Abstract
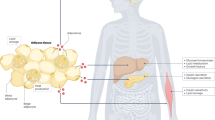
The gut microbiome has emerged as a key regulator of host metabolism. Here we review the various mechanisms through which the gut microbiome influences the energy metabolism of its host, highlighting the complex interactions between gut microbes, their metabolites and host cells. Among the most important bacterial metabolites are short-chain fatty acids, which serve as a direct energy source for host cells, stimulate the production of gut hormones and act in the brain to regulate food intake. Other microbial metabolites affect systemic energy expenditure by influencing thermogenesis and adipose tissue browning. Both direct and indirect mechanisms of action are known for specific metabolites, such as bile acids, branched chain amino acids, indole propionic acid and endocannabinoids. We also discuss the roles of specific bacteria in the production of specific metabolites and explore how external factors, such as antibiotics and exercise, affect the microbiome and thereby energy homeostasis. Collectively, we present a large body of evidence supporting the concept that gut microbiota-based therapies can be used to modulate host metabolism, and we expect to see such approaches moving from bench to bedside in the near future.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Cani, P. D. Human gut microbiome: hopes, threats and promises. Gut 67, 1716–1725 (2018).
Barr, J. J. A bacteriophages journey through the human body. Immunol. Rev. 279, 106–122 (2017).
Forde, A. & Hill, C. Phages of life—the path to pharma. Br. J. Pharmacol. 175, 412–418 (2018).
Sender, R., Fuchs, S. & Milo, R. Are we really vastly outnumbered? Revisiting the ratio of bacterial to host cells in humans. Cell 164, 337–340 (2016).
Li, J. et al. An integrated catalog of reference genes in the human gut microbiome. Nat. Biotechnol. 32, 834–841 (2014).
Qin, J. et al. A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464, 59–65 (2010).
Le Chatelier, E. et al. Richness of human gut microbiome correlates with metabolic markers. Nature 500, 541–546 (2013).
Cotillard, A. et al. Dietary intervention impact on gut microbial gene richness. Nature 500, 585–588 (2013).
Dao, M. C. et al. Akkermansia muciniphila and improved metabolic health during a dietary intervention in obesity: relationship with gut microbiome richness and ecology. Gut 65, 426–436 (2016).
Vandeputte, D. et al. Quantitative microbiome profiling links gut community variation to microbial load. Nature 551, 507–511 (2017).
Turnbaugh, P. J. et al. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 444, 1027–1031 (2006).
Bäckhed, F. et al. The gut microbiota as an environmental factor that regulates fat storage. Proc. Natl. Acad. Sci. USA 101, 15718–15723 (2004).
Wichmann, A. et al. Microbial modulation of energy availability in the colon regulates intestinal transit. Cell Host Microbe 14, 582–590 (2013).
Turnbaugh, P. J. et al. A core gut microbiome in obese and lean twins. Nature 457, 480–484 (2009).
Turnbaugh, P. J. et al. The effect of diet on the human gut microbiome: a metagenomic analysis in humanized gnotobiotic mice. Sci. Transl. Med. 1, 6ra14 (2009).
Dewulf, E. M. et al. Insight into the prebiotic concept: lessons from an exploratory, double blind intervention study with inulin-type fructans in obese women. Gut 62, 1112–1121 (2013).
Salazar, N. et al. Inulin-type fructans modulate intestinal Bifidobacterium species populations and decrease fecal short-chain fatty acids in obese women. Clin. Nutr. 34, 501–507 (2015).
Ridaura, V. K. et al. Gut microbiota from twins discordant for obesity modulate metabolism in mice. Science 341, 1241214 (2013).
Brahe, L. K., Astrup, A. & Larsen, L. H. Is butyrate the link between diet, intestinal microbiota and obesity-related metabolic diseases? Obes. Rev. 14, 950–959 (2013).
Neis, E. P. et al. Distal versus proximal intestinal short-chain fatty acid release in man. Gut https://doi.org/10.1136/gutjnl-2018-316161 (2018).
De Vadder, F. et al. Microbiota-generated metabolites promote metabolic benefits via gut–brain neural circuits. Cell 156, 84–96 (2014).
Rémésy, C., Demigné, C. & Chartier, F. Origin and utilization of volatile fatty acids in the rat. Reprod. Nutr. Dev. 20, 1339–1349 (1980).
Singh, V. et al. Microbiota-dependent hepatic lipogenesis mediated by stearoyl CoA desaturase 1 (SCD1) promotes metabolic syndrome in TLR5-deficient mice. Cell Metab. 22, 983–996 (2015).
Schwarzer, M. et al. Lactobacillus plantarum strain maintains growth of infant mice during chronic undernutrition. Science 351, 854–857 (2016).
Smythe, P. M. Changes in intestinal bacterial flora and role of infection in kwashiorkor. Lancet 2, 724–727 (1958).
Gupta, S. S. et al. Metagenome of the gut of a malnourished child. Gut Pathog. 3, 7 (2011).
Monira, S. et al. Gut microbiota of healthy and malnourished children in bangladesh. Front. Microbiol. 2, 228 (2011).
Ghosh, T. S. et al. Gut microbiomes of Indian children of varying nutritional status. PLoS One 9, e95547 (2014).
Subramanian, S. et al. Persistent gut microbiota immaturity in malnourished Bangladeshi children. Nature 510, 417–421 (2014).
De Filippo, C. et al. Impact of diet in shaping gut microbiota revealed by a comparative study in children from Europe and rural Africa. Proc. Natl. Acad. Sci. USA 107, 14691–14696 (2010).
Byndloss, M. X. et al. Microbiota-activated PPAR-γ signaling inhibits dysbiotic Enterobacteriaceae expansion. Science 357, 570–575 (2017).
Cani, P. D. Gut cell metabolism shapes the microbiome. Science 357, 548–549 (2017).
Waterson, M. J. & Horvath, T. L. Neuronal regulation of energy homeostasis: beyond the hypothalamus and feeding. Cell Metab. 22, 962–970 (2015).
Williams, K. W. & Elmquist, J. K. From neuroanatomy to behavior: central integration of peripheral signals regulating feeding behavior. Nat. Neurosci. 15, 1350–1355 (2012).
Suzuki, K., Jayasena, C. N. & Bloom, S. R. Obesity and appetite control. Exp. Diabetes Res. 2012, 824305 (2012).
Cani, P. D., Dewever, C. & Delzenne, N. M. Inulin-type fructans modulate gastrointestinal peptides involved in appetite regulation (glucagon-like peptide-1 and ghrelin) in rats. Br. J. Nutr. 92, 521–526 (2004).
Delzenne, N. M., Cani, P. D., Daubioul, C. & Neyrinck, A. M. Impact of inulin and oligofructose on gastrointestinal peptides. Br. J. Nutr. 93 (Suppl 1), S157–S161 (2005).
Cani, P. D., Neyrinck, A. M., Maton, N. & Delzenne, N. M. Oligofructose promotes satiety in rats fed a high-fat diet: involvement of glucagon-like Peptide-1. Obes. Res. 13, 1000–1007 (2005).
Cani, P. D., Joly, E., Horsmans, Y. & Delzenne, N. M. Oligofructose promotes satiety in healthy human: a pilot study. Eur. J. Clin. Nutr. 60, 567–572 (2006).
Cani, P. D., Hoste, S., Guiot, Y. & Delzenne, N. M. Dietary non-digestible carbohydrates promote L-cell differentiation in the proximal colon of rats. Br. J. Nutr. 98, 32–37 (2007).
Karaki, S. et al. Short-chain fatty acid receptor, GPR43, is expressed by enteroendocrine cells and mucosal mast cells in rat intestine. Cell Tissue Res. 324, 353–360 (2006).
Karaki, S. et al. Expression of the short-chain fatty acid receptor, GPR43, in the human colon. J. Mol. Histol. 39, 135–142 (2008).
Tazoe, H. et al. Expression of short-chain fatty acid receptor GPR41 in the human colon. Biomed. Res. 30, 149–156 (2009).
Tolhurst, G. et al. Short-chain fatty acids stimulate glucagon-like peptide-1 secretion via the G-protein-coupled receptor FFAR2. Diabetes 61, 364–371 (2012).
Psichas, A. et al. The short chain fatty acid propionate stimulates GLP-1 and PYY secretion via free fatty acid receptor 2 in rodents. Int. J. Obes. (Lond.) 39, 424–429 (2015).
Ronveaux, C. C., Tomé, D. & Raybould, H. E. Glucagon-like peptide 1 interacts with ghrelin and leptin to regulate glucose metabolism and food intake through vagal afferent neuron signaling. J. Nutr. 145, 672–680 (2015).
Hagemann, D. et al. Glucagon-like peptide 1 (GLP-1) suppresses ghrelin levels in humans via increased insulin secretion. Regul. Pept. 143, 64–68 (2007).
Murphy, K. G. & Bloom, S. R. Gut hormones and the regulation of energy homeostasis. Nature 444, 854–859 (2006).
Batterham, R. L. et al. Gut hormone PYY(3-36) physiologically inhibits food intake. Nature 418, 650–654 (2002).
Kanoski, S. E., Hayes, M. R. & Skibicka, K. P. GLP-1 and weight loss: unraveling the diverse neural circuitry. Am. J. Physiol. Regul. Integr. Comp. Physiol. 310, R885–R895 (2016).
Cani, P. D. & Knauf, C. How gut microbes talk to organs: the role of endocrine and nervous routes. Mol. Metab. 5, 743–752 (2016).
Rastelli, M., Knauf, C. & Cani, P. D. Gut microbes and health: a focus on the mechanisms linking microbes, obesity, and related disorders. Obes. (Silver Spring) 26, 792–800 (2018).
Lin, H. V. et al. Butyrate and propionate protect against diet-induced obesity and regulate gut hormones via free fatty acid receptor 3-independent mechanisms. PLoS One 7, e35240 (2012).
Chambers, E. S. et al. Effects of targeted delivery of propionate to the human colon on appetite regulation, body weight maintenance and adiposity in overweight adults. Gut 64, 1744–1754 (2015).
Li, Z. et al. Butyrate reduces appetite and activates brown adipose tissue via the gut-brain neural circuit. Gut 67, 1269–1279 (2018).
Goswami, C., Iwasaki, Y. & Yada, T. Short-chain fatty acids suppress food intake by activating vagal afferent neurons. J. Nutr. Biochem. 57, 130–135 (2018).
Frost, G. et al. The short-chain fatty acid acetate reduces appetite via a central homeostatic mechanism. Nat. Commun. 5, 3611 (2014).
Tennoune, N. et al. Bacterial ClpB heat-shock protein, an antigen-mimetic of the anorexigenic peptide α-MSH, at the origin of eating disorders. Transl. Psychiatry 4, e458 (2014).
Breton, J. et al. Gut commensal E. coli proteins activate host satiety pathways following nutrient-induced bacterial growth. Cell Metab. 23, 324–334 (2016).
Strandwitz, P. Neurotransmitter modulation by the gut microbiota. Brain Res. 1693 (Pt B), 128–133 (2018).
Yano, J. M. et al. Indigenous bacteria from the gut microbiota regulate host serotonin biosynthesis. Cell 161, 264–276 (2015).
Clarke, G. et al. The microbiome–gut–brain axis during early life regulates the hippocampal serotonergic system in a sex-dependent manner. Mol. Psychiatry 18, 666–673 (2013).
Kelly, J. R. et al. Transferring the blues: depression-associated gut microbiota induces neurobehavioural changes in the rat. J. Psychiatr. Res. 82, 109–118 (2016).
Zheng, P. et al. Gut microbiome remodeling induces depressive-like behaviors through a pathway mediated by the host’s metabolism. Mol. Psychiatry 21, 786–796 (2016).
De Palma, G. et al. Transplantation of fecal microbiota from patients with irritable bowel syndrome alters gut function and behavior in recipient mice. Sci. Transl. Med. 9, eaaf6397 (2017).
Sampson, T. R. et al. Gut microbiota regulate motor deficits and neuroinflammation in a model of parkinson’s disease. Cell 167, 1469–1480.e12 (2016).
Minter, M. R. et al. Antibiotic-induced perturbations in gut microbial diversity influences neuro-inflammation and amyloidosis in a murine model of Alzheimer’s disease. Sci. Rep. 6, 30028 (2016).
Berer, K. et al. Gut microbiota from multiple sclerosis patients enables spontaneous autoimmune encephalomyelitis in mice. Proc. Natl. Acad. Sci. USA 114, 10719–10724 (2017).
Hsiao, E. Y. et al. Microbiota modulate behavioral and physiological abnormalities associated with neurodevelopmental disorders. Cell 155, 1451–1463 (2013).
Castro, D. C., Cole, S. L. & Berridge, K. C. Lateral hypothalamus, nucleus accumbens, and ventral pallidum roles in eating and hunger: interactions between homeostatic and reward circuitry. Front. Syst. Neurosci. 9, 90 (2015).
Ding, L. & Zhang, J. Glucagon-like peptide-1 activates endothelial nitric oxide synthase in human umbilical vein endothelial cells. Acta Pharmacol. Sin. 33, 75–81 (2012).
Rotondo, A., Amato, A., Lentini, L., Baldassano, S. & Mulè, F. Glucagon-like peptide-1 relaxes gastric antrum through nitric oxide in mice. Peptides 32, 60–64 (2011).
Grasset, E. et al. A specific gut microbiota dysbiosis of type 2 diabetic mice induces GLP-1 resistance through an enteric no-dependent and gut–brain axis mechanism. Cell Metab. 25, 1075–1090.e75 (2017).
Catry, E. et al. Targeting the gut microbiota with inulin-type fructans: preclinical demonstration of a novel approach in the management of endothelial dysfunction. Gut 67, 271–283 (2018).
Brandl, K., Kumar, V. & Eckmann, L. Gut–liver axis at the frontier of host-microbial interactions. Am. J. Physiol. Gastrointest. Liver Physiol. 312, G413–G419 (2017).
Tilg, H., Cani, P. D. & Mayer, E. A. Gut microbiome and liver diseases. Gut 65, 2035–2044 (2016).
Schramm, C. Bile acids, the microbiome, immunity, and liver tumors. N. Engl. J. Med. 379, 888–890 (2018).
Chevre, R., Silvestre-Roig, C. & Soehnlein, O. Nutritional modulation of innate immunity: the fat–bile–gut connection. Trends Emdocrinol. Metab. 29, 686–698 (2018).
Pathak, P. et al. Intestine farnesoid X receptor agonist and the gut microbiota activate G-protein bile acid receptor-1 signaling to improve metabolism. Hepatology 68, 1574–1588 (2018).
Ma, C. et al. Gut microbiome–mediated bile acid metabolism regulates liver cancer via NKT cells. Science 360, eaan5931 (2018).
Fiorucci, S., Mencarelli, A., Palladino, G. & Cipriani, S. Bile-acid-activated receptors: targeting TGR5 and farnesoid-X-receptor in lipid and glucose disorders. Trends Pharmacol. Sci. 30, 570–580 (2009).
Watanabe, M. et al. Bile acids induce energy expenditure by promoting intracellular thyroid hormone activation. Nature 439, 484–489 (2006).
Anhê, F. F. et al. Treatment with camu camu (Myrciaria dubia) prevents obesity by altering the gut microbiota and increasing energy expenditure in diet-induced obese mice. Gut https://doi.org/10.1136/gutjnl-2017-315565 (2018).
Ziętak, M., Chabowska-Kita, A. & Kozak, L. P. Brown fat thermogenesis: Stability of developmental programming and transient effects of temperature and gut microbiota in adults. Biochimie 134, 93–98 (2017).
Chevalier, C. et al. Gut microbiota orchestrates energy homeostasis during cold. Cell 163, 1360–1374 (2015).
Ziętak, M. et al. Altered microbiota contributes to reduced diet-induced obesity upon cold exposure. Cell Metab. 23, 1216–1223 (2016).
Zhang, X. Y. et al. Huddling remodels gut microbiota to reduce energy requirements in a small mammal species during cold exposure. Microbiome 6, 103 (2018).
Derrien, M., Collado, M. C., Ben-Amor, K., Salminen, S. & de Vos, W. M. The Mucin degrader Akkermansia muciniphila is an abundant resident of the human intestinal tract. Appl. Environ. Microbiol. 74, 1646–1648 (2008).
Everard, A. et al. Cross-talk between Akkermansia muciniphila and intestinal epithelium controls diet-induced obesity. Proc. Natl. Acad. Sci. USA 110, 9066–9071 (2013).
Schneeberger, M. et al. Akkermansia muciniphila inversely correlates with the onset of inflammation, altered adipose tissue metabolism and metabolic disorders during obesity in mice. Sci. Rep. 5, 16643 (2015).
Cani, P. D. & de Vos, W. M. Next-generation beneficial microbes: the case of Akkermansia muciniphila. Front. Microbiol. 8, 1765 (2017).
Plovier, H. et al. A purified membrane protein from Akkermansia muciniphila or the pasteurized bacterium improves metabolism in obese and diabetic mice. Nat. Med. 23, 107–113 (2017).
Pierre, J. F. et al. Activation of bile acid signaling improves metabolic phenotypes in high-fat diet-induced obese mice. Am. J. Physiol. Gastrointest. Liver Physiol. 311, G286–G304 (2016).
Gao, X. et al. Polyphenol- and caffeine-rich postfermented pu-erh tea improves diet-induced metabolic syndrome by remodeling intestinal homeostasis in mice. Infect. Immun. 86, e00601–e00617 (2017).
Liu, J. et al. Gypenosides reduced the risk of overweight and insulin resistance in C57BL/6J mice through modulating adipose thermogenesis and gut microbiota. J. Agric. Food Chem. 65, 9237–9246 1 (2017).
Worthmann, A. et al. Cold-induced conversion of cholesterol to bile acids in mice shapes the gut microbiome and promotes adaptive thermogenesis. Nat. Med. 23, 839–849 (2017).
Kim, K. H. et al. Intermittent fasting promotes adipose thermogenesis and metabolic homeostasis via VEGF-mediated alternative activation of macrophage. Cell Res. 27, 1309–1326 (2017).
Mattson, M. P., Longo, V. D. & Harvie, M. Impact of intermittent fasting on health and disease processes. Ageing Res. Rev. 39, 46–58 (2017).
Anton, S. D. et al. Flipping the metabolic switch: understanding and applying the health benefits of fasting. Obes. (Silver Spring) 26, 254–268 (2018).
Li, G. et al. Intermittent fasting promotes white adipose browning and decreases obesity by shaping the gut microbiota. Cell Metab. 26, 672–685.e674 (2017).
Sonoyama, K. et al. Response of gut microbiota to fasting and hibernation in Syrian hamsters. Appl. Environ. Microbiol. 75, 6451–6456 (2009).
Remely, M. et al. Increased gut microbiota diversity and abundance of Faecalibacterium prausnitzii and Akkermansia after fasting: a pilot study. Wien. Klin. Wochenschr. 127, 394–398 (2015).
Costello, E. K., Gordon, J. I., Secor, S. M. & Knight, R. Postprandial remodeling of the gut microbiota in Burmese pythons. ISME J. 4, 1375–1385 (2010).
Belzer, C. & de Vos, W. M. Microbes inside—from diversity to function: the case of Akkermansia. ISME J. 6, 1449–1458 (2012).
Weitkunat, K. et al. Short-chain fatty acids and inulin, but not guar gum, prevent diet-induced obesity and insulin resistance through differential mechanisms in mice. Sci. Rep. 7, 6109 (2017).
Gao, Z. et al. Butyrate improves insulin sensitivity and increases energy expenditure in mice. Diabetes 58, 1509–1517 (2009).
Lu, Y. et al. Short chain fatty acids prevent high-fat-diet-induced obesity in mice by regulating G protein–coupled receptors and gut microbiota. Sci. Rep. 6, 37589 (2016).
Canfora, E. E., Jocken, J. W. & Blaak, E. E. Short-chain fatty acids in control of body weight and insulin sensitivity. Nat. Rev. Endocrinol. 11, 577–591 (2015).
Hanatani, S. et al. Acetate alters expression of genes involved in beige adipogenesis in 3T3-L1 cells and obese KK-Ay mice. J. Clin. Biochem. Nutr. 59, 207–214 (2016).
Sahuri-Arisoylu, M. et al. Reprogramming of hepatic fat accumulation and ‘browning’ of adipose tissue by the short-chain fatty acid acetate. Int. J. Obes. (Lond.) 40, 955–963 (2016).
Bouter, K. et al. Differential metabolic effects of oral butyrate treatment in lean versus metabolic syndrome subjects. Clin. Transl. Gastroenterol. 9, 155 (2018).
Canfora, E. E. et al. Colonic infusions of short-chain fatty acid mixtures promote energy metabolism in overweight/obese men: a randomized crossover trial. Sci. Rep. 7, 2360 (2017).
Chambers, E. S. et al. Acute oral sodium propionate supplementation raises resting energy expenditure and lipid oxidation in fasted humans. Diabetes Obes. Metab. 20, 1034–1039 (2018).
Rahat-Rozenbloom, S., Fernandes, J., Gloor, G. B. & Wolever, T. M. Evidence for greater production of colonic short-chain fatty acids in overweight than lean humans. Int. J. Obes. (Lond.) 38, 1525–1531 (2014).
Schwiertz, A. et al. Microbiota and SCFA in lean and overweight healthy subjects. Obes. (Silver Spring) 18, 190–195 (2010).
Pertwee, R. G. The pharmacology of cannabinoid receptors and their ligands: an overview. Int. J. Obes. 30(Suppl 1), S13–S18 (2006).
Geurts, L. et al. Adipose tissue NAPE-PLD controls fat mass development by altering the browning process and gut microbiota. Nat. Commun. 6, 6495 (2015).
Pacher, P., Bátkai, S. & Kunos, G. The endocannabinoid system as an emerging target of pharmacotherapy. Pharmacol. Rev. 58, 389–462 (2006).
Silvestri, C. & Di Marzo, V. The endocannabinoid system in energy homeostasis and the etiopathology of metabolic disorders. Cell Metab. 17, 475–490 (2013).
Mazier, W., Saucisse, N., Gatta-Cherifi, B. & Cota, D. The endocannabinoid system: pivotal orchestrator of obesity and metabolic disease. Trends Endocrinol. Metab. 26, 524–537 (2015).
Muccioli, G. et al. The endocannabinoid system links gut microbiota to adipogenesis. Mo. Syst. Biol. 6, 392 (2010).
Cluny, N. L., Keenan, C. M., Reimer, R. A., Le Foll, B. & Sharkey, K. A. Prevention of diet-induced obesity effects on body weight and gut microbiota in mice treated chronically with ∆9-tetrahydrocannabinol. PLoS One 10, e0144270 (2015).
Rousseaux, C. et al. Lactobacillus acidophilus modulates intestinal pain and induces opioid and cannabinoid receptors. Nat. Med. 13, 35–37 (2007).
Cani, P. D. et al. Endocannabinoids—at the crossroads between the gut microbiota and host metabolism. Nat. Rev. Endocrinol. 12, 133–143 (2016).
Mikkelsen, K. H., Allin, K. H. & Knop, F. K. Effect of antibiotics on gut microbiota, glucose metabolism and body weight regulation: a review of the literature. Diabetes Obes. Metab. 18, 444–453 (2016).
Cox, L. M. & Blaser, M. J. Antibiotics in early life and obesity. Nat. Rev. Endocrinol. 11, 182–190 (2015).
Rodrigues, R. R. et al. Antibiotic-induced alterations in gut microbiota are associated with changes in glucose metabolism in healthy mice. Front. Microbiol. 8, 2306 (2017).
Cani, P. D. et al. Changes in gut microbiota control metabolic endotoxemia-induced inflammation in high-fat diet–induced obesity and diabetes in mice. Diabetes 57, 1470–1481 (2008).
Cho, I. et al. Antibiotics in early life alter the murine colonic microbiome and adiposity. Nature 488, 621–626 (2012).
Cox, L. M. & Blaser, M. J. Pathways in microbe-induced obesity. Cell Metab. 17, 883–894 (2013).
Francino, M. P. Antibiotics and the human gut microbiome: dysbioses and accumulation of resistances. Front. Microbiol. 6, 1543 (2016).
Reijnders, D. et al. Effects of gut microbiota manipulation by antibiotics on host metabolism in obese humans: a randomized double-blind placebo-controlled trial. Cell Metab. 24, 63–74 (2016).
Nguyen, T. L., Vieira-Silva, S., Liston, A. & Raes, J. How informative is the mouse for human gut microbiota research? Dis. Model. Mech. 8, 1–16 (2015).
Gough, E. K. et al. The impact of antibiotics on growth in children in low and middle income countries: systematic review and meta-analysis of randomised controlled trials. Br. Med. J. 348, g2267 (2014).
Million, M., Diallo, A. & Raoult, D. Gut microbiota and malnutrition. Microb. Pathog. 106, 127–138 (2017).
Alcoba, G. et al. Do children with uncomplicated severe acute malnutrition need antibiotics? A systematic review and meta-analysis. PLoS One 8, e53184 (2013).
Jacobson, E. D., Chodos, R. B. & Faloon, W. W. An experimental malabsorption syndrome induced by neomycin. Am. J. Med. 28, 524–533 (1960).
Munukka, E. et al. Faecalibacterium prausnitzii treatment improves hepatic health and reduces adipose tissue inflammation in high-fat fed mice. ISME J. 11, 1667–1679 (2017).
Maurice, C. F., Haiser, H. J. & Turnbaugh, P. J. Xenobiotics shape the physiology and gene expression of the active human gut microbiome. Cell 152, 39–50 (2013).
Saad, R., Rizkallah, M. R. & Aziz, R. K. Gut pharmacomicrobiomics: the tip of an iceberg of complex interactions between drugs and gut-associated microbes. Gut Pathog. 4, 16 (2012).
Imhann, F. et al. The influence of proton pump inhibitors and other commonly used medication on the gut microbiota. Gut Microbes 8, 351–358 (2017).
Le Bastard, Q. et al. Systematic review: human gut dysbiosis induced by non-antibiotic prescription medications. Aliment. Pharmacol. Ther. 47, 332–345 (2018).
Falony, G. et al. Population-level analysis of gut microbiome variation. Science 352, 560–564 (2016).
Wang, Z. et al. Gut flora metabolism of phosphatidylcholine promotes cardiovascular disease. Nature 472, 57–63 (2011).
Lee, D. H., Porta, M., Jacobs, D. R. Jr & Vandenberg, L. N. Chlorinated persistent organic pollutants, obesity, and type 2 diabetes. Endocr. Rev. 35, 557–601 (2014).
Lu, K., Mahbub, R. & Fox, J. G. Xenobiotics: interaction with the intestinal microflora. ILAR J. 56, 218–227 (2015).
Caparrós-Martín, J. A. et al. Statin therapy causes gut dysbiosis in mice through a PXR-dependent mechanism. Microbiome 5, 95 (2017).
de la Cuesta-Zuluaga, J. et al. Metformin is associated with higher relative abundance of mucin-degrading Akkermansia muciniphila and several short-chain fatty acid-producing microbiota in the gut. Diabetes Care 40, 54–62 (2017).
Ma, W. et al. Metformin alters gut microbiota of healthy mice: implication for its potential role in gut microbiota homeostasis. Front. Microbiol. 9, 1336 (2018).
Ursell, L. K. & Knight, R. Xenobiotics and the human gut microbiome: metatranscriptomics reveal the active players. Cell Metab. 17, 317–318 (2013).
Zhao, X. et al. Response of gut microbiota to metabolite changes induced by endurance exercise. Front. Microbiol. 9, 765 (2018).
Monda, V. et al. exercise modifies the gut microbiota with positive health effects. Oxid. Med. Cell. Longev. 2017, 3831972 (2017).
Chen, J., Guo, Y., Gui, Y. & Xu, D. Physical exercise, gut, gut microbiota, and atherosclerotic cardiovascular diseases. Lipids Health Dis. 17, 17 (2018).
Cerdá, B. et al. Gut microbiota modification: another piece in the puzzle of the benefits of physical exercise in health? Front. Physiol. 7, 51 (2016).
Allen, J. M. et al. Exercise alters gut microbiota composition and function in lean and obese humans. Med. Sci. Sports Exerc. 50, 747–757 (2018).
Hsu, Y. J. et al. Effect of intestinal microbiota on exercise performance in mice. J. Strength Cond. Res. 29, 552–558 (2015).
Matsumoto, M. et al. Voluntary running exercise alters microbiota composition and increases n-butyrate concentration in the rat cecum. Biosci. Biotechnol. Biochem. 72, 572–576 (2008).
den Besten, G. et al. The role of short-chain fatty acids in the interplay between diet, gut microbiota, and host energy metabolism. J. Lipid Res. 54, 2325–2340 (2013).
Clark, A. & Mach, N. The crosstalk between the gut microbiota and mitochondria during exercise. Front. Physiol. 8, 319 (2017).
Newgard, C. B. et al. A branched-chain amino acid-related metabolic signature that differentiates obese and lean humans and contributes to insulin resistance. Cell Metab. 9, 311–326 (2009).
Pedersen, H. K. et al. Human gut microbes impact host serum metabolome and insulin sensitivity. Nature 535, 376–381 (2016).
Org, E. et al. Relationships between gut microbiota, plasma metabolites, and metabolic syndrome traits in the METSIM cohort. Genome Biol. 18, 70 (2017).
Liu, R. et al. Gut microbiome and serum metabolome alterations in obesity and after weight-loss intervention. Nat. Med. 23, 859–868 (2017).
Ben-Othman, N. et al. Long-term GABA administration induces alpha cell-mediated beta-like cell neogenesis. Cell 168, 73–85.e11 (2017).
de Vadder, F. & Mithieux, G. Gut–brain signaling in energy homeostasis: the unexpected role of microbiota-derived succinate. J. Endocrinol. 236, R105–R108 (2018).
De Vadder, F. et al. Microbiota-produced succinate improves glucose homeostasis via intestinal gluconeogenesis. Cell Metab. 24, 151–157 (2016).
Ley, R. E. Gut microbiota in 2015: Prevotella in the gut: choose carefully. Nat. Rev. Gastroenterol. Hepatol. 13, 69–70 (2016).
Scher, J. U. et al. Expansion of intestinal Prevotella copri correlates with enhanced susceptibility to arthritis. eLife 2, e01202 (2013).
Heianza, Y. et al. Gut microbiota metabolites, amino acid metabolites and improvements in insulin sensitivity and glucose metabolism: the POUNDS Lost trial. Gut https://doi.org/10.1136/gutjnl-2018-316155 (2018).
Zhu, W. et al. Gut microbial metabolite TMAO enhances platelet hyperreactivity and thrombosis risk. Cell 165, 111–124 (2016).
Gao, X. et al. Dietary trimethylamine N-oxide exacerbates impaired glucose tolerance in mice fed a high fat diet. J. Biosci. Bioeng. 118, 476–481 (2014).
Koeth, R. A. et al. γ-Butyrobetaine is a proatherogenic intermediate in gut microbial metabolism of L-carnitine to TMAO. Cell Metab. 20, 799–812 (2014).
Agus, A., Planchais, J. & Sokol, H. Gut microbiota regulation of tryptophan metabolism in health and disease. Cell Host Microbe 23, 716–724 (2018).
Zhang, L. S. & Davies, S. S. Microbial metabolism of dietary components to bioactive metabolites: opportunities for new therapeutic interventions. Genome Med. 8, 46 (2016).
Dodd, D. et al. A gut bacterial pathway metabolizes aromatic amino acids into nine circulating metabolites. Nature 551, 648–652 (2017).
Tuomainen, M. et al. Associations of serum indolepropionic acid, a gut microbiota metabolite, with type 2 diabetes and low-grade inflammation in high-risk individuals. Nutr. Diabetes 8, 35 (2018).
de Mello, V. D. et al. Indolepropionic acid and novel lipid metabolites are associated with a lower risk of type 2 diabetes in the Finnish Diabetes Prevention Study. Sci. Rep. 7, 46337 (2017).
Krishnan, S. et al. Gut microbiota–derived tryptophan metabolites modulate inflammatory response in hepatocytes and macrophages. Cell Rep. 23, 1099–1111 (2018).
Chimerel, C. et al. Bacterial metabolite indole modulates incretin secretion from intestinal enteroendocrine L cells. Cell Rep. 9, 1202–1208 (2014).
Beaumont, M. et al. The gut microbiota metabolite indole alleviates liver inflammation in mice. FASEB J. fj201800544 (2018).
Koh, A. et al. Microbially produced imidazole propionate impairs insulin signaling through mTORC1. Cell 175, 947–961.e17 (2018).
Klurfeld, D. M. et al. Considerations for best practices in studies of fiber or other dietary components and the intestinal microbiome. Am. J. Physiol. Endocrinol. Metab. https://doi.org/10.1152/ajpendo.00058.2018 (2018).
Routy, B. et al. The gut microbiota influences anticancer immunosurveillance and general health. Nat. Rev. Clin. Oncol. 15, 382–396 (2018).
Hempel, S. et al. Probiotics for the prevention and treatment of antibiotic-associated diarrhea: a systematic review and meta-analysis. J. Am. Med. Assoc. 307, 1959–1969 (2012).
Wei, D. et al. Probiotics for the prevention or treatment of chemotherapy- or radiotherapy-related diarrhoea in people with cancer. Cochrane Database Syst. Rev. 8, CD008831 (2018).
Goldenberg, J. Z. et al. Probiotics for the prevention of Clostridium difficile–associated diarrhea in adults and children. Cochrane Database Syst. Rev. 12, CD006095 (2017).
Zmora, N. et al. Personalized gut mucosal colonization resistance to empiric probiotics is associated with unique host and microbiome features. Cell 174, 1388–1405.e1321 (2018).
Suez, J. et al. Post-antibiotic gut mucosal microbiome reconstitution is impaired by probiotics and improved by autologous FMT. Cell 174, 1406–1423 e1416 (2018).
Kootte, R. S. et al. Improvement of insulin sensitivity after lean donor feces in metabolic syndrome is driven by baseline intestinal microbiota composition. Cell Metab. 26, 611–619.e616 (2017).
Kovatcheva-Datchary, P. et al. Dietary fiber-induced improvement in glucose metabolism is associated with increased Abundance of Prevotella. Cell Metab. 22, 971–982 (2015).
Zeevi, D. et al. Personalized nutrition by prediction of glycemic responses. Cell 163, 1079–1094 (2015).
Kondo, T., Kishi, M., Fushimi, T. & Kaga, T. Acetic acid upregulates the expression of genes for fatty acid oxidation enzymes in liver to suppress body fat accumulation. J. Agric. Food Chem. 57, 5982–5986 (2009).
Hong, Y. H. et al. Acetate and propionate short chain fatty acids stimulate adipogenesis via GPCR43. Endocrinology 146, 5092–5099 (2005).
Ge, H. et al. Activation of G protein–coupled receptor 43 in adipocytes leads to inhibition of lipolysis and suppression of plasma free fatty acids. Endocrinology 149, 4519–4526 (2008).
Jocken, J. W. E. et al. Short-chain fatty acids differentially affect intracellular lipolysis in a human white adipocyte model. Front. Endocrinol. (Lausanne) 8, 372 (2018).
Jia, Y. et al. Butyrate stimulates adipose lipolysis and mitochondrial oxidative phosphorylation through histone hyperacetylation-associated β3 -adrenergic receptor activation in high-fat diet-induced obese mice. Exp. Physiol. 102, 273–281 (2017).
van Nood, E. et al. Duodenal infusion of donor feces for recurrent Clostridium difficile. N. Engl. J. Med. 368, 407–415 (2013).
Chapman, B. C. et al. Fecal microbiota transplant in patients with Clostridium difficile infection: a systematic review. J. Trauma Acute Care Surg. 81, 756–764 (2016).
Carlucci, C., Petrof, E. O. & Allen-Vercoe, E. Fecal microbiota–based therapeutics for recurrent Clostridium difficile infection, ulcerative colitis and obesity. EBioMedicine 13, 37–45 (2016).
Vrieze, A. et al. Transfer of intestinal microbiota from lean donors increases insulin sensitivity in individuals with metabolic syndrome. Gastroenterology 143, 913–916.e17 (2012).
Cani, P. D. & Van Hul, M. Novel opportunities for next-generation probiotics targeting metabolic syndrome. Curr. Opin. Biotechnol. 32, 21–27 (2015).
Patel, R. & DuPont, H. L. New approaches for bacteriotherapy: prebiotics, new-generation probiotics, and synbiotics. Clin. Infect. Dis. 60(Suppl 2), S108–S121 (2015).
Garruti, G., Di Ciaula, A., Wang, H. H., Wang, D. Q. & Portincasa, P. Cross-talk between bile acids and gastro-intestinal and thermogenic hormones: clues from bariatric surgery. Ann. Hepatol. 16, s68–s82 (2017).
Liu, H., Hu, C., Zhang, X. & Jia, W. Role of gut microbiota, bile acids and their cross-talk in the effects of bariatric surgery on obesity and type 2 diabetes. J. Diabetes Investig. 9, 13–20 (2018).
Cani, P. D. Gut microbiota—at the intersection of everything? Nat. Rev. Gastroenterol. Hepatol. 14, 321–322 (2017).
Aron-Wisnewsky, J. et al. Major microbiota dysbiosis in severe obesity: fate after bariatric surgery. Gut 68, 70–82 (2018).
Cani, P. D. Severe obesity and gut microbiota: does bariatric surgery really reset the system? Gut 68, 5–6 (2018).
Wang, Z. et al. Non-lethal inhibition of gut microbial trimethylamine production for the treatment of atherosclerosis. Cell 163, 1585–1595 (2015).
Laurans, L. et al. Genetic deficiency of indoleamine 2,3-dioxygenase promotes gut microbiota-mediated metabolic health. Nat. Med. 24, 1113–1120 (2018).
Chen, Z. et al. Incorporation of therapeutically modified bacteria into gut microbiota inhibits obesity. J. Clin. Invest. 124, 3391–3406 (2014).
Acknowledgements
P.D.C. is a senior research associate at FRS-FNRS (Fonds de la Recherche Scientifique). A.E. is a research associate and MR research fellow at the FRS-FNRS. P.D.C. is a recipient of grants from FNRS, FRFS-WELBIO, under grant no. WELBIO-CR-2017-C02. This research was supported by the FRS-FNRS under The Excellence Of Science (EOS 30770923). This work is supported in part by the Funds Baillet Latour (Grant for Medical Research 2015). P.D.C. is a recipient of the POC ERC grant 2016 (European Research Council, Microbes4U_713547) and ERC Starting Grant 2013 (Starting grant 336452-ENIGMO).
Author information
Authors and Affiliations
Contributions
P.D.C. designed and conceived the outline of the Review. All authors have equally contributed to the writing.
Corresponding author
Ethics declarations
Competing interests
P.D.C. and A.E. are inventors on patent applications (PCT/EP2013/073972; PCT/EP2016/071327, PCT/EP2016/060033 filed in European Patent Office (EP), Australia (AU), Brazil (BR), Canada (CA), China (CN), Eurasian Patent Organization (EA), Israel (IL), India (IN), Hong Kong (HK), Japan (JP), South Korea (KR), Mexico (MX), New Zealand (NZ), and the United States (US)) about the therapeutic use of A. muciniphila and its components. P.D.C. is co-founder of A-Mansia biotech SA.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Cani, P.D., Van Hul, M., Lefort, C. et al. Microbial regulation of organismal energy homeostasis. Nat Metab 1, 34–46 (2019). https://doi.org/10.1038/s42255-018-0017-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-018-0017-4
This article is cited by
-
Gut microbiota in overweight and obesity: crosstalk with adipose tissue
Nature Reviews Gastroenterology & Hepatology (2024)
-
Progress in gut microbiota-host interaction
Science China Life Sciences (2024)
-
The effect of canagliflozin on gut microbiota and metabolites in type 2 diabetic mice
Genes & Genomics (2024)
-
Obese-associated gut microbes and derived phenolic metabolite as mediators of excessive motivation for food reward
Microbiome (2023)
-
The gut microbiome modifies the associations of short- and long-term physical activity with body weight changes
Microbiome (2023)