Abstract
Disruption of circadian rhythm during pregnancy produces adverse health outcomes in offspring; however, the role of maternal circadian rhythms in the immune system of infants and their susceptibility to inflammation remains poorly understood. Here we show that disruption of circadian rhythms in pregnant mice profoundly aggravates the severity of neonatal inflammatory disorders in both male and female offspring, such as necrotizing enterocolitis and sepsis. The diminished maternal production of docosahexaenoic acid (DHA) and the impaired immunosuppressive function of neonatal myeloid-derived suppressor cells (MDSCs) contribute to this phenomenon. Mechanistically, DHA enhances the immunosuppressive function of MDSCs via PPARγ-mediated mitochondrial oxidative phosphorylation. Transfer of MDSCs or perinatal supplementation of DHA relieves neonatal inflammation induced by maternal rhythm disruption. These observations collectively demonstrate a previously unrecognized role of maternal circadian rhythms in the control of neonatal inflammation via metabolic reprograming of myeloid cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The bulk RNA-seq data generated in this study have been deposited in the Gene Expression Omnibus under accession code GSE233545. Image source data were deposited in Figshare (https://doi.org/10.6084/m9.figshare.25237708)60. No third-party materials were included in this study. Source data are provided with this paper.
Code availability
No custom codes were used in this study.
References
Asher, G. et al. Crosstalk between components of circadian and metabolic cycles in mammals. Cell Metab. 13, 125–137 (2011).
Masri, S. et al. The emerging link between cancer, metabolism, and circadian rhythms. Nat. Med. 24, 1795–1803 (2018).
Zhang, E. E. et al. A genome-wide RNAi screen for modifiers of the circadian clock in human cells. Cell 139, 199–210 (2009).
Druzd, D. et al. Lymphocyte circadian clocks control lymph node trafficking and adaptive immune responses. Immunity 46, 120–132 (2017).
Cox, S. L. et al. Intertwining roles of circadian and metabolic regulation of the innate immune response. Semin. Immunopathol. 44, 225–237 (2022).
Fishbein, A. B. et al. Circadian disruption and human health. J. Clin. Invest. 131, e148286 (2021).
Hantsoo, L. et al. Inflammation: a proposed intermediary between maternal stress and offspring neuropsychiatric risk. Biol. Psychiatry 85, 97–106 (2019).
Deng, Y. et al. Prenatal inflammation exposure-programmed cardiovascular diseases and potential prevention. Pharmacol. Ther. 190, 159–172 (2018).
Cai, C. et al. The impact of occupational shift work and working hours during pregnancy on health outcomes: a systematic review and meta-analysis. Am. J. Obstet. Gynecol. 221, 563–576 (2019).
Salazar, E. R. et al. Gestational chronodisruption leads to persistent changes in the rat fetal and adult adrenal clock and function. J. Physiol. 596, 5839–5857 (2018).
Smarr, B. L. et al. Maternal and early-life circadian disruption have long-lasting negative consequences on offspring development and adult behavior in mice. Sci. Rep. 7, 3326 (2017).
Whelan, E. A. et al. Work schedule during pregnancy and spontaneous abortion. Epidemiology 18, 350–355 (2007).
Yu, J. C. et al. Innate immunity of neonates and infants. Front. Immunol. 9, 1759 (2018).
Gensollen, T. et al. How colonization by microbiota in early life shapes the immune system. Science 352, 539–544 (2016).
Brodin, P. Immune–microbe interactions early in life: a determinant of health and disease long term. Science 376, 945–950 (2022).
He, Y. M. et al. Transitory presence of myeloid-derived suppressor cells in neonates is critical for control of inflammation. Nat. Med. 24, 224–231 (2018).
Stewart, C. J. et al. Gut bacteria and necrotizing enterocolitis: cause or effect? Trends Microbiol. 23, 332–333 (2015).
Lassi, M. et al. Disruption of paternal circadian rhythm affects metabolic health in male offspring via nongerm cell factors. Sci. Adv. 7, eabg6424 (2021).
Levy, O. Innate immunity of the newborn: basic mechanisms and clinical correlates. Nat. Rev. Immunol. 7, 379–390 (2007).
Early, J. O. et al. Circadian clock protein BMAL1 regulates Il-1β in macrophages via NRF2. Proc. Natl Acad. Sci. USA 115, E8460–E8468 (2018).
Feng, Y. et al. Estrogen receptor-alpha expression in the mammary epithelium is required for ductal and alveolar morphogenesis in mice. Proc. Natl Acad. Sci. USA 104, 14718–14723 (2007).
Wagner, K. U. et al. Tsg101 is essential for cell growth, proliferation, and cell survival of embryonic and adult tissues. Mol. Cell. Biol. 23, 150–162 (2003).
Nguyen, K. D. et al. Circadian gene Bmal1 regulates diurnal oscillations of Ly6C(hi) inflammatory monocytes. Science 341, 1483–1488 (2013).
Liu, Y. et al. Lactoferrin-induced myeloid-derived suppressor cell therapy attenuates pathologic inflammatory conditions in newborn mice. J. Clin. Invest. 129, 4261–4275 (2019).
Bee, G. C. W. et al. Age-dependent differences in efferocytosis determine the outcome of opsonophagocytic protection from invasive pathogens. Immunity 56, 1255–1268 (2023).
Campion, A. et al. Use of enhanced nisin derivatives in combination with food-grade oils or citric acid to control Cronobacter sakazakii and Escherichia coli O157:H7. Food Microbiol. 65, 254–263 (2017).
Wang, W. et al. Protective effects and mechanism of a novel probiotic strain ligilactobacillus salivarius YL20 against Cronobacter sakazakii-induced necrotizing enterocolitis in vitro and in vivo. Nutrients 14, 3827 (2022).
Stanley, W. C. et al. Update on lipids and mitochondrial function: impact of dietary n-3 polyunsaturated fatty acids. Curr. Opin. Clin. Nutr. Metab. Care 15, 122–126 (2012).
Shahidi, F. et al. Omega-3 polyunsaturated fatty acids and their health benefits. Annu. Rev. Food Sci. Technol. 9, 345–381 (2018).
Abou El Fadl, D. K. et al. Impact of docosahexaenoic acid supplementation on proinflammatory cytokines release and the development of necrotizing enterocolitis in preterm neonates: a randomized controlled study. Saudi Pharm. J. 29, 1314–1322 (2021).
Chen, D. et al. Coupled analysis of transcriptome and BCR mutations reveals role of OXPHOS in affinity maturation. Nat. Immunol. 22, 904–913 (2021).
Basak, S. et al. Maternal docosahexaenoic acid status during pregnancy and its impact on infant neurodevelopment. Nutrients 12, 3615 (2020).
Lock, J. Y. et al. Impact of developmental age, necrotizing enterocolitis associated stress, and oral therapeutic intervention on mucus barrier properties. Sci. Rep. 10, 6692 (2020).
Helland, I. B. et al. Supplementation of n-3 fatty acids during pregnancy and lactation reduces maternal plasma lipid levels and provides DHA to the infants. J. Matern. Fetal Neonatal Med. 19, 397–406 (2006).
Imhoff-Kunsch, B. et al. Docosahexaenoic acid supplementation from mid-pregnancy to parturition influenced breast milk fatty acid concentrations at 1 month postpartum in Mexican women. J. Nutr. 141, 321–326 (2011).
Innis, S. M. Essential fatty acids in growth and development. Prog. Lipid Res. 30, 39–103 (1991).
Carlson, S. E. et al. DHA supplementation and pregnancy outcomes. Am. J. Clin. Nutr. 97, 808–815 (2013).
Meher, A. P. et al. Placental DHA and mRNA levels of PPARgamma and LXRα and their relationship to birth weight. J. Clin. Lipidol. 10, 767–774 (2016).
Calder, P. C. Docosahexaenoic acid. Ann. Nutr. Metab. 69, 7–21 (2016).
Miles, E. A. et al. Long-chain polyunsaturated fatty acids (LCPUFAs) and the developing immune system: a narrative review. Nutrients 13, 247 (2021).
Ohtsuka, Y. et al. Omega-3 fatty acids attenuate mucosal inflammation in premature rat pups. J. Pediatr. Surg. 46, 489–495 (2011).
Wijendran, V. et al. Long-chain polyunsaturated fatty acids attenuate the IL-1β-induced proinflammatory response in human fetal intestinal epithelial cells. Pediatr. Res. 78, 626–633 (2015).
Bernabe-Garcia, M. et al. Efficacy of docosahexaenoic acid for the prevention of necrotizing enterocolitis in preterm infants: a randomized clinical trial. Nutrients 13, 648 (2021).
Hsu, C. N. et al. Light and circadian signaling pathway in pregnancy: programming of adult health and disease. Int. J. Mol. Sci. 21, 2232 (2020).
Goldenberg, R. L. et al. Epidemiology and causes of preterm birth. Lancet 371, 75–84 (2008).
Varcoe, T. J. Timing is everything: maternal circadian rhythms and the developmental origins of health and disease. J. Physiol. 596, 5493–5494 (2018).
Astiz, M. et al. Feto-maternal crosstalk in the development of the circadian clock system. Front. Neurosci. 14, 631687 (2020).
Reiter, R. J. et al. Melatonin and stable circadian rhythms optimize maternal, placental and fetal physiology. Hum. Reprod. Update 20, 293–307 (2014).
Torres-Farfan, C. et al. Maternal melatonin effects on clock gene expression in a nonhuman primate fetus. Endocrinology 147, 4618–4626 (2006).
Tamura, H. et al. Fetal/placental regulation of maternal melatonin in rats. J. Pineal Res. 44, 335–340 (2008).
Ma, F. et al. Melatonin ameliorates necrotizing enterocolitis by preventing Th17/Treg imbalance through activation of the AMPK/SIRT1 pathway. Theranostics 10, 7730–7746 (2020).
Ryan, K. K. et al. A role for central nervous system PPAR-gamma in the regulation of energy balance. Nat. Med. 17, 623–626 (2011).
Bapat, S. P. et al. Obesity alters pathology and treatment response in inflammatory disease. Nature 604, 337–342 (2022).
Baregamian, N. et al. PPAR-γ agonist protects against intestinal injury during necrotizing enterocolitis. Biochem. Biophys. Res. Commun. 379, 423–427 (2009).
Torow, N. et al. The neonatal window of opportunity: setting the stage for life-long host-microbial interaction and immune homeostasis. J. Immunol. 198, 557–563 (2017).
Xu, H. et al. Maternal antibiotic exposure enhances ILC2 activation in neonates via downregulation of IFN1 signaling. Nat. Commun. 14, 8332 (2023).
Wang, Q. et al. Circadian rhythm-dependent and circadian rhythm-independent impacts of the molecular clock on type 3 innate lymphoid cells. Sci. Immunol. 4, eaay7501 (2019).
Egan, C. E. et al. Toll-like receptor 4-mediated lymphocyte influx induces neonatal necrotizing enterocolitis. J. Clin. Invest. 126, 495–508 (2016).
Rager, T. M. et al. Exosomes secreted from bone marrow-derived mesenchymal stem cells protect the intestines from experimental necrotizing enterocolitis. J. Pediatr. Surg. 51, 942–947 (2016).
Zhaohai, Cui. Source data. Figshare https://doi.org/10.6084/m9.figshare.25237708 (2024).
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (nos. 81925018 and 82130049 to J.Z.; 82321001 to Y.Y.; and 82225015 and 82171284 to Q.L.). This work was also supported by the New Cornerstone Science Foundation through the XPLORER PRIZE (to Q.L.), Natural Science Foundation of Tianjin (22JCQNJC01210 to H.X.).
Author information
Authors and Affiliations
Contributions
J.Z. conceived and supervised this study. Q.L. and Y.Y. jointly supervised this study. Z.C. performed the experiments, analysed the data and wrote the manuscript. H.X. participated in most of the experiments. X.Y. conducted bioinformatics analysis. F.W., J.C., Lin Zhu, Z. Shen, J.Y., C.J., L. Zhang and P.Z. participated in animal model and flow cytometry analysis. M.J.L., Lu Zhu, S.D. and Z.Y. provided suggestions in project design. J.Z. wrote the manuscript with inputs from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Metabolism thanks Lindsey Devisscher, Christoph Scheiermann and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Ashley Castellanos-Jankiewicz, in collaboration with the Nature Metabolism team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 General effects of maternal rhythm disruption on dams and pups.
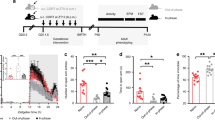
(a) Corticosterone content in serum non-targeted metabolomics at day 7 postpartum of dams in control and circadian rhythm disorder groups (n = 3). (b) The contents of melatonin in the serum of dams were measured by ELISA (n = 3/time point). (c) Average food intake per day of control and CRD dams (n = 5). (d) Body weights of pups were evaluated at birth (n = 8). (e) Milk consumption of 7-day-old was assessed (Ctrl: n = 7; CRD: n = 8). (f) The proportions of Th17 and Treg in small intestine were evaluated by flow cytometry after NEC induction (n = 3). (g) The survival rate of NEC mice at ZT0 and ZT6 (Ctrl-ZT0: n = 15; Ctrl-ZT6: n = 16; CRD-ZT0: n = 19; CRD-ZT6: n = 20). (h) Representative H&E staining and inflammation score of intestinal (scale bars: 100 μm, ZT0-Ctrl: n = 6; ZT6-Ctrl: n = 12; ZT0-CRD: n = 3; ZT6-CRD: n = 4). Data are representative of two independent experiments. Mean ± SEM were shown. Two-tailed unpaired Student’s t test for a, c, d, e, f, h. Log-rank (Mantel-Cox) test was used for g. Ns, not significant, *p < 0.05; **p < 0.01, exact P values are provided in the source data.
Extended Data Fig. 2 Effects of maternal rhythm disruption on neonatal MDSCs.
(a-c) WT pups cross-fostered by Bmal1fl/flWAPcre (n = 8) or Bmal1fl/fl (n = 8) mother were subjected to NEC induction. (a) The survival rate of pups. (b) Representative H&E staining of intestine and the Inflammation scores. scale bars, 100 μm (n = 3-6). (c) The expression of the pro-inflammation genes Il6 and Il1β in intestine was determined by qRT–PCR (n = 3-6). (d-e) The frequencies and absolute numbers of indicated immune cell types in spleen of 7-day-old pups were analysed by flow cytometry (n = 5). (f) The immunosuppressive function of splenic M-MDSC of pups (n = 3-5). (g) The immunosuppressive activity of neonatal PMN-MDSCs from control and CRD groups at different time points (n = 3/time point). (h) The mRNA expression of clock genes Cry2, Per3 and Nr1d2 in PMN-MDSCs from pups were determined by qRT–PCR (n = 3). Data are representative of two independent experiments. Mean ± SEM were shown. Log-rank (Mantel-Cox) test for a. Two-tailed unpaired Student’s t test for b, c, e, f and two-way ANOVA followed by Bonferroni’s multiple comparisons test for g. *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001, exact P values are provided in the source data.
Extended Data Fig. 3 Effects of maternal rhythm disruption on the transcriptome of neonatal MDSCs.
(a) Principal component analysis (PCA) of neonatal PMN-MDSCs transcriptome between control and CRD groups (n = 3). (b) Heat map of immunosuppressive related genes in PMN-MDSCs (n = 3). (c) WT pups were cross-fostered by Bmal1fl/fl or Bmal1fl/fl WAPcre (WT → Bmal1fl/fl: n = 3; WT → Bmal1fl/fl WAPcre: n = 3). The proliferation of CD8+ T from OT-I spleen were stimulated with OVA257-264 in the presence of neonatal PMN-MDSCs were shown. (d) Heat map of anti-bacterial, phagocytosis and chemokine related genes in PMN-MDSCs (n = 3). (e) The effect of DHA on the migration of PMN-MDSCs was evaluated in vitro by transwell migration assay (n = 5). Neonatal PMN-MDSCs from the spleens from the indicated group were seeded on the upper chamber of transwell. Medium containing chemokine CXCL1 was added to the bottom layer of the transwell. After 1 hour incubation, cells were counted at the bottom chamber. Data are representative of two independent experiments. Two-tailed unpaired Student’s t test for c, and one-way ANOVA followed by Bonferroni’s multiple comparisons test for e. *p < 0.05; **p < 0.01, exact P values are provided in the source data.
Extended Data Fig. 4 Effect of DHA supplementation on neonatal MDSCs.
(a) Principal component analysis (PCA) of non-targeted metabolomics from breast milk between CRD and control dams (n = 3). (b) Volcano plot showing changed metabolites in breast milk between two groups (n = 3). (c) Heat map displaying altered proteins in breast milk between control and CRD dams. (d) Breast milk was collected from control dams at different time points, followed by targeted metabolomics to evaluate the amounts of DHA (n = 3). (e) The expression of Elovl2 and Elovl5 mRNA in liver of dams at different time points was assessed by qRT–PCR (n = 3). (f) The mRNA expression of DHA synthesis genes in dams’ liver was evaluated by qRT–PCR (n = 3). (g) DHA was orally administered to CRD dams, the proportions of M-MDSC and PMN-MDSC in the spleen of pups were evaluated by flow cytometry (n = 6-7). (h-j) 2-day-old CRD-delivered pups were injected intraperitoneally with DHA at 20 mg/kg/day for 5 days. (i). The proportions of M-MDSC and PMN-MDSC in the spleen were evaluated by flow cytometry (n = 7). (j). The immunosuppressive function of total MDSCs was was evaluated T cell coculture experiment, T cell proliferation was indicated by CFSE labelling (n = 4-7). Data are representative of two independent experiments. Mean ± SEM were shown. Two-tailed unpaired Student’s t test for f, and one-way ANOVA followed by Bonferroni’s multiple comparisons test for g, i, j. Ns, not significant, p > 0.05; *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001, exact P values are provided in the source data.
Extended Data Fig. 5 Effects of maternal rhythm disruption on PPARγ signalling and mitochondrial function in neonatal PMN-MDSCs.
(a) Heat map of genes related to oxidative phosphorylation in PMN-MDSCs (n = 3). (b) Diurnal mRNA expression of genes involved in mitochondrial dynamics in PMN-MDSCs from control and CRD groups was determined by qRT–PCR (n = 3/time point). (c) Heat map of PPARγ pathway related genes in PMN-MDSCs (n = 3). (d) qRT–PCR analysis of mitochondrial related genes in PMN-MDSCs from pups of PPARγfl/fl and PPARγfl/fl Lysmcre (n = 3). Data are representative of two independent experiments. Mean ± SEM were shown. Two-tailed unpaired Student’s t test for d. *p < 0.05; ***p < 0.001, exact P values are provided in the source data.
Extended Data Fig. 6 Gating strategies for this paper.
(a) Gating strategy for MDSC. (b) The gating strategy for MDSC subsets. (c) Gating strategy for macrophage, natural killer cell, B cell, T cell and dendritic cell.
Supplementary information
Supplementary Data 1–4
1. Antibodies used for flow cytometry in this study. 2. Non-targeted metabolomics of dams. 3. Non-targeted metabolomics of neonates. 4. Primer sequences.
Source data
Source Data Fig. 1
Statistical Source Data.
Source Data Fig. 2
Statistical Source Data.
Source Data Fig. 3
Statistical Source Data.
Source Data Fig. 4
Statistical Source Data.
Source Data Fig. 5
Statistical Source Data.
Source Data Fig. 6
Statistical Source Data.
Source Data Fig. 7
Statistical Source Data.
Source Data Extended Data Fig. 1
Statistical Source Data.
Source Data Extended Data Fig. 2
Statistical Source Data.
Source Data Extended Data Fig. 3
Statistical Source Data.
Source Data Extended Data Fig. 4
Statistical Source Data.
Source Data Extended Data Fig. 5
Statistical Source Data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Cui, Z., Xu, H., Wu, F. et al. Maternal circadian rhythm disruption affects neonatal inflammation via metabolic reprograming of myeloid cells. Nat Metab (2024). https://doi.org/10.1038/s42255-024-01021-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42255-024-01021-y