Abstract
Homeostasis maintains serum metabolites within physiological ranges. For glucose, this requires insulin, which suppresses glucose production while accelerating its consumption. For other circulating metabolites, a comparable master regulator has yet to be discovered. Here we show that, in mice, many circulating metabolites are cleared via the tricarboxylic acid cycle (TCA) cycle in linear proportionality to their circulating concentration. Abundant circulating metabolites (essential amino acids, serine, alanine, citrate, 3-hydroxybutyrate) were administered intravenously in perturbative amounts and their fluxes were measured using isotope labelling. The increased circulating concentrations induced by the perturbative infusions hardly altered production fluxes while linearly enhancing consumption fluxes and TCA contributions. The same mass action relationship between concentration and consumption flux largely held across feeding, fasting and high- and low-protein diets, with amino acid homeostasis during fasting further supported by enhanced endogenous protein catabolism. Thus, despite the copious regulatory machinery in mammals, circulating metabolite homeostasis is achieved substantially through mass action-driven oxidation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All data and materials will be provided on reasonable request to the lead corresponding author (joshr@princeton.edu). Source data are provided with this paper.
References
Smith, D. A. & Dalvie, D. Why do metabolites circulate? Xenobiotica 42, 107–126 (2012).
Green, C. L. & Lamming, D. W. Regulation of metabolic health by essential dietary amino acids. Mech. Ageing Dev. 177, 186–200 (2019).
Balkau, B. et al. High blood glucose concentration is a risk factor for mortality in middle-aged nondiabetic men: 20-year follow-up in the Whitehall Study, the Paris Prospective Study, and the Helsinki Policemen Study. Diabetes Care 21, 360–367 (1998).
Bjørnholt, J. V. et al. Fasting blood glucose: an underestimated risk factor for cardiovascular death. Results from a 22-year follow-up of healthy nondiabetic men. Diabetes Care 22, 45–49 (1999).
Newgard, C. B. et al. A branched-chain amino acid-related metabolic signature that differentiates obese and lean humans and contributes to insulin resistance. Cell Metab. 9, 311–326 (2009).
Jervis, G. A. Studies on phenylpyruvic oligophrenia; the position of the metabolic error. J. Biol. Chem. 169, 651–656 (1947).
Gietzen, D. W. & Rogers, Q. R. Nutritional homeostasis and indispensable amino acid sensing: a new solution to an old puzzle. Trends Neurosci. 29, 91–99 (2006).
Bröer, S. & Bröer, A. Amino acid homeostasis and signalling in mammalian cells and organisms. Biochem. J. 474, 1935–1963 (2017).
McAninch, E. A. & Bianco, A. C. Thyroid hormone signaling in energy homeostasis and energy metabolism. Ann. N. Y. Acad. Sci. 1311, 77–87 (2014).
Mullur, R., Liu, Y.-Y. & Brent, G. A. Thyroid hormone regulation of metabolism. Physiol. Rev. 94, 355–382 (2014).
Kalra, S. & Gupta, Y. The insulin:glucagon ratio and the choice of glucose-lowering drugs. Diabetes Ther. 7, 1–9 (2016).
Aronoff, S. L., Berkowitz, K., Shreiner, B. & Want, L. Glucose metabolism and regulation: beyond insulin and glucagon. Diabetes Spectr. 17, 183–190 (2004).
Breckenridge, S. M., Cooperberg, B. A., Arbelaez, A. M., Patterson, B. W. & Cryer, P. E. Glucagon, in concert with insulin, supports the postabsorptive plasma glucose concentration in humans. Diabetes 56, 2442–2448 (2007).
Hayashi, Y. & Yusuke, S. Regulation of amino acid metabolism and α-cell proliferation by glucagon. J. Diabetes Investig. 9, 464–472 (2018).
Galsgaard, K. D. et al. Glucose and amino acid metabolism in mice depend mutually on glucagon and insulin receptor signaling. Am. J. Physiol. Endocrinol. Metab. 316, E660–E673 (2019).
Joshi, M. A. et al. Impaired growth and neurological abnormalities in branched-chain α-keto acid dehydrogenase kinase-deficient mice. Biochem. J. 400, 153–162 (2006).
Neinast, M. D. et al. Quantitative analysis of the whole-body metabolic fate of branched-chain amino acids. Cell Metab. 29, 417–429.e4 (2019).
Hui, S. et al. Glucose feeds the TCA cycle via circulating lactate. Nature 551, 115–118 (2017).
Donner, C. C., Fraze, E., Chen, Y. D. & Reaven, G. M. Quantitation of insulin-stimulated glucose disposal in patients with non-insulin-dependent diabetes mellitus. Diabetes 34, 831–835 (1985).
Chen, Y. D., Jeng, C. Y., Hollenbeck, C. B., Wu, M. S. & Reaven, G. M. Relationship between plasma glucose and insulin concentration, glucose production, and glucose disposal in normal subjects and patients with non-insulin-dependent diabetes. J. Clin. Invest. 82, 21–25 (1988).
Begg, D. P. & Woods, S. C. The endocrinology of food intake. Nat. Rev. Endocrinol. 9, 584 (2013).
Alberti, K. G., Johnston, D. G., Gill, A., Barnes, A. J. & Orskov, H. Hormonal regulation of ketone-body metabolism in man. Biochem. Soc. Symp. 43, 163–182 (1978).
Brosnan, J. T. & Brosnan, M. E. Branched-chain amino acids: enzyme and substrate regulation. J. Nutr. 136, 207S–211S (2006).
Tso, S.-C. et al. Benzothiophene carboxylate derivatives as novel allosteric inhibitors of branched-chain α-ketoacid dehydrogenase kinase. J. Biol. Chem. 289, 20583–20593 (2014).
Brown, R. P., Delp, M. D., Lindstedt, S. L., Rhomberg, L. R. & Beliles, R. P. Physiological parameter values for physiologically based pharmacokinetic models. Toxicol. Ind. Health 13, 407–484 (1997).
Chaix, A., Lin, T., Le, H. D., Chang, M. W. & Panda, S. Time-restricted feeding prevents obesity and metabolic syndrome in mice lacking a circadian clock. Cell Metab. 29, 303–319.e4 (2019).
Adamovich, Y., Ladeuix, B., Golik, M., Koeners, M. P. & Asher, G. Rhythmic oxygen levels reset circadian clocks through HIF1α. Cell Metab. 25, 93–101 (2017).
Chevalier, S., Gougeon, R., Kreisman, S. H., Cassis, C. & Morais, J. A. The hyperinsulinemic amino acid clamp increases whole-body protein synthesis in young subjects. Metabolism 53, 388–396 (2004).
Davis, T. A. et al. Stimulation of protein synthesis by both insulin and amino acids is unique to skeletal muscle in neonatal pigs. Am. J. Physiol. Endocrinol. Metab. 282, E880–E890 (2002).
Caso, G. et al. The increase in human muscle protein synthesis induced by food intake is similar when assessed with the constant infusion and flooding techniques. J. Nutr. 136, 1504–1510 (2006).
Sinturel, F. et al. Diurnal oscillations in liver mass and cell size accompany ribosome assembly cycles. Cell 169, 651–663.e14 (2017).
Hackett, S. R. et al. Systems-level analysis of mechanisms regulating yeast metabolic flux. Science 354, aaf2786 (2016).
Cornish-Bowden, A. & Cárdenas, M. L. Glucokinase: a monomeric enzyme with positive cooperativity. Front. Diabetes 16, 125–134 (2004).
Lowry, O. H., Carter, J., Ward, J. B. & Glaser, L. The effect of carbon and nitrogen sources on the level of metabolic intermediates in Escherichia coli. J. Biol. Chem. 246, 6511–6521 (1971).
Garland, P. B., Randle, P. J. & Newsholme, E. A. Citrate as an intermediary in the inhibition of phosphofructokinase in rat heart muscle by fatty acids, ketone bodies, pyruvate, diabetes and starvation. Nature 200, 169–170 (1963).
Balestri, F., Barsotti, C., Lutzemberger, L., Camici, M. & Ipata, P. L. Key role of uridine kinase and uridine phosphorylase in the homeostatic regulation of purine and pyrimidine salvage in brain. Neurochem. Int. 51, 517–523 (2007).
Egger, G., Liang, G., Aparicio, A. & Jones, P. A. Epigenetics in human disease and prospects for epigenetic therapy. Nature 429, 457–463 (2004).
Kerscher, O., Felberbaum, R. & Hochstrasser, M. Modification of proteins by ubiquitin and ubiquitin-like proteins. Annu. Rev. Cell Dev. Biol. 22, 159–180 (2006).
Dohmen, R. J. SUMO protein modification. Biochim. Biophys. Acta 1695, 113–131 (2004).
Scott, M., Gunderson, C. W., Mateescu, E. M., Zhang, Z. & Hwa, T. Interdependence of cell growth and gene expression: origins and consequences. Science 330, 1099–1102 (2010).
Schaechter, M., Maaløe, O. & Kjeldgaard, N. O. Dependency on medium and temperature of cell size and chemical composition during balanced grown of Salmonella typhimurium. Microbiology 19, 592–606 (1958).
Cayley, S., Lewis, B. A., Guttman, H. J. & Record, M. T. Jr Characterization of the cytoplasm of Escherichia coli K-12 as a function of external osmolarity: implications for protein–DNA interactions in vivo. J. Mol. Biol. 222, 281–300 (1991).
McGarry, J. D., Mannaerts, G. P. & Foster, D. W. A possible role for malonyl-CoA in the regulation of hepatic fatty acid oxidation and ketogenesis. J. Clin. Invest. 60, 265–270 (1977).
Rubio, V., Britton, H. G. & Grisolia, S. Mitochondrial carbamoyl phosphate synthetase activity in the absence of N-acetyl-L-glutamate. Mechanism of activation by this cofactor. Eur. J. Biochem. 134, 337–343 (1983).
Bouskila, M. et al. Allosteric regulation of glycogen synthase controls glycogen synthesis in muscle. Cell Metab. 12, 456–466 (2010).
Lu, W. et al. LC–MS and GC–MS based metabolomics platform for cancer research. Cancer Metab. 2, P41 (2014).
Su, X., Lu, W. & Rabinowitz, J. D. Metabolite spectral accuracy on orbitraps. Anal. Chem. 89, 5940–5948 (2017).
Acknowledgements
S.H. was supported by a National Institutes of Health (NIH) grant (no. 4R00DK117066). T.G.A. was supported by NIH grant no. DK109714 and a U.S. Department of Agriculture National Institute of Food and Agriculture grant no. NC1184-NJ14240. C.J. was supported by NIH grant no. 1R01AA02912. This work was supported by the NIH Pioneer (no. 1DP1DK113643) and Paul G. Allen Family Foundation grants (no. 0034665) and Ludwig Cancer Research.
Author information
Authors and Affiliations
Contributions
X.L., S.H., C.J. and J.D.R. designed the study. X.L. performed most of the experiments and data analysis. E.T.M. and T.G.A. contributed to the breeding and BCAA infusions on BCKDK knockout mice. W.O.J. performed the comprehensive laboratory animal monitoring system study on the BCKDK knockout mice. W.D.L. contributed to the hyperinsulinaemic–euglycaemic clamp study. X.Z. provided the portal vein data for the amino acids. X.L., C.J. and J.D.R. wrote the manuscript. All authors discussed and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
J.D.R. is a cofounder and stockholder in Toran and Serien Therapeutics and advisor to and stockholder in Agios Pharmaceuticals, Kadmon, Bantam Pharmaceutical, Colorado Research Partners, Rafael Pharmaceuticals, Barer Institute and L.E.A.F. Pharmaceuticals. The other authors declare no competing interests.
Peer review information
Nature Metabolism thanks Adam Rose and the other, anonymous, reviewers for their contribution to the peer review of this work. Alfredo Gimenez-Cassina and George Caputa were the primary handling editors.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Perturbative infusion outcomes for different regulatory mechanisms.
Green arrows reflect labeled metabolite fluxes including the experimenter-controlled influx from infusion. Blue arrows reflect unlabeled metabolite fluxes, including endogenous production. Green and blue circles are labeled and unlabeled metabolites, respectively. Red circles represent red blood cells. (a) Mass action. Labeled metabolites accumulate linearly with infusion rate, with unlabeled metabolite concentrations and fluxes not altered. (b) Active consumption induction. Labeled metabolites accumulate less than linearly with infusion rate, with unlabeled metabolite levels decreased but fluxes unaltered. (c) Consumption saturation. Labeled metabolites accumulate more than linearly with infusion rate, with unlabeled metabolites levels increased but fluxes unaltered. (d) Feedback inhibition of production. Unlabeled metabolite levels and fluxes are decreased.
Extended Data Fig. 2 Clearance of metabolite boluses is consistent with mass action kinetics.
(a) Mice were fasted from 9 AM to 5 PM (8 h fasting). At 5 PM, mice were injected with an intravenous bolus of the indicated [U-13C] metabolite at a low, medium, or high dose (as specified in the methods). Blood was taken 5, 15, 30, and 60 min after the injection, and the concentration of labeled metabolite in serum was measured by LC-MS. For experiments involving branched-chain amino acids, all three were given together, with only the indicated amino acid in labeled form. Labeled metabolite concentration was plotted against time post bolus. Lines are exponential decay curves fitted with mean value of each group. (b) Pseudo-first-order consumption rate constants from bolus and perturbative infusions. The slopes (α) calculated from the infusion experiments were plotted against the elimination constants (γ) calculated from the bolus experiments for each circulating metabolite. Line is linear regression fit.
Extended Data Fig. 3 Elevated portal vein alanine in fed mice.
(a) Circulating metabolite concentrations in the fasted and refed state. Fasting and feeding schedules were the same as Fig. 4a. Blood was taken at 5 PM for the fasted group and 11 PM for the refed group. Mean±SD. n = 4 mice. (b, c) Insulin does not alter concentrations or consumption fluxes of valine, lysine, and alanine. (b) Serum metabolite levels from hyperinsulinemic-euglycemic clamp (2.5 mU/kg/min insulin) and control (saline) experiments. Mice were fasted from 10 AM to 5 PM. The clamp was performed from 3 PM to 5 PM and blood was collected at 5 PM. Mean±s.d. n = 4 mice. (c) Consumption fluxes in the above clamp condition based on non-perturbative infusion of a mixture of 13C-valine, 13C-lysine, and 13C-alanine. The 13C-infusion was initiated 2.5 h prior to starting insulin to induce the hyperinsulinemic clamp and continued throughout the clamp experiment. Blood samples were taken immediately prior to or 120-min after initiation of the clamp. Mean±s.d. n = 4 mice. (d) Metabolite concentration ratios between the portal vein and tail vein of fasted (7.5 h fast starting at 9:30 AM, with sampling at 5 PM) or ad lib fed mice (with sampling at 11 PM). Mean±s.d. n = 4 mice. (e) Calculation of alanine consumption flux-concentration relationship using tail or portal vein data. Portal vein concentrations were calculated by multiplying alanine data from Fig. 4a by the portal/tail vein TIC ratio from (d). Lines are linear fits to each dataset.
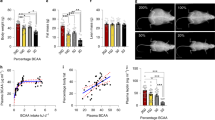
Extended Data Fig. 4 BCKDK whole-body KO mice are lethargic when fed low protein diet.
(a) Valine consumption flux versus circulating concentration in whole-body BCKDK knockout (KO) mice and littermate control mice (WT). Both groups were infused with [U-13C]valine as in Fig. 2. Lines are linear fits to each dataset. (b) KO and WT mice were fed either 20% or 5% protein diet for 7 days and blood was taken at 5 PM on the last day. Mean±SD. n = 5 WT and 4 KO mice. (c) Activity of KO and WT mice fed 5% protein diet as measured using metabolic chambers. Mean±SD. n = 6 mice.
Extended Data Fig. 5 TCA oxidation mediates mass action-driven consumption.
Tissue TCA labeling-concentration relationship for fasted perturbative infusions. Data are as in Fig. 5, except for (a) Use of succinate rather than malate to read out tissue TCA labeling or (b) Measurement of malate labeling across additional organs. Lines are linear fits to the data with intercept set to zero.
Extended Data Fig. 6 Protein synthesis rates are insensitive to branched-chain amino acid infusion.
(a) Calculation of protein synthesis rate. Mice were infused with [U-13C]valine as described in Fig. 6a (fasted group). Tissues were harvested and valine labeling fraction in hydrolyzed tissue proteins were plotted against time (left). Protein synthesis rate was then calculated based on the slopes (right) using the equation shown (below). Lines are linear fitting with mean values of each tissue. n = 2 mice per time point for each condition. (b) Tissue protein synthesis rates do not increase with infusion rate. n = 2 mice per time point for each condition. (c) Tissue protein synthesis rate does not change in response to perturbative valine infusion. Line is mean of the data as the slope is not significant. (d) Tissue protein labeling was consistent from different BCAAs. Mice were infused with [U-13C]valine, [U-13C]leucine, or [U-13C]isoleucine as in (a). n = 2 mice per time point for each condition.
Extended Data Fig. 7 Protein degradation rate does not change in response to perturbative valine infusion.
Data were from the experiments in Fig. 2 (valine panel). Line is mean of the data as the slope is not significant.
Extended Data Fig. 8 Raw data supporting the determination of protein synthesis rates after feeding.
Mice were infused with [U-13C]valine at the same condition as described in Fig. 6a (refed group). Lines are linear fitting with mean values of each tissue. n = 2 mice per timepoint.
Extended Data Fig. 9 Over 2 weeks, dietary protein fraction has little effect on body weight or food intake.
Mice were fed high- (HP), normal- (NP), and low-protein (LP) diets ad lib for 2 weeks. (a) Body weight gain. (b) Food intake. Mean ± SD, n = 6 mice.
Extended Data Fig. 10 TCA oxidation mediates mass action-driven valine consumption under high-, medium-, and low-protein diet.
Tissue malate labeling relative to serum valine labeling from non-perturbative infusion of [U-13C]valine, as in Fig. 7c. Lines are linear fits to the data with intercept set to zero.
Supplementary information
Supplementary Information
Supplementary Tables 1–7.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 10
Statistical source data.
Rights and permissions
About this article
Cite this article
Li, X., Hui, S., Mirek, E.T. et al. Circulating metabolite homeostasis achieved through mass action. Nat Metab 4, 141–152 (2022). https://doi.org/10.1038/s42255-021-00517-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-021-00517-1
This article is cited by
-
Trigonelline is an NAD+ precursor that improves muscle function during ageing and is reduced in human sarcopenia
Nature Metabolism (2024)
-
Metabolic mechanisms in physiological and pathological cardiac hypertrophy: new paradigms and challenges
Nature Reviews Cardiology (2023)
-
Comparison of fecal and blood metabolome reveals inconsistent associations of the gut microbiota with cardiometabolic diseases
Nature Communications (2023)
-
Metabolic balance—a masterclass in mass action
Nature Metabolism (2022)
-
Integrated landscape of cardiac metabolism in end-stage human nonischemic dilated cardiomyopathy
Nature Cardiovascular Research (2022)