Abstract
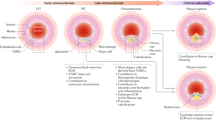
Atherosclerosis is a progressive vascular disease triggered by interplay between abnormal shear stress and endothelial lipid retention. A combination of these and, potentially, other factors leads to a chronic inflammatory response in the vessel wall, which is thought to be responsible for disease progression characterized by a buildup of atherosclerotic plaques. Yet molecular events responsible for maintenance of plaque inflammation and plaque growth have not been fully defined. Here we show that endothelial transforming growh factor β (TGF-β) signalling is one of the primary drivers of atherosclerosis-associated vascular inflammation. Inhibition of endothelial TGF-β signalling in hyperlipidemic mice reduces vessel wall inflammation and vascular permeability and leads to arrest of disease progression and regression of established lesions. These proinflammatory effects of endothelial TGF-β signalling are in stark contrast with its effects in other cell types and identify it as an important driver of atherosclerotic plaque growth and show the potential of cell-type-specific therapeutic intervention aimed at control of this disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
High-throughput sequencing data associated with this study have been deposited. The GEO accession number is GSE134557.
Code availability
The code and the version of the dependencies are available at https://github.com/wxncailab/NatureMetabolism_Endothelial.
References
Baeyens, N. & Schwartz, M. A. Biomechanics of vascular mechanosensation and remodeling. Mol. Biol. Cell 27, 7–11 (2016).
Tabas, I., Garcia-Cardena, G. & Owens, G. K. Recent insights into the cellular biology of atherosclerosis. J. Cell Biol. 209, 13–22 (2015).
Schwartz, M. A., Vestweber, D. & Simons, M. A unifying concept in vascular health and disease. Science 360, 270–271 (2018).
Chen, P.-Y. et al. Endothelial-to-mesenchymal transition drives atherosclerosis progression. J. Clin. Investig. 125, 4529–4543 (2015).
Evrard, S. M. et al. Endothelial to mesenchymal transition is common in atherosclerotic lesions and is associated with plaque instability. Nat. Commun. 7, 11853 (2016).
Feaver, R. E., Gelfand, B. D., Wang, C., Schwartz, M. A. & Blackman, B. R. Atheroprone hemodynamics regulate fibronectin deposition to create positive feedback that sustains endothelial inflammation. Circulation Res. 106, 1703–1711 (2010).
Rohwedder, I. et al. Plasma fibronectin deficiency impedes atherosclerosis progression and fibrous cap formation. EMBO Mol. Med. 4, 564–576 (2012).
Iwata, J. et al. Modulation of noncanonical TGF-beta signaling prevents cleft palate in Tgfbr2 mutant mice. J. Clin. Investig. 122, 873–885 (2012).
Yang, P. et al. Smooth muscle cell-specific Tgfbr1 deficiency promotes aortic aneurysm formation by stimulating multiple signaling events. Sci. Rep. 6, 35444 (2016).
Lutgens, E. et al. Deficient CD40-TRAF6 signaling in leukocytes prevents atherosclerosis by skewing the immune response toward an antiinflammatory profile. J. Exp. Med. 207, 391–404 (2010).
Virmani, R., Kolodgie, F. D., Burke, A. P., Farb, A. & Schwartz, S. M. Lessons from sudden coronary death: a comprehensive morphological classification scheme for atherosclerotic lesions. Arterioscler Thromb. Vasc. Biol. 20, 1262–1275 (2000).
Yu, P. et al. FGF-dependent metabolic control of vascular development. Nature 545, 224–228 (2017).
Dahlman, J. E. et al. In vivo endothelial siRNA delivery using polymeric nanoparticles with low molecular weight. Nat. Nanotechnol. 9, 648–655 (2014).
Sager, H. B. et al. RNAi targeting multiple cell adhesion molecules reduces immune cell recruitment and vascular inflammation after myocardial infarction. Sci. Transl. Med. 8, 342ra380 (2016).
Pardali, E. & Ten Dijke, P. TGFbeta signaling and cardiovascular diseases. Int. J. Biol. Sci. 8, 195–213 (2012).
Akhurst, R. J. & Hata, A. Targeting the TGFbeta signalling pathway in disease. Nat. Rev. Drug Discov. 11, 790–811 (2012).
McCaffrey, T. A. TGF-beta signaling in atherosclerosis and restenosis. Front. Biosci. 1, 236–245 (2009).
Schunkert, H. et al. Large-scale association analysis identifies 13 new susceptibility loci for coronary artery disease. Nat. Genet. 43, 333–338 (2011).
Toma, I. & McCaffrey, T. A. Transforming growth factor-beta and atherosclerosis: interwoven atherogenic and atheroprotective aspects. Cell Tissue Res. 347, 155–175 (2012).
Mallat, Z. et al. Inhibition of transforming growth factor-beta signaling accelerates atherosclerosis and induces an unstable plaque phenotype in mice. Circulation Res. 89, 930–934 (2001).
Lutgens, E. et al. Transforming growth factor-beta mediates balance between inflammation and fibrosis during plaque progression. Arterioscler Thromb. Vasc. Biol. 22, 975–982 (2002).
Lievens, D. et al. Abrogated transforming growth factor beta receptor II (TGFbetaRII) signalling in dendritic cells promotes immune reactivity of T cells resulting in enhanced atherosclerosis. Eur. heart J. 34, 3717–3727 (2013).
Gistera, A. et al. Transforming growth factor-beta signaling in T cells promotes stabilization of atherosclerotic plaques through an interleukin-17-dependent pathway. Sci. Transl. Med. 5, 196ra100 (2013).
Robertson, A. K. et al. Disruption of TGF-beta signaling in T cells accelerates atherosclerosis. J. Clin. Investig. 112, 1342–1350 (2003).
Chen, P. Y., Qin, L., Li, G., Tellides, G. & Simons, M. Smooth muscle FGF/TGFbeta cross talk regulates atherosclerosis progression. EMBO Mol. Med. 8, 712–728 https://doi.org/10.15252/emmm.201506181 (2016).
van Meeteren, L. A. & ten Dijke, P. Regulation of endothelial cell plasticity by TGF-beta. Cell tissue Res. 347, 177–186 (2012).
Deleavey, G. F. & Damha, M. J. Designing chemically modified oligonucleotides for targeted gene silencing. Chem. Biol. 19, 937–954 (2012).
Wang, Z. et al. A Non-canonical BCOR-PRC1.1 Complex Represses Differentiation Programs in Human ESCs. Cell Stem Cell 22, 235–251 e239 (2018).
Ramirez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic acids Res. 44, W160–W165 (2016).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform. 12, 323 (2011).
Robinson, M. D. & Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 11, R25 (2010).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Tomczak, J. M. & Welling, M. VAE with a VampPrior. Preprint at https://arxiv.org/abs/1705.07120 (2017).
Doersch, C. Tutorial on variational autoencoders. Preprint at https://arxiv.org/abs/1606.05908v2 (2016).
Zhao, S., Song, J. & Ermon, S. InfoVAE: Information maximizing variational autoencoders. Preprint at https://arxiv.org/abs/1706.02262 (2017).
Kharchenko, P. V., Silberstein, L. & Scadden, D. T. Bayesian approach to single-cell differential expression analysis. Nat. Methods 11, 740–742 (2014).
Risso, D., Perraudeau, F., Gribkova, S., Dudoit, S. & Vert, J. P. A general and flexible method for signal extraction from single-cell RNA-seq data. Nat. Commun. 9, 284 (2018).
Robinson, M. D. & Smyth, G. K. Small-sample estimation of negative binomial dispersion, with applications to SAGE data. Biostatistics 9, 321–332 (2008).
Srivastava, N., Hinton, G., Krizhevsky, A., Sutskever, I. & Salakhutdinov, R. Dropout: A simple way to prevent neural networks from overfitting. J. Mach. Learn. Res. 15, 1929–1958 (2014).
Levine, J. H. et al. Data-driven phenotypic dissection of AML Reveals progenitor-like cells that correlate with prognosis. Cell 162, 184–197 (2015).
Shekhar, K. et al. Comprehensive classification of retinal bipolar neurons by single-cell transcriptomics. Cell 166, 1308–1323 e1330 (2016).
Strehl, A. & Ghosh, J. Cluster ensembles—a knowledge reuse framework for combining multiple partitions. J. Mach. Learn. Res. 3, 583–617 (2002).
Shi, J. & Malik, J. Normalized cuts and image segmentation. IEEE Trans. Pattern Anal. Mach. Intell. 22, 888–905 (2000).
Young, M. D., Wakefield, M. J., Smyth, G. K. & Oshlack, A. Gene ontology analysis for RNA-seq: accounting for selection bias. Genome Biol. 11, R14 (2010).
Chittenden, T. W. et al. nEASE: a method for gene ontology subclassification of high-throughput gene expression data. Bioinformatics 28, 726–728 (2012).
Acknowledgements
We are grateful to J. Fang for aortic endothelial cell isolation used in scRNA-seq. We thank K. Hirschi for helpful discussions, R. Webber and L. Coon for maintaining mice colonies used in this study. This work was supported, in part, by National Nature Science Foundation of China grant no. 81600365 (G.L.) and NIH grant no. R01 HL135582 (M.S. and M.A.S.).
Author information
Authors and Affiliations
Contributions
P.Y.C., L.Q., G.L., Z.W., X.Z., B.A., A.C.D., R.L., P.K. and Y.J. performed experiments and generated data. L.S. and H.S. performed sequencing. J.M.L., S.G., N.A.C., J.G., T.W.C. carried out bioinformatics studies. J.E.D., K.J.K., A.S. and D.G.A. designed and synthesized nanoparticles and RNA chemical modifications for delivery studies. C.F.H., E.L., M.A.S. and J.S.P. assisted with studies of inflammation. P.Y.C. and M.S. wrote the manuscript. M.A.S., G.T., T.W.C. and M.S. supervised the project and provided funding.
Corresponding author
Ethics declarations
Competing interests
M.S. and P.Y.C. are holders of a provisional US Patent Application 62/311,086 and 62/406,732 dealing with endothelial-specific treatment of atherosclerosis. M.S., P.Y.C. and D.G.A. are scientific founders of VasoRx, Inc. M.S. and D.G.A. are members of VasoRx, Inc. Scientific Advisory Board. The other authors declare no competing interests.
Additional information
Peer review information: Primary Handling Editor: Pooja Jha.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–11 and Tables 1–3
Supplementary Dataset 1
Genes used in the analysis of the single-cell RNA-seq data. The values at every mouse genotype correspond to the total number of cells on which the gene was expressed.
Supplementary Dataset 2
Nested-GO terms functional enrichment. Accession number, nested-GO-term and their corresponding functional annotated genes for which the nested GOseq functional enrichment analysis was conducted (see Methods).
Supplementary Dataset 3
Cells used in the analysis of the single-cell RNA-seq data. For each cell barcode with its corresponding genotype we have the following information: VAE_ClusterLabel, cluster label as obtained from the single-cell RNA-seq analysis; LibraryCount, total cell’s library UMI count; FeatureCount, total number of genes expressed at the cell.
Rights and permissions
About this article
Cite this article
Chen, PY., Qin, L., Li, G. et al. Endothelial TGF-β signalling drives vascular inflammation and atherosclerosis. Nat Metab 1, 912–926 (2019). https://doi.org/10.1038/s42255-019-0102-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-019-0102-3
This article is cited by
-
Dysregulated cellular metabolism in atherosclerosis: mediators and therapeutic opportunities
Nature Metabolism (2024)
-
Influence of arsenic exposure and TGF-β gene single nucleotide polymorphisms (gene-environment interaction) on cardiovascular risk biomarkers levels in Mexican people from San Luis Potosi, Mexico
Toxicology and Environmental Health Sciences (2024)
-
CAFs vs. TECs: when blood feuds fuel cancer progression, dissemination and therapeutic resistance
Cellular Oncology (2024)
-
Vascular organoids: unveiling advantages, applications, challenges, and disease modelling strategies
Stem Cell Research & Therapy (2023)
-
The CLDN5 gene at the blood-brain barrier in health and disease
Fluids and Barriers of the CNS (2023)