Abstract
Left-handed Z-DNA/Z-RNA is bound with high affinity by the Zα domain protein family that includes ADAR (a double-stranded RNA editing enzyme), ZBP1 and viral orthologs regulating innate immunity. Loss-of-function mutations in ADAR p150 allow persistent activation of the interferon system by Alu dsRNAs and are causal for Aicardi-Goutières Syndrome. Heterodimers of ADAR and DICER1 regulate the switch from RNA- to protein-centric immunity. Loss of DICER1 function produces age-related macular degeneration, a different type of Alu-mediated disease. The overlap of Z-forming sites with those for the signal recognition particle likely limits invasion of primate genomes by Alu retrotransposons.
Similar content being viewed by others
Z-DNA is the left-handed conformer of double-stranded DNA that normally exists in the right-handed Watson-Crick B-form. The flip from the B-form to the Z-form occurs when processive enzymes such as polymerases and helicases generate underwound DNA in their wake. The existence of Z-DNA was unexpected and its discovery accidental, the structure trapped in the first synthetic DNA ever crystallized. Initially the biological importance of Z-DNA was overestimated, after which it has been underappreciated (“We tend to overestimate the effect of a technology in the short run and underestimate the effect in the long run” - Roy Anara). An important inflection point has been the identification of the Z-DNA binding domain named Zα1,2 from the dsRNA editing protein ADAR3,4. This domain’s specificity for the left-handed conformation of Z-DNA was shown in a series of high-resolution NMR and X-ray studies5,6. The interactions between Zα and Z-DNA are conformation-specific, with no base-specific contacts.
Over the past 25 years, work by many outstanding scientists has established a clearer view of the roles played by ADAR, along with other Z-binding proteins, in the interface between the RNA and protein worlds in health and disease. Central to these findings were genetic studies revealing an essential role for the interferon-induced p150 isoform of ADAR that includes the Zα domain in the negative regulation of immune responses induced by dsRNA. In addition, Zα was found to bind the left-handed Z-RNA conformation of dsRNA without sequence specificity, providing a mechanism for targeting ADAR to dsRNA editing substrates independently of the three ADAR dsRNA binding domains that recognize the more common right-handed A-form dsRNA conformation. Mutations that reduce p150 Z-binding along with those that impair enzymatic activity cause interferonopathies such as Aicardi-Goutières Syndrome. The dsRNAs that induce disease arise from transcripts with inverted repeats, which fold back and base pair with each other. Most commonly, these dsRNAs derive from Alu retroelements but they are also generated during viral infections.
The Z-conformation, both as Z-DNA and Z-RNA, has likely played an essential role in limiting Alu retroelement invasion of primate genomes during evolution. Besides ADAR, key partners in this battle have been the signal recognition particle proteins SRP9 and SRP14 (on which Alu retrotransposition depends) and DICER1 (an endoribonuclease that heterodimerizes with ADAR and initiates RNA interference against single copy Alu transcripts, which are unlikely to form the long dsRNA required either for editing by ADAR or for the formation of Z-RNA). SRP9/14 dimers bind to Alu sequences capable of Z-formation, with stronger Z-formers found in those elements most successful at invading the genome. Indeed, loss of Z-forming potential is associated with loss of SRP9/14 binding and diminished Alu invasion. Recognition of Z-formation by the Zα domain targets Alu dsRNA for editing by ADAR, while recognition of Z-DNA by ADAR recruits DICER1 machinery to single copy genomic insertions, both enzymes serving to limit further retrotransposition. Loss of DICER1 function is associated with the accumulation of Alumers and inflammasome activation, leading to age-related macular degeneration (AMRD). The need to defend against retroelements provides a rationale for maintaining Z-forming segments in the genome. Other roles for the left-handed conformation in the readout of genomic information exist and these too show an association with disease. A genome-wide analysis reveals that both dsRNA editing and known disease genes are enriched for long Z-DNA forming segments.
In this review, I discuss the properties of Z-DNA and Z-RNA, and detail how the Zα domain of ADAR limits Alu retrotransposition and protects against human disease. I examine the impact of the Z-conformation on primate evolution, and outline key questions that remain in the field.
Z-DNA and Zα domains
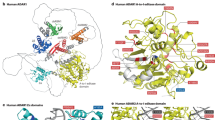
The discovery of Z-DNA occurred when an unusual DNA conformation was observed upon placing poly(dC-dG) in 5 M NaCl7. Its structure was revealed when the crystal of d(CG)6 was solved8. The left-handed helix was built from a dinucleotide repeat where the usual anti conformation of bases alternated with the unusual syn form (where the purine or pyrimidine base projects over the (deoxy)ribose ring, perpendicular to its plane, rather than pointing away from it as it does in anti), giving rise to a zig-zag backbone structure, features captured by the name Z-DNA (Fig. 1).
The B−Z junction. Z-DNA is a conformer of B-DNA stabilized both by negative superhelical stress and by binding of the Zα domain of ADAR1. Zα is conformation specific, contacting the DNA backbone through the α3 helix and its carboxy-terminal β Hairpin but not making any sequence-specific contacts with bases. Formation of the B−Z junction is driven entropically by the eversion of two bases from the helix, a process further favored when non-Watson-Crick basepairs, such as mismatches, are present at this position (PDB structure 2ACJ)
The demonstration that B-DNA could be flipped to form Z-DNA by negative superhelical stress without strand cleavage brought the left-handed conformation into the realm of biology9. The ease of Z-formation varied with sequence—d(CG)n flips better than d(TG)n and d(GGGC)n with less torsion than d(TA)n—reflecting the energetic cost of pushing one base of each pair into syn. The major barrier to the initiation of Z-formation was the additional energy required to create two B−Z junctions9,10. Once nucleated, the transition from B- to Z-DNA is cooperative. Z-DNA formation could be driven by processive enzymes, such as polymerases and helicases, that leave underwound DNA in their wake.
Structural studies of the Zα domain of ADAR confirmed that it is specific to the Z-conformation without any base-specific contacts. This domain has a helix-turn-helix motif similar to that found in B-DNA binding proteins and a binding site of 6 basepairs6,11. Cocrystallization of the Cyprinid herpes virus 3 ORF11 Zα with d(T(CG)9) DNA identified a guanosine base-specific contact outside the core-binding site that may help stabilize the B−Z transformation12. Further structural studies, enabled by Zα, detailed B−Z and Z−Z junctions at atomic resolution. The B−Z junction extruded a basepair from the helix where the phosphate backbone reverses direction13 (Fig. 1) and rendered each base susceptible to damage by mutagens or to modification by enzymes14. The Z−Z DNA junction, where out-of-phase Z-helices meet, was found unstacked and open to intercalation15.
Biophysical studies have demonstrated that Zα does not induce the Z-DNA conformation but rather is recruited after formation16. Dissociation from Z-DNA is slow (measured in hours), a feature that most likely enabled the initial purification of this domain2. Further studies revealed that the Zα domain stabilizes Z-DNA formed by G:T mismatches, by triplet d(GAC)4 repeats with A:A mismatches17, and in sequences with as many as three consecutive d(CC) dinucleotide steps18. Zα binds to Z-RNA in a manner very similar to its interaction with Z-DNA but with differences in solvation due to the 2′-OH group of RNA19. Unexpectedly, formation of a Zα-complex with double-stranded nucleic acids is most rapid with DNA−RNA hybrid duplexes, reflecting the lower energetic cost of junction formation, not a higher affinity of Zα for this structure20. The winged-helix-turn-helix Zα motif was found in 182 other proteins (SMART Domain SM00550) representing orthologs of ADAR1, ZBP1, PKZ, E3L and ORF112 from different species. Structural studies have confirmed that many of these bind Z-DNA, including a number of viral proteins, that, like ADAR, play a role in the innate immune response and are essential for viral infectivity12,21,22,23,24,25,26. The related Zβ domain of human ZBP1 also binds to Z-DNA but uses a different set of contact residues22, suggesting that there are even more divergent members of the winged-helix-turn-helix Z-DNA binding family. Zα also binds the parallel strand G-quadruplex formed by the MYC promoter. The Zα residues engaging the G-quadruplex were shown by NMR to differ from those contacting the Z-DNA surface27. In a separate study, a left-handed quadruplex with a syn-anti dinucleotide step was described28. Another motif with a Z-like dinucleotide syn-anti conformation, called a Z-turn, was found in RNA junctions, ribosome−protein interactions29, the CUG splicing protein30 and the IFIT5 RNA protein complex31. So far, none of the interactions have been base-specific.
ADAR and dsRNA
ADAR was first identified as an enzyme that deaminated adenosines in regions of dsRNA to produce inosine32,33. The edits made by ADAR in codons alter protein sequence since inosine is translated as guanosine, while those modifying splice sites and untranslated regions (UTR) change the isoform mix and transcript stability. Editing of miRNA dsRNA precursors also occurs and alters expression of miRNAs and the genes that they regulate34. Altering an A:C mismatch to an I:C basepair favors dsRNA formation, while editing of an A:U basepair to I:U is destabilizing34.
ADAR is expressed as two isoforms, p150 and p110. Both have a Zβ domain of unknown function, three dsRNA binding domains, and the catalytic domain35. Only the p150 isoform has the Zα domain. The p150 isoform is induced by interferon and is present predominantly in the cytoplasm, a localization modulated by nucleoytoplasmic shuffling36. p110 is constitutively produced and nuclear37. ADAR p110 regulates 3′ UTR stability in stress responses38, affecting translation efficiency34. The ADAR catalytic domain is fully functional without the attachment to the Zα, Zβ or dsRNA binding domains and edits hairpins with stems as short as 15 basepairs39. Like Zα, it also binds DNA/RNA hybrids, producing edits in both the RNA and DNA strands, a process that can lead to somatic mutation of elements within the genome and one with potential application in base-modifying therapeutics40.
Genetic studies revealed that ADAR deletion was embryonic lethal in mice due in part to a failure in generation of the hematopoietic system26. The phenotype is rescued, without any apparent developmental abnormalities, by deletion of the IFIH1 gene that encodes the MDA5 pattern sensor for long dsRNA. MDA5 acts through the MAVS protein to initiate transcription of interferon-stimulated genes (ISGs), including p150 and type 1 interferons. Interferon amplifies the response by further increasing its own production41. Editing creates clusters of I:U basepairs that inhibit further MDA5 activation42 and destabilizes ISG transcripts, which undergo Staufen-mediated decay34. Critically, negative regulation of the MDA5 pathway in mice43, along with other interferon-induced responses44, is completely dependent on the ADAR p150 isoform. These findings place p150 and Zα at the eye of the interferon storm, which, left unchecked, exacerbates inflammatory diseases.
dsRNA leading to induction of interferon and p150 can arise during viral infection. Pox viral homologs of Zα such as E3L inhibit this response and are essential for viral virulence45. A major source of dsRNA is from endogenous retroelements, which constitute more than half the human genome46. These include SINEs and LINEs that are primate-specific and differ from those in mice47. Of these, Alu SINE sequences represent about 10% of the genome. Most Alu’s are dimeric, about 280 bases long, with a characteristic fold that has a left and right arm (Fig. 2a, b). Different Alu clades harbor distinct mutations that reflect the history and age of origin. The distribution of Alu’s within the genome is highly correlated with the density of genes48.
The properties of Alu repeats. a Alu repeats consist of a right and left arm derived originally from the 7SL RNA present in the signal recognition protein (SRP). Transcription is driven by the A and B-boxes of the left arm that are promoters for RNA polymerase III. Alu retrotransposition requires binding of the SRP9/14 heterodimer, using sites on both arms (purple box in upper panel). The site on the left arm overlaps the A-Box. b Each Alu arm folds independently with SRP9/14 binding sites as visualized with VARNA. c The left-hand SRP9/14 sequence is constrained by the interaction of the A-Box with Pol III, with very little variation apparent in the WebLogo motif for the Alu family RepeatMasker consensus sequences. The right-hand site shows more variation and has a consensus logo favoring a 6 bairpair Z-forming alternating C−G motif that can be extended to a full turn of the Z-helix by flipping the out-of-alternation cytosine residue highlighted with a purple dot. The alternating syn-anti (SA) of the Z-conformation is annotated
Alu’s are heavily edited by ADAR, commonly in regions where there are inverted repeats separated only by a few thousand bases49,50,51. Transcripts from the inverted repeats fold back on each other and pair to create long dsRNA editing substrates52 (Fig. 3, RADAR database53). Failure to clear endogenous Alu dsRNA by editing activates MDA5 and the type 1 interferon system, causing the Aicardi-Goutières Syndrome54. This disease is associated with mutations in ADAR p150. A recurrent causal proline to alanine mutation, P193A, lies in the Zα domain of ADAR and diminishes binding of ADAR to the Z-conformation. In vitro cell expression systems confirm that this mutation also reduces dsRNA editing55, confirming an essential role for Z-DNA in this disease.
Mapping of Z-formation to editing sites. a The CTSSgene region chr1:150703475-150704757 (hg19) was analyzed for Z-DNA formation using the ZHUNT3 algorithm that generates the probability of Z-formation based on statistical mechanical calculations10. Z-sites map to a 5′ inverted AluSx repeat and a 3′ AluJo57. b The dsRNA foldback structure with editing sites indicated by red arrows contains an alternating syn-anti (SA) Z-forming segment (box with dashed lines). A dot marks the one cytosine out of alternation. The Z-site lies adjacent to the consensus ADAR binding site found by CLIP-seq (box with dotted lines)59. Editing sites extend 150 basepairs on either side of it. c Map of IRF3 with editing site and H3K12Ac status. d Nonsynonymous edit of exon2 of IRF3 changes readout of codon 45 from Glutamate (D45) to Glycine, the residue in loop 1 that hydrogen bonds to R345 of ATF2(PDB structure 1T2K). e A proposed minimal editing substrate. The Z-stem (box with dashes), formed by Z1 and Z2 sequences, incorporates the splice donor site (sd, bases circled in red) from exon 2. The edited A (red arrow) lies in the left arm of a 20 base long helix of similar structure to other ncRNA substrates
Z-forming sequences are contained in Alu elements (Table 1, Fig. 2c). An example is from the cathepsin-S (CTSS) gene (Fig. 3a), which encodes a protease associated with vascular inflammation, atherosclerotic plaque rupture, and aneurysm. Deletion of the CTSS gene protects against vascular disease in mice but disrupts normal repair processes by reducing angiogenesis56. The CTSS gene contains an AluSx1 element 515 bases away from an AluJo element, which is in the reverse orientation. When the CTSS gene is transcribed, the Alu repeats fold back on each other and basepair to create a long dsRNA editing substrate for ADAR that contains a Z-RNA forming region adjacent to the consensus ADAR binding site (Fig. 3b). The single-stranded short uridine repeat sequences unmasked by editing are bound by HuR, increasing the stability and the amount of CTSS message57. CTSS mRNA editing is induced by interferon, which is produced during vascular inflammation, implicating binding of the Z-RNA forming region by ADAR p150 and its Zα domain in the pathogenesis of atherosclerosis.
Z-DNA forming sequences in genes like CTSS can be predicted computationally with the ZHUNT3 program58. A score of 500 corresponds to a single turn of a Z-helix composed of d(AC)6 that adopts Z-DNA experimentally under reasonable levels of negative superhelical stress10,58. Higher scores imply a better Z-forming sequence and a higher likelihood of capture by Zα. In CTSS and many other RNAs, the Z-DNA forming elements lie within the two inverted Alu repeats (Fig. 3a). They align to form Z-RNA when the transcript folds back on itself. Editing can occur 150 bases either side of the Z-RNA region (Fig. 3b). The consensus binding site for ADAR, derived from whole-cell CHIP-seq analysis (underlined in Fig. 3b), lies adjacent to the Z-forming segment59 (Fig. 3b). The CTSS gene is just one example where structural motifs like dsRNA and Z-formation combine with sequence-specific binding proteins, like HuR, to regulate transcript levels.
Binding of Zα to Z-DNA permits localization of ADAR to transcribed Alu elements. Transfer from Z-DNA to Z-RNA can occur once the dsRNA substrate forms. The slow dissociation of Zα from Z-DNA increases time for mRNA folding by providing a temporary barrier to the passage of the next RNA polymerase through the region60 and provides time to complete the editing process. The association of Zα with Z-RNA further depends on the nature of the helicase(s) involved in straightening out local RNA kinks that compete with the formation of an extended dsRNA editing substrate. Processive helicases will promote Z-RNA formation by generating underwound dsRNA in their wash, more so with long substrates when the ends are fixed and unable to rotate freely.
Alu left and right arms
It is natural to ask why Alu and why Z-DNA? Alu sequences are present mostly in GC-rich regions of the genome and consist of many families initially derived from 7SL RNA, a noncoding RNA found in the signal recognition peptide (SRP). SRP has an S-domain that binds near the peptide exit tunnel of the ribosome and an Alu domain that can stall translation61. Retrotransposons derived from the Alu domain arose first as monomers, then as dimers. Some of them were more invasive than others (Table 1). Transposition requires the Alu elements to hijack the Line L1 copy and paste machinery. This feat is performed by the left arm of the Alu dimer61,62,63. Crystal structures reveal that the left arm inhibits ribosomal translation by filling the gap between the two ribosomal subunits, near the tRNA A-site where translation elongation factors attach61,62,63. Like the SRP protein, the fit is mediated through the SRP9/14 protein pair.
The SRP9/14 binding site in the left Alu arm overlaps with the RNA Polymerase III A-box promoter, constraining sequence variation (Fig. 2a). The right arm sequence, lacking such restrictions, shows more variation64. It has a potential Z-forming (CG)4 core that can be extended to form a 12 bp Z-helix by flipping the an out-of-alternation cytosine (Fig. 2c). The Z-DNA forming motif is maintained in the different Alu family consensus sequences despite a high mutation rate in this region (Table 1)65,66, hinting that there is selection for this motif in active Alu elements . Mutations that lower transposition also lower SRP9/14 binding64 and are expected to lower Z-formation. The right Alu arm, but not the left arm, increases translation of newly transcribed mRNA67; it strips SRP9/14 proteins from the preinitiation complex, preventing the reuse of existing templates68,69. Mutations of the right arm that diminish SRP9/14 binding also diminish effects on translation initiation. A dimeric Alu is thus able to promote translation of recently synthesized L1 mRNA and then capture of the L1 transposase. The site of binding of SRP9/14 to the small ribosome is unknown, but the highly conserved 18S RNA sequence tgcacgcgcgc in helix 30 (H-30) is similar to the Alu Z-forming motif of the right arm. H-30 is solvent-exposed and contacts the anti-codon loop in the ribosomal P-site both during initiation and elongation70,71. H-30 is bound by uS9 (Rsp16) protein, which extends through the 40S core to contact the scaffold protein RACK1 that binds many regulators of translation initiation72. These include eIF3d, a cap binding protein that promotes translation of an mRNA subset when the general factor eIF4E is inactivated by stress or nutrient deprivation73. H-30 is thus strategically placed to choreograph the reinitiation complex. The H-30 sequence is also predicted computationally to form Z-DNA. Experimental studies of E. coli ribosome showing Zα domain cross-links to H-30 (at base 1227). The binding site is close to that for uS974. Whether Zα captures H-30 in a Z-conformation is an open question. The interaction of SRP9/14 with the preinitiation complex would likely be sufficient to force Alu’s to mirror sequences like H-30 to guarantee efficient retrotransposition. The sequences need not be Z-forming in the ribosome, yet an Alu sequence similar to H-30, like that in the right arm, will flip to Z under physiological conditions when transcribed.
Alu and Z-element evolution
An overlap between SRP9/14 binding sites and Z-forming sequences is a potential weakness in the Alu retrotransposition strategy, one exploitable by ADAR to protect the host during periods of Alu retroelement invasion. Those Alu sequences with high affinity for SRP9/14 become targets for Zα and substrates for ADAR editing when they induce formation of dsRNA substrates. Persistence of the Z-DNA binding domain in ADAR1 diminishes the presence of active Alu elements. Within the primate lineage, Alu elements and ADAR evolved in tandem, one dependent upon the other. As Alu sequences mutate, they lose their ability to form Z-DNA, bind SRP9/14, regulate translation, and finally, their power to transpose, leading to their silencing as invasive agents. As seen in Table 1, the taming of Alu transposition has been very effective and left few exact matches for the proposed Z-forming consensus sequence in the Alu right arm. A trace of this history does still remain (Table 1). The fraction of each Alu family that is associated with editing varies. Of those edited, the majority form Z-DNA very poorly (Z-score < 250), suggesting that they are preferred substrates for ADAR p110 rather than p150. For those Alu’s with a Z-score > 250, the total number with edits increases with the mean Z-score for each family, the Z-score histogram of the best (AluSx) family being right-shifted compared to the worst (AluSx1) (Fig. 4a, Supplementary Data 1). The Pearson correlation, weighted by count, between editing ratio (Z-score > 250/Z-score < 250) and mean Z-score is 0.69. The Z-scores observed are consistent with transient Z-formation under physiological conditions, enabling a pause and scan mechanism where transcription is halted long enough for dsRNA editing substrates to form.
Edited genes are enriched for higher Z-scores. a Histogram of Z-scores less than 10,000 for AluSc annotated sequences from hg19 is right-shifted compared to AluSx1 sequences. b Histogram of the length of Z-elements with scores >10,000, showing enrichment of longer segments in edited genes in both exons (5′ UTR, Coding Sequences and 3′ UTR) and introns compared to nonedited genes
While Alu’s provide a challenge to the host, their spread generates genetic diversity and empowers natural selection47. Variability is further expanded by editing and other mutational mechanisms. Sequence constraints are lessened when transcription is driven by RNA Pol II rather than RNA Pol III promoters. Many examples exist where Alu sequences have been coopted to create new forms of gene regulation and novel combinations of features75. Added to these outcomes is the spread of Z-DNA elements that assume roles in transcription76, chromatin remodeling77,78 and in recombination14. When Alu elements cluster, new regulatory mechanisms evolve, such as those proposed for alternative splicing when 5′ and 3′ spice sites are brought together by pairing of Alu’s on either side of an exon. Splicing then excludes the exon from the transcript, yielding an RNA circle with no free ends. Circles are very stable and act as sponges to bind noncoding RNAs and proteins, making them unavailable for regulation of gene expression79. In contrast, editing by ADAR of the Alu stems allows splicing of trapped exons into other transcripts. Editing of isoforms can be selective. For example, the SRP9 gene produces two RNA isoforms. Nonsynonymous edits are only present in isoform 2, which cannot form an Alu SRP because it lacks the residues in helix 2 to bind SRP14. SRP9 has Alu Z-elements that can time when edits occur. Z-dependent editing will be greatest after p150 is induced by interferon. The switch to isoform 2, most likely Z-dependent, will prevent Alu transposition events by downregulating SRP9.
Alternative splicing and Z-formation is enriched in genes with Alu repeats and dsRNA editing of transcripts (Supplementary Data 1, 2, 3 and below). One example is the KRAB-domain family of transcriptional repressors that expanded in response to endogenous retroviruses80. Editing generates additional diversity to protect against novel invaders faster than protein evolution. It is of direct selective benefit for the host.
Not all edits are associated with Alu repeats. Certainly, editing can be performed independently of either the dsRNA or Z-domains39. An example of a minimal Z-dependent editing domain where the substrate binding is enhanced by the Zα domain with no involvement of dsRNA binding domains may be provided by exon 2 of IRF3, where a nonsynonymous edit of codon 45 (Aspartic Acid → Glycine) alters a loop 1 contact of IRF3 with ATF2, the consequences of which are unknown (Fig. 3c). The foldback dsRNA has a potential 10 bp Z-RNA stem that also incorporates the exon 2 splice donor site. The alternative IRF3 (IRF3a) transcript is initiated downstream from this element. The protein product lacks key residues for DNA binding and so acts as a negative modulator of IRF3 dimers, enabling fine-tuning of the interferon response81. A similar clustering of Z-stems overlapping splice sites in 5′ exons is present in IRF7 and EIF5A genes. Binding of p150 to EIF5A RNA is supported by RIP-seq studies82.
Long Z-elements
During evolution, some Z-forming segments have expanded, particularly d(G−T)n:d(C−A)n repeats. Those with Z-scores greater than 10,000 are enriched in edited genes, both in exons and introns (Fig. 4 and Supplementary Data 2). Edited transcripts with genes with high Z-scores also show enrichment for alternative spicing. Around 70% of edited 3′ UTRs from genes with Z-scores greater than 10,000 are annotated as alternatively spliced compared with 60% of edited 3′ UTRs from genes with Z-scores less than 10,000. The frequencies for 5′ UTRs of edited transcripts are 66% for genes with Z-scores greater than 10,000 and 60% for genes with Z-scores less than 10,000.
d(G−T)n repeats on the sense strand are favored by a margin of 2:1, raising the question of whether dsRNA foldback structures with wobble G:U basepairs are also bound by Zα. These long Z-regions, like Alu elements, likely have evolved many different functions beyond localization of epigenetic and DNA repair machinery77,78. One role may be to stall RNA polymerases and block the read-through transcription induced by viruses such as HSV that disrupt normal termination signals83. Stalling may also provide time for splice sites far apart to be transcribed and paired before the next polymerase enters the region. Stalling at Z-elements may optimize transcription of genes with overlapping reading frames, ensuing that each can be readout without interference from the other. Long Z-elements may block the use of an upstream transcription start site or a downstream termination signal, favoring a subset of transcripts. An example is provided by the interferon IFNAR2 receptor gene where a long Z-element is associated with editing of intron 2. The Z-element placement favors transcription downstream from the start site used for the full-length receptor. The isoform produced encodes the soluble form of the receptor, one that modulates IFNβ stability and fine-tunes local interferon responses by signaling in trans through the interferon IFNAR1 receptor on adjacent cells84,85. Other genes with long Z-elements include genes in the RIGI/MDAS pathway with a role in regulation of the innate immune response (Reactome Pathway R-HSA-168928—adjusted p value = 0.02, Supplementary Data 3). Long-Z genes (containing segments with Z-scores > 10,000) are enriched for disease mutations (UP_KEYWORDS Disease Mutations, p value = 1.49 × 10−8), including those related to amino acid and vitamin metabolism, cancer, hypoxia, TGFβ, FGF and EGF signaling along with viral response pathways (Supplementary Data 3). These genes provide an experimental opportunity to map the variations in Z-element scores and locations to genetically defined disease phenotypes.
From RNA to protein immunity
Not all Alumers (fragments from a single Alu element) form dsRNA editing substrates (Table 1). Additional counter measures to destroy Alumers include miRNAs produced by the DICER1 containing microRNA processor complex65. ADAR and DICER1 form a heterodimer via a protein:protein interaction and are thereby targeted to the same transcripts86. Overall, ADAR increases the efficiency of miRNA production. During stress, editing of pri-miRNA substrates by p150 increases the production of mature miRNA by DICER159 (Fig. 5), restoring the RNA-based suppression of ISG protein production. By turning on RNA-world controls87, p150 switches off protein-based immunity.
The Alu cycle and disease. The Alu cycle of retrotransposition involves the LINE1 retrotransposase (L1 ORF2), the SRP9/14 dimer and Alu dimers. Some genomic insertions result in formation of Alu inverted repeats. Transcripts from these regions fold to form dsRNA. ADAR in partnership with DICER1 regulates protein-mediated immune responses to Alu transcripts through a series of RNA-based switches. In the resting state, repression of interferon-stimulated genes (ISG) by the RISC complex is enhanced through protein−protein interactions between ADAR and DICER1. ADAR also reduces PKR (protein kinase, dsRNA activated (EIF2AK2)) stress-related responses by editing dsRNA. ADAR p150 and PKR expression is stimulated by interferon. Decreased ADAR activity or increased production of dsRNA promotes translation of other ISG, leading to amplification of interferon responses through the dsRNA sensor MDA5 (encoded by IFIH1) that promotes the assembly of MAVS filaments on the mitochondrial (mt) surface. Loss of ADAR function is causal for Aicardi-Goutières Syndrome, a disease where the persistent activation of the interferon system is driven by endogenous dsRNA formed in part from Alu inverted repeats. Loss of DICER1 function in age-related macular degeneration leads to accumulation of Alumers, loss of mt integrity, release of mt nucleic acids, inflammasome activation and cell death
DICER1 is associated with different diseases than ADAR. Loss of DICER1 function produces ARMD88. AMRD is characterized by a high level of cytoplasmic Alumers that induce the release of DNA from mitochondria89. The Alumers may sponge up SRP9/14, causing mis-targeting of proteins to mitochondria, leading to a loss of mitochondrial integrity and leakage of their contents into the cytoplasm90. Mitochondrial nucleic acids are sensed by cGAS (MB21D1) and by ZBP1 in epithelium91. Both proteins signal cooperatively through the STING-TBK1-IRF3 pathway to induce interferon-beta release and NLRP3 inflammasome activation, initiating the FAS/FASL-dependent cell death of retinal pigmented epithelium seen in ARMD89,92,93. Activation of ZBP1 could also be by Z-RNA as the best Z-element, gcgcgtacacac is from helix 28 of the mitochondrially encoded 12S RNA.
ADAR and DICER1 may jointly induce other pathologies. For example, GAC triplet repeats can form hairpins with noncanonical basepairs, either by DNA slippage or RNA foldback. Binding of Zα to these structures would also localize DICER1. The small RNAs made by DICER1 from hairpin substrates would induce a dominant negative disease phenotype by interfering with transcripts from both normal and mutant copies of the gene94. The outcome does not depend on the editing by ADAR. Targeting of other enzymatic machinery to Z-elements marking actively transcribed domains can generate many different outcomes.
Roles for Zα independent of DNA
There are conditions where Zα may only target RNA. Stress granules form when dsRNA activates the kinase PKR95 and are more prone to form in ADAR-deficient cells due to the accumulation of dsRNA44. They are stabilized by dsRNA tangles caused by trans-RNA interactions between repetitive elements such as Alu96. Such tangles, trapped by topology, are capable of Z-formation97. Indeed, many Zα domain proteins localize to stress granules98. ADAR may not only prevent stress granule formation, but also may help resolve them by editing and destabilizing tangles.
The known unknowns
The structure and function of Z-DNA has and will continue to generate many surprises. A world based on structural motifs enables fine-tuning of many phenotypes, with outcomes fashioned both by the length and positioning of shape elements within a genome. The regulated expression of proteins like ADAR p150 scales innate immune responses adaptively by changing the way transcripts are produced and processed. Challenges to further understanding Z-biology remain, both small and big. One task is to identify Zα family members with more divergent sequences to facilitate discovery of other Z-scaled outcomes. One question is whether Z-binding proteins exist that are sequence-specific. They may not. Evolution by shape can proceed faster using nucleotide variation to time when and where a structure forms. There is no need to tailor the proteins that bind it. The process can be accelerated by exploiting retroelements to spread shape motifs throughout a genome. Another task is to structurally confirm and define the interactions between Zα and noncanonical conformations like single-stranded Z-turns, (GU)n dsRNA stems, triplet repeats and quadruplex forming sequences. Many sequences can form more than one shape. Competition for each alternative by structure-specific proteins will enable different outcomes. Also of great importance are functional studies to examine how variations in positioning and length of Z-forming segments alter responsiveness to environmental perturbations. Existing in vitro and in vivo assays of immune responses and tumor metabolism are suitable for such purposes. When combined, these approaches will provide insight into how genomes encode information by both shape and sequence to generate selectable phenotypic plasticity99. Conformations like Z-DNA are of special interest as they dynamically modify the readout of sequence information from the genome. By altering the location and timing of RNA processing events, they enable a range of rapid responses to environmental stress.
Conclusion
This review captures roles for the Z-conformation in RNA- and protein-based immunity and describes parts played by the Zα domain in RNA-mediated diseases such as Aicardi-Goutières Syndrome and ARMD. A related theme focuses on the importance of Z-formation to the defense of primate genomes against the hordes of Alu invaders. During the many skirmishes, both RNA editing and RNA interference became weaponized with Z-formation likely supplying precision targeting coordinates, enabling the enzymes to curtail further attacks. The system evolved to regulate protein-centric innate immune responses against more sophisticated invaders, like viruses, which counter-attacked by perfecting their own Zα proteins. Another theme explores the many different ways in which Z-formation alters the readout of genomic information over relatively short time-periods. The change is not as fast as observed with post-translational modifications, such as phosphorylation or ubiquitination. The time scale is better suited to feedback mechanisms where alternative transcripts from a single gene calibrate responses by encoding contrary outcomes. This mode of genetic regulation is likely to vary between individuals and segregate with differences in disease risk. Long Z-DNA containing genes enriched for disease-causing mutations are one place to look for such associations.
References
Herbert, A. et al. A Z-DNA binding domain present in the human editing enzyme, double-stranded RNA adenosine deaminase. Proc. Natl Acad. Sci. USA 94, 8421–8426 (1997).
Herbert, A. G., Spitzner, J. R., Lowenhaupt, K. & Rich, A. Z-DNA binding protein from chicken blood nuclei. Proc. Natl Acad. Sci. USA 90, 3339–3342 (1993).
Kim, U., Wang, Y., Sanford, T., Zeng, Y. & Nishikura, K. Molecular cloning of cDNA for double-stranded RNA adenosine deaminase, a candidate enzyme for nuclear RNA editing. Proc. Natl Acad. Sci. USA 91, 11457–11461 (1994).
Patterson, J. B., Thomis, D. C., Hans, S. L. & Samuel, C. E. Mechanism of interferon action: double-stranded RNA-specific adenosine deaminase from human cells is inducible by alpha and gamma interferons. Virology 210, 508–511 (1995).
Schade, M. et al. A 6 bp Z-DNA hairpin binds two Z alpha domains from the human RNA editing enzyme ADAR1. FEBS Lett. 458, 27–31 (1999).
Schwartz, T., Rould, M. A., Lowenhaupt, K., Herbert, A. & Rich, A. Crystal structure of the Zalpha domain of the human editing enzyme ADAR1 bound to left-handed Z-DNA. Science 284, 1841–1845 (1999).
Pohl, F. M. & Jovin, T. M. Salt-induced co-operative conformational change of a synthetic DNA: equilibrium and kinetic studies with poly (dG-dC). J. Mol. Biol. 67, 375–396 (1972).
Wang, A. H. et al. Molecular structure of a left-handed double helical DNA fragment at atomic resolution. Nature 282, 680–686 (1979).
Peck, L. J. & Wang, J. C. Energetics of B-to-Z transition in DNA. Proc. Natl Acad. Sci. USA 80, 6206–6210 (1983).
Ho, P. S., Ellison, M. J., Quigley, G. J. & Rich, A. A computer aided thermodynamic approach for predicting the formation of Z-DNA in naturally occurring sequences. EMBO J. 5, 2737–2744 (1986).
Schade, M. et al. The solution structure of the Zalpha domain of the human RNA editing enzyme ADAR1 reveals a prepositioned binding surface for Z-DNA. Proc. Natl Acad. Sci. USA 96, 12465–12470 (1999).
Kus, K. et al. The structure of the cyprinid herpesvirus 3 ORF112-Zalpha.Z-DNA complex reveals a mechanism of nucleic acids recognition conserved with E3L, a Poxvirus inhibitor of interferon response. J. Biol. Chem. 290, 30713–30725 (2015).
Ha, S. C., Lowenhaupt, K., Rich, A., Kim, Y. G. & Kim, K. K. Crystal structure of a junction between B-DNA and Z-DNA reveals two extruded bases. Nature 437, 1183–1186 (2005).
Vasquez, K. M. & Wang, G. The yin and yang of repair mechanisms in DNA structure-induced genetic instability. Mutat. Res. 743–744, 118–131 (2013).
de Rosa, M. et al. Crystal structure of a junction between two Z-DNA helices. Proc. Natl Acad. Sci. USA 107, 9088–9092 (2010).
Bae, S., Kim, D., Kim, K. K., Kim, Y. G. & Hohng, S. Intrinsic Z-DNA is stabilized by the conformational selection mechanism of Z-DNA-binding proteins. J. Am. Chem. Soc. 133, 668–671 (2011).
Kolimi, N., Ajjugal, Y. & Rathinavelan, T. A B-Z junction induced by an A… A mismatch in GAC repeats in the gene for cartilage oligomeric matrix protein promotes binding with the hZalphaADAR1 protein. J. Biol. Chem. 292, 18732–18746 (2017).
Bothe, J. R., Lowenhaupt, K. & Al-Hashimi, H. M. Incorporation of CC steps into Z-DNA: interplay between B-Z junction and Z-DNA helical formation. Biochemistry 51, 6871–6879 (2012).
Placido, D. et al. A left-handed RNA double helix bound by the Z alpha domain of the RNA-editing enzyme ADAR1. Structure 15, 395–404 (2007).
Bae, S. et al. Energetics of Z-DNA binding protein-mediated helicity reversals in DNA, RNA, and DNA-RNA duplexes. J. Phys. Chem. B 117, 13866–13871 (2013).
Schwartz, T., Behlke, J., Lowenhaupt, K., Heinemann, U. & Rich, A. Structure of the DLM-1-Z-DNA complex reveals a conserved family of Z-DNA-binding proteins. Nat. Struct. Biol. 8, 761–765 (2001).
Ha, S. C. et al. The crystal structure of the second Z-DNA binding domain of human DAI (ZBP1) in complex with Z-DNA reveals an unusual binding mode to Z-DNA. Proc. Natl Acad. Sci. USA 105, 20671–20676 (2008).
de Rosa, M., Zacarias, S. & Athanasiadis, A. Structural basis for Z-DNA binding and stabilization by the zebrafish Z-DNA dependent protein kinase PKZ. Nucleic Acids Res. 41, 9924–9933 (2013).
Subramani, V. K., Kim, D., Yun, K. & Kim, K. K. Structural and functional studies of a large winged Z-DNA-binding domain of Danio rerio protein kinase PKZ. FEBS Lett. 590, 2275–2285 (2016).
Ha, S. C. et al. A poxvirus protein forms a complex with left-handed Z-DNA: crystal structure of a Yatapoxvirus Zalpha bound to DNA. Proc. Natl Acad. Sci. USA 101, 14367–14372 (2004).
Hartner, J. C. et al. Liver disintegration in the mouse embryo caused by deficiency in the RNA-editing enzyme ADAR1. J. Biol. Chem. 279, 4894–4902 (2004).
Kang, H. J. et al. Novel interaction of the Z-DNA binding domain of human ADAR1 with the oncogenic c-Myc promoter G-quadruplex. J. Mol. Biol. 426, 2594–2604 (2014).
Chung, W. J. et al. Structure of a left-handed DNA G-quadruplex. Proc. Natl Acad. Sci. USA 112, 2729–2733 (2015).
D’Ascenzo, L., Vicens, Q. & Auffinger, P. Identification of receptors for UNCG and GNRA Z-turns and their occurrence in rRNA. Nucleic Acids Res. 46, 7989–7997 (2018).
Teplova, M., Song, J., Gaw, H. Y., Teplov, A. & Patel, D. J. Structural insights into RNA recognition by the alternate-splicing regulator CUG-binding protein 1. Structure 18, 1364–1377 (2010).
Abbas, Y. M., Pichlmair, A., Gorna, M. W., Superti-Furga, G. & Nagar, B. Structural basis for viral 5’-PPP-RNA recognition by human IFIT proteins. Nature 494, 60–64 (2013).
Bass, B. L. RNA editing by adenosine deaminases that act on RNA. Annu. Rev. Biochem. 71, 817–846 (2002).
Nishikura, K. A-to-I editing of coding and non-coding RNAs by ADARs. Nat. Rev. Mol. Cell Biol. 17, 83–96 (2016).
Solomon, O. et al. RNA editing by ADAR1 leads to context-dependent transcriptome-wide changes in RNA secondary structure. Nat. Commun. 8, 1440 (2017).
Athanasiadis, A. et al. The crystal structure of the Zbeta domain of the RNA-editing enzyme ADAR1 reveals distinct conserved surfaces among Z-domains. J. Mol. Biol. 351, 496–507 (2005).
Strehblow, A., Hallegger, M. & Jantsch, M. F. Nucleocytoplasmic distribution of human RNA-editing enzyme ADAR1 is modulated by double-stranded RNA-binding domains, a leucine-rich export signal, and a putative dimerization domain. Mol. Biol. Cell 13, 3822–3835 (2002).
George, C. X., Gan, Z., Liu, Y. & Samuel, C. E. Adenosine deaminases acting on RNA, RNA editing, and interferon action. J. Interferon Cytokine Res. 31, 99–117 (2011).
Sakurai, M. et al. ADAR1 controls apoptosis of stressed cells by inhibiting Staufen1-mediated mRNA decay. Nat. Struct. Mol. Biol. 24, 534–543 (2017).
Herbert, A. & Rich, A. The role of binding domains for dsRNA and Z-DNA in the in vivo editing of minimal substrates by ADAR1. Proc. Natl Acad. Sci. USA 98, 12132–12137 (2001).
Zheng, Y., Lorenzo, C. & Beal, P. A. DNA editing in DNA/RNA hybrids by adenosine deaminases that act on RNA. Nucleic Acids Res. 45, 3369–3377 (2017).
Honda, K., Takaoka, A. & Taniguchi, T. Type I interferon [corrected] gene induction by the interferon regulatory factor family of transcription factors. Immunity 25, 349–360 (2006).
Vitali, P. & Scadden, A. D. Double-stranded RNAs containing multiple IU pairs are sufficient to suppress interferon induction and apoptosis. Nat. Struct. Mol. Biol. 17, 1043–1050 (2010).
Pestal, K. et al. Isoforms of RNA-editing enzyme ADAR1 independently control nucleic acid sensor MDA5-driven autoimmunity and multi-organ development. Immunity 43, 933–944 (2015).
George, C. X., Ramaswami, G., Li, J. B. & Samuel, C. E. Editing of cellular self-RNAs by adenosine deaminase ADAR1 suppresses innate immune stress responses. J. Biol. Chem. 291, 6158–6168 (2016).
Cao, H. et al. Innate immune response of human plasmacytoid dendritic cells to poxvirus infection is subverted by vaccinia E3 via its Z-DNA/RNA binding domain. PLoS ONE 7, e36823 (2012).
de Koning, A. P., Gu, W., Castoe, T. A., Batzer, M. A. & Pollock, D. D. Repetitive elements may comprise over two-thirds of the human genome. PLoS Genet. 7, e1002384 (2011).
Batzer, M. A. & Deininger, P. L. Alu repeats and human genomic diversity. Nat. Rev. Genet. 3, 370–379 (2002).
Grover, D., Mukerji, M., Bhatnagar, P., Kannan, K. & Brahmachari, S. K. Alu repeat analysis in the complete human genome: trends and variations with respect to genomic composition. Bioinformatics 20, 813–817 (2004).
Kim, D. D. et al. Widespread RNA editing of embedded alu elements in the human transcriptome. Genome Res. 14, 1719–1725 (2004).
Athanasiadis, A., Rich, A. & Maas, S. Widespread A-to-I RNA editing of Alu-containing mRNAs in the human transcriptome. PLoS Biol. 2, e391 (2004).
Levanon, E. Y. et al. Systematic identification of abundant A-to-I editing sites in the human transcriptome. Nat. Biotechnol. 22, 1001–1005 (2004).
Kawahara, Y. & Nishikura, K. Extensive adenosine-to-inosine editing detected in Alu repeats of antisense RNAs reveals scarcity of sense-antisense duplex formation. FEBS Lett. 580, 2301–2305 (2006).
Ramaswami, G. & Li, J. B. RADAR: a rigorously annotated database of A-to-I RNA editing. Nucleic Acids Res. 42, D109–D113 (2014).
Ahmad, S. et al. Breaching self-tolerance to Alu duplex RNA underlies MDA5-mediated inflammation. Cell 172, 797–810 (2018).
Mannion, N. M. et al. The RNA-editing enzyme ADAR1 controls innate immune responses to RNA. Cell Rep. 9, 1482–1494 (2014).
Wu, H. et al. Cathepsin S activity controls injury-related vascular repair in mice via the TLR2-mediated p38MAPK and PI3K-Akt/p-HDAC6 signaling pathway. Arterioscler., Thromb., Vasc. Biol. 36, 1549–1557 (2016).
Stellos, K. et al. Adenosine-to-inosine RNA editing controls cathepsin S expression in atherosclerosis by enabling HuR-mediated post-transcriptional regulation. Nat. Med. 22, 1140–1150 (2016).
Champ, P. C., Maurice, S., Vargason, J. M., Camp, T. & Ho, P. S. Distributions of Z-DNA and nuclear factor I in human chromosome 22: a model for coupled transcriptional regulation. Nucleic Acids Res. 32, 6501–6510 (2004).
Bahn, J. H. et al. Genomic analysis of ADAR1 binding and its involvement in multiple RNA processing pathways. Nat. Commun. 6, 6355 (2015).
Ditlevson, J. V. et al. Inhibitory effect of a short Z-DNA forming sequence on transcription elongation by T7 RNA polymerase. Nucleic Acids Res. 36, 3163–3170 (2008).
Voorhees, R. M. & Hegde, R. S. Structures of the scanning and engaged states of the mammalian SRP-ribosome complex. eLife 4, e07975 (2015).
Ahl, V., Keller, H., Schmidt, S. & Weichenrieder, O. Retrotransposition and crystal structure of an Alu RNP in the ribosome-stalling conformation. Mol. Cell 60, 715–727 (2015).
Halic, M. et al. Structure of the signal recognition particle interacting with the elongation-arrested ribosome. Nature 427, 808–814 (2004).
Bennett, E. A. et al. Active Alu retrotransposons in the human genome. Genome Res. 18, 1875–1883 (2008).
Lehnert, S. et al. Evidence for co-evolution between human microRNAs and Alu-repeats. PLoS ONE 4, e4456 (2009).
Price, A. L., Eskin, E. & Pevzner, P. A. Whole-genome analysis of Alu repeat elements reveals complex evolutionary history. Genome Res. 14, 2245–2252 (2004).
Rubin, C. M., Kimura, R. H. & Schmid, C. W. Selective stimulation of translational expression by Alu RNA. Nucleic Acids Res. 30, 3253–3261 (2002).
Berger, A. et al. Direct binding of the Alu binding protein dimer SRP9/14 to 40S ribosomal subunits promotes stress granule formation and is regulated by Alu RNA. Nucleic Acids Res. 42, 11203–11217 (2014).
Ivanova, E., Berger, A., Scherrer, A., Alkalaeva, E. & Strub, K. Alu RNA regulates the cellular pool of active ribosomes by targeted delivery of SRP9/14 to 40S subunits. Nucleic Acids Res. 43, 2874–2887 (2015).
Lomakin, I. B. & Steitz, T. A. The initiation of mammalian protein synthesis and mRNA scanning mechanism. Nature 500, 307–311 (2013).
Leroy, M. et al. Rae1/YacP, a new endoribonuclease involved in ribosome-dependent mRNA decay in Bacillus subtilis. EMBO J. 36, 1167–1181 (2017).
Nielsen, M. H., Flygaard, R. K. & Jenner, L. B. Structural analysis of ribosomal RACK1 and its role in translational control. Cell Signal. 35, 272–281 (2017).
Cate, J. H. Human eIF3: from ‘blobology’ to biological insight. Philos. Trans. Roy. Soc. London, Ser. B Biol. Sci. 372, 20160176 (2017).
Feng, S. et al. Alternate rRNA secondary structures as regulators of translation. Nat. Struct. Mol. Biol. 18, 169–176 (2011).
Chen, L. L. & Yang, L. ALUternative regulation for gene expression. Trends Cell Biol. 27, 480–490 (2017).
Shin, S. I. et al. Z-DNA-forming sites identified by ChIP-Seq are associated with actively transcribed regions in the human genome. DNA Res. 23, 477–486 (2016).
Liu, R. et al. Regulation of CSF1 promoter by the SWI/SNF-like BAF complex. Cell 106, 309–318 (2001).
Maruyama, A., Mimura, J., Harada, N. & Itoh, K. Nrf2 activation is associated with Z-DNA formation in the human HO-1 promoter. Nucleic Acids Res. 41, 5223–5234 (2013).
Ebert, M. S. & Sharp, P. A. Roles for microRNAs in conferring robustness to biological processes. Cell 149, 515–524 (2012).
Lukic, S., Nicolas, J. C. & Levine, A. J. The diversity of zinc-finger genes on human chromosome 19 provides an evolutionary mechanism for defense against inherited endogenous retroviruses. Cell Death Differ. 21, 381–387 (2014).
Karpova, A. Y., Ronco, L. V. & Howley, P. M. Functional characterization of interferon regulatory factor 3a (IRF-3a), an alternative splice isoform of IRF-3. Mol. Cell Biol. 21, 4169–4176 (2001).
Galipon, J., Ishii, R., Suzuki, Y., Tomita, M. & Ui-Tei, K. Differential binding of three major human ADAR isoforms to coding and long non-coding transcripts. Genes 8, 68 (2017).
Rutkowski, A. J. et al. Widespread disruption of host transcription termination in HSV-1 infection. Nat. Commun. 6, 7126 (2015).
McKenna, S. D. et al. Formation of human IFN-beta complex with the soluble type I interferon receptor IFNAR-2 leads to enhanced IFN stability, pharmacokinetics, and antitumor activity in xenografted SCID mice. J. Interferon Cytokine Res. 24, 119–129 (2004).
Samarajiwa, S. A. et al. Soluble IFN receptor potentiates in vivo type I IFN signaling and exacerbates TLR4-mediated septic shock. J. Immunol. 192, 4425–4435 (2014).
Ota, H. et al. ADAR1 forms a complex with Dicer to promote microRNA processing and RNA-induced gene silencing. Cell 153, 575–589 (2013).
Maillard, P. V. et al. Antiviral RNA interference in mammalian cells. Science 342, 235–238 (2013).
Tarallo, V. et al. DICER1 loss and Alu RNA induce age-related macular degeneration via the NLRP3 inflammasome and MyD88. Cell 149, 847–859 (2012).
Kerur, N. et al. cGAS drives noncanonical-inflammasome activation in age-related macular degeneration. Nat. Med. 24, 50–61 (2018).
Costa, E. A., Subramanian, K., Nunnari, J. & Weissman, J. S. Defining the physiological role of SRP in protein-targeting efficiency and specificity. Science 359, 689–692 (2018).
Szczesny, B. et al. Mitochondrial DNA damage and subsequent activation of Z-DNA binding protein 1 links oxidative stress to inflammation in epithelial cells. Sci. Rep. 8, 914 (2018).
DeFilippis, V. R., Alvarado, D., Sali, T., Rothenburg, S. & Fruh, K. Human cytomegalovirus induces the interferon response via the DNA sensor ZBP1. J. Virol. 84, 585–598 (2010).
Ma, Z. & Damania, B. The cGAS-STING defense pathway and its counteraction by viruses. Cell Host Microbe 19, 150–158 (2016).
Krol, J. et al. Ribonuclease dicer cleaves triplet repeat hairpins into shorter repeats that silence specific targets. Mol. Cell 25, 575–586 (2007).
McCormick, C. & Khaperskyy, D. A. Translation inhibition and stress granules in the antiviral immune response. Nat. Rev. Immunol. 17, 647–660 (2017).
Van Treeck, B. et al. RNA self-assembly contributes to stress granule formation and defining the stress granule transcriptome. Proc. Natl Acad. Sci. USA 115, 2734–2739 (2018).
Mao, C., Sun, W. & Seeman, N. C. Assembly of Borromean rings from DNA. Nature 386, 137–138 (1997).
Ng, S. K., Weissbach, R., Ronson, G. E. & Scadden, A. D. Proteins that contain a functional Z-DNA-binding domain localize to cytoplasmic stress granules. Nucleic Acids Res. 41, 9786–9799 (2013).
Kelly, S. A., Panhuis, T. M. & Stoehr, A. M. Phenotypic plasticity: molecular mechanisms and adaptive significance. Compr. Physiol. 2, 1417–1439 (2012).
Acknowledgements
The author would like to thank Dr. P. Shing Ho, Ph.D. for providing source code for the ZHUNT3 program to enable the analyses presented.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The author declares no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Herbert, A. Z-DNA and Z-RNA in human disease. Commun Biol 2, 7 (2019). https://doi.org/10.1038/s42003-018-0237-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-018-0237-x
This article is cited by
-
Inverted Alu repeats: friends or foes in the human transcriptome
Experimental & Molecular Medicine (2024)
-
Fn-OMV potentiates ZBP1-mediated PANoptosis triggered by oncolytic HSV-1 to fuel antitumor immunity
Nature Communications (2024)
-
A shorter splicing isoform antagonizes ZBP1 to modulate cell death and inflammatory responses
The EMBO Journal (2024)
-
The human PTGR1 gene expression is controlled by TE-derived Z-DNA forming sequence cooperating with miR-6867-5p
Scientific Reports (2024)
-
Osteogenesis imperfecta type 10 and the cellular scaffolds underlying common immunological diseases
Genes & Immunity (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.