Abstract
The quantification of animal social behaviour is an essential step to reveal brain functions and psychiatric disorders during interaction phases. While deep learning-based approaches have enabled precise pose estimation, identification and behavioural classification of multi-animals, their application is challenged by the lack of well-annotated datasets. Here we show a computational framework, the Social Behavior Atlas (SBeA) used to overcome the problem caused by the limited datasets. SBeA uses a much smaller number of labelled frames for multi-animal three-dimensional pose estimation, achieves label-free identification recognition and successfully applies unsupervised dynamic learning to social behaviour classification. SBeA is validated to uncover previously overlooked social behaviour phenotypes of autism spectrum disorder knockout mice. Our results also demonstrate that the SBeA can achieve high performance across various species using existing customized datasets. These findings highlight the potential of SBeA for quantifying subtle social behaviours in the fields of neuroscience and ecology.
Similar content being viewed by others
Main
Animal modelling is instrumental in human social disorder studies. However, failures to capture their specific behavioural biomarkers impede our understanding1. The biggest challenge to deciphering animal social behaviour is intraspecific appearance resemblance2. One direct way to distinguish their identities is through body markers such as radio-frequency identification devices3,4. Another way is combining depth information with red-green-blue images to reduce the identification error caused by body occlusion5. Recently, deep learning-based multi-animal tracking approaches, such as multi-animal DeepLabCut6, SLEAP7 and AlphaTracker8, have been avoiding the dependency of body markers or depth information. They maintain animal identities by using big-data features of continuous locomotion or appearances. Although these advances in deep learning multi-animal pose estimation6,7, identity recognition6,7,9,10 and behaviour classification11 have shown good performance in social behaviour analysis, their application across various experimental scenarios is limited by the availability of high-quality benchmark datasets2,6,7,9,12.
The model’s performance of multi-animal pose estimation is decided by the number of labelled frames7. Although there are several well-annotated datasets for multi-animal pose estimation6,7, they cannot cover diverse social behaviour test models. The frequent occlusion of multiple animals is a challenge for manual data annotations. The model’s performance would decrease because manual labels of occluded frames are not precise. Combining a multiview camera array with three-dimensional (3D) reconstruction technology can improve the pose estimation precision when facing occlusions13, but these methods are designed for a single animal rather than for multiple animals13,14.
Performances of image-based animal identification methods are also restricted by data annotations9,10. Animals have similar appearances, making it difficult to distinguish their identities when annotating identity datasets9. Unsupervised tracking-based methods are the alternative solutions to animal identification6,7. They demonstrate high performance when the animals are a relatively long distance away from each other, but the close interaction of animals can cause an identity swap problem2. This frequent close interaction means these methods cannot maintain identities for a long time period2.
New abnormal social behaviour patterns from animal disease models cannot be covered by existing behavioural classification datasets. Some subsecond behaviours are casually ignored in labelling because they are too short13. This means supervised behaviour classification methods are not suitable for detecting unusual behaviours9. Recent advances in unsupervised behaviour classification methods are appropriate for revealing subtle behavioural differences13,15, but they are only designed for a single animal. AlphaTracker is designed for the unsupervised clustering of social behaviour using human-defined features12, but these features cannot distinguish the subtle interactions constructed by limbs and paws.
To address these challenges, we propose the Social Behavior Atlas (SBeA), a few-shot learning framework for multi-animal 3D pose estimation, identity recognition and social behaviour classification. We propose a continuous occlusion copy-and-paste algorithm (COCA) for data augmentation in SBeA, combined with a multiview camera, to achieve multi-animal 3D social pose estimation with a few data annotations (roughly 400 frames)16,17. We propose a bidirectional transfer learning identity recognition strategy, achieving zero-shot annotation of multi-animal identity recognition with an accuracy rate exceeding 90% (refs. 18,19,20). We extend the Behaviour Atlas, an unsupervised behaviour decomposition framework, from a single animal to multiple animals, which achieves unsupervised fine-grained social behaviour module clustering with a purity exceeding 80% (refs. 13,21,22). In a study of free-social behaviour between the autism model and normal animals, SBeA enables automatic identification of animals with social abnormalities and explores the precise characteristics of these abnormal social behaviours. It demonstrates that SBeA can be an availably quantitative tool for studying animal social behaviour. SBeA can be applied to mice, parrots and Belgian Malinois dogs, showcasing its generalization abilities suitable for various application scenarios.
Results
SBeA: multi-animal 3D pose tracking and social behaviour mapping
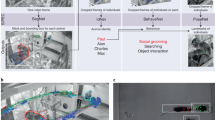
SBeA aims to quantify the behaviour of freely social animals comprehensively. It presents two substantial challenges: pose tracking and behaviour mapping. Pose tracking involves identifying key body parts of each animal and their identities, which is particularly challenging when animals look similar2. To address this issue, a free-social behaviour test model is developed that involves a multiview camera array (Fig. 1a). This approach covers more view angles of animals and helps to overcome the challenge of frequent occlusion13,14,21,22. The camera array is used to capture images of a chequerboard for camera calibration, followed by videos of two free-moving animals for the social behaviour test (video capture phase 1, Fig. 1a). Finally, the array captures videos of single free-moving animals for identification (video capture phase 2, Fig. 1a).
a, Video acquisition for the free-social behaviour test. The camera array is used for behavioural capturing and it is calibrated by the chessboard images. There are two phases for behavioural video capturing including social behaviour test and animal digital identity. Phase 1 captures the videos of free-social interactions of two mice. Phase 2 captures the identities of each mouse in phase 1. b, Data annotation for AI training. The SBeA needs the annotations of multi-animal contour and single-animal pose. c, The multistage artificial neural networks for 3D pose tracking. d, The outputs of 3D pose tracking. The left shows the outputs of AI including video instances, multi-animal poses and multi-animal identities. The centre shows the combination of video instances, multi-animal poses and multi-animal identities with camera calibration parameters for 3D reconstruction with identities. The right shows the visualization of 3D poses with identities. e, Parallel dynamic decomposition of body trajectories. Raw 3D trajectories of two animals can be decomposed into locomotion, non-locomotor movement and body distance. After dynamical temporal decomposition, these three parts are merged as social behaviour motifs for behavioural mapping. f, Social behaviour metric. Social behaviour motifs are clustered and phenotyped according to the distribution in social behaviour space. M1, mouse 1. M2, mouse 2. Mp, mouse with index p. Mq, mouse with index q. Mn, mouse with index n.
After the video acquisition, the multi-animal contour of video capture phase 1 and the single-animal pose of video capture phase 2 are manually annotated for the training of artificial intelligence (AI) to output the 3D poses with identities of animals (Fig. 1b,c). Through these multistage networks, the tasks of multi-animal video instance segmentation (VIS), pose estimation and identity recognition were achieved with a relatively small number of manual annotations (Fig. 1d, left). By incorporating camera parameters, the above results from various camera angles were matched on the basis of geometric constraints to reconstruct 3D pose trajectories with identities for each animal (Fig. 1d, centre and right).
The process of behaviour mapping involves breaking down the trajectories of animals into distinct behaviour modules and obtaining a low-dimensional representation of them13. 3D trajectories are separately decomposed into locomotion, non-locomotor movement and body distance components (Fig. 1e, top and middle). These parallel components are then divided into segments and subsequently merged into social behavioural modules using the dynamic behaviour metric (Fig. 1e, bottom). To gain insight into the distribution of features within social behavioural modules, it is necessary to convert them into low-dimensional representations (Fig. 1f). These representations incorporate both spatial and temporal aspects, with the spatial aspect being captured by low-dimensional embeddings of distance features in the SBeA framework (Fig. 1f, left). The temporal aspect is represented by the social ethogram (Fig. 1f, right). This approach allows for a more comprehensive understanding of the distribution of features within social behavioural modules.
A general augmenter for multi-animal pose estimation
The flexible social interactions among animals challenge the creating of a comprehensive training dataset for deep learning-based pose estimation methods. Inadequately trained deep neural networks tend to produce higher tracking errors, particularly in frames with close animal interactions2. To address this issue, we introduce a general data augmenter COCA (Fig. 2a) in SBeA. Previous studies show that image copy-paste can increase the precision of instance segmentation and multi-object tracking16,17, which inspires the development of COCA.
a, Concept diagram of COCA. From the raw scenario, the instances of background and animals can be synthesized with occlusion in a new combination. That achieves generation of big data from small data. b, Video capture of two free-moving animals. Two animals are put in the transparent circular open field and the video streams of behaviour are captured by a camera array. c, COCA as a general augmenter for multi-animal patching according to a little manually labelled data. Behavioural video streams are separated into backgrounds (top left), trajectories (medium left) and manually labelled masks (bottom left). Self-training instance segmentation model is used to predict more unlabelled masks from manually labelled masks. They are then combined with backgrounds and trajectories to generate new scenarios of two free-moving mice. d, Mask and pose prediction. Spatial-temporal learning is used for the new scenarios and to predict the masks of real mouse instances. Then, the single-animal pose estimation model can be used for each animal and, further, the 2D poses of them are merged to achieve multi-animal pose estimation. e, 3D pose reconstruction. The camera array is calibrated by chessboard images using Zhang’s calibration. Reprojection errors of all combination pairs of 2D poses of each animal are optimized for 3D reconstruction. The top right shows a 3D view of the 3D poses of two mice in this case. The bottom right shows a 2D view of the 3D poses of two mice. f, Comparison of the number of manually labelled points of SBeA and maDLC. g, Distance distribution of two free-moving mice. Pink stems are distance boundaries clustered by k-means (close 60.69, interim 195.03, far 327.47). h, Prediction error comparison of all validation data. The differences between all and close data are about ±2 pixels (two-way ANOVA followed by a Sidak multiple comparisons test, n1 (All) = 14,400, n2 (Close) = 4,602: the adjusted P values from nose to tip of tail are <0.0001, 0.0023, <0.0001, 0.0369, 0.1049, 0.0590, 0.0002, <0.0001, <0.0001, 0.2068, 0.0026, 0.0013, 0.4167, <0.0001, <0.0001 and <0.0001). Stems represent the mean values of each violin plot. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. NS, not significant.
Overlap of animals during social behaviour leads to loss of tracking in the single-view camera. To address this, SBeA uses a multiview camera array to capture video streams, enabling compensation for the visual field of cameras (Fig. 2b)13,14,22. Then, background and trajectories are extracted (Fig. 2c, left top and left middle), and frames with close social interactions are extracted for manual contour annotations (Fig. 2c, left bottom). YOLACT++ is trained by self-training using approximately 400–800 annotated contour frames (Fig. 2c, centre bottom), which enhances its performance while ensuring time-efficiency23,24. The well-trained YOLACT++ predicts masks and crops the animal instances from video streams. As some trajectories of multiple animals can overlap in the same spatial position across different periods, merging animal instances, backgrounds, trajectories and masks can generate virtual scenarios with various occlusion relationships (Fig. 2c, centre top and centre middle). The COCA increases the scale of the training dataset without vast manual annotations, producing a VIS dataset with successive frames of behaving animals and annotations. To capture the spatial-temporal patterns of occluded animals, the VIS with transformers (VisTR) method is modified and applied to the VIS dataset (Fig. 2c, right top)25. Well-trained VisTR can patch raw video streams to display only one animal per video (Fig. 2d, left top and left middle). Thus, pose estimation models trained for single animals can be used to predict single-animal poses (Fig. 2c, right bottom, and Fig. 2d, left bottom). Finally, the single-animal poses are merged into multi-animal poses (Fig. 2d, left top, middle and bottom).
The subsequent step is the 3D reconstruction (Fig. 2e). The MouseVenue3D system is used to acquire camera parameters (Fig. 2e, left top)14,22. On the basis of the epipolar constraint of camera parameters, the combination of each animal instance in each camera view is optimized to achieve minimum reprojection error (Fig. 2e, left bottom). In the 3D skeleton, the close contact between two animals can be quantified (Fig. 2e, right top and bottom).
The pose annotation strategy in SBeA linearly increases with body points and the number of animals compared with the square increase of maDLC6 and SLEAP7 (Fig. 2f). We then create a well-annotated dataset Social Black Mice for VIS (SBM-VIS) to compare the tracking performance of SBeA with other methods. The close interaction of the test dataset is separated according to the distance distribution (Fig. 2g, the left orange stem). The pixel root-mean-square error (r.m.s.e.) of all data is significantly lower than the close interaction of about 2 pixels of different body parts (Fig. 2h). But, compared with maDLC and SLEAP, SBeA still has significantly lower r.m.s.e. of animal close interaction (Extended Data Fig. 1 and Extended Data Fig. 2). For all the test data, SBeA achieves equivalent or lower r.m.s.e. (Extended Data Fig. 1a and Extended Data Fig. 2a). For the close contact part, most of the r.m.s.e. of SBeA are significantly lower than maDLC (Extended Data Fig. 1b), and SBeA has significantly lower r.m.s.e. than SLEAP except for the neck (Extended Data Fig. 2b). These results show that SBeA can get higher precision with fewer manual annotations than routine multi-animal pose estimation methods.
SBeA needs no annotations for multi-animal identification
Accurately distinguishing the identities of free-moving animals is crucial for social behaviour tests, particularly in studying treatment-induced behaviours in transgenic animal models13,26,27. However, their frequent occlusion leads to imprecise identification in manual labelling, especially for the same breed animals. To address these challenges, we propose bidirectional transfer learning in SBeA (Fig. 3a). Transfer learning allows artificial neural networks to use previous knowledge in new tasks19. For animal segmentation and identification tasks, the knowledge between them can be shared and transferred bidirectionally with each other. So, the bidirectional transfer learning of them avoids unnecessary manual data annotations.
a, Concept diagram of bidirectional transfer learning-based animal identification. The well-trained segmentation model on multi-animals can be transferred to the single-animal videos, and the well-trained identity recognition model on the single animal can also be transferred to multi-animals. The transfer learning of two models reduces unnecessary manual annotations of animal identities. b, Segmentation model reuse. The left shows an animal being put in the transparent circular open field and the video streams are captured by a camera array. The centre shows the well-trained VisTR is reused for the single animal. The right shows the output of well-trained VisTR on the single animal. c, Single-animal identification model training. The left shows the single-animal instances of multiview are cropped, cascaded and resized to an image. The centre shows the use of EfficientNet as the backbone to train the multi-animal identification classifier. The right shows the identity recognition pattern visualization by LayerCAM. d, Multi-animal segmentation with 3D reprojection. The left shows mask reprojection of each camera view. The right shows the crop, cascade and resize of two animal instances from matched camera view angles. e, Identification model reuse. The well-trained identification model on the single animal can be reused in multi-animal identification. f, Confusion matrix of single-animal identification. g, Feature representation of single-animal identification using t-SNE. h, The sorted validation precision of f. i, The sorted silhouette coefficient of g (mean ± s.d., one-way ANOVA with Dunn’s multiple comparisons test, n = 60, adjusted P values from bottom (M2) to top (M7) are 0.0684, 0.0415, >0.9999, <0.0001, >0.9999, <0.0001, <0.0001, <0.0001 and <0.0001). j, The manual validation precision of multi-animal identification. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
Well-trained VisTR can be used to segment single-animal instances from multiple view angles (Fig. 3b). These instances are then cropped, cascaded and resized to generate training data for an identification model (Fig. 3c, left and centre)28. After that, LayerCAM (where CAM stands for class activation maps) is used to evaluate the patterns for identification recognition (Fig. 3c, right)29. Before using the identification model in multi-animal instances, the cascaded and resized image frames were prepared (Fig. 3d, right). By using the geometric constraint of 3D poses, instances from each frame view angle of each animal were matched to construct input frames of the identification model (Fig. 3d, left). Finally, the well-trained model output the top prediction probabilities to append the identities of instances and 3D poses with the visualization of LayerCAM (Fig. 3e).
To evaluate the identification performance of SBeA, we conducted experiments with ten mice. The first 4 minutes of videos were used for training the identification model, and the last minute was used for validation. The validation confusion matrix demonstrated that the model can identify most of the mice (Fig. 3f). The t-SNE (t-distributed stochastic neighbour embedding) was used to create a two-dimensional (2D) feature representation of the identified mice (Fig. 3g). The features of mice with ID M4 and M5 were found to be mixed with other classes, as quantified by the silhouette coefficient (Fig. 3i). The statistical analysis of silhouette coefficient demonstrates that even the outlier could reduce the silhouette coefficient, such as M2 and M3, the precision would not be influenced too much (Fig. 3g–i).
To assess the identification model’s performance in multi-animal data, we recorded the free-social behaviours using the above mice. We manually verified their identities of mask reprojection images and 3D poses frame by frame (Fig. 3j). Although some of the single mouse identity precisions were lower (Fig. 3i), the overall precision in identifying pairs of mice could be higher than 0.85, as seen in the case of the pairs of M3 and M4 and M5 and M6.
Unsupervised learning reveals social behavioural structures
Following pose tracking, mapping the trajectories with animal identities to a low-dimensional space is necessary to gain insights into behaviour (Fig. 4a). We expand our previous work on the single-animal behaviour mapping framework, Behaviour Atlas (BeA), to encompass multiple animals13. The parallel and dynamic behaviour decomposition from BeA is adopted in SBeA (Fig. 4b,c). In the social process, the distance between animals is an essential component30, which models body position with non-locomotor movement and locomotion (Fig. 4b). Then, each component is decomposed by dynamic time alignment kernel (DTAK)13 to retain the dynamic structures of behaviour (Fig. 4c). To distinguish subtle structures of social behaviour, the temporal points of decomposition for each component are merged through logical addition (Fig. 4d). These steps enable the metric of social behaviour, resulting in the transformation of continuous pose trajectories into discrete social behaviour modules.
a, The 3D trajectories of two animals. b, The parallel decomposition of trajectories. The top shows non-locomotor movement, the middle shows locomotion and the bottom shows the distance. c, The dynamic decomposition after parallel decomposition using DTAK. The horizontal and the vertical axis of the DTAK matrix are the time. The values of the DTAK matrix are the similarities between the trajectories over time. The yellow boxes are the optimized results of the DTAK matrix, which groups trajectories with similar DTAK patterns as a dynamic segment. The sections of the grey bar indicate the duration of each dynamic segment. d–f, Social behaviour metric after dynamic decomposition. d, Decomposed segments merging. e, Feature representation of segments. The left shows dimensional reduction of distance dynamics. The right shows the ResMLP for feature refining. f, SBeA construction. The adaptive watershed is used for clustering. Coloured dots represent large clusters and areas enclosed by grey lines represent subclusters. g, Social behaviour cases clustered in the SBeA. h–l, The performance quantification of SBeA on the PAIR-R24M dataset. h, The visualization of two mice in the PAIR-R24M dataset. i, The SBeA of the PAIR-R24M dataset. The social classes of the PAIR-R24M dataset are separated in the SBeA. The ellipse is the Gaussian model fitting of the three classes. j, The SBeA of all the class labels of the PAIR-R24M dataset. The 11 classes of each mouse are combined into 121 classes, and the 121 classes are distributed with patterns. k, The distance map of the SBeA. The distance distribution of the distance map is coincident with labels in i. l, The cluster purity of social classes: mean ± s.d., n (from 1 to 14) = 26, 26, 26, 26, 26, 26, 26, 26, 26, 26, 26, 24, 23 and 18.
Next, the social behaviour modules are embedded in a low-dimensional space for behaviour representation (Fig. 4e,f). The distance component is chosen for the feature representation of social behaviour modules to keep the social information (Fig. 4e, left). The dimensionally reduced distance component by uniform manifold approximation and projection (UMAP) is beneficial to improve the separation of behaviour atlas13,14,21,22,31. However, with the increase in data scale, UMAP would be unacceptable because of limited memory space. The residual multilayer perceptron (ResMLP) is combined with UMAP for a common feature representation to solve the memory problem (Fig. 4e, right)32. The distance dynamics are embedded by DTAK and UMAP to construct the SBeA (Fig. 4f). To reveal the distributions of different social behaviour modules, we modify the watershed algorithm to automatically determine the best cluster density with upper and lower boundaries. Finally, the social behaviour modules of the same clusters are manually identified and defined (Fig. 4g).
We conduct supervised validation of SBeA using the PAIR-R24M dataset (Fig. 4h)33. We use SBeA to construct the SBeA for the dataset, and append the three social labels (close, chase and explore) defined in the PAIR-R24M dataset (Fig. 4i). The distributions of the three social labels are separated and match their similarity relationship. The 121 combinations of subject behaviour labels also show distribution patterns in the SBeA (Fig. 4j). The social labels such as close and explore are consistent with the close distance distribution in the distance map, and the chase label is consistent with the distance transition zone of the distance map (Fig. 4k). To quantify the clustering performance, we use the cluster purity of social and subject behaviour labels (Fig. 4l and Supplementary Fig. 7). For the upper boundary of clustering, 14 classes are clustered with a mean cluster purity of 0.77 ± 0.16 (Fig. 4l). For the lower boundary of clustering, 405 classes are clustered and the probability of cluster purities greater than 0.95 is significantly higher than for other purities (Supplementary Fig. 7). These results indicate that SBeA can classify the behaviour clusters with high cluster purity.
SBeA identifies free-social Shank3B knockout mice
Social behaviour can serve as an indicator of genetic variations that underlie neuropsychiatric disorders34. SBeA is well-suited for this purpose, as it allows for a detailed characterization of social behaviour at an atlas level. To test whether SBeA could detect genetic differences from social behaviour, we used an animal model of autism spectrum disorder: Shank3B knockout (KO) mice13,26. While abnormal individual behaviours of these mice have been previously identified, the limitations of existing techniques have made it difficult to fully understand their abnormal free-social behaviours13,26.
The SBeA with the distance map is shown in Supplementary Fig. 9b. The density map is calculated to compare the social behaviour distribution of each group (Supplementary Fig. 9c). The density map shows obvious differences across the three groups. The wild-type (WT)–WT group shows social behaviour phenotypes with flexible distances from close to far, the KO–KO group shows more abnormal social behaviours than the WT–WT group and the WT–KO group shows more close social interaction than the WT–WT group.
The 260 identified social behaviour modules were clustered to reveal their coincident patterns (Fig. 5a). Principal component analysis (PCA) was used to determine the percentage variability explained by each principal component to compare the three groups (Fig. 5b). The results indicated that three components could account for 90% of the variance, while 11 components could account for 99% of the variance. Further, UMAP was used to construct the phenotype space according to the social behaviour modules, with the dimension number set to three based on the 90% variance explanation, owing to the more robust feature representation of nonlinear dimensional reduction (Fig. 5c). The distributions of the three groups in the phenotype space were found to be segregated, matching the distribution of the density map (Supplementary Fig. 9c).
a, The fractions of social behavioural modules of three social groups. The fractions of each group are normalized, and they are clustered and re-sorted according to the dimension of social behaviour modules. b, Dimensional reduction of behaviour fractions using PCA after hypothesis testing (two-way ANOVA followed by the Tukey multiple comparisons test). In the three groups, 24 social behaviour modules show significant differences. Three components can explain more than 90% variances, and 11 components can explain more than 99% variances. c, The construction of phenotype space. UMAP is used to reduce the 260 dimensions of social behaviour modules to three dimensions according to e. Different coloured dots represent different social groups. The phenotypes of three social groups can be separated in phenotype space. d, The merging of social behaviour modules according to behavioural feature angles and b. First, 24 social behaviour modules with significant differences are mapped to PCA feature space, and then the angular separation is calculated to construct the angle spectrum. Further, hierarchical clustering is used to cluster the angle spectrum into 11 clusters according to b. e, The comparison of behavioural fractions of three social groups: 24 social behaviour modules with significant differences are manually identified (mean ± s.d., two-way ANOVA followed by Tukey multiple comparisons test, n = 20, adjusted P values from left to right (group A versus group B, group A versus group C and group B versus group C) are >0.9999, <0.0001, <0.0001, 0.9990, <0.0001, <0.0001, >0.9999, <0.0001, <0.0001, 0.9939, 0.0002, <0.0001, >0.9999, <0.0001, <0.0001, 0.9919, 0.0016, 0.0010, 0.8210, 0.0055, 0.0331, 0.0001, 0.0029, 0.7179, 0.8213, 0.0438, 0.1703, 0.9882, <0.0001, <0.0001, 0.2691, 0.0323, 0.6000, 0.5677, 0.0952, 0.0057, 0.6034, 0.0101, 0.1239, 0.0733, 0.0145, 0.8183, 0.2698, 0.0184, 0.4735, 0.1217, 0.0011, 0.2511, 0.4016, 0.0397, 0.4864, 0.5691, 0.0001, 0.0054, 0.0005, <0.0001, <0.0001, 0.6728, 0.0297, 0.2076, 0.0175, 0.7233, 0.1220, 0.0445, 0.2145, 0.7555, 0.0222, 0.4986, 0.2810, 0.1373, 0.8823 and 0.0454). f, The visualization of merged social behaviour modules. With the assistance of d, nine social behaviour modules are merged and identified from 24 social behaviour modules. The colour of mice represents the behaviour cases with the highest mean fraction in e. The orange 3D mice represent KO mice and green 3D mice represent WT mice. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
Further, SBeA was used to identify subtle social behaviour modules that distinguish KO and WT mice and 24 social behaviour modules showed significant differences (Fig. 5e). The angle spectrum clustering was proposed and used to reduce the redundancy of these results (Fig. 5d). The social behaviour modules were merged on the basis of their angular separation of features, resulting in the human identification of nine social behaviours (Fig. 5f and Extended Data Table 1).
The nine social behaviours highlighted significant differences among the three groups. The WT–WT group exhibited more allogrooming, a prosocial behaviour, than the WT–KO and KO–KO groups35. Conversely, allogrooming was rare in unstressed partners and even rarer in Shank3B KO mice, suggesting an antisocial behavioural phenotype36. The exploring behaviour of the WT–WT group was significantly higher than that of the KO–KO group, which displayed reduced motor ability or social novelty13,26. In the WT–KO group, social behaviour with significant differences was divided into two parts, namely peer sniffing and independent grooming. Peer sniffing was observed more frequently in the WT mouse, particularly when the KO mouse was grooming or in locomotion, indicating a behavioural phenotype of curiosity. Furthermore, the KO mouse could induce higher interest in the WT mouse than vice versa. Independent grooming could be an imitation of the WT mouse by the KO mouse, and in the KO–KO groups, the higher incidence of independent grooming could be attributed to the increased individual grooming of each mouse. In addition to increased independent grooming, two abnormal behaviour phenotypes, namely synchronous behaviours and two kinds of immobility, were observed. The synchronous behaviours displayed five subtypes, including grooming, hunching, rearing, sniffing and micromovement, indicating greater behaviour variability in free-social conditions compared to individual spontaneous behaviour of KO mice13. These findings demonstrate that SBeA can differentiate genetic mutant animals on the basis of social behaviour and identify genetic mutant-related subtle social behaviour modules.
SBeA is robust across species in different environments
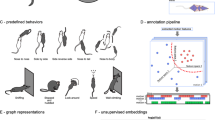
To assess the generalizability of SBeA to different animal species and experimental settings, the behaviours of birds and dogs were captured with varying device configurations22. The animals were prepared to have as similar appearances as possible (Fig. 6a,e, top), and it was difficult for human experimenters to separate two animals from the randomly selected frames. Videos were manually annotated to train the AI of the pose tracking component of SBeA (Fig. 6a,e, bottom), using 19 body parts for birds and 17 body parts for dogs, based on previous studies37,38 (Fig. 6b,f). Then, they were mapped to the social ethogram and behaviour atlas (Fig. 6c,g). In total, 34 and 15 social behaviour classes were identified for birds and dogs, respectively, and their typical cases were visualized in 3D (Fig. 6d,h). The 3D pose tracking of birds showed clear identification of their claw touching their rectrix, while the 3D pose tracking of dogs was robust to occlusion even in the lying posture.
a–d, SBeA is used for birds. a, The preparation of birds. Two parrots with inconspicuous appearance differences are used for the social behaviour test. After video recording of identity and free-social behaviour by camera array, the contours and poses are manually annotated, then 19 body parts are defined for 3D pose tracking. b, The social poses and identities outputs of SBeA. c, The social ethogram and SBeA of birds. d, The 3D social behaviour cases of birds. e–h, SBeA is used for dogs. e, The preparation of dogs. Two Belgian Malinois with inconspicuous appearance differences are used for the social behaviour test. After video recording of identity and free-social behaviour by camera array, the contours and poses are manually annotated: 17 body parts are defined for 3D pose tracking. f, The social poses and identities outputs of SBeA. g, The social ethogram and SBeA of dogs. h, The 3D social behaviour cases of dogs.
Discussion
SBeA is a few-shot learning framework for 3D pose estimation, identification and behaviour embedding of multiple free-social animals. It builds on the BeA framework, extending it to enable multi-animal pose estimation and social behaviour clustering13,14,21,22. SBeA reduces the labour required for annotation of pose estimation and identification6,7,9. It also overcomes the issue of occlusion and reconstructs 3D behaviours accurately using a camera array. SBeA resolves the challenge of animal identification over extended frames, facilitating the study of close social interactions2. The framework is versatile and has been successfully applied to Shank3B KO mice, where it has revealed abnormal social behaviours and a reduction in social interest. SBeA’s cross-species application has been verified in birds and dogs. In summary, SBeA represents a breakthrough in deep learning-based pose estimation and identification, offering numerous potential applications in animal behaviour research.
Although the benchmark datasets are critical to the advances in deep learning tools6, the large labelled data number could render them unfeasible20. SBeA gets rid of the dependency on large datasets and achieves results by only using hundreds of labelled frames to track 3D poses and identities of multiple animals in millions of new frames. Recent studies have shown the precision increasing of large transformer models in human pose estimation39,40, but the benchmark datasets of animals are still too small to apply them6. The data generation strategy in SBeA can be a bridge between small animal datasets and large models. The phenotypes of social behaviour are diverse, which are difficult to comprehensively predefine in a dataset13,33,41. The unsupervised clustering in SBeA provides an unbiased way to classify undefined social behaviour modules and supports the building of a comprehensive social behaviour dataset.
maDLC and SLEAP are two excellent tools that can be applied to many animal models6,7, but they do not include the mechanism for maintaining animal identities during long-term experiments, which influences the accuracy of building a behavioural representation relying on animal identities2. SBeA incorporates the identity recognition approach of idTracker.ai and TRex, using deep neural networks to directly learn the appearance features of animals10,42. This results in the alleviation of the identity swap problem, which can detect frames with higher error rates. Additionally, SBeA provides an extension of 2D tracking tools to 3D tracking, which is critical for making accurate inferences about animal behaviour2,14,22.
One potential area for future research to improve SBeA is to develop an end-to-end model that can reduce storage consumption. The identity videos available in this context may contain sufficient information to train a deep learning model for tasks such as multi-animal segmentation, identification and pose estimation. Furthermore, the behaviour atlas of a single animal could be combined with a SBeA of multiple animals. An algorithmic bridge from BeA to SBeA could facilitate not only social behaviour analysis, but also other forms of analysis within the field.
Methods
Experiments of mice, birds and dogs
There are four experiments in this study. The first is the free-social behaviour test of two WT mice for the program design of SBeA. In total, 32 adult male C57BL/6 mice (7–12 weeks old) are used for the free-social behaviour test. The mice were housed at 4–5 mice per cage under a 12 h light–dark cycle at 22–25 °C with 40–70% humidity, and were allowed to access water and food ad libitum (Shenzhen Institutes of Advanced Technology, Shenzhen, China). Before the social behaviour test, the mice had tail tags added using a black marker pen. The tail tags were constructed of horizontal and vertical lines. The horizontal line represented one, and the vertical line represented five. Using the combination of horizontal and vertical lines, the mice were marked according to the sequence of the experiment. After that, the mice were put into a circular open field made of a transparent acrylic wall and white plastic ground, with a base diameter of 50 or 20 cm and a height of 50 cm for 5 min or 15 min identity recording one by one using MouseVenue3D. Then, the mice were paired and put into the same circular open field for the free-social behaviour test.
The second test is the free-social behaviour test of mice with different genotypes. Five adult (8 weeks old) Shank3B KO (Shank3B−/−) mice on C57BL/6J genetic background and five adults (8 weeks old) male C57BL/6 mice, were used in the behavioural experiments. Shank3B−/− mice were obtained from the Jackson Laboratory (Jax catalogue no. 017688) and were described previously26. The mice were housed at 4–5 mice per cage under a 12 h light–dark cycle at 22–25 °C with 40–70% humidity, and were allowed to access water and food ad libitum (Shenzhen Institutes of Advanced Technology). The mice had tail tags added as mentioned above. After that, the mice were put into a circular open field with a base diameter of 20 cm introduced before for 5 min identity recording. Then the mice were paired in WT–WT, WT–KO and KO–KO groups and put into the same circular open field for the free-social behaviour test. The combinations of groups and the sequence of experiments were randomly generated by customized MATLAB code.
The third is the free-social behaviour test of two birds. One male and one female Melopsittacus undulatus (about 26 weeks old) were used in this experiment. They were housed in a conventional environment and fed regularly (Shenzhen Institutes of Advanced Technology, Shenzhen, China). The birds were first put into a circular open field with a base diameter of 20 cm for 5 min of identity recording one by one, and then put together for 15 min free-social behaviour test and recording.
The fourth is the free-social behaviour test of two dogs. Two female Belgian Malinois dogs (13 weeks old) were used in this experiment. They were housed in Kunming Police Dog Base of the Chinese Ministry of Public Security, and their behaviour test was finished in the State Key Laboratory of Genetic Resources and Evolution, Kunming Institute of Zoology, Chinese Academy of Sciences. The dogs were first put into a 2 × 2 m2 open field made by fences one by one for the identity recording. Restricted by the locomotion of the animals, only 6 and 11 min identity frames were captured by MouseVenue3D, and both of them were used for identification. Then, they were both put into the open field for a 15 min free-social behaviour test.
All husbandry and experimental procedures of mice and birds in this study were approved by the Animal Care and Use Committees at the Shenzhen Institute of Advanced Technology, Chinese Academy of Sciences. All husbandry and experimental procedures of dogs in this study were approved by the Animal Care and Use Committees at the Kunming Institute of Zoology, Chinese Academy of Sciences.
SBM-VIS dataset
The free-social behaviour of two C57BL/6 mice introduced above is captured by the first version of MouseVenue3D. The first 1 min frames of four cameras are annotated as the SBM-VIS dataset, which is 7,200 frames in total. To accelerate the data annotation, we used deep learning for assistance. Here, 30% of the contours are manually labelled, and the rest are labelled by YOLACT++ after being trained by the manually labelled 30% contours and then checked by humans. Next, the single-animal DeepLabCut is used to predict the poses of masked frames with a human check. Groups of 18 frames are gathered for a video instance and saved in YouTubeVIS format43, and the poses are saved as a .csv file. The identities across different cameras are corrected by human annotators. This SBM-VIS dataset is available in figshare44, and other data for method comparison reproduction are also available45.
New scenario generation for VIS
The new scenario generation for VIS is divided into several steps: contour extraction, trajectory extraction, dataset labelling, background calculation, model self-training and video dataset generation. After that, it can be input into the instance segmentation model for large-scale training. Suppose the number of animals in the video is n. Conda virtual environment configuration includes OpenCV v.4.5.5.62, Python v.3.8.12 and Pytorch v.1.10.1. The computer was configured with Intel(R) Xeon(R) Silver 4210 R CPU at 2.40 GHz and NVIDIA RTX3090 graphical processing unit (GPU).
In the animal contour step, image thresholding is first carried out and then the contour in the image is extracted. The following formula is used to determine whether the frame is social or not, where i stands for a frame, Ri stands for the judgement result of this frame and numi stands for the number of contours in this frame:
When extracting the animal trajectory, due to the influence of noise, all the contour centre points are recorded as the candidates of the animal frame centre point, and the closest point to each animal in the previous frame is selected from multiple centre points as the true centre point of this frame. Then, the Hungarian matching idea is used to remove the matching points successfully to optimize the animal trajectory.
For dataset annotation, different manually annotated datasets were used for different animals. We manually annotated 272 images in the 50 cm mice open field experiment, 805 images in the 20 cm mice open field experiment, 600 images in the birds experiment and 800 images in the dogs experiment.
For background calculation, the non-mask position (the background) of each image is extracted and fused into the final background image using the labelled dataset. The above operation is repeated for all datasets to obtain a clean background image.
The labelled dataset is used for YOLACT++ round training, and the trained model is used to predict video frames. The predicted high-quality frames will be added to the original dataset for the next round of training. Among them, the selection method for high-quality frames is as follows: i represents a certain frame, fi is the segmentation result of the frame i, \({f}_{i-1}\) is the segmentation result of the frame \(i-1\), F is the calculation process of scoring matrix of all segmentation results in two frames where the calculation idea refers to the Hungarian matching idea and the calculation result is Gi:
Then, all Gi are merged and clustered, and the class with the higher overall matrix score is selected as the high-quality frame class and added to the training dataset. YOLACT++ selects the ResNet50 model as the pretraining model, and the maximum number of iterations is 150,000 generations. The training process takes about 5 h. After YOLACT++ finishes training, its final model is used to predict the results for all frames.
The video dataset required for instance segmentation training is subsequently generated. The dataset is divided into three parts, which are the real dataset, the social area dataset and the randomly generated dataset. The real dataset is the continuous high-quality frames predicted and filtered by YOLACT++, which are written into the video dataset after data enhancement, where the data enhancement is performed by flipping the image left and right. Because there are many occlusions during social interaction and the performance of the model decreases, it is necessary to generate multiple datasets in the social area. Here, consecutive frames of animals in the social area are selected and augmented to generate the social area dataset, where N forms of enhancement are generated by data augmentation, as shown in equation (3), where C represents combination (that is, the combination of different masks is selected for flipping in each frame). A stands for alignment (that is, all masks are aligned to occlusion):
As the number of real data and social area datasets may be far from enough to complete the model training task, some datasets in the animal activity area are randomly generated after this step. In this part, the real animal trajectory in the video, the obtained animal mask and the background calculated in the previous step are used for data collection, and the video dataset is written after data enhancement. Here, 14,940 video datasets were generated for the 50 cm mice open field experiment, 15,130 for the 20 cm mice open field experiment, 5,970 for the bird experiment and 41,755 for the dog experiment.
3D pose reconstruction of multi-animals
Here, we use the multiview geometry method in computer vision for the 3D reconstruction of multiple animals. The basic projection formula between 2D points and 3D space points is as follows.
Here, s represents the scaling factor, x and y are the points in the image, K is the camera internal reference, R is the rotation matrix, t is the translation matri, and X, Y and Z represent the coordinates of the 3D points. First, all two-dimensional skeleton information about the multi-animal and multiview is read, and the points in the two-dimensional file with too low a confidence rate are directly set to null value. Then, the relative position parameters between multiple cameras are read and the triangulation algorithm is used for the 3D reconstruction of a single animal. The basic principle is as follows:
Here, \({\alpha }_{1}\) to \({\alpha }_{n}\) represent the two-dimensional points with the same content in different cameras, K1 to Kn represent the internal parameter matrix of different cameras, R1 to Rn represent the rotation matrix of different cameras, t1 to tn represent the translation matrix of different cameras and the three-dimensional point P can be solved by combining these equations, so we use the singular value decomposition to solve the least-squares regression problem.
Next, as the appearance of animals in different views is very similar, the identities of instance segmentation may be swapped and the wrong 3D point coordinates may be calculated. Therefore, we first obtain the full permutation index list of all 2D points of multiple animals in each view angle, and then obtain the 3D point coordinates in all cases. Eventually, the point with the smallest error is selected as the final multi-animal 3D skeleton point.
Pattern visualization of animal identification by LayerCAM
LayerCAM can generate the CAMs of each layer of convolutional neural network-based models29. The LayerCAM of each layer of the EfficientNet-based identity recognition network is averaged to output a global visualization pattern of animal identities. To further compare the feature weights of different body parts of animals, the 2D poses are used for the body part location of identity frames. From the 2D poses to identity frames, there is a coordinate transformation. The transformed 2D poses on identity frames Pt can be calculated as:
where Kr is the resized matrix of cascade frames, Kb is the scale matrix of the bounding box of single camera view, Pcam is the raw 2D poses, Bb is the bias matrix of the bounding box of single camera view and the index cam is the camera number. The Kb is decided by the size of frames and the bounding box size of the cropped animal instance. To reduce the disturbance of 2D pose estimation, a box centred on Pt of each transformed 2D pose crops the LayerCAM value. And the mean value of them represents the CAM weights of each body part.
Parallel decomposition of trajectories
The parallel decomposition of trajectories includes three parts.
The first part is the decomposition of non-locomotor movement. Let \({X}_{ij}^{m}\) be the behaviour trajectories of animals m with i frames and j dimensions, so the non-locomotor movement component YNM can be calculated as follows:
where J is all one vector, and N is the number of frames. After this step, the centre of the body of the animals can be aligned together.
The second part is the decomposition of locomotion. The locomotion component YL can be calculated as follows:
The third part is the decomposition of distance. The distance component YD can be calculated as follows:
Feature representation of distance dynamics
The distance dynamics YDD can be calculated as follows:
where fUMAP(·) is the UMAP mapping including the parameters n_neighbors set to 199, and min_dist set to 0.3, Ithres is the threshold of frames set to 200,000 and fResMLP(·) is the feature representation including ResMLP. For fResMLP(·), first, the YD is randomly sampled to YDs according to Ithres. And the rest of YD is YDr. Then, YDs and YDDs = fUMAP(YDs), the UMAP of YDs, is used to train ResMLP for feature encoding. After the training, the ResMLP predicts the YDDr from YDr, and the YDD can be recombined by YDDs and YDDr according to the sample point.
The ResMLP is based on the residual module and multilayer perceptron46,47. The residual block is constructed by a multilayer perceptron with two layers. Each layer has 64 neurons, and two residual blocks are stacked to construct the residual part. The head of ResMLP is one 1D convolution layer and one global max pooling layer for the feature encoding of distance dynamics48. The output part of ResMLP is constructed by one fully connected layer with one sigmoid layer for the continuous value representation49. The activation function of ResMLP uses rectified linear unit layers49. The optimizer of ResMLP is Adam, the initial learning rate is set to 0.001, the mini-batch size is set to 2,000 and the epoch number is set to 100 (ref. 50). The final r.m.s.e. of validation is 0.02–0.06, and the training time of ResMLP is about 4 min on NVIDIA GeForce RTX3090 GPU.
The time consumption comparison of ResMLP
After the manual time consumption test of UMAP, the quadratic function is used for the estimation time comparison. The coefficient of the quadratic function is 0.00002. The time consumption of ResMLP is estimated as a linear function with a slope set to 0.000008 and an intercept set to 240 based on the training and prediction time of ResMLP. The functions of the time consumption are as follows:
where TUMAP is the time consumption of UMAP, kUMAP is the coefficient of quadratic function, yD is the number of distance components, TResMLP is the time consumption of ResMLP, kResMLP is the slope of ResMLP and bResMLP is the intercept.
The distance map
Let YE be the low-dimensional embedding of the SBeA, and YDM be the distance of YE. The YDM can be calculated as follows:
where j is one of the points in YDM, p is the start time point of \({Y}_{\mathrm{DM}}^{\,j}\) and q is the end time point of \({Y}_{\mathrm{DM}}^{\,j}\).
The map to body distance
The body distance is equivalent to YDM. The map distance YEM can be calculated as follows:
where yE is one point of YE. The map to body distance YMB can be calculated as follows:
The adaptive watershed clustering
The variable of watershed clustering on 2D embeddings is the kernel bandwidth kb, which decides the density. The adaptive watershed clustering is designed to automatically choose the best density. The best density is determined by the stable number of clusters cst. To get cst, the clusters under certain kb are first calculated as:
where fWC(·) is the watershed clustering and cn is the number of clusters. Then, the cst is calculated as:
where fMode(·) is the mode function. The cs is the lower bound of watershed clustering with a larger kernel bandwidth. To improve the sensitivity of watershed clustering for the subtle differences of social behaviour, a threshold uthres is set to 0.9 to restrict kb in more fine grain. So, the number of sensitivity clusters cse can be calculated as:
where fMax(·) is the maximum function and fMin(·) is the minimum function. The cst and cse together determine the lower and upper bound of watershed clustering.
The cluster purity
The cluster purity is an indicator quantifying the uniformity of a cluster. Let \(P=\{{p}_{1},{p}_{2},\ldots ,{p}_{N}\}\) be the ground truth indexes of all data, the \(Q=\{{q}_{1},{q}_{2},\ldots ,{q}_{N}\}\) is the cluster indexes of all data and N is the number of clusters, so the cluster purity CP can be calculated as:
The cluster gram of grouped mice
To reveal the inherent patterns of behaviour fractions of each group, the cluster gram is first stacked group by group. Then, all the behaviour fractions are normalized according to the dimension of the subject and sorted by hierarchical clustering according to the dimension of the social behaviour module. The clustering tree is normalized for better visualization. Further, the behaviour fractions of each group are sorted according to Euclidean distance for the similarity metric. The initial row of each group for sorting is chosen by the maximum change rate Rm. The Rm can be calculated as:
where sm is the sorted social behaviour fractions by hierarchical clustering.
The angle spectrum clustering
The angle spectrum clustering is proposed and used to merge similar subclusters of behaviour in feature vector space. Let V be the feature vector matrix of social behaviour modules in PCA space, the angle spectrum As can be calculated as:
where v is one of the feature vectors in V. Then, the As is clustered by hierarchical clustering according to the 11 components of 99% variance explanation.
Statistics
Before hypothesis testing, data were first tested for normality by the Shapiro–Wilk normality test and for homoscedasticity by the F test. For normally distributed data with homogeneous variances, parametric tests were used; otherwise, non-parametric tests were used. All the analyses of variance (ANOVA) have been corrected by the recommended options of Prism v.8.0. No data in this work have been removed. All related data are included in the analysis.
The usage of ChatGPT
ChatGPT was used to improve the language of this paper. The authors confirm that all changes were carefully reviewed to ensure that no changes to the content of the paper occurred in this process.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Source data for result reproduction are provided with this paper. They are available in figshare with the hyperlink to the dataset (https://figshare.com/projects/Social_behavior_atlas/162718), and DOI of the dataset (https://doi.org/10.6084/m9.figshare.22314994.v1). The SBM-VIS dataset44 is available under https://doi.org/10.6084/m9.figshare.24597111.v1. The PAIR-R24M dataset33 is available under https://doi.org/10.6084/m9.figshare.14754374.v2
Code availability
We provide a code repository of SBeA at https://github.com/YNCris/SBeA_release (https://doi.org/10.5281/zenodo.8238067)51. This repository includes SBeA_tracker and SBeA_mapper. SBeA_tracker achieves 3D pose tracking, which has a software interface to guide its usage. SBeA_mapper achieves the atlas mapping of social behaviours from 3D pose trajectories with different configurations. It also contains the code to replicate the figures of this paper.
References
Stanley, D. A. & Adolphs, R. Toward a neural basis for social behavior. Neuron https://doi.org/10.1016/j.neuron.2013.10.038 (2013).
Agezo, S. & Berman, G. J. Tracking together: estimating social poses. Nat. Methods 19, 410–411 (2022).
Peleh, T., Bai, X., Kas, M. J. H. & Hengerer, B. RFID-supported video tracking for automated analysis of social behaviour in groups of mice. J. Neurosci. Methods 325, 108323 (2019).
de Chaumont, F. et al. Real-time analysis of the behaviour of groups of mice via a depth-sensing camera and machine learning. Nat. Biomed. Eng. 3, 930–942 (2019).
Ebbesen, C. L. & Froemke, R. C. Automatic mapping of multiplexed social receptive fields by deep learning and GPU-accelerated 3D videography. Nat. Commun. 13, 593 (2022).
Lauer, J. et al. Multi-animal pose estimation, identification and tracking with DeepLabCut. Nat. Methods 19, 496–504 (2022).
Pereira, T. D. et al. SLEAP: a deep learning system for multi-animal pose tracking. Nat. Methods 19, 486–495 (2022).
Chen, Z. et al. AlphaTracker: a multi-animal tracking and behavioral analysis tool. Front. Behav. Neurosci. 17, 1111908 (2023).
Marks, M. et al. Deep-learning-based identification, tracking, pose estimation and behaviour classification of interacting primates and mice in complex environments. Nat. Mach. Intell. 4, 331–340 (2022).
Romero-Ferrero, F., Bergomi, M. G., Hinz, R. C., Heras, F. J. H. & de Polavieja, G. G. idtracker.ai: tracking all individuals in small or large collectives of unmarked animals. Nat. Methods 16, 179–182 (2019).
Ro, S. et al. Simple Behavioral Analysis (SimBA) – an open source toolkit for computer classification of complex social behaviors in experimental animals. Preprint at bioRxiv https://doi.org/10.1101/2020.04.19.049452 (2020).
Chen, Z. et al. AlphaTracker: a multi-animal tracking and behavioral analysis tool. Front Behav Neurosci. 17, 1111908 (2023).
Huang, K. et al. A hierarchical 3D-motion learning framework for animal spontaneous behavior mapping. Nat. Commun. 12, 2784 (2021).
Han, Y., Huang, K., Chen, K., Wang, L. & Wei, P. An automatic three dimensional markerless behavioral tracking system of free-moving mice. In Proc. 2021 IEEE 11th Annual International Conference on CYBER Technology in Automation, Control, and Intelligent Systems (ed. Chen, H.) 306–310 (IEEE, 2021).
Marshall, J. D. et al. Continuous whole-body 3D kinematic recordings across the rodent behavioral repertoire. Neuron 109, 420–437.e8 (2021).
Ghiasi, G. et al. Simple copy-paste is a strong data augmentation method for instance segmentation. GitHub https://cocodataset.org/ (2021).
Xu, Z. et al. Continuous copy-paste for one-stage multi-object tracking and segmentation. In Proc. 2021 IEEE/CVF Conference on Computer Vision and Pattern Recognition (ed. Berg, T.) 15323–15332 (IEEE, 2021)
Weiss, K., Khoshgoftaar, T. M. & Wang, D. D. A survey of transfer learning. J. Big Data 3, 9 (2016).
Zhuang, F. et al. A comprehensive survey on transfer learning. Proc. IEEE https://doi.org/10.1109/JPROC.2020.3004555 (2021).
Mathis, A. et al. DeepLabCut: markerless pose estimation of user-defined body parts with deep learning. Nat. Neurosci. 21, 1281–1289 (2018).
Liu, N. et al. Objective and comprehensive re-evaluation of anxiety-like behaviors in mice using the Behavior Atlas. Biochem. Biophys. Res. Commun. 559, 1–7 (2021).
Han, Y. et al. MouseVenue3D: a markerless three-dimension behavioral tracking system for matching two-photon brain imaging in free-moving mice. Neurosci. Bull. 38, 303–317 (2022).
Bolya, D., Zhou, C., Xiao, F. & Lee, Y. J. YOLACT++ better real-time instance segmentation. IEEE Trans. Pattern Anal. Mach. Intell. 44, 1108–1121 (2022).
Bolya, D., Fanyi, C. Z., Yong, X. & Lee, J. YOLACT. Real-time instance segmentation. GitHub https://github.com/dbolya/yolact (2019).
Wang, Y. et al. VisTR. End-to-end video instance segmentation with transformers. GitHub https://git.io/VisTR (2021).
Peça, J. et al. Shank3 mutant mice display autistic-like behaviours and striatal dysfunction. Nature 472, 437–442 (2011).
Mei, Y. et al. Adult restoration of Shank3 expression rescues selective autistic-like phenotypes. Nature 530, 481–484 (2016).
Tan, M. & Le, Q. V. EfficientNet: rethinking model scaling for convolutional neural networks. In Proc. International Conference on Machine Learning (ed. Lawrence, N.) 6105–6114 (PMLR, 2019)
Jiang, P. T., Zhang, C., bin, Hou, Q., Cheng, M. M. & Wei, Y. LayerCAM: exploring hierarchical class activation maps for localization. IEEE Trans. Image Process. 30, 5875–5888 (2021).
Bzdok, D. & Dunbar, R. I. M. The neurobiology of social distance. Trends Cogn. Sci. 24, 717–733 (2020).
McInnes, L., Healy, J. & Melville, J. UMAP: uniform manifold approximation and projection for dimension reduction. Preprint at arXiv https://doi.org/10.48550/arXiv.1802.03426 (2018).
Shi, S., Wang, Y., Dong, H., Gui, G. & Ohtsuki, T. Smartphone-aided human activity recognition method using residual multi-layer perceptron. In Proc. INFOCOM, IEEE Conference on Computer Communications Workshops (ed. Misra, S.) 1–6 (IEEE, 2022).
Marshall, J. D. et al. The PAIR-R24M dataset for multi-animal 3D pose estimation. Preprint at bioRxiv https://doi.org/10.1101/2021.11.23.469743 (2021).
Day, F. R., Ong, K. K. & Perry, J. R. B. Elucidating the genetic basis of social interaction and isolation. Nat. Commun. 9, 2457 (2018).
Wu, Y. E. & Hong, W. Neural basis of prosocial behavior. Trends Neurosci. https://doi.org/10.1016/J.TINS.2022.06.008 (2022).
Wu, Y. E. et al. Neural control of affiliative touch in prosocial interaction. Nature 599, 262–267 (2021).
Dunn, T. W. et al. Geometric deep learning enables 3D kinematic profiling across species and environments. Nat. Methods 18, 564–573 (2021).
Mathis, A. et al. Pretraining boosts out-of-domain robustness for pose estimation. In Proc. 2021 IEEE Winter Conference on Applications of Computer Vision (eds Medioni, G. & Bowyer, K.) 1859–1868 (IEEE, 2021).
Li, W. et al. Exploiting temporal contexts with strided transformer for 3D human pose estimation. IEEE Trans. Multimedia https://doi.org/10.1109/TMM.2022.3141231 (2022).
Vaswani, A. et al. Attention Is All You Need. In Advances in Neural Information Processing Systems 30 (NeurIPS, 2017).
Sun, J. J. et al. The multi-agent behavior dataset: mouse dyadic social interactions. Preprint at arXiv https://doi.org/10.48550/arXiv.2104.02710 (2021).
Walter, T. & Couzin, I. D. Trex, a fast multi-animal tracking system with markerless identification, and 2D estimation of posture and visual elds. eLife 10, 64000 (2021).
Yang, L., Fan, Y. & Xu, N. Video instance segmentation. In Proc. IEEE International Conference on Computer Vision (ed. Lee, K.) 5188–5197 (IEEE, 2019).
Han, Y. SBM-VIS Dataset. figshare https://doi.org/10.6084/m9.figshare.24597111.v1 (2023).
Han, Y. SBeA Upload Data. figshare https://doi.org/10.6084/m9.figshare.22314994.v1 (2023).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. In Proc. IEEE Conference on Computer Vision and Pattern Recognition (ed. Bajcsy, R.) 770–778 (IEEE, 2016).
Kruse, R., Mostaghim, S., Borgelt, C., Braune, C. & Steinbrecher, M. in Computational Intelligence (eds Hazzan, O. & Maurer, F.) 53–124 (Springer, 2022).
Kiranyaz, S. et al. 1D convolutional neural networks and applications: a survey. Mech. Syst. Signal Process. 151, 107398 (2021).
Lecun, Y., Bengio, Y. & Hinton, G. Deep learning. Nature https://doi.org/10.1038/nature14539 (2015).
Zhang, Z. Improved Adam optimizer for deep neural networks. In Proc. 2018 IEEE/ACM 26th International Symposium on Quality of Service (eds Wu, C. & Li, Z.) 1–2 (IEEE/ACM, 2019).
Han, Y. SBeA_release. Zenodo https://doi.org/10.5281/zenodo.8238067 (2023).
Acknowledgements
This work was supported in part by STI2030-Major Projects (grant no. 2021ZD0203900 to P.W.), National Natural Science Foundation of China (grant no. 32222036 to P.W.), the Youth Innovation Promotion Association of the Chinese Academy of Sciences (grant no. Y2021100 to P.W.), the National Key R&D Program of China (grant no. 2018YFA0701403 to P.W.), CAS Key Laboratory of Brain Connectome and Manipulation (grant no. 2019DP173024 to L.W.), and Guangdong Provincial Key Laboratory of Brain Connectome and Behavior (grant no. 2017B030301017 to LP.W.). We thank J. Zhang, Y. Wu and G. Gao for the preliminary tests and experiments, and Z. Jiang for his comments on our paper.
Author information
Authors and Affiliations
Contributions
Conceptualization was done by Y. Han, K.C. and P.W. Code was written by Y. Han, K.C., Z.W, and S.C. The user interface was developed by Z.W. The algorithms were designed by Y. Han and K.C. Animal data were gathered by Y. Han, W.L., X.W., Y. Huang, C.H., J. Li, Y.S., N.W., J. Li, G.-D.W., Y.Z. and J. Liao. Hardware was set up by Y. Han and K.H. Data analysis was done by Y. Han. Experiments were supported by L.W. and P.W. The article was written by Y. Han, K.C., Y.W. and P.W. with input from all authors. P.W. supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Machine Intelligence thanks Minmin Luo, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Performance comparison of SBeA and maDLC.
a, Prediction error comparison of all test data. The RMSE of most of the body parts of SBeA is significantly lower than maDLC (two-way ANOVA followed by Sidak multiple comparisons test, N = 14400, adjusted P values from Nose to Tip Tail are <0.0001, <0.0001, <0.0001, 0.3810, >0.9999, 0.9972, 0.9975, >0.9999, <0.0001, 0.0025, >0.9999, 0.9928, <0.0001, <0.0001, <0.0001, and <0.0001). b, Prediction error comparison of close contact. The RMSE of all of the body parts of SBeA is significantly lower than maDLC or even with maDLC (two-way ANOVA followed by Sidak multiple comparisons test, N = 4602, adjusted P values from Nose to Tip Tail are <0.0001, <0.0001, <0.0001, 0.3060, <0.0001, 0.0040, 0.0267, 0.9775, 0.7650, 0.9838, 0.0002, 0.2037, >0.9999, <0.0001, <0.0001, and <0.0001). Stems represent the mean values of each violin plot. RMSE: root-mean-squared error, n.s.: no significant difference, *: P < 0.05, **: P < 0.01, ***: P < 0.001, ****: P < 0.0001.
Extended Data Fig. 2 Performance comparison of SBeA and SLEAP.
a, Prediction error comparison of all test data (two-way ANOVA followed by Sidak multiple comparisons test, N = 14400, adjusted P values from Nose to Tip Tail are <0.0001, <0.0001, <0.0001, 0.8054, <0.0001, <0.0001, 0.0030, <0.0001, 0.4651, <0.0001, <0.0001, <0.0001, <0.0001, <0.0001, <0.0001, and <0.0001). b, Prediction error comparison of close contact (two-way ANOVA followed by Sidak multiple comparisons test, N = 4602, adjusted P values from Nose to Tip Tail are <0.0001, <0.0001, <0.0001, <0.0001, <0.0001, <0.0001, <0.0001, <0.0001, <0.0001, <0.0001, <0.0001, <0.0001, >0.9999, <0.0001, <0.0001, and <0.0001). Stems represent the mean values of each violin plot. RMSE: root-mean-squared error, n.s.: no significant difference, *: P < 0.05, **: P < 0.01, ***: P < 0.001, ****: P < 0.0001.
Supplementary information
Supplementary Information
Supplementary Figs. 1–11, Table 1, Notes and Methods.
Source data
Source Data Fig. 1
3D_mice_mouse1, 3D_mice_mouse2, Social_behavior_atlas.
Source Data Fig. 2
3D_poses_reconstruction_error, Comparison_of_annotations, Distance_distribution; Precision_comparison_all, Precision_comparison_close.
Source Data Fig. 3
Confusion_matrix, Representation, Precision, Silhouette_coefficient, Precision_images, Precision_3D.
Source Data Fig. 4
Social_behavior_atlas, Atlas_with_all_labels, Distance_map, Cluster_purity.
Source Data Fig. 5
Cluster gram, Variance_explained, Phenotype space, Angle_spectrum, Fractions.
Source Data Fig. 6
Ethogram_birds, Atlas_birds, 3D_birds, Ethogram_dogs, Atlas_dogs, 3D_dogs.
Source Data Extended Data Fig. 1
sbea_all, dlc_all, sbea_close, dlc_close.
Source Data Extended Data Fig. 2
sbea_all, sleap_all, sbea_close, sleap_close.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Han, Y., Chen, K., Wang, Y. et al. Multi-animal 3D social pose estimation, identification and behaviour embedding with a few-shot learning framework. Nat Mach Intell 6, 48–61 (2024). https://doi.org/10.1038/s42256-023-00776-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42256-023-00776-5