Abstract
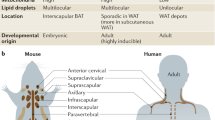
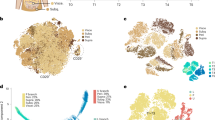
The perception of adipose tissue, both in the scientific community and in the general population, has changed dramatically in the past 20 years. While adipose tissue was thought for a long time to be a rather simple lipid storage entity, it is now recognized as a highly heterogeneous organ and a critical regulator of systemic metabolism, composed of many different subtypes of cells, with important endocrine functions. Additionally, adipose tissue is nowadays recognized to contribute to energy turnover, due to the presence of specialized thermogenic adipocytes, which can be found in many adipose depots. This review discusses the unprecedented insights that we have gained into the heterogeneity of thermogenic adipocytes and their respective precursors due to the technical developments in single-cell and nucleus technologies. These methodological advances have increased our understanding of how adipose tissue catabolic function is influenced by developmental and intercellular communication events.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Gesner, K. Historiae Animalium (Apud Christ. Froschouerum, 1551).
Hatai, S. On the presence in human embryos of an interscapular gland corresponding to the so-called hibernating gland of lower mammals. Anat. Anz. 21, 369–373 (1902).
Young, P., Arch, J. R. S. & Ashwell, M. Brown adipose tissue in the parametrial fat pad of the mouse. FEBS Lett. 167, 10–14 (1984).
Rondini, E. A. & Granneman, J. G. Single cell approaches to address adipose tissue stromal cell heterogeneity. Biochem. J. 477, 583–600 (2020).
Deutsch, A., Feng, D., Pessin, J. E. & Shinoda, K. The impact of single-cell genomics on adipose tissue research. Int. J. Mol. Sci. 21, 4773 (2020).
Hull, D. & Segall, M. M. Distinction of brown from white adipose tissue. Nature 212, 469–472 (1966).
Cadrin, M. et al. Immunohistochemical identification of the uncoupling protein in rat brown adipose tissue. J. Histochem. Cytochem. 33, 150–154 (1985).
Petrovic, N. et al. Chronic peroxisome proliferator-activated receptor γ (PPARγ) activation of epididymally derived white adipocyte cultures reveals a population of thermogenically competent, UCP1-containing adipocytes molecularly distinct from classic brown adipocytes. J. Biol. Chem. 285, 7153–7164 (2010).
Wu, J. et al. Beige adipocytes are a distinct type of thermogenic fat cell in mouse and human. Cell 150, 366–376 (2012).
Zingaretti, M. C. et al. The presence of UCP1 demonstrates that metabolically active adipose tissue in the neck of adult humans truly represents brown adipose tissue. FASEB J. 23, 3113–3120 (2009).
Lee, Y.-H., Kim, S.-N., Kwon, H.-J. & Granneman, J. G.Metabolic heterogeneity of activated beige/brite adipocytes in inguinal adipose tissue. Sci. Rep. 7, 39794 (2017).
Shinoda, K. et al. Genetic and functional characterization of clonally derived adult human brown adipocytes. Nat. Med. 21, 389–394 (2015).
Xue, R. et al. Clonal analyses and gene profiling identify genetic biomarkers of the thermogenic potential of human brown and white preadipocytes. Nat. Med. 21, 760–768 (2015).
Lee, K. Y. et al. Developmental and functional heterogeneity of white adipocytes within a single fat depot. EMBO J. 38, e99291 (2019).
Min, S. Y. et al. Diverse repertoire of human adipocyte subtypes develops from transcriptionally distinct mesenchymal progenitor cells. Proc. Natl Acad. Sci. USA 116, 17970–17979 (2019).
Rosenwald, M., Perdikari, A., Rülicke, T. & Wolfrum, C. Bi-directional interconversion of brite and white adipocytes. Nat. Cell Biol. 15, 659–667 (2013).
Lee, Y.-H., Petkova, A. P., Konkar, A. A. & Granneman, J. G. Cellular origins of cold-induced brown adipocytes in adult mice. FASEB J. 29, 286–299 (2015).
Wang, Q. A., Tao, C., Gupta, R. K. & Scherer, P. E. Tracking adipogenesis during white adipose tissue development, expansion and regeneration. Nat. Med. 19, 1338–1344 (2013).
Shao, M. et al. Cellular origins of beige fat cells revisited. Diabetes 68, 1874–1885 (2019).
Chen, Y. et al. Thermal stress induces glycolytic beige fat formation via a myogenic state. Nature 565, 180–185 (2019).
Tang, F. et al. mRNA-Seq whole-transcriptome analysis of a single cell. Nat. Methods 6, 377–382 (2009).
Cao, J. et al. A human cell atlas of fetal gene expression. Science 370, eaba7721 (2020).
Ngara, M. et al. Exploring parasite heterogeneity using single-cell RNA-seq reveals a gene signature among sexual stage Plasmodium falciparum parasites. Exp. Cell Res. 371, 130–138 (2018).
Brennecke, P. et al. Accounting for technical noise in single-cell RNA-seq experiments. Nat. Methods 10, 1093–1095 (2013).
Gierahn, T. M. et al. Seq-Well: portable, low-cost RNA sequencing of single cells at high throughput. Nat. Methods 14, 395–398 (2017).
Zheng, G. X. Y. et al. Haplotyping germline and cancer genomes with high-throughput linked-read sequencing. Nat. Biotechnol. 34, 303–311 (2016).
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
Klein, A. M. et al. Droplet barcoding for single-cell transcriptomics applied to embryonic stem cells. Cell 161, 1187–1201 (2015).
Rosenberg, A. B. et al. Single-cell profiling of the developing mouse brain and spinal cord with split-pool barcoding. Science 360, 176–182 (2018).
Rajbhandari, P. et al. Single cell analysis reveals immune cell–adipocyte crosstalk regulating the transcription of thermogenic adipocytes. eLife 8, e49501 (2019).
Sun, W. et al. snRNA-seq reveals a subpopulation of adipocytes that regulates thermogenesis. Nature 587, 98–102 (2020).
Schwalie, P. C. et al. A stromal cell population that inhibits adipogenesis in mammalian fat depots. Nature 559, 103–108 (2018).
Merrick, D. et al. Identification of a mesenchymal progenitor cell hierarchy in adipose tissue. Science 364, eaav2501 (2019).
Song, A. et al. Low- and high-thermogenic brown adipocyte subpopulations coexist in murine adipose tissue. J. Clin. Invest. 130, 247–257 (2020).
Karlina, R. et al. Identification and characterization of distinct brown adipocyte subtypes in C57BL/6J mice. Life Sci. Alliance 4, e202000924 (2020).
Henriques, F. et al. Single-cell RNA profiling reveals adipocyte to macrophage signaling sufficient to enhance thermogenesis. Cell Rep. 32, 107998 (2020).
Oguri, Y. et al. CD81 controls beige fat progenitor cell growth and energy balance via FAK signaling. Cell 182, 563–577.e20 (2020).
Ramirez, A. K. et al. Single-cell transcriptional networks in differentiating preadipocytes suggest drivers associated with tissue heterogeneity. Nat. Commun. 11, 2117 (2020).
Cao, J. et al. Comprehensive single-cell transcriptional profiling of a multicellular organism. Science 357, 661–667 (2017).
Hepler, C. et al. Identification of functionally distinct fibro-inflammatory and adipogenic stromal subpopulations in visceral adipose tissue of adult mice. eLife 7, e39636 (2018).
Vijay, J. et al. Single-cell analysis of human adipose tissue identifies depot and disease specific cell types. Nat. Metab. 2, 97–109 (2020).
Jaitin, D. A. et al. Massively parallel single-cell RNA-seq for marker-free decomposition of tissues into cell types. Science 343, 776–779 (2014).
Spallanzani, R. G. et al. Distinct immunocyte-promoting and adipocyte-generating stromal components coordinate adipose tissue immune and metabolic tenors. Sci. Immunol. 4, eaaw3658 (2019).
Burl, R. B. et al. Deconstructing adipogenesis induced by β3-adrenergic receptor activation with single-cell expression profiling. Cell Metab. 28, 300–309.e4 (2018).
Lacar, B. et al. Nuclear RNA-seq of single neurons reveals molecular signatures of activation. Nat. Commun. 7, 11022 (2016).
Habib, N. et al. Massively-parallel single nucleus RNA-seq with DroNc-seq. Nat. Methods 14, 955–958 (2017).
Habib, N. et al. Div-Seq: single-nucleus RNA-seq reveals dynamics of rare adult newborn neurons. Science 353, 925–928 (2016).
Krishnaswami, S. R. et al. Using single nuclei for RNA-seq to capture the transcriptome of postmortem neurons. Nat. Protoc. 11, 499–524 (2016).
Dos Santos, M. et al. Single-nucleus RNA-seq and FISH identify coordinated transcriptional activity in mammalian myofibers. Nat. Commun. 11, 5102 (2020).
Petrany, M. J. et al. Single-nucleus RNA-seq identifies transcriptional heterogeneity in multinucleated skeletal myofibers. Nat. Commun. 11, 6374 (2020).
Sárvári, A. K. et al. Plasticity of epididymal adipose tissue in response to diet-induced obesity at single-nucleus resolution. Cell Metab. 33, 437–453.e5 (2020).
Slyper, M. et al. A single-cell and single-nucleus RNA-seq toolbox for fresh and frozen human tumors. Nat. Med. 26, 792–802 (2020). Slyper et al. describe multiple practical workflows for performing single-cell and single-nucleus transcriptome analyses.
Bakken, T. E. et al. Single-nucleus and single-cell transcriptomes compared in matched cortical cell types. PLoS ONE 13, e0209648 (2018).
Grindberg, R. V. et al. RNA-sequencing from single nuclei. Proc. Natl Acad. Sci. USA 110, 19802–19807 (2013).
Lake, B. B. et al. A comparative strategy for single-nucleus and single-cell transcriptomes confirms accuracy in predicted cell-type expression from nuclear RNA. Sci. Rep. 7, 6031 (2017).
Bagchi, M. et al. Vascular endothelial growth factor is important for brown adipose tissue development and maintenance. FASEB J. 27, 3257–3271 (2013).
Drokhlyansky, E. et al. The human and mouse enteric nervous system at single-cell resolution. Cell 182, 1606–1622.e23 (2020).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181 (2014).
Hagemann-Jensen, M. et al. Single-cell RNA counting at allele and isoform resolution using Smart-seq3. Nat. Biotechnol. 38, 708–714 (2020).
Hashimshony, T., Wagner, F., Sher, N. & Yanai, I. CEL-Seq: single-cell RNA-seq by multiplexed linear amplification. Cell Rep. 2, 666–673 (2012).
Hashimshony, T. et al. CEL-Seq2: sensitive highly-multiplexed single-cell RNA-Seq. Genome Biol. 17, 77 (2016).
Keren-Shaul, H. et al. MARS-seq2.0: an experimental and analytical pipeline for indexed sorting combined with single-cell RNA sequencing. Nat. Protoc. 14, 1841–1862 (2019).
Carninci, P. et al. High-efficiency full-length cDNA cloning by biotinylated CAP trapper. Genomics 37, 327–336 (1996).
Kouno, T. et al. C1 CAGE detects transcription start sites and enhancer activity at single-cell resolution. Nat. Commun. 10, 360 (2019).
Murata, M. et al. Detecting expressed genes using CAGE. Methods Mol. Biol. 1164, 67–85 (2014).
Batut, P. & Gingeras, T. R. RAMPAGE: promoter activity profiling by paired-end sequencing of 5′-complete cDNAs. Curr. Protoc. Mol. Biol. 104, 25B.11 (2013).
Policastro, R. A., Raborn, R. T., Brendel, V. P. & Zentner, G. E. Simple and efficient profiling of transcription initiation and transcript levels with STRIPE-seq. Genome Res. 30, 910–923 (2020).
Yamashita, R. et al. Genome-wide characterization of transcriptional start sites in humans by integrative transcriptome analysis. Genome Res. 21, 775–789 (2011).
Core, L. J. et al. Analysis of nascent RNA identifies a unified architecture of initiation regions at mammalian promoters and enhancers. Nat. Genet. 46, 1311–1320 (2014).
Svensson, V. et al. Power analysis of single-cell RNA-sequencing experiments. Nat. Methods 14, 381–387 (2017).
Natarajan, K. N. et al. Comparative analysis of sequencing technologies for single-cell transcriptomics. Genome Biol. 20, 70 (2019).
Zheng, G. X. Y. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Wang, X., He, Y., Zhang, Q., Ren, X. & Zhang, Z. Direct comparative analyses of 10X Genomics Chromium and smart-seq2. Genomics Proteomics Bioinformatics https://doi.org/10.1016/j.gpb.2020.02.005 (2021).
Xi, N. M. & Li, J. J.Benchmarking computational doublet-detection methods for single-cell RNA sequencing data. Cell Syst. 12, 176–194.e6 (2020).
McGinnis, C. S., Murrow, L. M. & Gartner, Z. J. DoubletFinder: doublet detection in single-cell RNA sequencing data using artificial nearest neighbors. Cell Syst. 8, 329–337.e4 (2019).
Wolock, S. L., Lopez, R. & Klein, A. M. Scrublet: computational identification of cell doublets in single-cell transcriptomic data. Cell Syst. 8, 281–291.e9 (2019).
DePasquale, E. A. K. et al. DoubletDecon: deconvoluting doublets from single-cell RNA-sequencing data. Cell Rep. 29, 1718–1727.e8 (2019).
Bernstein, N. J. et al. Solo: doublet identification in single-cell RNA-seq via semi-supervised deep learning. Cell Syst. 11, 95–101.e5 (2020).
Goldstein, L. D. et al. Massively parallel nanowell-based single-cell gene expression profiling. BMC Genomics 18, 519 (2017).
Tran, H. T. N. et al. A benchmark of batch-effect correction methods for single-cell RNA sequencing data. Genome Biol. 21, 12 (2020).
Welch, J. D. et al. Single-cell multi-omic integration compares and contrasts features of brain cell identity. Cell 177, 1873–1887.e17 (2019).
Hie, B., Bryson, B. & Berger, B. Efficient integration of heterogeneous single-cell transcriptomes using Scanorama. Nat. Biotechnol. 37, 685–691 (2019).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902.e21 (2019).
Chazarra-Gil, R., van Dongen, S., Kiselev, V. Y. & Hemberg, M. Flexible comparison of batch correction methods for single-cell RNA-seq using BatchBench. Nucleic Acids Res. 49, e42 (2021).
Ding, J. & Regev, A. Deep generative model embedding of single-cell RNA-Seq profiles on hyperspheres and hyperbolic spaces. Nat. Commun. 12, 2554 (2021).
Jolliffe, I. T. Principal Component Analysis (Springer, 2002).
Hinton, G. E. & Roweis, S. Stochastic neighbor embedding. Adv. Neural Inf. Process. Syst. 15, 857–864 (2002).
McInnes, L., Healy, J. & Melville, J. UMAP: uniform manifold approximation and projection for dimension reduction. Preprint at https://arxiv.org/abs/1802.03426 (2020).
Aran, D. et al. Reference-based analysis of lung single-cell sequencing reveals a transitional profibrotic macrophage. Nat. Immunol. 20, 163–172 (2019).
Fleming, S. J., Marioni, J. C. & Babadi, M. CellBender remove-background: a deep generative model for unsupervised removal of background noise from scRNA-seq datasets. Preprint at bioRxiv https://doi.org/10.1101/791699 (2019). Fleming et al. develop a machine-learning algorithm to remove ambient RNA and exclude empty droplets in scRNA-seq datasets.
Spaethling, J. M. et al. Single-cell transcriptomics and functional target validation of brown adipocytes show their complex roles in metabolic homeostasis. FASEB J. 30, 81–92 (2016).
Hagberg, C. E. et al. Flow cytometry of mouse and human adipocytes for the analysis of browning and cellular heterogeneity. Cell Rep. 24, 2746–2756.e5 (2018).
Tran, K.-V. et al. The vascular endothelium of the adipose tissue gives rise to both white and brown fat cells. Cell Metab. 15, 222–229 (2012).
Cattaneo, P. et al. Parallel lineage-tracing studies establish fibroblasts as the prevailing in vivo adipocyte progenitor. Cell Rep. 30, 571–582.e2 (2020).
Roh, H. C. et al. Warming induces significant reprogramming of beige, but not brown, adipocyte cellular identity. Cell Metab. 27, 1121–1137.e5 (2018).
Sun, W. et al. Cold-induced epigenetic programming of the sperm enhances brown adipose tissue activity in the offspring. Nat. Med. 24, 1372–1383 (2018).
Park, J. et al. Progenitor-like characteristics in a subgroup of UCP1+ cells within white adipose tissue. Dev. Cell 56, 985–999.e4 (2021). Park et al. identify that mature brite/beige adipocytes are proliferative and contribute to WAT browning.
Betz, M. J. & Enerbäck, S. Targeting thermogenesis in brown fat and muscle to treat obesity and metabolic disease. Nat. Rev. Endocrinol. 14, 77–87 (2018).
Müller, S. et al. Proteomic analysis of human brown adipose tissue reveals utilization of coupled and uncoupled energy expenditure pathways. Sci. Rep. 6, 30030 (2016).
Shamsi, F. et al. Vascular smooth muscle-derived Trpv1+ progenitors are a source of cold-induced thermogenic adipocytes. Nat. Metab. 3, 485–495 (2021). Shamsi et al. describe a novel type of thermogenic adipocyte progenitor cell.
Angueira, A. R. et al. Defining the lineage of thermogenic perivascular adipose tissue. Nat. Metab. 3, 469–484 (2021). Angueira et al. identify a novel type of adipogenic SMC.
Tang, W. et al. White fat progenitor cells reside in the adipose vasculature. Science 322, 583–586 (2008).
Chabowska-Kita, A. & Kozak, L. P. The critical period for brown adipocyte development: genetic and environmental influences. Obesity 24, 283–290 (2016).
Xue, B., Coulter, A., Rim, J. S., Koza, R. A. & Kozak, L. P. Transcriptional synergy and the regulation of Ucp1 during brown adipocyte induction in white fat depots. Mol. Cell Biol. 25, 8311–8322 (2005).
Sanchez-Gurmaches, J. et al. PTEN loss in the Myf5 lineage redistributes body fat and reveals subsets of white adipocytes that arise from Myf5 precursors. Cell Metab. 16, 348–362 (2012).
Berry, R. & Rodeheffer, M. S. Characterization of the adipocyte cellular lineage in vivo. Nat. Cell Biol. 15, 302–308 (2013).
Lee, Y.-H., Petkova, A. P., Mottillo, E. P. & Granneman, J. G. In vivo identification of bipotential adipocyte progenitors recruited by β3-adrenoceptor activation and high-fat feeding. Cell Metab. 15, 480–491 (2012).
Seale, P. et al. PRDM16 controls a brown fat/skeletal muscle switch. Nature 454, 961–967 (2008).
Sanchez-Gurmaches, J. & Guertin, D. A. Adipocytes arise from multiple lineages that are heterogeneously and dynamically distributed. Nat. Commun. 5, 4099 (2014).
Sebo, Z. L. & Rodeheffer, M. S.Assembling the adipose organ: adipocyte lineage segregation and adipogenesis in vivo. Development 146, dev172098 (2019).
Xia, C., Fan, J., Emanuel, G., Hao, J. & Zhuang, X. Spatial transcriptome profiling by MERFISH reveals subcellular RNA compartmentalization and cell cycle-dependent gene expression. Proc. Natl Acad. Sci. USA 116, 19490–19499 (2019).
Rodriques, S. G. et al. Slide-seq: a scalable technology for measuring genome-wide expression at high spatial resolution. Science 363, 1463–1467 (2019).
Stickels, R. R. et al. Sensitive spatial genome wide expression profiling at cellular resolution. Preprint at bioRxiv https://doi.org/10.1101/2020.03.12.989806 (2020).
Ståhl, P. L. et al. Visualization and analysis of gene expression in tissue sections by spatial transcriptomics. Science 353, 78–82 (2016).
Vickovic, S. et al. High-definition spatial transcriptomics for in situ tissue profiling. Nat. Methods 16, 987–990 (2019).
Moncada, R. et al. Integrating microarray-based spatial transcriptomics and single-cell RNA-seq reveals tissue architecture in pancreatic ductal adenocarcinomas. Nat. Biotechnol. 38, 333–342 (2020).
Zhou, Y. et al. Human and mouse single-nucleus transcriptomics reveal TREM2-dependent and TREM2-independent cellular responses in Alzheimer’s disease. Nat. Med. 26, 131–142 (2020).
Mathys, H. et al. Single-cell transcriptomic analysis of Alzheimer’s disease. Nature 570, 332–337 (2019).
Ferrante, A. W. Macrophages, fat, and the emergence of immunometabolism. J. Clin. Invest. 123, 4992–4993 (2013).
Fischer, K. et al. Alternatively activated macrophages do not synthesize catecholamines or contribute to adipose tissue adaptive thermogenesis. Nat. Med. 23, 623–630 (2017).
Pirzgalska, R. M. et al. Sympathetic neuron-associated macrophages contribute to obesity by importing and metabolizing norepinephrine. Nat. Med. 23, 1309–1318 (2017).
Camell, C. D. et al. Inflammasome-driven catecholamine catabolism in macrophages blunts lipolysis during ageing. Nature 550, 119–123 (2017).
Hill, D. A. et al. Distinct macrophage populations direct inflammatory versus physiological changes in adipose tissue. Proc. Natl Acad. Sci. USA 115, E5096–E5105 (2018).
Winer, S. et al. Normalization of obesity-associated insulin resistance through immunotherapy. Nat. Med. 15, 921–929 (2009).
Feuerer, M. et al. Lean, but not obese, fat is enriched for a unique population of regulatory T cells that affect metabolic parameters. Nat. Med. 15, 930–939 (2009).
Bapat, S. P. et al. Depletion of fat-resident Treg cells prevents age-associated insulin resistance. Nature 528, 137–141 (2015).
Vasanthakumar, A. et al. Sex-specific adipose tissue imprinting of regulatory T cells. Nature 579, 581–585 (2020).
LaMarche, N. M. et al. Distinct iNKT cell populations use IFNγ or ER stress-induced IL-10 to control adipose tissue homeostasis. Cell Metab. 32, 243–258.e6 (2020).
Cryer, A., Riley, S. E., Williams, E. R. & Robinson, D. S. Effect of nutritional status on rat adipose tissue, muscle and post-heparin plasma clearing factor lipase activities: their relationship to triglyceride fatty acid uptake by fat-cells and to plasma insulin concentrations. Clin. Sci. Mol. Med. 50, 213–221 (1976).
Davies, B. S. J. et al. GPIHBP1 is responsible for the entry of lipoprotein lipase into capillaries. Cell Metab. 12, 42–52 (2010).
Fischer, A. W. et al. Lysosomal lipoprotein processing in endothelial cells stimulates adipose tissue thermogenic adaptation. Cell Metab. 33, 547–564.e7 (2021).
Hagberg, C. E. et al. Vascular endothelial growth factor B controls endothelial fatty acid uptake. Nature 464, 917–921 (2010).
Crewe, C. et al. An endothelial-to-adipocyte extracellular vesicle axis governed by metabolic state. Cell 175, 695–708.e13 (2018).
Lamalice, L., Le. Boeuf, F. & Huot, J. Endothelial cell migration during angiogenesis. Circ. Res. 100, 782–794 (2007).
Xue, Y. et al. Hypoxia-independent angiogenesis in adipose tissues during cold acclimation. Cell Metab. 9, 99–109 (2009).
Cao, Y. Angiogenesis and vascular functions in modulation of obesity, adipose metabolism, and insulin sensitivity. Cell Metab. 18, 478–489 (2013).
Gupta, R. K. et al. Zfp423 expression identifies committed preadipocytes and localizes to adipose endothelial and perivascular cells. Cell Metab. 15, 230–239 (2012).
Seki, T. et al. Endothelial PDGF-CC regulates angiogenesis-dependent thermogenesis in beige fat. Nat. Commun. 7, 12152 (2016).
Bae, C.-R. et al. Overexpression of C-type natriuretic peptide in endothelial cells protects against insulin resistance and inflammation during diet-induced obesity. Sci. Rep. 7, 9807 (2017).
Acknowledgements
We thank members of the laboratory of C.W. and Ian Mitchell for helpful discussions. Research in the laboratory of C.W. was supported by the Swiss National Science Foundation (grant 185011).
Author information
Authors and Affiliations
Contributions
W.S. and C.W. wrote the main text. S.M. contributed to illustrating the figures. H.D. contributed Fig. 3.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Metabolism thanks James G. Granneman and Patrick Seale for their contribution to the peer review of this work. Primary Handling Editors: Christoph Schmitt; Isabella Samuelson.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Sun, W., Modica, S., Dong, H. et al. Plasticity and heterogeneity of thermogenic adipose tissue. Nat Metab 3, 751–761 (2021). https://doi.org/10.1038/s42255-021-00417-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-021-00417-4
This article is cited by
-
The novel importance of miR-143 in obesity regulation
International Journal of Obesity (2023)
-
A ‘replace me’ signal from dying brown fat fires up weight loss
Nature (2022)
-
Adipocyte IRE1α promotes PGC1α mRNA decay and restrains adaptive thermogenesis
Nature Metabolism (2022)