Abstract
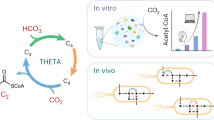
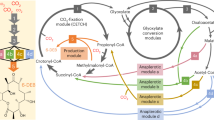
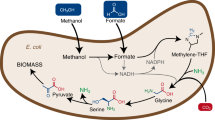
Metabolic engineering often entails concurrent engineering of substrate utilization, central metabolism and product synthesis pathways, inevitably creating interdependency with native metabolism. Here we report an alternative approach using synthetic pathways for C1 bioconversion that generate multicarbon products directly from C1 units and hence are orthogonal to the host metabolic network. The engineered pathways are based on formyl-CoA elongation (FORCE) reactions catalysed by the enzyme 2-hydroxyacyl-CoA lyase. We use thermodynamic and stoichiometric analyses to evaluate FORCE pathway variants, including aldose elongation, α-reduction and aldehyde elongation. Promising variants were prototyped in vitro and in vivo using the non-methylotrophic bacterium Escherichia coli. We demonstrate the conversion of formate, formaldehyde and methanol into various products including glycolate, ethylene glycol, ethanol and glycerate. FORCE pathways also have the potential to be integrated with the host metabolism for synthetic methylotrophy by the production of native growth substrates as demonstrated in a two-strain co-culture system.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
All data supporting the findings of this study are included in the paper and its supplementary information as well as the following public databases: MetaCyc (https://metacyc.org/), eQuilibrator (https://equilibrator.weizmann.ac.il/) and Uniprot (https://www.uniprot.org/). Uniprot accession numbers for enzymes involved in the study are available in Supplementary Table 1.
Code availability
The scripts used to perform the analyses in the study are available at https://github.com/ahc7/FORCE_manuscript/.
References
Nielsen, J. & Keasling, J. D. Engineering cellular metabolism. Cell 164, 1185–1197 (2016).
Csete, M. & Doyle, J. Bow ties, metabolism and disease. Trends Biotechnol. 22, 446–450 (2004).
Bar-Even, A., Noor, E. & Milo, R. A survey of carbon fixation pathways through a quantitative lens. J. Exp. Bot. 63, 2325–2342 (2012).
Kalyuzhnaya, M. G., Puri, A. W. & Lidstrom, M. E. Metabolic engineering in methanotrophic bacteria. Metab. Eng. 29, 142–152 (2015).
Bogorad, I. W. et al. Building carbon–carbon bonds using a biocatalytic methanol condensation cycle. Proc. Natl Acad. Sci. USA 111, 15928–15933 (2014).
Siegel, J. B. et al. Computational protein design enables a novel one-carbon assimilation pathway. Proc. Natl Acad. Sci. USA 112, 3704–3709 (2015).
Schwander, T., Schada von Borzyskowski, L., Burgener, S., Cortina, N. S. & Erb, T. J. A synthetic pathway for the fixation of carbon dioxide in vitro. Science 354, 900–904 (2016).
Lu, X. et al. Constructing a synthetic pathway for acetyl-coenzyme A from one-carbon through enzyme design. Nat. Commun. 10, 1378 (2019).
Kim, S. et al. Growth of E. coli on formate and methanol via the reductive glycine pathway. Nat. Chem. Biol. 16, 538–545 (2020).
Clomburg, J. M., Crumbley, A. M. & Gonzalez, R. Industrial biomanufacturing: the future of chemical production. Science 355, aag0804 (2017).
Liu, C. C., Jewett, M. C., Chin, J. W. & Voigt, C. A. Toward an orthogonal central dogma. Nat. Chem. Biol. 14, 103–106 (2018).
Pandit, A. V., Srinivasan, S. & Mahadevan, R. Redesigning metabolism based on orthogonality principles. Nat. Commun. 8, 1–11 (2017).
Costello, A. & Badran, A. H. Synthetic biological circuits within an orthogonal central dogma. Trends Biotechnol. https://doi.org/10.1016/j.tibtech.2020.05.013 (2020).
Natarajan, A. et al. Engineering orthogonal human O-linked glycoprotein biosynthesis in bacteria. Nat. Chem. Biol. 16, 1062–1070 (2020).
Black, W. B. et al. Engineering a nicotinamide mononucleotide redox cofactor system for biocatalysis. Nat. Chem. Biol. 16, 87–94 (2020).
Chou, A., Clomburg, J. M., Qian, S. & Gonzalez, R. 2-Hydroxyacyl-CoA lyase catalyzes acyloin condensation for one-carbon bioconversion. Nat. Chem. Biol. 15, 900–906 (2019).
Burgener, S., Cortina, N. S. & Erb, T. J. Oxalyl-CoA decarboxylase enables nucleophilic one-carbon extension of aldehydes to chiral α-hydroxy acids. Angew. Chem. Int. Ed. Engl. https://doi.org/10.1002/anie.201915155 (2020).
Müller, J. E. N. et al. Engineering Escherichia coli for methanol conversion. Metab. Eng. 28, 190–201 (2015).
Wu, T. Y. et al. Characterization and evolution of an activator-independent methanol dehydrogenase from Cupriavidus necator N-1. Appl. Microbiol. Biotechnol. 100, 4969–4983 (2016).
Whitaker, W. B. et al. Engineering the biological conversion of methanol to specialty chemicals in Escherichia coli. Metab. Eng. 39, 49–59 (2017).
Jonsson, S., Ricagno, S., Lindqvist, Y. & Richards, N. G. J. Kinetic and mechanistic characterization of the formyl-CoA transferase from Oxalobacter formigenes. J. Biol. Chem. 279, 36003–36012 (2004).
Sly, W. W. S. & Stadtman, E. R. Formate metabolism II. Enzymatic synthesis of formyl phosphate and formyl coenzyme a in Clostridium cylindrosporum. J. Biol. Chem. 238, 2639–2647 (1963).
Singh, R. K. et al. Insights into cell-free conversion of CO2 to chemicals by a multienzyme cascade reaction. ACS Catal. 8, 11085–11093 (2018).
Schuchmann, K. & Muller, V. Direct and reversible hydrogenation of CO2 to formate by a bacterial carbon dioxide reductase. Science 342, 1382–1385 (2013).
Roger, M., Brown, F., Gabrielli, W. & Sargent, F. Efficient hydrogen-dependent carbon dioxide reduction by Escherichia coli. Curr. Biol. 28, 140–145 (2018).
Sirajuddin, S. & Rosenzweig, A. C. Enzymatic oxidation of methane. Biochemistry 54, 2283–2294 (2015).
Felnagle, E. A., Chaubey, A., Noey, E. L., Houk, K. N. & Liao, J. C. Engineering synthetic recursive pathways to generate non-natural small molecules. Nat. Chem. Biol. 8, 518–526 (2012).
Kim, J., Hetzel, M., Boiangiu, C. D. & Buckel, W. Dehydration of (R)-2-hydroxyacyl-CoA to enoyl-CoA in the fermentation of α-amino acids by anaerobic bacteria. FEMS Microbiol. Rev. 28, 455–468 (2004).
Kandasamy, V. et al. Engineering Escherichia coli with acrylate pathway genes for propionic acid synthesis and its impact on mixed-acid fermentation. Appl. Microbiol. Biotechnol. 97, 1191–1200 (2013).
Clomburg, J. M., Vick, J. E., Blankschien, M. D., Rodríguez-Moyá, M. & Gonzalez, R. A synthetic biology approach to engineer a functional reversal of the β-oxidation cycle. ACS Synth. Biol. 1, 541–554 (2012).
Vick, J. E. et al. Escherichia coli enoyl-acyl carrier protein reductase (FabI) supports efficient operation of a functional reversal of the β-oxidation cycle. Appl. Environ. Microbiol. 81, 1406–1416 (2014).
Cheong, S., Clomburg, J. M. & Gonzalez, R. Energy-and carbon-efficient synthesis of functionalized small molecules in bacteria using non-decarboxylative Claisen condensation reactions. Nat. Biotechnol. 34, 556–561 (2016).
Buckel, W. et al. Enzyme catalyzed radical dehydrations of hydroxy acids. Biochim. Biophys. Acta 1824, 1278–1290 (2012).
Obradors, N., Cabiscol, E., Aguilar, J. & Ros, J. Site-directed mutagenesis studies of the metal-binding center of the iron-dependent propanediol oxidoreductase from Escherichia coli. Eur. J. Biochem. 258, 207–213 (1998).
Tobimatsu, T. et al. Heterologous expression, purification, and properties of diol dehydratase, an adenosylcobalamin-dependent enzyme of Klebsiella oxytoca. Arch. Biochem. Biophys. 347, 132–140 (1997).
Jain, R., Sun, X., Yuan, Q. & Yan, Y. Systematically engineering Escherichia coli for enhanced production of 1,2-propanediol and 1-propanol. ACS Synth. Biol. 4, 746–756 (2015).
Wang, J. et al. Rational engineering of diol dehydratase enables 1,4-butanediol biosynthesis from xylose. Metab. Eng. 40, 148–156 (2017).
Lennen, R. M. & Pfleger, B. F. Engineering Escherichia coli to synthesize free fatty acids. Trends Biotechnol. 30, 659–667 (2012).
Dellomonaco, C., Clomburg, J. M., Miller, E. N. & Gonzalez, R. Engineered reversal of the β-oxidation cycle for the synthesis of fuels and chemicals. Nature 476, 355–359 (2011).
Kunjapur, A. M. & Prather, K. L. J. Microbial engineering for aldehyde synthesis. Appl. Environ. Microbiol. 81, 1892–1901 (2015).
Noor, E. et al. Pathway thermodynamics highlights kinetic obstacles in central metabolism. PLoS Comput. Biol. 10, e1003483 (2014).
Mitsui, R., Omori, M., Kitazawa, H. & Tanaka, M. Formaldehyde-limited cultivation of a newly isolated methylotrophic bacterium, Methylobacterium sp. MF1: enzymatic analysis related to C1 metabolism. J. Biosci. Bioeng. 99, 18–22 (2005).
Meyer, F. et al. Methanol-essential growth of Escherichia coli. Nat. Commun. 9, 1508 (2018).
Bang, J. & Lee, S. Y. Assimilation of formic acid and CO2 by engineered Escherichia coli equipped with reconstructed one-carbon assimilation pathways. Proc. Natl Acad. Sci. USA 115, E9271–E9279 (2018).
De Graef, M. R., Alexeeva, S., Snoep, J. L. & Teixeira De Mattos, M. J. The steady-state internal redox state (NADH/NAD) reflects the external redox state and is correlated with catabolic adaptation in Escherichia coli. J. Bacteriol. 181, 2351–2357 (1999).
Berríos-Rivera, S. J., Bennett, G. N. & San, K. Y. The effect of increasing NADH availability on the redistribution of metabolic fluxes in Escherichia coli chemostat cultures. Metab. Eng. 4, 230–237 (2002).
San, K. Y. et al. Metabolic engineering through cofactor manipulation and its effects on metabolic flux redistribution in Escherichia coli. Metab. Eng. 4, 182–192 (2002).
Dudley, Q. M., Anderson, K. C. & Jewett, M. C. Cell-free mixing of Escherichia coli crude extracts to prototype and rationally engineer high-titer mevalonate synthesis. ACS Synth. Biol. 5, 1578–1588 (2016).
Rodriguez, G. M. & Atsumi, S. Toward aldehyde and alkane production by removing aldehyde reductase activity in Escherichia coli. Metab. Eng. 25, 227–237 (2014).
Caballero, E., Baldomá, L., Ros, J., Boronat, A. & Aguilar, J. Identification of lactaldehyde dehydrogenase and glycolaldehyde dehydrogenase as functions of the same protein in Escherichia coli. J. Biol. Chem. 258, 7788–7792 (1983).
Nattermann, M. et al. Engineering a highly efficient carboligase for synthetic one-carbon metabolism. ACS Catal. https://doi.org/10.1021/acscatal.1c01237 (2021).
Monk, J. M. et al. iML1515, a knowledgebase that computes Escherichia coli traits. Nat. Biotechnol. 35, 904–908 (2017).
Mak, W. S. et al. Integrative genomic mining for enzyme function to enable engineering of a non-natural biosynthetic pathway. Nat. Commun. 6, 10005 (2015).
Yishai, O., Bouzon, M., Döring, V. & Bar-Even, A. In vivo assimilation of one-carbon via a synthetic reductive glycine pathway in Escherichia coli. ACS Synth. Biol. 7, 2023–2028 (2018).
Schada von Borzyskowski, L. et al. Marine proteobacteria metabolize glycolate via the β-hydroxyaspartate cycle. Nature 575, 500–504 (2019).
Yamane, T. & Hirano, S. Semi-batch culture of microorganisms with constant feed of substrate: a mathematical simulation. J. Ferment. Technol. 55, 156–165 (1977).
Rudroff, F. Whole-cell based synthetic enzyme cascades—light and shadow of a promising technology. Curr. Opin. Chem. Biol. 49, 84–90 (2019).
Burg, J. M. et al. Large-scale bioprocess competitiveness: the potential of dynamic metabolic control in two-stage fermentations. Curr. Opin. Chem. Eng. 14, 121–136 (2016).
Dinh, C. V. & Prather, K. L. Layered and multi-input autonomous dynamic control strategies for metabolic engineering. Curr. Opin. Biotechnol. 65, 156–162 (2020).
Shen, X., Wang, J., Li, C., Yuan, Q. & Yan, Y. Dynamic gene expression engineering as a tool in pathway engineering. Curr. Opin. Biotechnol. 59, 122–129 (2019).
Kim, S. & Gonzalez, R. Selective production of decanoic acid from iterative reversal of β-oxidation pathway. Biotechnol. Bioeng. 115, 1311–1320 (2018).
Hernández Lozada, N. J. et al. Highly active C8-acyl-ACP thioesterase variant isolated by a synthetic selection strategy. ACS Synth. Biol. 7, 2205–2215 (2018).
Yan, Q. & Pfleger, B. F. Revisiting metabolic engineering strategies for microbial synthesis of oleochemicals. Metab. Eng. 58, 35–46 (2020).
Rohwerder, T., Rohde, M., Jehmlich, N. & Purswani, J. Actinobacterial degradation of 2-hydroxyisobutyric acid proceeds via acetone and Formyl-CoA by employing a thiamine-dependent lyase reaction. Front. Microbiol 11, 00691 (2020).
Caspi, R. et al. The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of pathway/genome databases. Nucleic Acids Res. 42, D459–D471 (2014).
Hannum, G. et al. Creation and analysis of biochemical constraint-based models using the COBRA Toolbox v.3.0. Nat. Protoc. 2, 727–738 (2019).
Sambrook, J., Fritsch, E. F., Maniatis, T. & Russell, D. W. Molecular Cloning: A Laboratory Manual (Cold Spring Harbor Laboratory Press, 2001).
Neidhardt, F. C., Bloch, P. L. & Smith, D. F. Culture medium for enterobacteria. J. Bacteriol. 119, 736–747 (1974).
Acknowledgements
We thank S. Qian for assistance with genetic methods. We thank M. Nattermann, S. Burgener and T. J. Erb (Max Planck Institute for Terrestrial Microbiology) for the suggestion and sequence of wild-type CaAbfT. We thank N. Paczia at the Max Planck Institute for Terrestrial Microbiology for informative suggestions on LC–MS analysis. We thank the University of South Florida Chemical Purification Analysis and Screening core facility for access to the Agilent 6540 Q-TOF LC–MS system with the help of L. Calcul. BsmHACL was identified and the DNA was provided as part of a grant from the US Department of Energy Joint Genome Institute DNA Synthesis programme. This work was supported by grants from the US National Science Foundation (CBET-1605999), the US Department of Energy (DE-EE0008499), and BASF Corporation. The funders had no role in study design, data collection and analysis, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
R.G. conceptualized the research and supervised the project. A.C., J.M.C., S.H.L., F.Z. and R.G. designed the methodology. A.C. performed thermodynamic analysis, flux balance analysis and cell-free experiments. A.C., F.Z. and J.M.C. performed resting-cell experiments. S.H.L. and J.M.C. performed methanol experiments. F.Z. performed purified enzyme experiments. S.H.L., F.Z. and J.M.C. performed the two-strain growth experiments. A.C., S.H.L. and F.Z. constructed E. coli strains. A.C., J.M.C., F.Z. and S.H.L. analysed the data. A.C., S.H.L., J.M.C., F.Z. and R.G. prepared the manuscript with feedback from all authors.
Corresponding author
Ethics declarations
Competing interests
A.C., J.M.C. and R.G. are co-inventors and assignees on a patent application (PCT/US2015/058121), which relates to the reported research. The remaining authors declare no competing interests.
Additional information
Peer review information Nature Metabolism thanks Marina Kalyuzhnaya, Steven Singer and the other, anonymous, reviewers for their contribution to the peer review of this work. Primary Handling Editor: Isabella Samuelson.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Canonical (a) and orthogonal, synthetic (b) architectures for biological C1 utilization.
a, ‘Bowtie’ architecture of metabolism in which carbon substrates are consolidated into central metabolites from which a host of products can be produced through fermentative and biosynthetic pathways. Metabolic engineering typically operates within this framework by manipulating either one or all of the three components of the bowtie. b, The orthogonal FORCE pathways serve as a platform for both product synthesis and for providing substrates/metabolites for growth. This is an alternative framework to the traditional approach, which feeds all carbon through central metabolism, and from which both products and biomass are derived.
Extended Data Fig. 2 An alternative FORCE pathway based on dehydration of the 2-hydroxyacyl-CoA and α-reduction.
The pathway resembles β-oxidation reversal (β-reduction)39. This pathway is also a potential route for the production of unsaturated products. HACL: 2-hydroxyacyl-CoA lyase; HACD: 2-hydroxyacyl-CoA dehydratase; TER: trans-2-enoyl-CoA reductase; ACR: acyl-CoA reductase.
Extended Data Fig. 3 The impact of NADH/NAD+ ratio on formaldehyde (top) and methanol (bottom) conversion to glycolate or acetate via FORCE pathways.
Vertical dotted lines indicate the physiological range of NADH/NAD+ ratios.
Extended Data Fig. 4 The impact of termination on the iterative aldose elongation pathway.
Termination by hydrolysis of the acyl-CoA to produce sugar acids increases the driving force of the pathway for low numbers of iterations, but the driving forces converge as the number of iterations increase.
Extended Data Fig. 5 Production of glycolate from formate by E. coli engineered with a formate-activating pathway.
Resting cell experiments were performed with a strain expressing CaAbfT and BsmHACL (blue bars) and the corresponding control lacking BsmHACL (orange bars). Cultures (2.5 OD600 = 2.5*109 CFU/mL) were incubated at 30 °C for 24 hours in 25 mL flasks shaking at 200 rpm using 10 mM formate (plus 1 mM formaldehyde) as carbon source (control cultures with 1 mM formaldehyde and no formate also shown). All data points are shown for n = 6 replicates. Bars are drawn to the mean values.
Extended Data Fig. 6 Predicted biomass electron and carbon yields from various C1 substrates by the implementation of select pathways enabling methylotrophy.
Abbreviations: Formald: formaldehyde, FORCE-Glycerald: FORCE pathway with reactions enabling glyceraldehyde production, RuMP: Ribulose monophosphate pathway, FORCE-Ac: FORCE pathway with reactions enabling acetate production, SACA: Synthetic Acetyl-CoA pathway, FORCE-Glycolate: FORCE pathway with reactions enabling glycolate production. The scenarios in bold correspond to the predicted flux maps illustrated in Fig. 8.
Extended Data Fig. 7 Paraformaldehyde solubilization rate and resting cell bioconversion with paraformaldehyde.
a, Solubilization rate of commercially available paraformaldehyde (pFALD) with different particle sizes. Solubilization rates are measured in 10 mL M9 media in a 25 mL flask at 30 °C shaking at 200 rpm. b, Resting cell bioconversion of strains expressing BsmHACL, LmACR and AldA induced with 40 μM cumate and 100 μM IPTG. 3 mg prilled paraformaldehyde is added to 20 mL M9 media (2.5 mM formaldehyde equivalent) in a 25 mL flask at 30 °C shaking at 200 rpm. Formaldehyde accumulates only at sub-millimolar concentrations under these conditions.
Extended Data Fig. 8 Time course profiles for glycolate, formate, and formaldehyde concentration and cell-growth of the sensor strain in the two-strain system with 5 mM paraformaldehyde.
a, Time course in which the producer strain did not express an HACL. b, Plates from a representative experiment of the time course shown in panel a. c, Time course in which the producer strain expresses HACL. d, Plates from a representative experiment of the time course shown in panel c. 50 μL of cultures (5 × 10−3 dilution) at various time points plated on minimal media plates containing 2.5 g/L glycolate. All data is shown for duplicate replicates (n = 2). Lines are drawn to the mean values.
Extended Data Fig. 9 Time course profiles for glycolate, formate, and formaldehyde concentration and cell-growth of the sensor strain in the two-strain system with 500 mM methanol.
a, Time course in which the producer strain did not express an HACL. b, Plates from a representative experiment of the time course shown in panel a. c, Time course in which the producer strain expresses HACL. d, Plates from a representative experiment of the time course shown in panel c. 50 μL of cultures (5×10−3 dilution) at various time points plated on minimal media plates containing 2.5 g/L glycolate. All data is shown for duplicate replicates (n = 2). Lines are drawn to the mean values.
Extended Data Fig. 10 Time course profiles for glycolate, and formaldehyde concentration in the two-strain system with 1 mM formaldehyde and 10 mM formate.
a, Time course in which the producer strain expresses BsmHACL. b, Plates from a representative experiment of the time course shown in panel a. 50 μL of cultures (5 × 10−3 dilution) at various time points plated on minimal media plates containing 2.5 g/L glycolate. All data is shown for duplicate replicates (n = 2). Lines are drawn to the mean values.
Supplementary information
Supplementary Information
Supplementary Tables 1–3 and Refs. 68–71.
Rights and permissions
About this article
Cite this article
Chou, A., Lee, S.H., Zhu, F. et al. An orthogonal metabolic framework for one-carbon utilization. Nat Metab 3, 1385–1399 (2021). https://doi.org/10.1038/s42255-021-00453-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-021-00453-0
This article is cited by
-
Revealing reaction intermediates in one-carbon elongation by thiamine diphosphate/CoA-dependent enzyme family
Communications Chemistry (2024)
-
Engineering a synthetic energy-efficient formaldehyde assimilation cycle in Escherichia coli
Nature Communications (2023)
-
Engineering a new-to-nature cascade for phosphate-dependent formate to formaldehyde conversion in vitro and in vivo
Nature Communications (2023)
-
Characterisation of acetogen formatotrophic potential using Eubacterium limosum
Applied Microbiology and Biotechnology (2023)
-
Rescuing yeast from cell death enables overproduction of fatty acids from sole methanol
Nature Metabolism (2022)