Abstract
The presence of ERG gene fusion; from developing prostatic intraepithelial neoplasia (PIN) lesions to hormone resistant high grade prostate cancer (PCa) dictates disease progression, altered androgen metabolism, proliferation and metastasis1,2,3. ERG driven transcriptional landscape may provide pro-tumorigenic cues in overcoming various strains like hypoxia, nutrient deprivation, inflammation and oxidative stress. However, insights on the androgen independent regulation and function of ERG during stress are limited. Here, we identify PGC1α as a coactivator of ERG fusion under various metabolic stress. Deacetylase SIRT1 is necessary for PGC1α-ERG interaction and function. We reveal that ERG drives the expression of antioxidant genes; SOD1 and TXN, benefitting PCa growth. We observe increased expression of these antioxidant genes in patients with high ERG expression correlates with poor survival. Inhibition of PGC1α-ERG axis driven transcriptional program results in apoptosis and reduction in PCa xenografts. Here we report a function of ERG under metabolic stress which warrants further studies as a therapeutic target for ERG fusion positive PCa.
Similar content being viewed by others
Introduction
E-Twenty Six transcription factor ERG; exists as a fusion oncogene; in many cancers including myeloid leukemia, Ewing sarcoma, and prostate cancer (PCa)4,5,6,7. Physiologically, temporal tissue-specific expression of ERG is important for hematopoiesis, angiogenesis, and valve development8,9,10. Although reported in ~50% of PCa with connection to aggressiveness and castration resistance (CrPC), the role of TMPRSS2-ERG fusion gene in driving CrPC is not addressed thoroughly1,2,3,7,11. Recent studies of androgen receptor (AR) independent PCa with altered transcriptional circuits strengthen the possibility of ERG fusion-driven programs12,13,14. Like many cancers, metabolic programs associated with altered transcription is an emerging hallmark for PCa and accentuate the requirement of specific transcription factors and nutrients during disease progression15,16,17. In this context, metabolic profiling of ERG fusion-positive PCa patients exhibits altered metabolism of glucose, citrate, and fatty acid associated with high Gleason score15,18,19,20. However, the signals that drive ERG in absence of androgen or under nutrient stress are not well known. Recently, PPARGC1A (PPAR gamma Coactivator 1 alpha/PGC1α) a well-known coactivator has been implicated in PCa oncogenesis via coactivation of important transcription factors like AR, ERR-α21,22,23. PGC1α is a metabolically inducible coactivator that can transactivate various transcription factors in a tissue-dependent manner under metabolic stress24 Interestingly, PGC1α can coactivate a plethora of transcription factors in cancers including PCa, thereby regulating cancer progression25,26,27. Nonetheless, the behavior of PGC1α under nutrient deprivation has not been studied in the context of PCa progression. We speculate that metabolically induced PGC1α may transactivate ERG fusion-driven programs under metabolic stress.
In this study, we found that PGC1α can interact and coactivate ERG under various metabolic stress in the presence of SIRT1 deacetylase and drives an antioxidant function. We found antioxidant target genes of ERG fusion; namely SOD1 and TXN resulting in reactive oxygen species (ROS) clearance and survival. Abrogation of PGC1α, ERG, or antioxidant genes SOD1 and TXN results in ROS-mediated apoptosis during metabolic stress. PCa xenografts show tumor regression and apoptosis in absence of PGC1α. Together our study identifies ERG- PGC1α axis during metabolic stress which might be crucial for PCa progression.
Results
PGC1α acts as a coactivator for ERG fusion under metabolic stress
PGC1α is known to get induced and activated under various metabolic stress and hence act as a coactivator24,27,28,29. We tested if PGC1α affects ERG function under metabolic stress; like glucose or serum deprivation. We deprived VCaP cells harboring ERG fusion of glucose (5 mM) as described previously30 and checked for expression of ERG, PGC1α and verified ERG targets SPP131 (secreted phosphoprotein 1/Osteopontin) and PLAU3 (plasminogen activator, urokinase). We found increased expression of PGC1α without any alteration of ERG. Interestingly, we observed an increased expression of ERG target genes, SPP1 and PLAU under glucose stress (Fig. 1a, b). We have used, AICAR (N1-(β-d-Ribofuranosyl)-5-aminoimidazole-4-carboxamide), to mimic glucose deprivation in VCaP cells. Post-treatment with AICAR showed similar alterations in the expression of ERG and its target genes (Fig. S1a, b). To test the transactivation function of PGC1α over ERG during metabolic stress, we used luciferase reporter tagged SPP1 promoter (SPP1-Luc), in PC3 cells that lack ERG fusion31. Introducing exogenous ERG along with an increasing amount of PGC1α in PC3 resulted in increased SPP1-Luc activity (Fig. 1c). We speculate that overexpression of PGC1α possibly mimics induced PGC1α levels found during metabolic stress resulting in SPP1 luciferase activity. However, in absence of ERG, induced or exogenous levels of PGC1α couldn’t rescue SPP1 luciferase activity indicating PGC1α works by transactivating ERG (Fig. S1c). We verified three different PGC1α shRNA and used VCaP cells with stably silenced PGC1α expression containing shRNA#130 to check the effects of PGC1α over ERG fusion under metabolic stress (Fig. S1d). Glucose deprivation was able to increase SPP1-Luc activity in VCaP control cells but not in VCaP PGC1αKD cells, indicating PGC1α is important for the transactivation of endogenous ERG (Fig. 1d). Basal glucose concentration (25 mM) was unable to induce PGC1α and hence fail to activate ERG-mediated SPP1 luciferase activity (Fig. 1d). We observed a similar PGC1 mediated transactivation function only under glucose deprivation in another ERG containing cell line Colo320 but not in PGC1αKD Colo320 cells or in basal conditions (Fig. S1e). Additionally, AICAR treatment showed coactivation of SPP1-Luc promoter in VCaP control cells in contrast to PGC1αKD cells (Fig. S1f). We further checked the expression of ERG target genes in control and PGC1αKD; VCaP and Colo320 cells. As reported previously24, PGC1α expression is induced under glucose deprivation resulting in activation of ERG target genes SPP1 and PLAU. Glucose deprivation in VCaP and Colo320 PGC1αKD cells did not show any induction of ERG target genes (Fig. S1g). AR-mediated signaling is an important regulator of ERG fusion mediated transcriptional function. To understand the effects of AR modulation during metabolic stress, we treated glucose-deprived VCaP cells with AR antagonist; Flutamide and measured SPP1-Luc activity. As compared to control cells, we observe minimal decrement in SPP1 activity in the presence of Flutamide; indicating PGC1α can trigger ERG transactivation independent of androgen response (Fig. S1h). We also checked if the PGC1α coactivation function over ERG is affected by the androgen analog; R1881. Interestingly, we found that R1881 addition to VCaP control cells further enhanced SPP1-Luc activity suggesting a possible synergistic effect as reported by others (Fig. S1i)21. However, R1881 stimuli in VCaP PGC1αKD cells couldn’t activate SPP1-Luc at 24 h, indicating PGC1α is important for the coactivation of ERG during metabolic stress (Fig. S1i). As reported by others32, we did not observe any upregulation of PGC1α mRNA levels in VCap cells upon treatment with R1881 at normal glucose concentration (Fig. S1j). We checked the effects of other metabolic stress like serum deprivation and cold stress in the coactivation of ERG. Indeed, serum deprivation and cold stress-induced PGC1 coactivation function subsequently resulted in ERG-mediated activation of the SPP1-Luc promoter (Figs. 1e and S1k). The basal condition of 10% serum couldn’t induce PGC1 and subsequently failed to show any SPP1 luciferase activity (Fig. 1e). We checked for ERG target gene expression in VCaP cells (Control or PGC1αKD) under serum deprivation or cold stress and observed induction in control cells but not in PGC1αKD cells (Fig. 1f, g). We further tested if PGC1-β, a homologous PGC1 family member; has overlapping functions towards transactivation of ERG. Co-transfection with PGC1-β did not alter the transactivation of SPP1-Luc, indicating PGC1-β to be non-essential for ERG transactivation functions (Fig. S1l). Together, the results indicate that only under metabolic stress PGC1α is induced and may act as a coactivator for ERG transcription factor.
a VCap Cells were treated for 24 h. Relative transcript levels of genes were analyzed using qRT-PCR. b VCaP cells were treated for 24 h and the whole-cell extract was immunoblotted using indicated antibodies. c PC3 cells were cotransfected as indicated. Luciferase assay was performed post 24 h of transfection and relative firefly luciferase activity was plotted. d VCaP control (Scrambled shRNA) or VCAP PGC1α KD (PGC1α shRNA#1) stable cells transfected with SPP1 luciferase were glucose deprived(5 mM) or cultured under basal condition(25 mM) post-transfection as indicated. Luciferase assay was performed and relative firefly luciferase activity was plotted. e VCaP control or VCAP PGC1α KD stable cells transfected with SPP1 luciferase were serum-deprived or cultured under basal condition post-transfection as indicated. Luciferase assay was performed and relative firefly luciferase activity was plotted. f VCaP control or VCAP PGC1α KD stable cells were serum-starved (Left panel) or received cold stress (4 °C, Right panel) as indicated for 24 h and the whole-cell extract was immunoblotted using indicated antibodies. g VCaP control or VCAP PGC1α KD stable cells were serum-starved (Left panel) or received cold stress (4 °C, Right panel) as indicated for 24 h. Relative transcript levels of genes were analyzed using qRT-PCR. Experiments were performed as biological triplicates (mean ± SD). Two-way ANOVA with multiple comparison test used for statistical significance of 1a, 1g. For 1c, d, e; one-way ANOVA with Dunnett’s multiple comparison test used for statistical significance where ****p < .0001, ***p = 0.0001, **p = .001, *p = .01, n.s not significant.
PGC1α interacts and transactivates ERG fusion
In order to act as a coactivator, PGC1α needs to interact with the ERG transcription factor. PGC1 family of proteins are known to interact with various transcription factors under nonbasal conditions, resulting in transactivation of such factors25,26,27. Under metabolic stress, PGC1α is activated by deacetylation and is induced at mRNA and protein levels for interaction27,28,29,33. For endogenous interaction, VCaP cells were glucose deprived for 24 h and immunoprecipitation of PGC1α protein was performed. We found that induced PGC1α interacted with endogenous ERG during glucose starvation but not under basal conditions (Fig. 2a). A reverse IP on glucose starved VCaP cells with ERG antibody showed that endogenous ERG interacts with PGC1α during metabolic stress (Fig. 2b). Interestingly, we observed that ERG interacts with a deacetylated form of PGC1α but not with the acetylated PGC1α, suggesting that an induced and activate form of PGC1 formed during metabolic stress is capable of interaction (Fig. 2a, b). We confirmed the interaction using overexpression studies in VCaP and PC3 cells and found that ERG interacts with a deacetylated form of exogenously expressed PGC1α (Fig. S2a, b). We found similar endogenous interaction during serum deprivation in VCaP cells between ERG and PGC1α (Fig. S2c). To understand if the PGC1α transactivation domain (1–180 amino acid)34 is necessary for interaction with ERG protein, we performed interaction studies using ectopic ΔN-PGC1α FLAG (PGC1 lacking 1–180 amino acid) in VCaP cells. We did not observe any pull-down of ERG protein with ΔN-PGC1α FLAG (Fig. 2c). Moreover, ΔN-PGC1α FLAG could not transactivate SPP1-Luc in PC3 cells (Fig. 2d). We tried to rescue the transactivation function using the PGC1α 1–180 mutant but did not observe a significant restoration of the PGC1α function. To our surprise, a 1–400 amino acid long PGC1α mutant containing both transactivation domain and deacetylase binding domain was able to completely restore transactivation function (Fig. 2d). We found that PGC1α (1–400) was able to interact with ERG protein (Fig. S2d) and rescue SPP1 luciferase activity under glucose stress in absence of full-length PGC1α (Fig. S2e). As reported earlier34,35, our findings indicate that both PGC1α (1–180) and PGC1α (200–400) domains are necessary for interaction and transactivation function. We speculate that 1–400 is also an activated form of PGC1α as the full-length PGC1’s inhibitory constrains are lacking34,35. Further PGC1α (1–400) domain was able to induce ERG target genes in PGC1α KD cells during glucose starvation (Fig. S2f). Together, the results indicate PGC1α (1–400) domain that includes the N-terminal transactivation domain and deacetylase binding domain is necessary for interaction with ERG fusion protein and results in the transactivation of ERG during metabolic stress.
a VCaP cells were treated as indicated. Cells were harvested after 24 h and subjected to immunoprecipitations using anti-PGC1α antibody or anti-IgG antibody, and western blots were performed for the indicated proteins. b VCaP cells were treated as indicated. Cells were harvested after 24 h and subjected to immunoprecipitations using anti-ERG antibody or anti-IgG antibody, and western blots were performed for the indicated proteins. c VCaP cells were transfected with either vector control(EV), PGC1α Flag, or ΔN-PGC1α Flag as indicated. Cells were harvested 24 h post-transfection and subjected to immunoprecipitations using anti-FLAG antibody and western blots were performed for the indicated proteins. d PC3 cells were cotransfected as indicated. Luciferase assay was performed 24 h post-transfection and relative firefly luciferase activity was plotted. Experiments were performed as biological triplicates (mean ± SD). One-way ANOVA with Dunnett’s multiple comparison test was used for statistical significance of 2d where **p = 0.001.
SIRT1 mediated deacetylation of PGC1α is necessary for ERG interaction and function
Interaction studies indicate that PGC1α deacetylation was crucial for the interaction and function of ERG. In this regard, SIRT1, an important regulator of PGC1α function under metabolic stress had been shown to deacetylate and activate PGC1α29,33,35. Deacetylated PGC1α becomes accessible for binding to various protein partners during metabolic stress27,28. Since inhibition of SIRT1 has been reported to suppress prostate cancer growth36,37, we silenced SIRT1 in VCaP cells to interrogate its role in ERG-PGC1α interaction (Fig. S3a). As reported earlier29, we found that SIRT1 interacts and deacetylates PGC1α under glucose deprivation (Fig. 3a). We observed ERG interaction with PGC1α in the presence of SIRT1 which was absent in SIRT1 silenced cells during glucose deprivation (Fig. 3a). SIRT1 specific inhibitor38; EX-527 also potentiated similar loss of interaction between ERG and PGC1α proteins under glucose stress (Fig. 3b). The SPP1-Luc assay showed no transactivation during glucose or serum starvation in the presence of an SIRT1 inhibitor (Figs. 3c and S3b). These observations indicate that SIRT1 deacetylates PGC1α under stress and promotes its coactivation function towards ERG. We further observed that stress-induced PGC1α could not increase ERG target genes expression during metabolic stress in the absence of SIRT1 (Fig. S3c). Exogenous ERG expression that shall mimic strong ERG activation was able to restore SPP1-Luc activity in the presence of SIRT1 inhibitor confirming ERG transactivation as a downstream event (Fig. 3d). Together these results suggest that SIRT1 mediated deacetylation of PGC1α is necessary for the transactivation of ERG under metabolic stress.
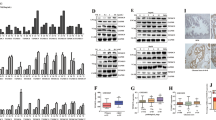
a VCaP cells were transfected with either SiRNA Negative control (SiNeg) or SiRNA Sirt1. Post 24 h of transfection, cells were glucose deprived as indicated, harvested, and subjected to immunoprecipitations using anti- PGC1α antibody. Western blots were performed for the indicated proteins. b VCaP cells were glucose deprived and subjected to either 0.1% DMSO or 10 μM of Sirt1 inhibitor (EX-527) treatment for 24 h. Cells were harvested and subjected to immunoprecipitations using an anti-ERG antibody, and western blots were performed for the indicated proteins. c SPP1 luciferase transfected VCaP cells were glucose deprived and subjected to either 0.1% DMSO or 10 μM of SIRT1 inhibitor as indicated. Luciferase assay was performed and relative firefly luciferase activity was plotted. d SPP1 luciferase transfected VCaP cells were glucose deprived post-ERG transfection and subjected to either 0.1%DMSO or 10 μM of SIRT1 inhibitor as indicated. Luciferase assay was performed and relative firefly luciferase activity was plotted. e ROS levels in treated cells were measured by FACS analysis after staining with fluorescent dye DCFDA. f ROS levels of treated cells were measured by flow cytometry after staining with the fluorescent dye DCFDA. g ROS levels in glucose deprived VCaP cells treated with either 0.1% DMSO or 10 μM of Sirt1 inhibitor were measured by flow cytometry after staining with the fluorescent dye DCFDA. h ROS levels in treated cells post-transfection with indicated constructs were measured by FACS analysis after staining with fluorescent dye DCFDA. i Relative caspase3/7 activity of glucose-deprived VCaP control or VCAP PGC1α KD stable cells were measured at the indicated time. ROS scavenger; Tiron (5 mM) was added for the last 12 h as indicated. j Relative caspase3/7 activity of glucose-deprived VCaP cells treated with either DMSO or 10 μM SIRT1 inhibitor for an indicated time were measured. ROS scavenger; Tiron (5 mM) was added for the last 12 h as indicated. Experiments were performed as biological triplicates (mean ± SD). For 3c–j one-way ANOVA with Dunnett’s multiple comparison test used for statistical significance were performed; where ****p < 0.0001, ***p = 0.0001, **p = 0.001, *p = 0.01, n.s not significant.
Physiological role of ERG- PGC1α axis during metabolic stress
Metabolic stress results in the generation of reactive oxygen species (ROS) that affect cell fate39,40. ROS clearance is an effective mechanism that prostate cancer tumors might utilize to overcome nutrient burden while acquiring castration resistance41,42. The role of ERG in regulating ROS is not well understood during PCa development. We observed increased ROS levels in ERG silenced VCaP cells (VCaP siERG) undergoing metabolic stress (Figs. 3e and S3d). Abrogation of PGC1α using shRNA or PGC1α-inhibitor SR18292 mediated a similar increase of ROS during metabolic stress (Figs. 3f and S3e, f). We manipulated SIRT1 using inhibitor; EX-527 and observed an increase in ROS levels of VCaP cells undergoing metabolic stress (Fig. 3g). We further verified if the increased ROS was a direct effect of SIRT1-PGC1α-ERG axis perturbations by performing rescue experiments using the PGC1α (1–400) mutant which was able to function similarly to full-length PGC1α in our interaction and transactivation assays. Interestingly, the PGC1α (1–400) domain was able to curtail ROS generation during glucose starvation in VCaP PGC1αKD cells but not in ERG silenced VCaP cells indicating PGC1 acts via ERG for ROS clearance (Fig. 3h). In the case of SIRT1 inhibition, exogenous ERG which shall mimic strong ERG activation (Fig. 3d) was able to completely reduce ROS generated during glucose stress (Fig. S3g). However, the PGC1α (1–400) domain could partially curtail ROS (~20% decrease) in cells with SIRT1 inhibition (Fig. S3g) suggesting SIRT1 regulates functional forms of PGC1 to some extent. We checked for apoptosis in VCaP cells undergoing energy stress. Concomitant with increased ROS, we observed increased caspase3/7 activity in VCaP cells where ERG, PGC1α, or SIRT1 was abrogated using RNAi or inhibitors (Figs. 3i, j and S3h, i). ROS scavenger; tiron was able to neutralize oxidative stress and abrogate subsequent apoptosis, suggesting that apoptosis was due to increased levels of ROS (Figs. 3i, j and S3h, i). We envisage that ERG fusion-positive prostate cancer cells are susceptible to energy stress and the PGC1α-Sirt1 axis might provide survival benefits to tumors by activating antioxidant and metabolic functions of ERG fusion.
PGC1α mediated transactivation of ERG fusion revealed antioxidant target genes
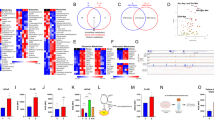
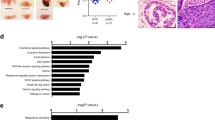
Since our data revealed increased ROS in cells upon ERG-PGC1α axis modulation, we were interested in ERG inducible target genes that might affect ROS clearance. In order to understand the metabolic signatures of ERG fusion-positive prostate cancer, we interrogated the transcriptome of VCaP cells having perturbations of ERG (GSE16671, GSE110656 ERG knockdown or GSE14595; GSE164859 ERG overexpression). Gene set enrichment analysis identified a cluster of antioxidant genes that showed negative enrichment upon knockdown of ERG (ERG KD) in VCaP cells (Fig. 4a Top panel). We analyzed the data set with overexpression of ERG (ERG OE) in fusion negative cells and found the same cluster of antioxidant genes belonging to the reactive oxygen species pathway to be upregulated (Fig. 4a Bottom panel). Next, we picked the significant genes which show core enrichment of ROS pathways in more than one ERG perturbation (Fig. 4b). ERG binding at promoter regions of these core enriched genes were verified by revalidating ERG ChIP Seq datasets (GSE110655) from VCaP cells with ERG knockdown. Interestingly we found four antioxidant genes; TXN, SOD1, LAMTOR5, and ATOX1 with ERG binding signatures at their promoters which were abolished upon knockdown of ERG (Fig. 4c). We verified two other datasets for ERG ChIP (GSE14092 and GSE28950) in VCaP cells and found ERG binding at the promoters of the same genes (Fig. S4a). We did not observe any AR binding at promoters of these genes, indicating a possibility of AR independent induction of these targets by ERG (Figs. 4c and S4a). We further confirmed the presence of a previously reported ERG binding consensus motif in the promoters of these genes43,44 (Fig. S4b).
a Enrichment plot from GSEA was conducted with datasets of ERG Knockdown (top panel) and ERG overexpression (bottom panel). Plots represent statistically significant enrichment (FDR <0.25) of “reactive oxygen species pathway” genes. b Heatmap of core enriched genes in “reactive oxygen species pathway” from GSEA analysis as indicated. c Aligned chromosomal peak regions and enriched ChIP signals (ERG ChIP left and AR ChIP right) in bigwig format were visualized using the UCSC browser for indicated samples. d ChIP assay performed with either ERG or control IgG antibody on glucose-deprived cells. The fold enrichment of co-precipitating DNA was determined by qPCR for indicated promoters. e VCaP control or VCAP PGC1α KD cells treated for 24 h. Relative transcript levels of indicated genes were analyzed using qRT-PCR. f ROS levels of treated VCaP cells were measured by flow cytometry after staining with the fluorescent dye DCFDA. g Relative caspase3/7 activity of treated VCaP cells were measured. h Representative cleaved-caspase3 staining (20X top, 40X bottom) in VCaP control and VCaP PGC1α KD xenograft following STF on day 21. i VCaP (control) and VCaP(PGC1α KD) xenograft tumor volume (mm3) following STF on days 4, 21, and 42. Lines indicate the mean ± SD (n = 6 per group). j Violin plots for transcript level in PRAD patient data set (TCGA targeted GTEx) were accessed using the Xena browser. Data were Z-score normalized and p value calculated using Welch’s t-test. k Kaplan–Meier curves for survival probability of TCGA PRAD data separated into high- and low-risk groups. Survival analysis was performed using Surv Express software. Experiments were performed as biological triplicates (mean ± SD). Two-way ANOVA with multiple comparison tests used for statistical significance of 4e. For 4 d, f, g, i one-way ANOVA with Dunnett’s multiple comparison test used for statistical significance were performed; where ****p < 0.0001, ***p = 0.0001, **p = 0.001, *p = 0.01, n.s not significant.
To understand how the ERG-PGC1α axis relates to these genes under metabolic stress, we performed chromatin immunoprecipitation studies on VCaP PGC1αKD cells undergoing glucose deprivation. We found a loss of ERG binding at promoters of these genes in the absence of PGC1α (Fig. 4d). We checked the expression of these genes under various metabolic stress. Interestingly, TXN, SOD1, and LAMTOR5 were significantly upregulated in VCaP control cells during glucose deprivation. No significant changes were observed in the absence of PGC1α or ERG during glucose stress indicating functions to be mediated via the ERG-PGC1α complex (Figs. 4e and S4c). We checked for modulation of these ERG target genes under serum stress and found them to be induced by ERG in a PGC1α dependent manner (Fig. S4d). Together these data indicate that PGC1α might regulate direct transcriptional target genes of ERG during metabolic stress.
In order to access the functional importance of these antioxidant genes, we checked ROS levels and caspase3/7 activity of VCap cells undergoing glucose starvation by silencing TXN and SOD1 genes selectively. We found that silencing of TXN or SOD1 genes abrogated ROS clearance in VCaP cells under metabolic stress (Fig. 4f). We also observed increased caspase3/7 activity in cells under metabolic stress in absence of these genes (Fig. 4g). Simultaneous knockdown of SOD1 and TXN further elevated ROS and Caspase activity under metabolic stress. We observed similar results for Caspase3/7 activity in ERG expressing COLO320 cells during glucose stress (Fig. S4e). Under basal/unstressed conditions, we did not observe any significant caspase activity associated with the silencing of these genes alone (Fig. S4f). To corroborate our finding in vivo, we used intermittent short-term fasting (STF) on PCa xenografts as described previously45,46. We had subjected SCID mice (NOD.Cg-Prkdcscid) (The Jackson Laboratory, Bar Harbor, USA) bearing VCaP control or VCaP PGC1α knockdown xenografts to STF, as described(details in supplementary methods) previously46. Although no significant changes in animal body weight subjected to STF was observed (Fig. S4g). We found that Caspase3/7 activity increased in PGC1α silenced xenografts as compared to control which correlated with decreased tumor growth observed in PGC1α silenced xenograft-containing animals (Fig. 4h, i). Additionally, the expression of ERG target genes SOD1, TXN, and PLAT went down in PGC1α silenced xenograft as compared to control animals under STF (Fig. S4h). Further tumor growth between VCaP control or VCaP PGC1α knockdown xenograft animals not subjected to STF were comparable, indicating that PGC1α’s tumor-promoting function is activated only under starvation resulting in ERG-mediated ROS clearance and anti-apoptotic functions in control animals but fails in absence of PGC1α (Figs. S4i and 4i). We conclude that PGC1α silenced animals fail to initiate an ERG-mediated antioxidant response to metabolic stress resulting in ROS-mediated apoptosis and decreased tumor growth. Our in vitro results during metabolic stress were further verified using an SIRT1 inhibitor on VCAP xenografts containing animals. We observed significant Caspase3/7 staining on xenografts of animals treated with a combination of STF and SIRT1 inhibitor but not with SIRT1 inhibitor alone indicating SIRT1-PGC1 axis activity only during metabolic stress (Fig. S4j). Together, our in vivo studies supports the role of SIRT1 mediated deacetylation of PGC1α resulting in the coactivation of ERG antioxidant and protumorigenic functions during metabolic stress. Interestingly, we found high expression of antioxidant genes; SOD1, TXN, LAMTOR5, and ATOX1 in prostate cancer patients having higher ERG fusion expression (Figs. 4j and S4k). Multivariant survival analysis predicts poor survival and higher tumor recurrence of PCa patients in correlation with high expression of these antioxidant genes (Figs. 4k and S4l). In conclusion, our study identified the ERG-PGC1 axis which provides antioxidant functions to PCa cells undergoing metabolic perturbations.
Discussion
Interestingly metabolic stress including glucose, glutamine, and serum deprivation has been reported to inhibit PCa growth via altering the redox balance of cells47,48,49,50. Categorically, carbohydrate restriction in mouse retarded PCa tumor growth indicates adaptive glucose metabolism51,52. However, in depth understanding of mechanisms maintaining redox balance thereby providing adaptive benefits under nutrient-deprived state for PCa is not well understood. Our study on metabolic stress and ERG fusion-driven antioxidant functions is supportive of PCa metabolic program. We report that PGC1α acts as a coactivator of ERG fusion during metabolic stress resulting in the activation of antioxidant target genes of ERG. SIRT1 mediated deacetylation of PGC1α is necessary for ERG and PGC1α interaction and prevention of ROS-dependent apoptosis. Given high SIRT1 levels have been associated with PCa initiation, progression, and metastasis, our study suggests that modulation of SIRT1 can be an important vulnerability36,37,53.
Our study in PCa is corollary to oncogenic roles of PGC1α coactivator in various cancers21,32,54,55. Earlier reports suggest that PGC1α supports PCa progression by activating the PGC1α-AR program under androgen stimulation21,32. Our results with AR antagonist flutamide didn’t affect PGC1α mediated transactivation of ERG fusion suggesting an AR independent mechanism under metabolic stress. Additionally, we observed a synergistic increase in ERG transcriptional activity upon androgen stimulation in presence of PGC1α but not in its absence; indicating PGC1α is necessary for transactivation of ERG fusion under metabolic stress. As reported by others, we anticipate prolonged androgen stimuli to activate the PGC1α-AR-dependent program which is absent in our studies and is in support of a low PGC1α state (basal levels) or AR low environment21,22,32. On the contrary, PGC1α has been recently shown to have tumor suppressor functions in PCa22,23. In this context, the transcription factor ERRα is reported to cooperate with PGC1α and suppress PCa metastasis22,23. Interestingly, Torrano et al. report low endogenous levels of PGC1α in basal cellular context while cautioning about nonbasal conditions like stress to alter expression levels; which might explain our contradictory observation with respect to induced endogenous levels of PGC1α under metabolic stress22. Opposingly, ERRα is reported to form a reciprocal regulatory loop with ERG fusion which can steer PCa progression56. We contemplate that the ERRα-PGC1α studies exogenously overexpress PGC1α in ERG fusion negative PC3 and DU145 cells thus unable to acknowledge ERG fusion’s regulation over ERRα22,23,56. PGC1α has been reported to alter promoter specificity of various transcription factors under nonbasal conditions of metabolic stress27,28,30. We envisage that PGC1α might alter promoter specificity of ERG during nonbasal conditions thus overriding ERRα function. Overall, our study highlights a prominent oncogenic role of the PGC1α-ERG axis during metabolic stress, where elevated PGC1α levels resulted in a pro-survival response.
In recent years, intermittent caloric restriction in combination with chemotherapy has gained importance and clinical trials are being proposed for cancers including PCa with dietary restrictions57 (NCT02126449, NCT04288336, NCT02710721, NCT04292041, and NCT03478904). The previous finding on dietary restrictions alone in PCa xenograft-containing mice had opposing results on survival and tumor growth45,46. Under short-term fasting, we observed tumor inhibition in mice bearing PGC1α silenced xenografts in comparison to ones having intact PGC1α - ERG axis. Our findings on intermittent caloric restrictions with abolished ERG-PGC1 axis might indicate a positive outcome for patients with low PGC1α levels.
Oxidative stress has been insinuated into PCa initiation, aggressiveness, castration resistance, and yet clinical interventions with antioxidant agents is not very promising58,59,60,61,62. Although antioxidant genes like SOD3, TXN, and SOD1 has been associated with PCa aggressiveness and metastasis, none have shown a direct relationship with ERG fusion63,64,65. Our study identified the previously unknown antioxidant function of ERG fusion under metabolic stress which is mitigated through its direct transcriptional target genes SOD1 and TXN. We speculate that PCa initiation under a nutrient-rich environment would show high ROS levels and cancer growth as reported previously61,62. While nutrient deficiency at later stages of the disease would activate the PGC1α-ERG axis for prolonged survival via ROS clearance.
SOD1 and TXN has been previously shown to have a profound role in the progression of various cancers66,67. We observed that simultaneous inhibition of SOD1 and TXN genes during metabolic stress caused significant apoptosis in PCa cells. Our study showed high expression levels of these genes in ERG fusion-positive PCa patients and their associated survival risk. Given the recurrence of ERG fusion in ~50% of patients, these newly found antioxidant functions might provide a therapeutic target for ERG positive Pca patients2,7. We surmise that PGC1α mediated coactivation of ERG fusion is largely tumor type and context-dependent and might be advantageous during the transition to castration-resistant phenotype.
Methods
Cell lines and culture conditions
We obtained VCaP (ATCC:CRL-2876™) from ATCC (Manassas, VA). PC3 and Colo320 cells was purchased from the National center for cell sciences (Pune, India). We cultured PC3 in RPMI medium (Thermo Fisher Scientific) and Colo320, VCaP cells in DMEM medium (Thermo Fisher Scientific). We supplemented the growth medium with 10% fetal bovine serum (Thermo Fisher Scientific), and cells were maintained at 37 °C in 5% CO2 for all experiments. The identity of all cell lines purchased were confirmed by STR analysis. All cell lines were determined to be mycoplasma free using the MycoAlert Plus system and kit (Lonza). For inducing glucose starvation, cells were grown in glucose-free DMEM (Thermo Fisher Scientific) containing dialyzed FBS (Thermo Fisher Scientific) and 5 mM glucose (Sigma) as described previously30. Control cells for starvation experiments were also grown in the same glucose-free DMEM containing dialyzed FBS and 25 mM glucose for basal conditions. For inducing serum starvation, cells were grown in serum-free DMEM (Thermo Fisher Scientific). For cold stress, cells were exposed to 4 °C as described previously30.
Compounds, siRNAs, plasmids, and transfections
See Supplementary Table 1 for compounds and siRNAs used in this study. All compounds were dissolved in DMSO at room temperature except glucose and AICAR that were dissolved in Phosphate buffered saline. All siRNAs were transfected using RNAi-Max (Thermo Fisher Scientific). All Plasmids used are listed in Supplementary Table 2. The SPP1 gene promoter with ERG binding site as described previously31; was cloned in the PGL4.24 luciferase reporter vector (Promega). PGC1alpha shRNAs were cloned in the pBABE-U6-shRNA plasmid (see Supplementary Table 2). pcDNA3FlagERG was a gift from Christopher Vakoc (Addgene plasmid # 66977). pcDNA-f:PGC1alpha, pcDNAflagPGC1alpha (180–797), and pcDNA-f:PGC1b were gifts from Bruce Spiegelman (Addgene plasmid # 1026, Addgene plasmid # 8943, Addgene plasmid # 1031). For transgene expression studies, plasmid DNA was transfected using Lipofectamine 3000 (Thermo Fisher Scientific). We used puromycin (2 µg/ml) to select for stably expressing cells (Thermo Fisher Scientific).
Gene expression
For real-time PCR analysis, total RNA from cells and tumor tissues were extracted using an RNA extraction kit (Ambion, Life technologies) and cDNA synthesis using the iScript cDNA synthesis kit (Bio-Rad Laboratories) was performed following the manufacturer’s protocol. qPCR was performed using the primer sets listed in Supplementary Table 2. Relative gene expression was assessed using three biological replicas and data are shown as the mean ± SD. 18sRNA was used as an internal control for all samples.
Immunoblotting and immunoprecipitation analysis
Whole-cell lysates from cells were prepared using lysis buffer (Thermo Fisher Scientific) supplemented with protease inhibitor and PhosphoSTOP tablets (Merck Millipore). An equal amount of total cellular proteins per sample was subjected to SDS–PAGE, transferred to a PVDF membrane (Merck Millipore) and probed with required antibodies. See Supplementary Table 3 for antibodies used in this study.
Immunoprecipitations were performed using 500 μg cell extracts as described previously68 and incubated with antibodies and precleared Protein A/G plus agarose (Santacruz Biotechnology). Subsequent immunoblots were performed as described above.
Chromatin Immunoprecipitation analysis
Chromatin immunoprecipitations were performed using ChIP Kit (Millipore) according to the manufacturer’s instructions. Briefly, 1 × 107 cells were fixed using 1% formaldehyde and then lysed using SDS lysis buffer. We incubated 500 µl of sonicated chromatin with either anti-ERG antibody or normal rabbit IgG antibody overnight at 4 °C followed by 2 h incubation in Agarose bead Slurry. Precipitated chromatin DNA cross-links were reversed by incubation at 65 °C for 4 h and immunoprecipitated DNA was analyzed by qPCR. We normalized values to those derived from normal IgG-precipitated. qPCR primer sequences for ChIP are listed in Supplementary Table 2. Fold-enrichment was assessed using three biological replicas of each treatment and data are shown as the mean ± SD.
Functional assays
For luciferase assays, cells were treated as indicated 24 h post-transfection. Post experiment, luciferase activity was determined using the Dual Glo Luciferase Reporter assay system (Promega) as per the manufacturer’s instruction. The difference in transfection efficiency across samples was normalized by co-transfecting pRL-TK which expresses Renilla luciferase. Caspase activity was measured using Caspase Glo 3/7 assay system (Promega) per the manufacturer’s instruction. For all functional assays, three biological replicas were used and data were shown as mean ± SD.
ROS generation was measured using CH2- DCFDA (Life Technologies). The cells were incubated with 10 mM DCFDA at 37 °C for 30 min. The cells were then harvested and resuspended in culture media. Samples were analyzed by flow cytometry (BD Biosciences) and data analyzed using Flo Jo software.
Statistics and reproducibility
One-way analysis of variance (one-way ANOVA) was performed in Prism (GraphPad) using Dunnett’s multiple comparisons test to correct for multiple comparisons to the same control group; a p value < 0.05 was considered significant. Brown Forsythe Welch corrections were performed for variation in SD. Two-way analysis of variance (two-way ANOVA) was performed in Prism (GraphPad) using Sidak’s multiple comparisons test to correct for multiple comparisons to the same control group; a p value < 0.05 was considered significant. All experiments were performed in triplicate and mean ± SD was used to represent data.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The following datasets are used for GSEA analysis GSE16671, GSE110656, GSE14595; GSE164859. For ChIP-sequencing data analysis GSE110655, GSE14092, and GSE28950 were used. All datasets used for the study are listed in Supplementary Table 4. Details of methods and tools used are described in Supplementary Methods. All raw data used in the study are provided in Supplementary Data 1 and Supplementary Data 2. Other online tools like UCSC XENA browser used TCGA PRAD and GTex data, SURV express used TCGA prostate cancer and GSE40272 datasets.
References
Klezovitch, O. et al. A causal role for ERG in neoplastic transformation of prostate epithelium. Proc. Natl Acad. Sci. USA 105, 2105–2110 (2008).
Mehra, R. et al. Characterization of TMPRSS2-ETS gene aberrations in androgen-independent metastatic prostate cancer. Cancer Res. 68, 3584–3590 (2008).
Tomlins, S. A. et al. Role of the TMPRSS2-ERG gene fusion in prostate cancer. Neoplasia 10, 177–188 (2008).
Shimizu, K. et al. An ets-related gene, ERG, is rearranged in human myeloid leukemia with t(16;21) chromosomal translocation. Proc. Natl Acad. Sci. USA 90, 10280–10284 (1993).
Sorensen, P. H. et al. A second Ewing’s sarcoma translocation, t(21;22), fuses the EWS gene to another ETS-family transcription factor, ERG. Nat. Genet. 6, 146–151 (1994).
Tomlins, S. A. et al. Distinct classes of chromosomal rearrangements create oncogenic ETS gene fusions in prostate cancer. Nature 448, 595–599 (2007).
Tomlins, S. A. et al. Recurrent fusion of TMPRSS2 and ETS transcription factor genes in prostate cancer. Science 310, 644–648 (2005).
Birdsey, G. M. et al. Transcription factor Erg regulates angiogenesis and endothelial apoptosis through VE-cadherin. Blood 111, 3498–3506 (2008).
Loughran, S. J. et al. The transcription factor Erg is essential for definitive hematopoiesis and the function of adult hematopoietic stem cells. Nat. Immunol. 9, 810–819 (2008).
Vijayaraj, P. et al. Erg is a crucial regulator of endocardial-mesenchymal transformation during cardiac valve morphogenesis. Development 139, 3973–3985 (2012).
Gopalan, A. et al. TMPRSS2-ERG gene fusion is not associated with outcome in patients treated by prostatectomy. Cancer Res 69, 1400–1406 (2009).
Bluemn, E. G. et al. Androgen receptor pathway-independent prostate cancer is sustained through FGF signaling. Cancer Cell 32, 474–489.e476 (2017).
Roudier, M. P. et al. Characterizing the molecular features of ERG-positive tumors in primary and castration resistant prostate cancer. Prostate 76, 810–822 (2016).
Shukla, S. et al. Aberrant activation of a gastrointestinal transcriptional circuit in prostate cancer mediates castration resistance. Cancer Cell 32, 792–806.e797 (2017).
Bader, D. A. & McGuire, S. E. Tumour metabolism and its unique properties in prostate adenocarcinoma. Nat. Rev. Urol. 17, 214–231 (2020).
Reina-Campos, M. et al. Increased serine and one-carbon pathway metabolism by PKClambda/iota deficiency promotes neuroendocrineprostate cancer. Cancer Cell 35, 385–400.e389 (2019).
Shao, Y. et al. Metabolomics and transcriptomics profiles reveal the dysregulation of the tricarboxylic acid cycle and related mechanisms in prostate cancer. Int. J. Cancer 143, 396–407 (2018).
Eidelman, E., Twum-Ampofo, J., Ansari, J. & Siddiqui, M. M. The metabolic phenotype of prostate cancer. Front. Oncol. 7, 131 (2017).
Hansen, A. F. et al. Presence of TMPRSS2-ERG is associated with alterations of the metabolic profile in human prostate cancer. Oncotarget 7, 42071–42085 (2016).
Meller, S. et al. Integration of tissue metabolomics, transcriptomics and immunohistochemistry reveals ERG- and gleason score-specific metabolomic alterations in prostate cancer. Oncotarget 7, 1421–1438 (2016).
Shiota, M. et al. Peroxisome proliferator-activated receptor gamma coactivator-1alpha interacts with the androgen receptor (AR) and promotes prostate cancer cell growth by activating the AR. Mol. Endocrinol. 24, 114–127 (2010).
Torrano, V. et al. The metabolic co-regulator PGC1alpha suppresses prostate cancer metastasis. Nat. Cell Biol. 18, 645–656 (2016).
Valcarcel-Jimenez, L. et al. PGC1alpha suppresses prostate cancer cell invasion through ERRalpha transcriptional control. Cancer Res. 79, 6153–6165 (2019).
Finck, B. N. & Kelly, D. P. PGC-1 coactivators: inducible regulators of energy metabolism in health and disease. J. Clin. Invest 116, 615–622 (2006).
Girnun, G. D. The diverse role of the PPARgamma coactivator 1 family of transcriptional coactivators in cancer. Semin. Cell Dev. Biol. 23, 381–388 (2012).
Luo, C., Widlund, H. R. & Puigserver, P. PGC-1 coactivators: shepherding the mitochondrial biogenesis of tumors. Trends Cancer 2, 619–631 (2016).
Tan, Z. et al. The role of PGC1alpha in cancer metabolism and its therapeutic implications. Mol. Cancer Ther. 15, 774–782 (2016).
Lin, J., Handschin, C. & Spiegelman, B. M. Metabolic control through the PGC-1 family of transcription coactivators. Cell Metab. 1, 361–370 (2005).
Canto, C. & Auwerx, J. PGC-1alpha, SIRT1 and AMPK, an energy sensing network that controls energy expenditure. Curr. Opin. Lipido. 20, 98–105 (2009).
Sen, N., Satija, Y. K. & Das, S. PGC-1alpha, a key modulator of p53, promotes cell survival upon metabolic stress. Mol. Cell 44, 621–634 (2011).
Flajollet, S. et al. Abnormal expression of the ERG transcription factor in prostate cancer cells activates osteopontin. Mol. Cancer Res. 9, 914–924 (2011).
Tennakoon, J. B. et al. Androgens regulate prostate cancer cell growth via an AMPK-PGC-1alpha-mediated metabolic switch. Oncogene 33, 5251–5261 (2014).
Rodgers, J. T. et al. Nutrient control of glucose homeostasis through a complex of PGC-1alpha and SIRT1. Nature 434, 113–118 (2005).
Puigserver, P. et al. Activation of PPARgamma coactivator-1 through transcription factor docking. Science 286, 1368–1371 (1999).
Rodgers, J. T. et al. Metabolic adaptations through the PGC-1 alpha and SIRT1 pathways. FEBS Lett. 582, 46–53 (2008).
Byles, V. et al. SIRT1 induces EMT by cooperating with EMT transcription factors and enhances prostate cancer cell migration and metastasis. Oncogene 31, 4619–4629 (2012).
Huang, S. B. et al. Androgen deprivation-induced elevated nuclear SIRT1 promotes prostate tumor cell survival by reactivation of AR signaling. Cancer Lett. 505, 24–36 (2021).
Solomon, J. M. et al. Inhibition of SIRT1 catalytic activity increases p53 acetylation but does not alter cell survival following DNA damage. Mol. Cell Biol. 26, 28–38 (2006).
Ahmad, I. M. et al. Mitochondrial O2*- and H2O2 mediate glucose deprivation-induced stress in human cancer cells. J. Biol. Chem. 280, 4254–4263 (2005).
Aykin-Burns, N., Ahmad, I. M., Zhu, Y., Oberley, L. W. & Spitz, D. R. Increased levels of superoxide and H2O2 mediate the differential susceptibility of cancer cells versus normal cells to glucose deprivation. Biochem. J. 418, 29–37 (2009).
Han, C. et al. Roles of reactive oxygen species in biological behaviors of prostate cancer. Biomed. Res. Int. 2020, 1269624 (2020).
Luo, X. H. et al. KLF14 potentiates oxidative adaptation via modulating HO-1 signaling in castrate-resistant prostate cancer. Endocr. Relat. Cancer 26, 181–195 (2019).
Martens, J. H. et al. ERG and FLI1 binding sites demarcate targets for aberrant epigenetic regulation by AML1-ETO in acute myeloid leukemia. Blood 120, 4038–4048 (2012).
Nhili, R. et al. Targeting the DNA-binding activity of the human ERG transcription factor using new heterocyclic dithiophene diamidines. Nucleic Acids Res. 41, 125–138 (2013).
Bonorden, M. J. et al. Intermittent calorie restriction delays prostate tumor detection and increases survival time in TRAMP mice. Nutr. Cancer 61, 265–275 (2009).
Thomas, J. A., II et al. Effect of intermittent fasting on prostate cancer tumor growth in a mouse model. Prostate Cancer Prostatic Dis. 13, 350–355 (2010).
Bueno De Paiva, L. et al. Effects of RhoA and RhoC upon the sensitivity of prostate cancer cells to glutamine deprivation. Small GTPases 12, 20–26, https://doi.org/10.1080/21541248.2018.1546098.
Gonzalez-Menendez, P. et al. GLUT1 protects prostate cancer cells from glucose deprivation-induced oxidative stress. Redox Biol. 17, 112–127 (2018).
Thomas, R. & Kim, M. H. HIF-1 alpha: a key survival factor for serum-deprived prostate cancer cells. Prostate 68, 1405–1415 (2008).
White, E. Z. et al. Serum deprivation initiates adaptation and survival to oxidative stress in prostate cancer cells. Sci. Rep. 10, 12505 (2020).
Caso, J. et al. The effect of carbohydrate restriction on prostate cancer tumor growth in a castrate mouse xenograft model. Prostate 73, 449–454 (2013).
Freedland, S. J. et al. Carbohydrate restriction, prostate cancer growth, and the insulin-like growth factor axis. Prostate 68, 11–19 (2008).
Bader, D. A. et al. Mitochondrial pyruvate import is a metabolic vulnerability in androgen receptor-driven prostate cancer. Nat. Metab. 1, 70–85 (2019).
Sancho, P. et al. MYC/PGC-1alpha balance determines the metabolic phenotype and plasticity of pancreatic cancer stem cells. Cell Metab. 22, 590–605 (2015).
Vazquez, F. et al. PGC1alpha expression defines a subset of human melanoma tumors with increased mitochondrial capacity and resistance to oxidative stress. Cancer Cell 23, 287–301 (2013).
Xu, Z. et al. Nuclear receptor ERRalpha and transcription factor ERG form a reciprocal loop in the regulation of TMPRSS2:ERG fusion gene in prostate cancer. Oncogene 37, 6259–6274 (2018).
Palomar-Cros, A. et al. The association of nighttime fasting duration and prostate cancer risk: results from the multicase-control (MCC) study in Spain. Nutrients 13, 2662 (2021).
Chaiswing, L., Zhong, W., Liang, Y., Jones, D. P. & Oberley, T. D. Regulation of prostate cancer cell invasion by modulation of extra- and intracellular redox balance. Free Radic. Biol. Med. 52, 452–461 (2012).
Gaziano, J. M. et al. Vitamins E and C in the prevention of prostate and total cancer in men: the Physicians’ Health Study II randomized controlled trial. JAMA 301, 52–62 (2009).
Klein, E. A. et al. Vitamin E and the risk of prostate cancer: the selenium and vitamin E cancer prevention trial (SELECT). JAMA 306, 1549–1556 (2011).
Kumar, B., Koul, S., Khandrika, L., Meacham, R. B. & Koul, H. K. Oxidative stress is inherent in prostate cancer cells and is required for aggressive phenotype. Cancer Res 68, 1777–1785 (2008).
Paschos, A., Pandya, R., Duivenvoorden, W. C. & Pinthus, J. H. Oxidative stress in prostate cancer: changing research concepts towards a novel paradigm for prevention and therapeutics. Prostate Cancer Prostatic Dis 16, 217–225 (2013).
Kim, J. et al. SOD3 acts as a tumor suppressor in PC-3 prostate cancer cells via hydrogen peroxide accumulation. Anticancer Res. 34, 2821–2831 (2014).
Vance, T. M. et al. Thioredoxin 1 in Prostate Tissue Is Associated with Gleason Score, Erythrocyte Antioxidant Enzyme Activity, and Dietary Antioxidants. Prostate Cancer 2015, 728046, https://doi.org/10.1155/2015/728046.
Young, B., Purcell, C., Kuang, Y. Q., Charette, N. & Dupre, D. J. Superoxide dismutase 1 regulation of CXCR4-mediated signaling in prostate cancer cells is dependent on cellular oxidative state. Cell Physiol. Biochem. 37, 2071–2084 (2015).
Harris, I. S. et al. Glutathione and thioredoxin antioxidant pathways synergize to drive cancer initiation and progression. Cancer Cell 27, 211–222 (2015).
Papa, L., Manfredi, G. & Germain, D. SOD1, an unexpected novel target for cancer therapy. Genes Cancer 5, 15–21 (2014).
Groisman, R. et al. The ubiquitin ligase activity in the DDB2 and CSA complexes is differentially regulated by the COP9 signalosome in response to DNA damage. Cell 113, 357–367 (2003).
Acknowledgements
Apologies to those whose related publications were not cited due to space limitations. The authors acknowledge the Rajiv Gandhi Center for Biotechnology, Trivandrum, University of Kalyani, West Bengal, and the Department of Zoology, Burdwan University; West Bengal for providing facilities to carry out experiments. The authors acknowledge the Department of Bioinformatics & Applied Sciences, Indian Institute of Information Technology, Allahabad for providing computational help. N.S. was funded by DBT—Ramalingaswami Fellowship (Award sanction No: BT/RLF/Re-entry/27/2016) from the Department of Biotechnology, India. A.D. worked under a fellowship from SERB-DST (Award sanction No: ECR/2018/000595) awarded to N.S. from the Department of Science and Technology, India. The authors acknowledge the TCGA Research Network: https://www.cancer.gov/tcga.The authors have no financial conflict of interest to report.
Author information
Authors and Affiliations
Contributions
A.D. and N.S. designed the study, performed the experiments, and analyzed the data. N.S. was responsible for conceptualization, funding acquisition, investigation, methodology, project administration, resources, supervision, writing, and editing. I.A. and P.K.V. were responsible for the investigation, data curation, and visualization related to bioinformatics analysis. A.P. and S.K.H. were responsible for animal tissue-related work, resources, supervision, writing, and editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Ping Mu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Maralice Conacci Sorrell and Christina Karlsson Rosenthal.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Dhara, A., Aier, I., Paladhi, A. et al. PGC1 alpha coactivates ERG fusion to drive antioxidant target genes under metabolic stress. Commun Biol 5, 416 (2022). https://doi.org/10.1038/s42003-022-03385-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-022-03385-x
This article is cited by
-
Peroxisome proliferator-activated receptor gamma coactivator-1 (PGC-1) family in physiological and pathophysiological process and diseases
Signal Transduction and Targeted Therapy (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.