Abstract
Different organs undergo distinct transcriptional, epigenetic and physiological alterations that guarantee their functional maturation after birth. However, the roles of epitranscriptomic machineries in these processes have remained elusive. Here we demonstrate that expression of RNA methyltransferase enzymes Mettl3 and Mettl14 gradually declines during postnatal liver development in male mice. Liver-specific Mettl3 deficiency causes hepatocyte hypertrophy, liver injury and growth retardation. Transcriptomic and N6-methyl-adenosine (m6A) profiling identify the neutral sphingomyelinase, Smpd3, as a target of Mettl3. Decreased decay of Smpd3 transcripts due to Mettl3 deficiency results in sphingolipid metabolism rewiring, characterized by toxic ceramide accumulation and leading to mitochondrial damage and elevated endoplasmic reticulum stress. Pharmacological Smpd3 inhibition, Smpd3 knockdown or Sgms1 overexpression that counteracts Smpd3 can ameliorate the abnormality of Mettl3-deficent liver. Our findings demonstrate that Mettl3–N6-methyl-adenosine fine-tunes sphingolipid metabolism, highlighting the pivotal role of an epitranscriptomic machinery in coordination of organ growth and the timing of functional maturation during postnatal liver development.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The transcriptomic dataset of livers from C57BL/6J mice across 12 time points, from late embryonic stage (E17.5) to adulthood (day 60), were obtained from the National Center for Biotechnology Information GEO database (accession no. GSE103322). RNA-seq and Me–RIP–seq data reported in this study have been deposited in the GEO database under accession no. GSE197799. Source data are provided with this paper.
Code availability
All codes and scripts used for association studies are available on request.
References
Plagge, A. et al. The imprinted signaling protein XL alpha s is required for postnatal adaptation to feeding. Nat. Genet. 36, 818–826 (2004).
Uddin, M. et al. Distinct genomic signatures of adaptation in pre- and postnatal environments during human evolution. Proc. Natl Acad. Sci. USA 105, 3215–3220 (2008).
Zamboulis, D. E. et al. Postnatal mechanical loading drives adaptation of tissues primarily through modulation of the non-collagenous matrix. eLife 9, e58075 (2020).
Reese, J. et al. Coordinated regulation of fetal and maternal prostaglandins directs successful birth and postnatal adaptation in the mouse. Proc. Natl Acad. Sci. USA 97, 9759–9764 (2000).
Willers, M. et al. S100A8 and S100A9 are important for postnatal development of gut microbiota and immune system in mice and infants. Gastroenterology 159, 2130–2145 (2020).
Rossant, J., Vijh, K. M., Grossi, C. E. & Cooper, M. D. Clonal origin of haematopoietic colonies in the postnatal mouse liver. Nature 319, 507–511 (1986).
Lotto, J. et al. Single-cell transcriptomics reveals early emergence of liver parenchymal and non-parenchymal cell lineages. Cell 183, 702–716 (2020).
Gunewardena, S. S. et al. Deciphering the developmental dynamics of the mouse liver transcriptome. PLoS ONE 10, e0141220 (2015).
Li, T. et al. Multi-stage analysis of gene expression and transcription regulation in C57/B6 mouse liver development. Genomics 93, 235–242 (2009).
Costa, R. H., Kalinichenko, V. V., Holterman, A. X. L. & Wang, X. H. Transcription factors in liver development, differentiation, and regeneration. Hepatology 38, 1331–1347 (2003).
Sen, S., Jumaa, H. & Webster, N. J. G. Splicing factor SRSF3 is crucial for hepatocyte differentiation and metabolic function. Nat. Commun. 4, 1336 (2013).
Waterland, R. A. et al. Epigenomic profiling indicates a role for DNA methylation in early postnatal liver development. Hum. Mol. Genet. 18, 3026–3038 (2009).
Mu, T. et al. Embryonic liver developmental trajectory revealed by single-cell RNA sequencing in the Foxa2(eGFP) mouse. Commun. Biol. 3, 642 (2020).
Gordillo, M., Evans, T. & Gouon-Evans, V. Orchestrating liver development. Development 142, 2094–2108 (2015).
Frye, M., Harada, B. T., Behm, M. & He, C. RNA modifications modulate gene expression during development. Science 361, 1346–1349 (2018).
Shi, H. L., Wei, J. B. & He, C. Where, when, and how: context-dependent functions of RNA methylation writers, readers, and erasers. Mol. Cell 74, 640–650 (2019).
Wang, C. X. et al. METTL3-mediated m6A modification is required for cerebellar development. PLoS Biol. 16, e2004880 (2018).
Wang, Y. et al. METTL3 is essential for postnatal development of brown adipose tissue and energy expenditure in mice. Nat. Commun. 11, 1648 (2020).
Li, H. B. et al. m(6)A mRNA methylation controls T cell homeostasis by targeting the IL-7/STAT5/SOCS pathways. Nature 548, 338–342 (2017).
Yao, Y. et al. METTL3-dependent m(6)A modification programs T follicular helper cell differentiation. Nat. Commun. 12, 1333 (2021).
Kyrmizi, I. et al. Plasticity and expanding complexity of the hepatic transcription factor network during liver development. Genes Dev. 20, 2293–2305 (2006).
Karagianni, P., Moulos, P., Schmidt, D., Odom, D. T. & Talianidis, I. Bookmarking by non-pioneer transcription factors during liver development establishes competence for future gene activation. Cell Rep. 30, 1319–1328 (2020).
Barajas, J. M. et al. METTL3 regulates liver homeostasis, hepatocyte ploidy, and circadian rhythm–controlled gene expression in mice. Am. J. Pathol. 192, 56–71 (2022).
Xu, Y. et al. Mettl3-mediated mRNA m6A modification controls postnatal liver development by modulating the transcription factor Hnf4a. Nat. Commun. 13, 4555 (2022).
Li, X. et al. The methyltransferase METTL3 negatively regulates nonalcoholic steatohepatitis (NASH) progression. Nat. Commun. 12, 7213 (2021).
Batista, P. J. et al. m(6)A RNA modification controls cell fate transition in mammalian embryonic stem cells. Cell Stem Cell 15, 707–719 (2014).
Zhang, C. et al. m(6)A modulates haematopoietic stem and progenitor cell specification. Nature 549, 273–276 (2017).
Postic, C. et al. Dual roles for glucokinase in glucose homeostasis as determined by liver and pancreatic beta cell-specific gene knock-outs using Cre recombinase. J. Biol. Chem. 274, 305–315 (1999).
Green, C. D., Maceyka, M., Cowart, L. A. & Spiegel, S. Sphingolipids in metabolic disease: the good, the bad, and the unknown. Cell Metab. 33, 1293–1306 (2021).
Deng, X. et al. Ceramide biogenesis is required for radiation-induced apoptosis in the germ line of C. elegans. Science 322, 110–115 (2008).
Turpin, S. M. et al. Obesity-induced CerS6-dependent C16:0 ceramide production promotes weight gain and glucose intolerance. Cell Metab. 20, 678–686 (2014).
Geula, S. et al. m6A mRNA methylation facilitates resolution of naïve pluripotency toward differentiation. Science 347, 1002–1006 (2015).
Wang, X. et al. N-6-methyladenosine-dependent regulation of messenger RNA stability. Nature 505, 117–120 (2014).
Zaccara, S. & Jaffrey, S. R. A unified model for the function of YTHDF proteins in regulating m(6)A-modified mRNA. Cell 181, 1582–1595 (2020).
Lasman, L. et al. Context-dependent functional compensation between Ythdf m(6)A reader proteins. Genes Dev. 34, 1373–1391 (2020).
Moylan, J. S. et al. Neutral sphingomyelinase-3 mediates TNF-stimulated oxidant activity in skeletal muscle. Redox Biol. 2, 910–920 (2014).
Heinrich, M. et al. Cathepsin D links TNF-induced acid sphingomyelinase to Bid-mediated caspase-9 and-3 activation. Cell Death Differ. 11, 550–563 (2004).
Stith, J. L., Velazquez, F. N. & Obeid, L. M. Advances in determining signaling mechanisms of ceramide and role in disease. J. Lipid Res. 60, 913–918 (2019).
Hammerschmidt, P. et al. CerS6-derived sphingolipids interact with Mff and promote mitochondrial fragmentation in obesity. Cell 177, 1536–1552 (2019).
Ogura, Y. et al. Postnatal changes in gene expression of retinal dehydrogenase and retinoid receptors in liver of rats. Life Sci. 74, 1519–1528 (2004).
Barajas, J. M. et al. METTL3 regulates liver homeostasis, hepatocyte ploidy, and Circadian rhythm-controlled gene expression in mice. Am. J. Pathol. 192, 18–20 (2022).
Poh, H. X., Mirza, A. H., Pickering, B. F. & Jaffrey, S. R. Alternative splicing of METTL3 explains apparently METTL3-independent m6A modifications in mRNA. PLoS Biol. 20, e3001683 (2022).
Ivanova, I. et al. The RNA m6A reader YTHDF2 is essential for the post-transcriptional regulation of the maternal transcriptome and oocyte competence. Mol. Cell 67, 1059–1067 (2017).
Zhao, B. S. & He, C. ‘Gamete On’ for m6A: YTHDF2 exerts essential functions in female fertility. Mol. Cell 67, 903–905 (2017).
Holland, W. L. et al. Lipid-induced insulin resistance mediated by the proinflammatory receptor TLR4 requires saturated fatty acid-induced ceramide biosynthesis in mice. J. Clin. Invest. 121, 1858–1870 (2011).
Holland, W. L. et al. An FGF21-adiponectin-ceramide axis controls energy expenditure and insulin action in mice. Cell Metab. 17, 790–797 (2013).
Jiang, C. T. et al. Intestinal farnesoid X receptor signaling promotes nonalcoholic fatty liver disease. J. Clin. Invest. 125, 386–402 (2015).
Salisbury, D. A. et al. Transcriptional regulation of N(6)-methyladenosine orchestrates sex-dimorphic metabolic traits. Nat. Metab. 3, 940–953 (2021).
Mizutani, Y., Kihara, A. & Igarashi, Y. Mammalian Lass6 and its related family members regulate synthesis of specific ceramides. Biochem. J. 390, 263–271 (2005).
Teng, Y. et al. Novel function of SART1 in HNF4α transcriptional regulation contributes to its antiviral role during HBV infection. J. Hepatol. 75, 1072–1082 (2021).
Li, W. C., Ralphs, K. L. & Tosh, D. Isolation and culture of adult mouse hepatocytes. Methods Mol. Biol. 633, 185–196 (2010).
Vosahlikova, M. et al. Effect of therapeutic concentration of lithium on live HEK293 cells; increase of Na+/K+-ATPase, change of overall protein composition and alteration of surface layer of plasma membrane. Biochim. Biophys. Acta Gen. Subj. 1861, 1099–1112 (2017).
Zhou, Y. et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 10, 1523 (2019).
Dominissini, D., Moshitch-Moshkovitz, S., Salmon-Divon, M., Amariglio, N. & Rechavi, G. Transcriptome-wide mapping of N(6)-methyladenosine by m(6)A-seq based on immunocapturing and massively parallel sequencing. Nat. Protoc. 8, 176–189 (2013).
Liu, Y. et al. N (6)-methyladenosine RNA modification-mediated cellular metabolism rewiring inhibits viral replication. Science 365, 1171–1176 (2019).
Dong, L. et al. Relaxed initiation pausing of ribosomes drives oncogenic translation. Sci. Adv. 7, eabd6927 (2021).
Dixit, D. et al. The RNA m6A reader YTHDF2 maintains oncogene expression and is a targetable dependency in glioblastoma stem cells. Cancer Discov. 11, 480–499 (2021).
Acknowledgements
This study was supported by the National Key Research and Development Program of China to S.L. and D.Y. (no. 2020YFA0804400); the National Natural Science Foundation of China (nos. 82270621, 81802799, 82071854, 81971936, 32100590, 32270820 and 32000857); Shandong Provincial Natural Science Foundation, China (nos. ZR2019BH002, ZR2020QH038, ZR2020MH269, ZR2021QB012 and ZR2022MH150); Jiangsu Provincial Natural Science Foundation, China (no. BK20180222); Funds for Youth Interdisciplinary and Innovation Research Groups of Shandong University (nos. 2020QNQT003 and 2020QNQT009); The Fundamental Research Funds for the Central Universities (nos. 2022JC008 and 2042021gf0013); Hubei Province’s Outstanding Medical Academic Leader Program; Basic and Clinical Medical Research Joint Fund of Zhongnan Hospital, Wuhan University; and Taishan Scholars Programme of Shandong Province. M.H. was supported by an ERC consolidator grant and Rainer Hoenig Stiftung. We thank S. Huang (Center for Reproductive Medicine, Shandong University) and the Core Facility of Advanced Medical Research Institute of Shandong University for technical support.
Author information
Authors and Affiliations
Contributions
D.Y., M.H., S.L. and Y.X. designed experiments, interpreted data and wrote the manuscript. S.W. and S.C. contributed to experimental design and performed in vivo animal studies. S.W., P.H., J.S., B.X., X.L., L.L. and H.Z. performed in vitro experiments. Y.Z. and Z.X. performed AAV8 packaging. P.Z., P.M. and C.Z. contributed to histological analysis. All authors provided input and reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Metabolism thanks B. Finck, W. Holland and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Isabella Samuelson, in collaboration with the Nature Metabolism team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Gene expression patterns in mouse liver development.
a, Eight k-means clusters showing different expression trends in murine livers at different stages of development determined by the normalized gene expression using the Z-score transformation method. b, The average temporal expression patterns of genes in the eight clusters. c, Heatmap of enriched GO and KEGG terms colored by p-values.
Extended Data Fig. 2 Dynamic changes of Mettl3 expression during postnatal liver development.
a, qRT-PCR of livers from WT C57BL/6J mice at different ages for indicated genes (n = 4 for Afp, Apoe, Fabp1, Fasn, Ppara, Fbp1 and Pck1; n = 6 for Ythdf2, Ythdf3, Alkbh5 and Ythdf1). Data are shown in mean ± SEM. b, IHC of Mettl3 in livers from WT C57BL/6J mice at different ages as indicated. Scale bar, 20 μm.
Extended Data Fig. 3 Hepatic Mettl3 deficiency induces hepatocyte hypertrophy and liver injury during postnatal development.
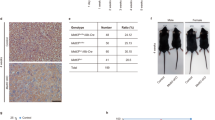
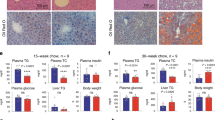
a, Schematic representation of genomic Mettl3 (top), floxed Mettl3 and deleted Mettl3 (bottom) alleles. b, H&E staining of liver sections from 5-week-old WT and Mettl3ΔHep mice (n = 3 biological independent samples). Scale bar, 50 μm. c, H&E staining of liver sections from 4-week-old WT and Mettl3ΔHep mice (n = 3 biological independent samples). Scale bar, 100 μm. d, p-Akt levels in livers from WT and Mettl3ΔHep mice in response to fasting and refeeding (n = 3 biological independent samples). e, IHC of CD45, CD3 and B220 in livers from WT and Mettl3ΔHep mice. Scale bar, 20 μm. (n = 3 biological independent samples). f, qRT-PCR of livers from WT (n = 5) and Mettl3ΔHep (n = 5) mice for indicated genes. g, Timeline of AAV8-induced Cre expression in adult Mettl3fl/fl mice and immunoblotting for Mettl3 showing the Knock-out effect. h, IHC of Mettl3 in livers from AAV8-Mock and AAV8-Cre Mettl3fl/fl mice. Black arrowheads, hepatocytes; Red arrowheads, non-parenchymal cells. n = 3 biological independent samples. Scale bar, 50 μm. i, Representative picture of AAV8-Mock and AAV8-Cre Mettl3fl/fl mice 2 months after i.v. injection (n = 3 biological independent samples). Scale bar, 5cm. j, Serum ALT and AST in AAV8-Mock (n = 3) and AAV8-Cre Mettl3fl/fl mice (n = 3). k, Gross appearance of livers in AAV8-Mock and AAV8-Cre Mettl3fl/fl mice 2 months after i.v. injection (n = 3 biological independent samples). The scale bar represents 1 cm. l, Representative H&E staining of livers from AAV8-Mock and AAV8-Cre Mettl3fl/fl mice (n = 3 biological independent samples). Scale bar, 50 μm. Data are shown in mean ± SEM; ns, not significant, *p < 0.05, **p < 0.01, ***p < 0.001 by unpaired two-tailed Student’s t test.
Extended Data Fig. 4 Livers of AAV8-Cre mice show no signs of damage.
Representative H&E staining of livers from AAV8-Mock, AAV8-Cre and Mettl3ΔHep mice at different time points (n = 3 biological independent samples). Scale bar, 20 μm.
Extended Data Fig. 5 meRIP and mRNA stability assay for sphingolipid metabolism-related genes.
a, The m6A motif and m6A modification for sphingolipid metabolism-related genes. b, mRNA stability analysis in primary hepatocytes isolated from Mettl3ΔHep (n = 3) versus WT (n = 3) mice treated with actinomycin D (5 μg/mL) for the indicated times. The residual RNAs were normalized to the value of time 0. Data are shown in mean ± SEM; ns, not significant, *p < 0.05, **p < 0.01 by two-way ANOVA statistics (b).
Extended Data Fig. 6 METTL3 regulates SMPD3 expression in HEK293T cells and H1 ESCs.
a, The GGAC motif for human SMPD3. b, Immunoblotting for METTL3 and SMPD3 in HEK293T cells. c, Immunoblotting for METTL3 and SMPD3 in H1 ESCs. d, m6A enrichment of the SMPD3 mRNAs in sgREN (n = 4) versus sgMETTL3 clone #1 (n = 4) and clone #5 (n = 4) in H1 cells by m6A-RIP-qPCR. Data are shown in mean ± SEM; ****p < 0.0001 by unpaired two-tailed Student’s t test.
Extended Data Fig. 7 Mettl3 deficiency in hepatocytes causes mitochondrial damage and ER stress.
a, Membrane fluidity of primary hepatocytes from WT and Mettl3ΔHep mice determined by TMA-DPH (n = 7 biological independent samples). b, Electron microscopy of WT and Mettl3ΔHep livers. Arrowheads indicate the perinuclear space. Scale bar, 5 μm. c, Mitochondrial membrane potential assessment of primiary hepatocytes from WT and Mettl3ΔHep mice 24 hours after isolation with the mitochondria-specific probe JC-1. Red and green fluorescence indicate J-aggregates and JC-1 monomers, respectively (n = 3 biological independent samples). Scale bar, 100 μm. d, Mitochondrial membrane potential assessment of primiary hepatocytes from WT and Mettl3ΔHep mice 48 hours after isolation with the mitochondria-specific probe JC-1. Red and green fluorescence indicate J-aggregates and JC-1 monomers, respectively (n = 3 biological independent samples). Scale bar, 100 μm. **p < 0.01 by unpaired two-tailed Student’s t test.
Extended Data Fig. 8 Inhibition of ceramide synthesis by GW4869 ameliorated mitochondrial dysfunction, apoptosis and hepatocyte injury in Mettl3ΔHep mice.
a, H&E of WT and Mettl3ΔHep livers treated with vehicle or different inhibitors as indicated (n = 3 biological independent samples). Scale bar, 100 μm. b, MDA, Cyt-c, BIP and p-eIF2α IHC in Mettl3ΔHep livers treated with vehicle versus GW4869 (n = 4 biological independent samples). Scale bar, 50 μm. c-g, IHC of cl-Casp3 (c), TUNEL (d), CD45 (e), F480 (f) and Ki67 (g) in Mettl3ΔHep livers treated with vehicle versus GW4869 (n = 4 biological independent samples). Scale bar, 50 μm. h, qRT-PCR of Mettl3ΔHep livers treated with vehicle versus GW4869 for indicated genes (n = 4 biological independent samples). Data are shown in mean ± SEM; ns, not significant, *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001 by unpaired two-tailed Student’s t test.
Extended Data Fig. 9 The liver damage in Mettl3ΔHep mice were reduced by Smpd3 knockdown.
a,b, Silencing efficiency of siRNAs targeting Smpd3 was detected by qRT-PCR (n = 4) (a) and immunoblotting (b) in mouse liver cell line AML12. c, qRT-PCR detection of sphingolipid metabolism-related genes in livers from WT (n = 4) and Mettl3ΔHep mice treated with (n = 4) or siSmpd3 (n = 5). d, IHC of MDA, p-eIF2α, CD45 and Epcam from siNC- or siSmpd3-treated livers. (n = 3 biological independent samples). e, Quantification for Atf3, Bax, Mettl3, cl-Casp3 and PCNA proteins in Fig. 8k (n = 3). f, qRT-PCR of livers from WT (n = 4) and Mettl3ΔHep mice treated with siNC (n = 4) or siSmpd3 (n = 5) for indicated genes. Data are shown in mean ± SEM; ns, not significant, *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001 by unpaired two-tailed Student’s t test.
Extended Data Fig. 10 Sgms1 overexpression attenuated liver injury in Mettl3ΔHep mice.
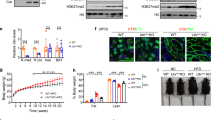
a, Timeline of AAV8-Sgms1 and AAV8-EGFP treatment in Mettl3ΔHep mice. b, qRT-PCR was used to detect the expression of Sgms1 in AAV-EGFP (n = 5) and AAV-Sgms1 (n = 4) livers. c, Representative IHC of Sgms1 in AAV-EGFP and AAV-Sgms1 livers. Scale bar, 20 μm. d, Mouse body weight gain over time post AAV8 injection (n = 4 for each group). e, Liver injury was assessed by serum AST and ALT from mice in (b) (n = 4/5/4). f, Representative liver macroscopy. Scale bar, 1 cm. g, Representative H&E staining of livers from mice in (f). Scale bar, 50 μm. h, Western blot of liver lysates from mice in (b) for indicated proteins. i, Quantification for the indicated proteins in (h) (n = 3). j, qRT-PCR of livers from mice in (b) for indicated genes (n = 4/5/4). k, IHC of cl-Casp3, Cyt-c, CD45 and Ki67 from AAV-EGFP and AAV-Sgms1 livers, and quantification for cl-Casp3 and Ki67 staining (n = 5/4). Scale bar, 20 μm. l, p-eIF2a, MDA, BIP, Epcam and CK19 IHC in livers from Mettl3ΔHep mice injected with AAV-EGFP or AAV-Sgms1. Data are shown in mean ± SEM; *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001 by unpaired two-tailed Student’s t test (b, d, e, I, j and k) or by two-way ANOVA statistics (d).
Supplementary information
Supplementary Table 1
Downregulating genes for functional enrichment analysis and network construction.
Supplementary Table 2
Upregulating genes for functional enrichment analysis and network construction.
Supplementary Table 3
Neonatal-enriched genes for functional enrichment analysis and network construction.
Supplementary Table 4
Primers for qPCR.
Source data
Source Data Fig. 1
Statistical Source Data.
Source Data Fig. 2
Statistical Source Data.
Source Data Fig. 3
Statistical Source Data.
Source Data Fig. 4
Statistical Source Data.
Source Data Fig. 5
Statistical Source Data.
Source Data Fig. 6
Statistical Source Data.
Source Data Fig. 7
Statistical Source Data.
Source Data Fig. 8
Statistical Source Data.
Source Data Extended Data Fig. 2
Statistical Source Data.
Source Data Extended Data Fig. 3
Statistical Source Data.
Source Data Extended Data Fig. 5
Statistical Source Data.
Source Data Extended Data Fig. 6
Statistical Source Data.
Source Data Extended Data Fig. 7
Statistical Source Data.
Source Data Extended Data Fig. 8
Statistical Source Data.
Source Data Extended Data Fig. 9
Statistical Source Data.
Source Data Extended Data Fig. 10
Statistical Source Data.
Source data Figs. 2, 4 and 6–8 and Extended Data Figs. 3, 7 and 10
Unprocessed immunoblots.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, S., Chen, S., Sun, J. et al. m6A modification-tuned sphingolipid metabolism regulates postnatal liver development in male mice. Nat Metab 5, 842–860 (2023). https://doi.org/10.1038/s42255-023-00808-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-023-00808-9
This article is cited by
-
Writing and erasing ceramides to alter liver disease
Nature Metabolism (2023)