metab. 26, 633–647 (2017).
Ramsey, H. E. et al. A novel MCL1 inhibitor combined with venetoclax rescues venetoclax-resistant acute myelogenous leukemia. Cancer Discov. 8, 1566–1581 (2018).
Kotschy, A. et al. The MCL1 inhibitor S63845 is tolerable and effective in diverse cancer models. Nature 538, 477–482 (2016).
Shlush, L. I. et al. Tracing the origins of relapse in acute myeloid leukaemia to stem cells. Nature 547, 104–108 (2017).
Ho, T. C. et al. Evolution of acute myelogenous leukemia stem cell properties after treatment and progression. Blood 128, 1671–1678 (2016).
Padanad, M. S. et al. Fatty acid oxidation mediated by Acyl-CoA synthetase long chain 3 Is required for mutant KRAS lung tumorigenesis. Cell Rep. 16, 1614–1628 (2016).
Gouw, A. M. et al. Oncogene KRAS activates fatty acid synthase, resulting in specific ERK and lipid signatures associated with lung adenocarcinoma. Proc. Natl Acad. Sci. USA 114, 4300–4305 (2017).
Samudio, I. et al. Pharmacologic inhibition of fatty acid oxidation sensitizes human leukemia cells to apoptosis induction. J. Clin. Invest. 120, 142–156 (2010).
Farge, T. et al. Chemotherapy-resistant human acute myeloid leukemia cells are not enriched for leukemic stem cells but require oxidative metabolism. Cancer Discov. 7, 716–735 (2017).
Tabe, Y. et al. Inhibition of FAO in AML co-cultured with BM adipocytes: mechanisms of survival and chemosensitization to cytarabine. Sci. Rep. 8, 16837 (2018).
Nemkov, T., D’Alessandro, A. & Hansen, K. C. Three-minute method for amino acid analysis by UHPLC and high-resolution quadrupole orbitrap mass spectrometry. Amino Acids 47, 2345–2357 (2015).
Nemkov, T., Reisz, J. A., Gehrke, S., Hansen, K. C. & D’Alessandro, A. High-throughput metabolomics: isocratic and gradient mass spectrometry-based methods. Methods Mol. Biol. 1978, 13–26 (2019).
Gehrke, S. et al. Red blood cell metabolic responses to torpor and arousal in the hibernator Arctic ground squirrel. J. Proteome Res. 18, 1827–1841 (2019).
Brunetti, L., Gundry, M. C., Kitano, A., Nakada, D. & Goodell, M. A. Highly efficient gene disruption of murine and human hematopoietic progenitor cells by CRISPR/Cas9. J. Vis. Exp. https://doi.org/10.3791/57278 (2018).
Acknowledgements
We thank the University of Colorado Hematology Clinical Trials Unit and the University of Colorado Health Apheresis team for essential help in acquisition of patient samples. We also acknowledge the Molecular and Cellular Analytical Core within the Colorado Nutrition and Obesity Research Center for use of the Seahorse analyzer. This work was supported by the Evans MDS Foundation young investigator award (to B.M.S.); the Leukemia and Lymphoma Society, American Cancer Society (no. 25A5072) and the Cancer League of Colorado (to C.L.J.); the University of Colorado Department of Medicine Outstanding Early Career Scholar Program and the Leukemia and Lymphoma Society Clinical Scholars award (to D.A.P.); the Webb-Waring Early Career Award by the Boettcher Foundation (RM1GM131968), the National Institute of General and Medical Sciences (R01HL146442 and R01HL148151) and the National Heart, Lung and Blood Institutes (to A.D.); the St. Baldrick’s Fellow award (to A.W.); the Ruth and Ralph Seligman Chair in Hematology (to C.S.); National Institutes of Health National Cancer Institute grants R01 CA200707, R01 CA243452 and P30CA046934 (to C.T.J.); and a Leukemia and Lymphoma Society Specialized Center of Research grant (principal investigator, C.T.J.). C.T.J. is supported by the Nancy Carroll Allen Endowed Chair.
Author information
Authors and Affiliations
Contributions
B.M.S. and C.L.J. designed and performed the research, collected, analyzed and interpreted the data, performed the statistical analysis and wrote the manuscript. D.A.P. directed all clinical research, designed and directed the research, analyzed and interpreted data and wrote the manuscript. R.C.-H. and A.D. performed metabolomics and lipidomics experiments, collected, analyzed and interpreted metabolomics data and wrote the manuscript. S.P. assisted with figure design and wrote the manuscript. A.W., A.K., H.Y., A.E.G. and M.G. performed experiments. M.W.B. and M.R.S. designed the research. C.S. wrote the manuscript. D.A. performed statistical analysis. C.T.J. designed and directed the research, analyzed and interpreted data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
D.A.P. receives research funding from Abbvie and has served as a consultant for Abbvie.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Metabolic alternations in venetoclax resistant LSCs.
Metabolite analysis of sensitive versus resistant LSCs. a. Global LC/MS metabolite screen of sensitiveandresistant LSCs from Fig. 2.Heatmap of differential metabolites from samples. Values are normalized by metabolite to median expression of row. Each data point represents the median calculated from four technical replicates. b. Metabolite pathway analysis shows increased fatty acid metabolism pathways in resistant LSCs as determined by Metaboanalyst software analysis of global metabolite levels in LSCs. P value represents adjusted P value after multiple comparison Holm test. c. Heatmap of carnitine and fatty acid levels. The samples in a–c are from the same sensitive (N = 3 patient specimens) and resistant (N = 7 patient specimens) AML patient samples.
Extended Data Fig. 2 PTPN11 confers metabolic alterations leading to venetoclax resistance.
Effects of PTPN11 mutation on metabolism and venetoclax sensitivity. a. Measurement of fatty acids in ven/aza sensitive and resistant LSCs from lipodomics analysis shown as box and whiskers plot with min and max represented. N = 3 sensitive patient specimens versus N = 3 resistant patient specimens measured across 4 technical replicates per patient sample. Significance was measured by paired two-tailed Student’s t-test. b. Venetoclax and ven/aza viability of primary AML cells of patients with or without a PTPN11 mutation. N = 3 independent patient wild-type specimens versus N = 2 (venetoclax + aza) or 3 (venetoclax alone). PTPN11 resistant patient specimens measured across 3 technical replicates per patient sample. Data are presented as mean values. c, d. Basal respiration and glycolytic rate of primary AML cells transduced with lentiviral PTPN11 E76A mutation. N = 1 independent patient wild-type specimens versus N = 1 PTPN11 mutant patient specimens measured across 4-5 technical replicates per sample. Individual values in glycolytic rate consolidated in instrument software and mean value shown. Data are presented as mean values. Patient sample run as single experiment with single transduction.
Extended Data Fig. 3 Fatty acid metabolism can be inhibited by MCL-1 inhibitors.
MCL-1 inhibition decreases fatty acid metabolism in ven/aza resistant LSCs. a. Relative expression of ACADVL in siRNA knockdown samples N = 4 independent patient samples. Data are presented as mean values + /- SD. Significance was measured by paired two-tailed Student’s t-test. b. Basal respiration in ACADVL siRNA knockdown cells with or without VU and VU + Aza. N = 3 patient specimens with N = 5 technical replicates for each experiment. Data are presented as mean values. Patient samples run as single experiment with 3 unique patients. c. SiRNA knockdown of ACADVL and viability with or without VU and VU + Aza. N = 3 independent patient samples for each experiment. Data are presented as mean values + /- SD. Significance was measured by unpaired two-tailed Student’s t-test. d. Global metabolite measurement in sensitive and resistant LSCs after VU + Aza treatment relative to control. N = 6 independent patient specimens versus N = 3 sensitive patient specimens measured across 4 technical replicates per patient sample. Heat map shows the ratio of VU660103 treated to untreated control. e. Global carnitine and fatty acid levels after 4 h of VU + Aza in ven/aza resistant (R) and sensitive (S) specimens. Metabolites that show selective decrease upon drug treatment in ven/aza resistant specimens are indicated by a red X. N = 6 independent patient specimens versus N = 3 sensitive patient specimens measured in across 4 technical replicates per patient sample.
Extended Data Fig. 4 MCL-1 inhibitors alter resistant LSC metabolism.
a. Global amino acid metabolite levels after 4 h of VU + Azacitidine. N = 3 independent patient specimens measured across 4 technical replicates per patient sample. b. Global TCA cycle metabolite levels after 4 h of VU + Azacitidine. N = 3 independent patient specimens versus N = 3 sensitive patient specimens measured across 4 technical replicates per patient sample. Data are presented as median values + /- SD. Significance was measured by unpaired two-tailed Student’s t-test. c. Resistant LSCs exhibit increased palmitate contribution to fatty acid transport metabolites and TCA cycle intermediates as measured through stable isotope labelled palmitate flux. N = 2 resistant specimens measured across 4 technical replicates per specimen per condition. Data are presented as median. d. Palmitate abundance in TCA metabolites after 1 h or 8 h of VU660103 treatment. Stable isotope labeled amino acid flux into TCA cycle intermediates after 1 h or 8 h of treatment. N = 1 independent patient sample measured across 4 technical replicates. Patient sample run as single experiment..e. Engraftment of resistant LSCs is decreased after ex vivo treatment with VU/Aza as measured by human CD45 labelled cells per femur. N = 8 independent animals in vehicle and N = 6 independent animals in VU treated for 1 independent patient derived xenograft. Data are presented as mean values + /- SD. Significance was measured by unpaired two-tailed Student’s t-test.
Extended Data Fig. 5 Fatty acid transport can be targeted to sensitize LSCs to venetoclax.
Quantification of CD36, CPT1A, CPT1C after siRNA. a. Relative MFI from flow cytometry of CD36 after siRNA. N = 4 independent patient samples b. RNA levels of CPT1A, CPT1C, and CD36 24 h post transfection in primary AML specimens. N = 4 independent patient samples. Data are presented as mean values + /- SD. c. Western blot quantification of CPT1A, CPT1C or CPT1A and CPT1C after siRNA in 4 specimens. N = 4 independent patient samples. Western blots representative of at least 2 experimental replicates per patient sample. d. SiRNA for CPT1A and CPT1C alone and in combination and the effects on viability after 24 h treatment with ven+aza. N = 3 independent patient samples. Data are presented as mean values + /- SD. Significance was measured by paired two-tailed Student’s t-test. e. Etomoxir addition in resistant LSCs and normal CD34 HSC/progenitor cells. e. Viability measured after etomoxir treatment in the presence or absence of amino acids. N = 4 independent patient samples. Data are presented as mean values + /- SD. Significance was measured by paired two-tailed Student’s t-testF. Oxygen consumption rate measured after etomoxir treatment in the presence or absence of amino acids via Seahorse analysis. N = 5 independent patient samples. Data are presented as mean values + /- SD. Significance was measured by paired two-tailed Student’s t-test.. g, h Viability and oxygen consumption rate of LSCs after ven/aza, etomoxir, and octanoic acid addition. N = 3 independent patient samples. Data are presented as mean values + /- SD. Significance was measured by paired two-tailed Student’s t-test. i. Viability of LSCs after cytrabine, etomoxir, and octanoic acid addition. N = 3 independent patient samples. Data are presented as mean values + /- SD. Significance was measured by paired two-tailed Student’s t-test. j, k Viability and oxygen consumption rate of LSCs after ven, etomoxir, or ven+ etomoxir. N = 3 independent patient samples. Data are presented as mean values + /- SD. Significance was measured by paired two-tailed Student’s t-test.
Extended Data Fig. 6 Etomoxir reverses resistance to venetoclax in LSCs.
Fold change of amino acid from control after 4 h treatment with ven+aza, VU + aza, VU + ven+ aza, or Eto+ ven+ aza. N = 3 independent patient specimens versus N = 3 sensitive patient specimens measured across 4 technical replicates per patient sample. Data are presented as median values. b. Fold change over control of TCA metabolites after 4 h treatment with ven+aza, VU + aza, VU + ven+ aza, or Eto+ ven+ aza. N = 3 independent patient specimens versus N = 3 sensitive patient specimens measured across 4 technical replicates per patient sample. Data are presented as median values c. Viability normal human CD34 + cells after ven/aza and etomoxir addition. N = 3 independent patient samples. Data are presented as mean values + /- SD. Significance was measured by paired two-tailed Student’s t-test. d. Colony forming units of normal human CD34 + cells after ven/aza and etomoxir addition. N = 3 independent patient samples. Data are presented as mean values + /- SD. Significance was measured by paired two-tailed Student’s t-test. e. Etomoxir and ven+aza treatment in patient derived xenograft. Counts per femur measured with human CD45 + and human CD45 + / CD34 + / CD38-/CD123 via flow cytometry. N = 3 mice for the vehicle, venetoclax + azacitidine, and etomoxir groups and N = 4 for the venetoclax + azacitidine + etomoxir group for 1 independent patient sample derived xenograft. Data are presented as mean values + /- SD. Significance was measured by unpaired two-tailed Student’s t-test. f. Mouse weight average across in vivo ven/aza/eto combination therapy experiment. N = 12 control, N = 11 eto, N = 12 ven+aza, and N = 12 ven+aza+eto mice per dose for 1 independent patient sample derived xenograft. Data are presented as mean values + /- SD. Significance was measured by unpaired two-tailed Student’s t-test. g. Hematoxylin and eosin staining of mouse liver sections from ven/aza/eto combination compared to vehicle control - 10x and 20x magnification. Images representative of 3 individual mice per treatment group.
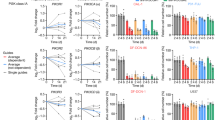
Extended Data Fig. 7 Transcriptomic analysis reveals fatty acid alterations in venetoclax resistant cells.
Transcriptional analysis of CPT1 and fatty acid metabolism in multiple data sets and patients that progress on ven/aza. a, b. Analysis of CPT1 in paired diagnosis versus relapse LSCs. N = 11 paired patient specimens. a. Overall expression compared between diagnosis and relapse. b. Expression between paired specimens with lines linking same patient. Data are presented as mean values + /- SD. Data are presented as median with lower hinges and upper hinges at 25th and 75th percentile with upper whisker to largest value and lower whisker to smallest value. c. Analysis of diagnosis and relapse paired LSCs for CPT1a. N = 35 sorted cell populations from 12 patients with following subpopulation numbers: N = 6 HSC cell populations, N = 5 non -lsc cell populations, N = 9 DX LSC cell populations, and N = 15 RI LSC cell populations. Data are presented as median with lower hinges and upper hinges at 25th and 75th percentile with upper whisker to largest value and lower whisker to smallest value d. Flow cytometric measurement of ACADVL in RAS mutant and RAS WT patient samples. N = 4 mutant versus N = 4 wildtype patient specimens e. Transcript levels of CD36 and ACADVL from Beat AML database for RAS mutant and RAS WT patient samples. N = 21 mutant patient specimens versus N = 256 wildtype patient specimens. Data are presented as mean values + /- SEMF. Flow cytometry of ven/aza sensitive and resistant LSCs for CD36. N = 3 sensitive versus N = 5 resistant patient specimens. Data are presented as mean values + /- SD. g. Enrichment plot of fatty acid metabolism from University of Colorado ven/aza patient. N = 9 patients (6 responders/3 progressors). LSCs shown to be enriched in patients that progress on therapy. h. UMAP projection of CITE Seq data of 6 patient specimens including n = 3 ven+aza clinical non responders and n = 3 ven+aza responders and blast cluster identification on UMAP projection. i. UMAP projection colored for CD36 antibody level of 6 patient specimens.
Supplementary information
Supplementary Tables
Supplementary Tables 1 and 2.
Supplementary Data 1
Gating strategy.
Source data
Source Data Fig. 1
Numerical data.
Source Data Fig. 2
Numerical data.
Source Data Fig. 3
Numerical data.
Source Data Fig. 4
Numerical data.
Source Data Fig. 5
Numerical data.
Source Data Extended Data Fig. 1
Numerical data.
Source Data Extended Data Fig. 2
Numerical data.
Source Data Extended Data Fig. 3
Numerical data.
Source Data Extended Data Fig. 4
Numerical data.
Source Data Extended Data Fig. 5
Numerical data.
Source Data Extended Data Fig. 5
Uncut westerns
Source Data Extended Data Fig. 6
Numerical data.
Source Data Extended Data Fig. 7
Numerical data.
Rights and permissions
About this article
Cite this article
Stevens, B.M., Jones, C.L., Pollyea, D.A. et al. Fatty acid metabolism underlies venetoclax resistance in acute myeloid leukemia stem cells. Nat Cancer 1, 1176–1187 (2020). https://doi.org/10.1038/s43018-020-00126-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43018-020-00126-z
This article is cited by
-
C/EBPα-p30 confers AML cell susceptibility to the terminal unfolded protein response and resistance to Venetoclax by activating DDIT3 transcription
Journal of Experimental & Clinical Cancer Research (2024)
-
Venetoclax efficacy on acute myeloid leukemia is enhanced by the combination with butyrate
Scientific Reports (2024)
-
ATP1A1/BCL2L1 predicts the response of myelomonocytic and monocytic acute myeloid leukemia to cardiac glycosides
Leukemia (2024)
-
SMS121, a new inhibitor of CD36, impairs fatty acid uptake and viability of acute myeloid leukemia
Scientific Reports (2024)
-
Hematopoietic stem cells with granulo-monocytic differentiation state overcome venetoclax sensitivity in patients with myelodysplastic syndromes
Nature Communications (2024)