Abstract
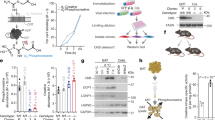
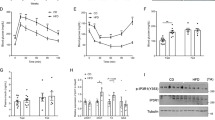
Depleting creatine levels in thermogenic adipocytes by inhibiting creatine biosynthesis reduces thermogenesis and causes obesity. However, whether creatine import from the circulation affects adipocyte thermogenesis is unknown. Here we show that deletion of the cell-surface creatine transporter (CrT) selectively in fat (AdCrTKO) substantially reduces adipocyte creatine and phosphocreatine levels, and reduces whole-body energy expenditure in mice. AdCrTKO mice are cold intolerant and become more obese than wild-type animals when fed a high-fat diet. Loss of adipocyte creatine transport blunts diet- and β3-adrenergic-induced thermogenesis, whereas creatine supplementation during high-fat feeding increases whole-body energy expenditure in response to β3-adrenergic agonism. In humans, CRT expression in purified subcutaneous adipocytes correlates with lower body mass index and increased insulin sensitivity. Our data indicate that adipocyte creatine abundance depends on creatine sequestration from the circulation. Given that it affects whole-body energy expenditure, enhancing creatine uptake into adipocytes may offer an opportunity to combat obesity and obesity-associated metabolic dysfunction.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All proteomic data generated or analysed during this study are included in this published article (and its supplementary information files). Additional data that support the findings of this study are available from the corresponding authors on reasonable request.
References
Lengyel, E., Makowski, L., DiGiovanni, J. & Kolonin, M. G. Cancer as a matter of fat: the crosstalk between adipose tissue and tumors. Trends Cancer 4, 374–384 (2018).
Twig, G. et al. Body-mass index in 2.3 million adolescents and cardiovascular death in adulthood. N. Engl. J. Med. 374, 2430–2440 (2016).
Ravussin, E. et al. Reduced rate of energy expenditure as a risk factor for body-weight gain. N. Engl. J. Med. 318, 467–472 (1988).
Jung, R. T., Shetty, P. S., James, W. P., Barrand, M. A. & Callingham, B. A. Reduced thermogenesis in obesity. Nature 279, 322–323 (1979).
Hofmann, W. E., Liu, X., Bearden, C. M., Harper, M. E. & Kozak, L. P. Effects of genetic background on thermoregulation and fatty acid-induced uncoupling of mitochondria in UCP1-deficient mice. J. Biol. Chem. 276, 12460–12465 (2001).
Liu, X. et al. Paradoxical resistance to diet-induced obesity in UCP1-deficient mice. J. Clin. Invest. 111, 399–407 (2003).
Mottillo, E. P. et al. Coupling of lipolysis and de novo lipogenesis in brown, beige, and white adipose tissues during chronic beta3-adrenergic receptor activation. J. Lipid Res. 55, 2276–2286 (2014).
Muller, S. et al. Proteomic analysis of human brown adipose tissue reveals utilization of coupled and uncoupled energy expenditure pathways. Sci. Rep. 6, 30030 (2016).
Rowland, L. A., Maurya, S. K., Bal, N. C., Kozak, L. & Periasamy, M. Sarcolipin and uncoupling protein 1 play distinct roles in diet-induced thermogenesis and do not compensate for one another. Obesity (Silver Spring) 24, 1430–1433 (2016).
Ukropec, J., Anunciado, R. P., Ravussin, Y., Hulver, M. W. & Kozak, L. P. UCP1-independent thermogenesis in white adipose tissue of cold-acclimated Ucp1-/- mice. J. Biol. Chem. 281, 31894–31908 (2006).
Ikeda, K. et al. UCP1-independent signaling involving SERCA2b-mediated calcium cycling regulates beige fat thermogenesis and systemic glucose homeostasis. Nat. Med. 23, 1454–1465 (2017).
Bertholet, A. M. et al. Mitochondrial patch clamp of beige adipocytes reveals UCP1-positive and UCP1-negative cells both exhibiting futile creatine cycling. Cell Metab. 25, 811–822 e814 (2017).
Kazak, L. et al. A creatine-driven substrate cycle enhances energy expenditure and thermogenesis in beige fat. Cell 163, 643–655 (2015).
Wakatsuki, T. et al. Thermogenic responses to high-energy phosphate contents and/or hindlimb suspension in rats. Jpn J. Physiol. 46, 171–175 (1996).
Yamashita, H. et al. Increased growth of brown adipose tissue but its reduced thermogenic activity in creatine-depleted rats fed beta-guanidinopropionic acid. Biochim. Biophys. Acta 1230, 69–73 (1995).
Kazak, L. et al. Genetic depletion of adipocyte creatine metabolism inhibits diet-induced thermogenesis and drives obesity. Cell Metab. 26, 660–671 e663 (2017).
Fitch, C. D., Shields, R. P., Payne, W. F. & Dacus, J. M. Creatine metabolism in skeletal muscle. 3. Specificity of the creatine entry process. J. Biol. Chem. 243, 2024–2027 (1968).
Berlet, H. H., Bonsmann, I. & Birringer, H. Occurrence of free creatine, phosphocreatine and creatine phosphokinase in adipose tissue. Biochim. Biophys. Acta 437, 166–174 (1976).
Skelton, M. R. et al. Creatine transporter (CrT; Slc6a8) knockout mice as a model of human CrT deficiency. PLoS ONE 6, e16187 (2011).
Lee, J., Choi, J., Aja, S., Scafidi, S. & Wolfgang, M. J. Loss of adipose fatty acid oxidation does not potentiate obesity at thermoneutrality. Cell Rep. 14, 1308–1316 (2016).
Kazak, L. et al. UCP1 deficiency causes brown fat respiratory chain depletion and sensitizes mitochondria to calcium overload-induced dysfunction. Proc. Natl Acad. Sci. USA 114, 7981–7986 (2017).
Eguchi, J. et al. Transcriptional control of adipose lipid handling by IRF4. Cell Metab. 13, 249–259 (2011).
Speakman, J. R., Krol, E. & Johnson, M. S. The functional significance of individual variation in basal metabolic rate. Physiol. Biochem. Zool. 77, 900–915 (2004).
Bloom, J. D. et al. Disodium (R,R)-5-[2-[[2-(3-chlorophenyl)-2-hydroxyethyl]-amino] propyl]-1,3-benzodioxole-2,2-dicarboxylate (CL 316,243). A potent beta-adrenergic agonist virtually specific for beta 3 receptors. A promising antidiabetic and antiobesity agent. J. Med. Chem. 35, 3081–3084 (1992).
Himms-Hagen, J., Hogan, S. & Zaror-Behrens, G. Increased brown adipose tissue thermogenesis in obese (ob/ob) mice fed a palatable diet. Am. J. Physiol. 250, E274–E281 (1986).
Bachman, E. S. et al. betaAR signaling required for diet-induced thermogenesis and obesity resistance. Science 297, 843–845 (2002).
Rothwell, N. J. & Stock, M. J. A role for brown adipose tissue in diet-induced thermogenesis. Nature 281, 31–35 (1979).
Leibel, R. L. & Hirsch, J. Diminished energy requirements in reduced-obese patients. Metabolism 33, 164–170 (1984).
Eringa, E. C. et al. Regulation of vascular function and insulin sensitivity by adipose tissue: focus on perivascular adipose tissue. Microcirculation 14, 389–402 (2007).
Singhal, A. et al. Influence of leptin on arterial distensibility: a novel link between obesity and cardiovascular disease? Circulation 106, 1919–1924 (2002).
Shimizu, I. et al. Vascular rarefaction mediates whitening of brown fat in obesity. J. Clin. Invest. 124, 2099–2112 (2014).
Ernande, L. et al. Relationship of brown adipose tissue perfusion and function: a study through beta2-adrenoreceptor stimulation. J. Appl. Physiol. (1985) 120, 825–832 (2016).
Hankir, M. K. & Klingenspor, M. Brown adipocyte glucose metabolism: a heated subject. EMBO Rep. 19, e46404 (2018).
Cereijo, R. et al. CXCL14, a brown adipokine that mediates brown-fat-to-macrophage communication in thermogenic adaptation. Cell Metab. 28, 750–763.e6 (2018).
Rosell, M. et al. Brown and white adipose tissues: intrinsic differences in gene expression and response to cold exposure in mice. Am. J. Physiol. Endocrinol. Metab. 306, E945–E964 (2014).
Svensson, P. A. et al. Gene expression in human brown adipose tissue. Int. J. Mol. Med. 27, 227–232 (2011).
Gerngross, C., Schretter, J., Klingenspor, M., Schwaiger, M. & Fromme, T. Active brown fat during (18)F-FDG PET/CT imaging defines a patient group with characteristic traits and an increased probability of brown fat redetection. J. Nucl. Med. 58, 1104–1110 (2017).
Din, M. U. et al. Postprandial oxidative metabolism of human brown fat indicates thermogenesis. Cell Metab. 28, 207–216.e3 (2018).
Fischer, A. W., Cannon, B. & Nedergaard, J. Optimal housing temperatures for mice to mimic the thermal environment of humans: an experimental study. Mol. Metab. 7, 161–170 (2018).
Speakman, J. R. & Keijer, J. Not so hot: Optimal housing temperatures for mice to mimic the thermal environment of humans. Mol. Metab. 2, 5–9 (2012).
Wada, S. et al. The tumor suppressor FLCN mediates an alternate mTOR pathway to regulate browning of adipose tissue. Genes Dev. 30, 2551–2564 (2016).
Perna, M. K. et al. Creatine transporter deficiency leads to increased whole body and cellular metabolism. Amino Acids 48, 2057–2065 (2016).
Streijger, F. et al. Mice lacking brain-type creatine kinase activity show defective thermoregulation. Physiol. Behav. 97, 76–86 (2009).
Fuhrer, T., Heer, D., Begemann, B. & Zamboni, N. High-throughput, accurate mass metabolome profiling of cellular extracts by flow injection-time-of-flight mass spectrometry. Anal. Chem. 83, 7074–7080 (2011).
Katz, A. et al. Quantitative insulin sensitivity check index: a simple, accurate method for assessing insulin sensitivity in humans. J. Clin. Endocrinol. Metab. 85, 2402–2410 (2000).
Matthews, D. R. et al. Homeostasis model assessment: insulin resistance and beta-cell function from fasting plasma glucose and insulin concentrations in man. Diabetologia 28, 412–419 (1985).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Elias, J. E. & Gygi, S. P. Target-decoy search strategy for increased confidence in large-scale protein identifications by mass spectrometry. Nat. Methods 4, 207–214 (2007).
Huttlin, E. L. et al. A tissue-specific atlas of mouse protein phosphorylation and expression. Cell 143, 1174–1189 (2010).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Smyth, G. K. Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat. Appl. Genet. Mol. Biol. 3, Article3 (2004).
Kammers, K., Cole, R. N., Tiengwe, C. & Ruczinski, I. Detecting significant changes in protein abundance. EuPA Open Proteom. 7, 11–19 (2015).
Eden, E., Navon, R., Steinfeld, I., Lipson, D. & Yakhini, Z. GOrilla: a tool for discovery and visualization of enriched GO terms in ranked gene lists. BMC Bioinformatics 10, 48 (2009).
Acknowledgements
This work was supported by the Canadian Institutes of Health Research (CIHR; grant PJT-159529), Goodman Cancer Research Centre and McGill University New Investigator Program, and DK114528-01 NIH/NIDDK K99 Pathway to Independence award (to L.K.). We acknowledge funding from a Canderel Fellowship (to J.F.R.). We acknowledge technical assistance from the McGill/GCRC Metabolomics core facility. The GCRC Metabolomics Core Facility is funded by the Dr. John R and Clara M. Fraser Memorial Trust, the Terry Fox Foundation, the Québec Breast Cancer Foundation and McGill University. We acknowledge funding from NIH R01HL 85744 and U24DK100469 Mayo Clinic Metabolomics Resource Core (to P.D.), AHA 13POST14540015 and NIH/NIDDK P30 DK057521 (to L.T.), NIH/NIDDK P30 DK057521, NIH/NIDDK R01 DK102173 and R01 ES017690 (to E.D.R.), and NIH DK31405 and JPB Foundation (to B.M.S.).
Author information
Authors and Affiliations
Contributions
L.K. conceptualized the study, designed research, performed biochemical, cellular and in vivo experiments, analysed data and wrote the paper. J.F.R., B.S., G.Z.L. and F.Y.D. performed in vivo experiments. M.P.J. performed proteomics experiments. M.L., L.C.R. and I.R.W. analysed proteomics data. S.Z. performed and analysed NMR experiments. E.T.C., P.D. and E.D.R. provided resources. L.T. recruited human subjects and isolated adipocytes, and D.T. performed RNA-seq experiments. L.K. and B.M.S. co-wrote the paper, with assistance from co-authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–3 and Supplementary Table 1
Supplementary Data 1
Proteomics inventory from AdCrTKO and CrTlox/y controls at 30 °C (CrTlox/y, n = 5; AdCrTKO, n = 5).
Supplementary Data 2
Proteomics inventory from AdCrTKO and CrTlox/y controls at 22 °C (CrTlox/y, n = 5; AdCrTKO, n = 5).
Supplementary Data 3
Cross-referenced proteomics inventory from AdCrTKO and CrTlox/y controls at 30 °C and 22 °C (CrTlox/y, n = 10; AdCrTKO, n = 10).
Rights and permissions
About this article
Cite this article
Kazak, L., Rahbani, J.F., Samborska, B. et al. Ablation of adipocyte creatine transport impairs thermogenesis and causes diet-induced obesity. Nat Metab 1, 360–370 (2019). https://doi.org/10.1038/s42255-019-0035-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-019-0035-x
This article is cited by
-
CCE and EODF as two distinct non-shivering thermogenesis models inducing weight loss
Pflügers Archiv - European Journal of Physiology (2023)
-
The muscle-enriched myokine Musclin impairs beige fat thermogenesis and systemic energy homeostasis via Tfr1/PKA signaling in male mice
Nature Communications (2023)
-
Impaired phosphocreatine metabolism in white adipocytes promotes inflammation
Nature Metabolism (2022)
-
The evolving view of thermogenic fat and its implications in cancer and metabolic diseases
Signal Transduction and Targeted Therapy (2022)
-
ATP-consuming futile cycles as energy dissipating mechanisms to counteract obesity
Reviews in Endocrine and Metabolic Disorders (2022)