Abstract
Ovarian cancer is the most lethal gynecologic malignancy, mainly due to late-stage diagnosis, frequent recurrences, and eventually therapy resistance. To identify potentially actionable genetic variants, sequencing data of 351 Belgian ovarian cancer patients were retrospectively captured from electronic health records. The cohort included 286 (81%) patients with high-grade serous ovarian cancer, 17 (5%) with low-grade serous ovarian cancer, and 48 (14%) with other histotypes. Firstly, an overview of the prevalence and spectrum of the BRCA1/2 variants highlighted germline variants in 4% (11/250) and somatic variants in 11% (37/348) of patients. Secondly, application of a multi-gene panel in 168 tumors revealed a total of 214 variants in 28 genes beyond BRCA1/2 with a median of 1 (IQR, 1–2) genetic variant per patient. The ten most often altered genes were (in descending order): TP53, BRCA1, PIK3CA, BRCA2, KRAS, ERBB2 (HER2), TERT promotor, RB1, PIK3R1 and PTEN. Of note, the genetic landscape vastly differed between the studied histotypes. Finally, using ESCAT the clinical evidence of utility for every genetic variant was scored. Only BRCA1/2 pathogenic variants were classified as tier-I. Nearly all patients (151/168; 90%) had an ESCAT tier-II variant, most frequently in TP53 (74%), PIK3CA (9%) and KRAS (7%). In conclusion, our findings imply that although only a small proportion of genetic variants currently have direct impact on ovarian cancer treatment decisions, other variants could help to identify novel (personalized) treatment options to address the poor prognosis of ovarian cancer, particularly in rare histotypes.
Similar content being viewed by others
Introduction
Epithelial ovarian cancer (OC) is an umbrella term for distinct malignancies affecting the ovaries, each characterized by its own histological and molecular profile. OC is the most lethal gynecological malignancy and the eighth most prevalent female cancer worldwide, with an annual abysmal mortality rate of two million fatalities worldwide1. The existence of late symptom onset and multiple overlapping symptoms with other gastrointestinal, genitourinary, and gynecological diseases often leads to late-stage diagnosis with already extensive extra-ovarian metastasis present, whereby the 5 year survival plummets to <30%2,3,4,5. Despite their inherent differences, most OCs are generally treated in the same manner: the cornerstones are cytoreductive surgery and platinum-based chemotherapy, with or without Poly (ADP-Ribose) Polymerase (PARP) inhibitors (PARPis) and/or bevacizumab6. Unfortunately, up to 70% of patients will experience recurrence7. Thus, it is well established that a “one-size-fits-all” treatment approach should no longer be applied. Tailored treatment options based on individualized molecular tumor data are warranted to enable practitioners to more effectively treat specific OC subtypes. For instance, EMA and FDA have approved PARPis targeting germline or somatic BRCA1/2 mutated advanced OCs. Given their efficacy, mutational screening of BRCA1/2 genes in OC has become of great importance8,9,10,11. Besides PARPis, targeted therapies are increasingly available for other gene alterations, including KRAS, BRAF and TRK inhibitors12. Therefore, implementation of companion diagnostic tests, which go beyond somatic and germline BRCA1/2 testing is highly warranted.
To help clinicians with interpreting genomic reports, the European Society for Medical Oncology (ESMO), Translational Research and Precision Medicine Working Group published a systemic framework in 2018 to rank molecular targets based on available evidence supporting their value as clinical targets13. Its main goal was to implement a harmonized vocabulary and to facilitate communication between academia, pharmaceutical industry, healthcare professionals and patients. The ESMO scale for Clinical Actionability of Molecular Targets (ESCAT) was previously applied across multiple cancer types, but OC has never been the focus of such studies and histopathologic subtypes were never distinguished14,15,16,17.
In the present retrospective cohort study, we aim to characterize (potentially targetable) genetic variants in patients with OC across various histotypes, using the ESCAT framework.
Results
Patients and samples
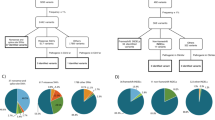
HE slides from 358 OC patients (median age at molecular analysis was 68 year, range 29–90 years) who received surgery between 2014 and 2022 were reviewed by an expert pathologist; seven borderline or non-epithelial tumors were excluded for further analysis (n = 7). From the final cohort of 351 patients, sequencing data were available from 348 tumor samples and 250 blood samples. Patient matched tumor and blood samples were available for 247 patients (Fig. 1a). The most common histotypes were high-grade serous ovarian cancer (HGSOC) (n = 286) and low-grade serous ovarian cancer (LGSOC) (n = 17), with other (rare) histotypes accounting for 48 patients: 15 clear cell carcinoma of the ovaries (CCOC), 14 endometrioid ovarian cancers (EOC), 11 mixed malignant mullerian tumors (MMMT), 6 mucinous ovarian cancers (MOC) and 2 mesonephric-like adenocarcinomas of the ovary (MLAOC) (Fig. 1b).
Genetic variants in BRCA1/2
Among the 351 OC patients analyzed, 49 (14%) patients harbored BRCA1 (9%) and/or BRCA2 (5%) variants. In one HGSOC patient, both a somatic BRCA1 and a germline BRCA2 variant were detected. Eleven variants were of germline origin (six in BRCA1 and five in BRCA2). BRCA1/2 variants were most prevalent in the HGSOC (16%) subgroup; notably, all germline variants were found in this histotype. The remaining somatic BRCA1/2 variants were found in CCOC (13%), MMMT (9%) and EOC (7%) histotypes. No BRCA1/2 variants were found in the LGSOC, MOC, and MLAOC histotypes (Table 1).
To evaluate if the variants cluster in specific regions, all (likely) pathogenic variants are depicted in a lollipop plot, illustrating that the variants were clearly spread over the entire coding sequence of BRCA1 and BRCA2 genes, without particular hotspot regions (Fig. 2). Less than half of the variants were found in the ovarian cluster regions (OCCR): sixteen in the BRCA1 OCCR (13/27 somatic, 3/6 germline), four in BRCA2 OCCR1 and one in BRCA2 OCCR2 (all somatic). Twenty-two (6% of patients) and eleven (3% of patients) variants occurred in a functional domain of BRCA1 and BRCA2, respectively. In BRCA1, this concerns seven variants in the Really Interesting New Gene (RING) domain, twelve in the DNA Binding Domain (DBD) and three in the BRCA1 C-Terminal (BRCT) domain. In BRCA2, four variants clustered in the RAD51-Binding Domain (RAD51-BD) and seven in the DBD. A detailed overview of all germline and somatic BRCA1/2 variants identified in the different histotypes is provided in Supplementary Table 1.
The diagrams linearly represent BRCA1/2 protein domains (x-axis). BRCA1 domains: green, C3HC4-type RING finger; red, DBD (DNA-binding domain of BRCA1); blue, BRCT (BRCA1 C-terminal) domain. BRCA2 domains: orange, RAD51-BD (RAD51-binding domain); red, DBD (DNA-binding domain). Each variant is represented by a single lollipop; the stick heights indicate the mutation frequency (y-axis), and dots are color-coded according to histotype: pink dots, HGSOC; green dots, LGSOC; orange dots, CCOC; blue dots, MMMT; yellow dots, EOC. Germline variants are indicated by an arrow. The deletion of exons 5 & 6 in BRCA1 is marked by a horizontal lollipop under the plot. OCCR (ovarian cancer cluster regions) are indicated by a thick black line under the plot. The graphs are adapted from MutationMapper tool, cBioportal (http://www.cbioportal.org/mutation_mapper).
Genetic variants beyond BRCA1/2
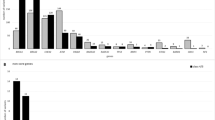
A multi-gene panel was introduced in 2019 for solid cancers and revealed a wide range of genetic variants in 168 OCs. Beyond BRCA1/2, 214 variants across 27 genes were detected in 134 patients (80%) with a median of 1 variant (interquartile range (IQR), 1–2) per patient. The majority of these variants were substitutions (164/214, 77%), small deletions (33/214, 15%), or duplications (9/214, 4%). In a limited number of samples, coverage analysis revealed gene amplifications: PIK3CA (2%), ERBB2 (2%), FGFR2 (1%), FGFR3 (1%), MET (1%), CCND1 (1%). Sixty-six tumors harbored more than one (likely) pathogenic variant in the same gene or multiple genes. Table 2 displays an overview of the most frequently altered genes per histotype. A complete list of all variants found in our cohort is provided in Supplementary Table 2.
Summarized, in HGSOC, the most frequently altered gene was TP53 in 94% of patients. Other affected genes include PIK3CA (4.8%), RB1 (3.2%), KRAS (2.4%), and PIK3R1 (2.4%). Sporadically altered genes (only 1 or 2 tumors) in this subtype are RNF43, FGFR2, ERBB2, MET, AKT1, PTEN, GNAS, FGFR1, FGFR3, SMARCA4, VHL and SPOP. In fifteen CCOC tumors, variants were identified in eleven genes with PIK3CA being mutated in six (40%) and the TERT promotor in four (27%) tumors, respectively. Other genes (TP53, ATM, PIK3R1, PTEN, KRAS, CCND1, CDKN2A, SMARCB1 and POLE) were only sporadically affected in CCOC. Ten out of eleven LGSOC tumors had variants identified in NRAS (27%), ERBB2 (27%) and sporadically in BAP1, KRAS, CDKN2A and BRAF. Six out of seven EOC tumors harbored variants in TP53, AKT1, PIK3CA, PTEN, KRAS, CCND1 and CTNNB1. In three EOC tumors multiple variants were detected (one tumor with six variants, another had 3 variants and a third had two variants). Out of six carcinosarcomas, three had variants in TP53. Both mucinous and mesonephric-like adenocarcinomas tumors harbored the same KRAS p.(Gly12Asp) variant. Additionally, one mucinous tumor harbored a TP53 variant and another had a PIK3CA variant.
Affected signaling pathways
We investigated if specific molecular pathways were involved per histotype (Fig. 3). The p53 signaling pathway was most frequently altered regardless of the histotype (127/168). Almost all HGSOC tumors (117/125) had genetic variants in the TP53 signaling pathway. Additionally, the HR (21/125), followed by PI3K/AKT/MTOR (11/125), RTK/RAS/MAPK (9/125), cell cycle (4/154), ERBB2 (2/125), and WNT (2/125) pathways were altered in this subtype. In contrast, nearly half of CCOC tumors (7/15) displayed an aberrant P13/AKT/MTOR pathway, while TP53 (2/15), HR (2/15), RTK/RAS/MAPK (2/15), and cell cycle (2/15) were less frequently altered. In LGSOC tumors alterations in the RTK/RAS/MAPK (4/11) and ERBB2 (3/11) pathways were most prevalent. The HR and cell cycle pathways were each affected in only one LGSOC tumors each. Four out of seven EOC tumors displayed an aberrant PI3K/AKT/MTOR signaling pathway, besides the TP53 (3/7), WNT (2/7), and cell cycle (1/7) pathways. Four out of six adenocarcinomas had an altered TP53 pathway, and in one a pathogenic variant in the HR pathway was detected. The RTK/RAS/MAPK pathway was affected in both mucinous tumors that underwent multi-gene panel testing, in addition to the PI3K/AKT/MTOR pathway or TP53 pathway. Both mesonephric-like adenocarcinomas were altered in the RTK/RAS/MAPK pathway but no other pathways were affected. Although there are some trends, no significantly enriched pathways between the subgroups could be obtained, as the number of some histotypes is rather small.
Drug-target matches were annotated based on their clinical actionability according to ESCAT and patients were assigned to the highest level of actionability. The oncoplot shows a summary of the genes affected (rows) in individual patients (columns). Genes are grouped by pathways and patients are grouped according to histotype.
Classification of genetic variants using ESCAT
To determine the clinical utility of the variants identified by the multi-gene cancer panel, ESCAT scores were assigned for all genetic variants detected in 168 patients who underwent multi-gene panel testing. A comprehensive list of all clinical evidence leading to ESCAT ranking is described in Supplementary Table 3. In 156/168 (93%) OC tumors, at least one actionable alteration was detected. Noteworthy, only 17% (28/168) were classified as ESCAT tier-I (BRCA1 and BRCA2). Nearly all patients (151/168) had an ESCAT tier-II classified variant, most frequently in TP53 (127/168), followed by PIK3CA (15/168) and KRAS (11/168). Rare tier-II variants, affecting <3% of the OC tumors include: ATM, AKT1, PIK3R1, CDKN2A, PTEN, ERBB2, NRAS, BRAF, SMARCA4, and SMARCB1. All other variants were scored as ESCAT tier-IIIA, tier-IIIB, tier-IV or tier-X. The oncoplot in Fig. 3 provides a detailed overview of all genetic variants affecting individual patients across different histotypes. ESCAT-scores correlated with the age of the patient are summarized in a bar chart in Supplementary Fig. 1.
Discussion
This study unraveled the genomic landscape from a unique cohort of 351 Belgian OC patients across different histotypes using real-world data. Rapid evolvement of next-generation sequencing currently allows the medical community to produce a myriad of genetic information; the clinical utility, however, is not always apparent. Here, we demonstrated that a cancer gene panel offers a focused approach allowing the medical community to capture the most relevant genetic variants and that scoring their clinical utility is feasible.
The first objective of this study was to give an overview of the BRCA1/2 mutational status in epithelial OC. In total, eleven (4%) patients were found to have a germline, and 39 (11%) had a somatic (likely) pathogenic BRCA1/2 variant. Compared to other reports from European populations (15–20%), the frequency of BRCA1/2 variants in our cohort (14%) is at the lower limit11,18,19. As expected, the majority (94%) of BRCA1/2 variants were detected in HGSOC tumors. Only two out of fifteen CCOC tumors, one out of fourteen EOC and one out of eleven MMMT had a BRCA1/2 variant. Due to relatively small numbers of rare histotypes in our study and in literature, it is hard to compare our findings with those of others. Overall, our prevalence of BRCA1/2 variants appears to be somewhat lower, except for the CCOC histotype. Similar to other reports, no BRCA1/2 variants were found in MOC tumors20. Another difference is the higher occurrence of somatic versus germline variants in our cohort. A recent review reported that 13%–21% of epithelial OC patients have germline BRCA1/2 variants and only 6% harbor somatic BRCA1/2 variants18. Historically, BRCA1/2 testing was biased towards patients with a family history of breast/ovarian cancer and focused on patients with early-onset disease, which may explain higher mutation detection rates in other studies11. However, we cannot exclude a bias in referral for our cohort, as tumors from patients with known germline variants may not have been referred for tumor testing, especially before 2019.
As the majority of the BRCA1/2 variants are of somatic origin, the mutation spectrum is more diverse than previously described for germline variants in the Belgian OC population. In 2004, Claes et al. reported six major recurrent BRCA1/2 variants accounting for 60% of all identified variants in a Belgian cohort. Only two of these recurrent germline variants (BRCA2 c.6275_6276del & c.8904del) were found in two patients of our cohort21.
About 50% of all somatic BRCA1/2 variants, 50% of BRCA1 and 25% of BRCA2 germline variants were detected in the OCCRs, displaying only a small enrichment of variants in the OC cluster regions. Although, the RING-domain is associated with a lower OC risk compared to breast cancer, still one germline and six somatic BRCA1 variants were detected in this region. An exploratory analysis of the PAOLA-1 trial suggested that the location of mutation in BRCA1/2 could influence the magnitude of benefit from platinum salts and/or olaparib (plus bevacizumab): patients with mutations in the DBD region of BRCA2 might be extremely sensitive to platinum, whereas those with mutations in the DBD region of BRCA1 might be less sensitive to platinum but extremely sensitive to olaparib plus bevacizumab. Currently, the biological mechanisms underlying domain-related sensitivity to PARPi are unknown. However, if this intriguing finding could be confirmed in independent studies, treatments may not only be guided by the mutated gene but also by the location of the variant22.
The second objective of this study was to evaluate the prevalence of additional actionable genetic variants in epithelial OC beyond BRCA1/2. In 157/168 OCs that underwent multi-gene panel testing, 242 (likely) pathogenic alterations were detected. These variants were ranked according to ESCAT into six levels of evidence ranging from ‘ready for routine use’ (tier-I) to ‘lack of evidence’ (tier-X). Tier-I variants (BRCA1 and BRCA2) were found in nearly 17% of the OC tumors. Several randomized clinical trials have demonstrated improved oncological outcomes with PARPi maintenance treatment over standard of care in newly diagnosed OC patients with BRCA1/2 genetic variants, leading to EMA and FDA approvals8,10,22. OC patients harboring tier-IIB variants can be enrolled in several ongoing clinical studies that test the efficacy of a drug matched to genetic variants, without data being available on overall or disease-free survival. Genes in this category include: TP53, ATM, AKT1, PIK3CA, PIK3R1, PTEN, ERBB2, KRAS, NRAS and BRAF. One notable example is the BOUQUET trial (NCT04931342) that evaluates the efficacy and safety of biomarker-driven therapy in patients with platinum-resistant OC of a rare (non-HGSOC) histotype. In BOUQUET, PIK3CA, AKT1, PTEN, ERBB2, KRAS, NRAS and BRAF, are being used as biomarkers to plan individualized study treatment23,24. Other large ongoing studies on the effectiveness of targeted anti-cancer drugs and immunotherapy, in patients where the tumor is known to have specific genetic variants are NCI-MATCH (NCT02465060), DRUP (NCT02925234) and TAPUR (NCT02693535)25,26,27,28. Concerning tier-IIIA, OC tumors with FGFR1/2/3 pathogenic variants could be treated with FGFR-inhibitors. These are classified as tier-IA in other tumor types (like urothelial carcinoma and biliary tract cancer14) and the ongoing NCI-MATCH trial is investigating, amongst others, the efficacy of FGFRi in additional tumor types29. However, to date no supportive clinical data is yet available in OC. ERBB2 amplified tumors are matched to anti-HER2 antibody, trastuzumab, treatment and are classified as tier I-A in breast cancer and are currently investigated in multiple clinical trials in OC patients (e.g. DESTINY: phase 2. NCT04482309). MET inhibitors have shown significant benefit for treatment in MET amplified lung cancers (tier-II), however, data for OC is currently still lacking. BAP1 was categorized as tier-IIIB, as this is a protein that interacts with BRCA1/2 to mediate homologous recombination during DNA repair. Testing PARPis in patients with a deficient BAP1 protein would be conceptually reasonable. Several clinical trials targeting BAP1 alterations in multiple cancer types (not OC) are ongoing (e.g. NCT03207347, NCT04666740, NCT03654833, NCT02925234). Two groups investigated whether silencing of Cyclin D1 (CCND1) could lead to a BRCAm-like phenotype and thus improve the efficacy of PARPis. Micro-RNA and short hairpin-RNA were used in vitro and in vivo in mice to downregulate CCND1 resulting in a substantial benefit in OC management (tier-IVA)30,31. Other genes like, POLE, RB1, TERT, GNAS, CTNNB1, RNF43 and SPOP have prognostic values or are frequently altered/upregulated in certain histologic subtypes, but are currently not considered as targets matched to specific treatments (tier-X)32,33,34,35,36,37,38.
To our knowledge, the present study is the most comprehensive study classifying alterations in a large epithelial OC cohort using ESCAT. Previously, Lapke et al. reported actionable variants in a smaller group of OCs (n = 85). In their study, eligibility for targeted therapy was determined by a pathway-based approach; however, no classification according to degrees of clinical evidence was applied39. ESCAT classification has been applied in other cancer types, including head and neck cancer, breast cancer and tumor-agnostic studies; these reported that implementation of ESCAT for clinical-decision making is feasible and beneficial for personalized treatment14,15,16,17,40,41,42,43,44. ESCAT scoring in our cohort only identified a limited number of patients (17%) with tier-I genetic variants; however, tier-II actionable variants were detected in 90% of the patients. This implies that only a small proportion of reported genetic variants have a direct impact on treatment decisions. Nevertheless, many patients are potentially eligible to specific clinical trials tailored to their cancer’s genome and could benefit from drugs targeting a tier-II variant. We hypothesize that one reason why OC is lagging behind other cancer types towards targeted treatments is conventional trial design. Trials in which all patients with a specific cancer type receive the same treatment limit the possible benefits of precision oncology. From our data it is clear that rare histologic subtypes have distinct molecular features that are possibly actionable. Since the most common histotype is HGSOC, it is often the only subtype included in clinical trials. Recently, clinical research has evolved from using cancer type-centered to biomarker-directed and histology-specific trials. These ‘master protocol’ designs are tailored to molecular profiles of the patients.
To improve the concept of personalized treatment, the strategy should be implemented earlier in the course of the disease and tumors should have comprehensive tumor profiling at each stage of the disease45,46. Another major challenge for oncologists remains how to integrate patients’ biomarker profiles into therapeutic decision-making as literature is complex, rapidly evolving, and voluminous. Next, local drug access is also critical for clinicians seeking therapeutic options for their patients. To aid clinicians, several knowledge bases have been developed. Recently, a curated version specific to the Australian healthcare setting was developed: TOPOGRAPH—Therapy-oriented precision oncology guidelines for recommendation of anticancer pharmaceuticals47. But utility studies on such an electronic compendium with respect to treatment recommendations are warranted.
Strengths of our study include a relatively large sample size that covered a large spectrum of histotypes in OC, the participation of a single expert center with uniform management procedures and NGS testing, including variant interpretation, a small proportion of patients excluded for analysis, full access to all relevant pathology specimens for rigorous review, and the extensive literature search for assessing strength of evidence using ESCAT. However, several limitations should be considered when interpreting the results of this study. First, the retrospective and anonymized design precluded the collection of additional data or samples types. Therefore, we could neither assess patients’ survival time, nor determine whether or not patients received matched drugs to genetic variants. Second, some concerns may be raised that genomic instability evaluating the consequence of HRD beyond BRCAm (e.g., loss of heterozygosity, telomeric allelic imbalance and/or large-scale state transitions) was not assessed in our study. Although our panel did contain two genes involved in HRD beyond BRCA1/2 (i.e., BAP1 and ATM), it has been demonstrated that mutations in non-BRCA homologous recombination repair genes (HRRm) did not predict benefit from olaparib (a PARPi) plus bevacizumab in PAOLA-1; hence, non-BRCA HRRm in multigene panels do not substitute for HRD determined by genomic instability testing (and BRCA mutation status)48. Third, clear deletions or amplifications are picked up by coverage analysis but smaller exon-spanning deletions or duplications, gene fusions, mutations in regulatory sequences or deep-intronic mutations could not be detected by the applied molecular analysis. Bearing this in mind, some highly druggable targets are missed (e.g. NTRK fusions)49. Similarly (fourth), our methods did not yet assess microsatellite instability and tumor mutational burden (too small panel size around 100 kb), although both have been shown to be predictive biomarkers of immunotherapy in several solid tumors (albeit with limited evidence in OC). Fifth, the ESCAT is a validated and well-respected instrument; however, tumor heterogeneity (i.e., spatial differences) and evolutionary dynamics of the cancer genome (i.e., temporal differences) may hinder the more extensive use of molecular profiling-based strategies. Sixth, as our cohort comprised mainly Caucasian women, representative of the Flemish population, validations in other ethnicities are warranted. Therefore, in a similar prospective study design most of these limitations will disappear. Other limitations like MSI status and tools for detecting exon-spanning deletions/amplifications should be tested and validated. Spatial differences and temporal differences could be addressed using cfDNA mutational analysis at different time points during a patients’ treatment course, although, this will cost more and implementing this into standard clinical practice may take some time.
In conclusion, we unveiled a diverse landscape of potentially actionable genetic variants in this histologically and genetically well-defined cohort of OC patients. Unfortunately, directly actionable (ESCAT tier-I) variants accounted only for a small proportion. Multiple biomarker-driven clinical trials recruiting OC patients are currently ongoing; these trials generate information that could ultimately lead to the dawn of precision oncology in OC patients.
Methods
Patients and samples
This retrospective cohort study included 358 patients diagnosed with OC between January 2014 and December 2022 in 28 hospitals across East-Flandres and West-Flandres. These patients underwent molecular testing of blood and/or tumor samples at the Center for Medical Genetics Ghent (Ghent University Hospital, Belgium). The study was approved by the ethics committee of Ghent University Hospital (ONZ-2023-0389); the need for informed consent of patients was waived because of the retrospective nature of the study and anonymized data was transferred to the investigators. The study was conducted in accordance with the Declaration of Helsinki and followed the ‘Strengthening the Reporting of Observational Studies in Epidemiology’ (STROBE) guideline. Histopathologic information and molecular sequencing data were extracted from the institutional electronic health records. A gynecological pathologist (KV) reviewed all available tumor slides (using both HE and immunohistochemistry) in a blind manner to classify them into distinct ovarian cancer histotypes: high-grade serous ovarian cancer (HGSOC), low-grade serous ovarian cancer (LGSOC), clear cell carcinoma of the ovaries (CCOC), endometrioid ovarian cancer (EOC), mucinous ovarian cancer (MOC), mixed malignant mullerian tumor (MMMT) and mesonephric-like adenocarcinoma of the ovary (MLAOC). All borderline or non-epithelial tumors were excluded for further analysis.
Molecular profiling
Tumor samples obtained before January 2019 were only examined for somatic genetic variants in BRCA1 and BRCA2 using the PCR-based BRCA Tumor Mastr Plus Dx kit (Multiplicom). In parallel, germline testing was performed for BRCA1, BRCA2, TP53 and PALB2 using an in-house developed amplicon-based target enrichment method, followed by next generation sequencing (Miseq, Illumina) as previously described50. Starting from 2019, a capture-based approach was applied for target enrichment. A detailed description of the methods can be found in Supplementary Methods 1. Both tumor and blood samples were examined using a customized cancer gene panel that has been updated over time (all panel versions can be found in Supplementary Tables 4–6). Only variants that were classified as pathogenic or likely pathogenic were considered for further analysis.
Functional domains BRCA1 and BRCA2
Functional domains were defined as:
BRCA1: RING (Really Interesting New Gene) domain: amino acids (AA) 8–96, DBD (DNA-Binding Domain): AA 452–1092, BRCT (BRCA1 C-Terminal) domain: AA 1646-1736 and AA 1760-1855.
BRCA2: RAD51-BD (RAD51-Binding Domain): AA 900-2000, DBD: AA 2459-3190.
The Ovarian Cancer Cluster Regions (OCCRs) were defined for BRCA1: AA 460-1354; and for BRCA2: OCCR1 - AA 1083-1894 and OCCR2- AA 2215-249020,51.
Level of actionability
Genetic variants were ranked according to the ESMO Scale for Clinical Actionability of molecular Targets (ESCAT) into six levels of evidence ranging from ‘ready for routine use’ (tier-I) to ‘lack of evidence’ (tier-X), previously described by Mateo et al.13. To rank the specific variants, the results of clinical trials (ClinicalTrials.gov) were summarized and classified based on several characteristics including prospectiveness, randomization, and availability of results. When no clinical trials in OC were available, basket trials or trials in other cancer types were examined. All listed drug-target matches were investigated for FDA or EMA approvals and for treatment recommendations by NCCN. OncoKB and publications which already describe ESCAT scoring systems in other cancer types were used to propose a score in case of doubt about drug-target matches13,14,15,16,17,52,53,54. The proposed ranking was validated by the co-authoring medical oncologists, pathologists and clinical laboratory geneticists until consensus. Patients with multiple genetic variants were ranked by the highest level of evidence. A comprehensive report of all genetic variants and matched drugs accompanied with clinical evidence and ESCAT ranking is included in Supplementary Table 313.
Signaling pathways
For the most frequently mutated genes the canonical pathways in which they are playing a role, were determined. Five signaling pathways were previously curated by TCGA PanCancer Atlas54: (1) cell cycle pathway (CCND1, CDKN2A, RB1), (2) RTK/RAS/MAPK pathway (KRAS, NRAS, BRAF, FGFR1, FGFR2, FGFR3, MET), (3) p53 pathway (TP53), (4) PI3K/AKT/mTOR pathway (AKT1, PIK3CA, PIK3R1, PTEN), (5) WNT pathway (CTNNB1, RNF43). Two additional pathways included the HR pathway (BRCA1, BRCA2, BAP1, ATM) and ErbB kinases (ERBB2). Seven additional genes with at least one genetic variant could not be assigned to any of these seven pathways (SMARCA4, SMARCB1, GNAS, POLE, SPOP, TERT, VHL).
Statistical analysis
All analyses and visualizations were performed using Excel version 2307 and R Studio version R4.3.1 (R program for Statistical Computing). To summarize all actionable variants for the individual patients, an oncoprint was generated in R Studio using the Complex Heatmap package. Patients were grouped according to histotype; genes were grouped in their respective pathway. The lollipop plot for BRCA1 and BRCA2 variants was adapted from cBioportal (genome build GRCh38).
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
All data analyzed in this study are included in the published article and its supplementary information files. Raw data are available from the corresponding author. Qualified researchers can apply for access to these datasets via a collaboration or data usage agreement. The data are not publicly available due to privacy and ethical restrictions.
Code availability
Relevant R script codes are available from the corresponding author upon request.
References
Sung, H. et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185x countries. CA Cancer J. Clin. 71, 209–249 (2021).
Siegel, R. L., Miller, K. D., Fuchs, H. E. & Jemal, A. Cancer statistics, 2021. CA Cancer J. Clin. 71, 7–33 (2021).
Sambasivan, S. Epithelial ovarian cancer: review article. Cancer Treat. Res. Commun. 33, 100629 (2022).
Chen, V. W. et al. Pathology and classification of ovarian tumors. Cancer 97, 2631–2642 (2003).
Reid, B. M., Permuth, J. B. & Sellers, T. A. Epidemiology of ovarian cancer: a review. Cancer Biol. Med. 14, 9–32 (2017).
Alvarez Secord, A., O’Malley, D. M., Sood, A. K., Westin, S. N. & Liu, J. F. Rationale for combination PARP inhibitor and antiangiogenic treatment in advanced epithelial ovarian cancer: a review. Gynecol. Oncol. 162, 482–495 (2021).
Lorusso, D. et al. Newly diagnosed ovarian cancer: which first-line treatment? Cancer Treat. Rev. 91, 102111 (2020).
Kyo, S. et al. Clinical landscape of PARP inhibitors in ovarian cancer: molecular mechanisms and clues to overcome resistance. Cancers 14, 2504 (2022).
Fujiwara, K. et al. Olaparib plus bevacizumab as maintenance therapy in patients with newly diagnosed, advanced ovarian cancer: Japan subset from the PAOLA-1/ENGOT-ov25 trial. J. Gynecol. Oncol. 32, e82 (2021).
DiSilvestro, P. et al. Overall survival with maintenance olaparib at a 7-year follow-up in patients with newly diagnosed advanced ovarian cancer and a BRCA mutation: the SOLO1/GOG 3004 trial. J. Clin. Oncol. https://doi.org/10.1200/JCO.22.01549 (2022).
Huepenbecker, S. P. et al. Temporal patterns and adoption of germline and somatic BRCA testing in ovarian cancer. Obstet. Gynecol. 140, 758–767 (2022).
ElNaggar, A. et al. Genomic profiling in low grade serous ovarian cancer: identification of novel markers for disease diagnosis and therapy. Gynecol. Oncol. 167, 306–313 (2022).
Mateo, J. et al. A framework to rank genomic alterations as targets for cancer precision medicine: the ESMO scale for clinical actionability of molecular targets (ESCAT). Ann. Oncol. 29, 1895–1902 (2018).
Martin-Romano, P. et al. Implementing the European society for medical oncology scale for clinical actionability of molecular targets in a comprehensive profiling program: impact on precision medicine oncology. JCO Precis. Oncol. 6, e2100484 (2022)
Mosele, F. et al. Recommendations for the use of next-generation sequencing (NGS) for patients with metastatic cancers: a report from the ESMO precision medicine working group. Ann. Oncol. 31, 1491–1505 (2020).
Marret, G. et al. Genomic alterations in head and neck squamous cell carcinoma: level of evidence according to ESMO scale for clinical actionability of molecular targets (ESCAT). JCO Precis. Oncol. 5, 215–226 (2021).
Condorelli, R. et al. Genomic alterations in breast cancer: level of evidence for actionability according to ESMO scale for clinical actionability of molecular targets (ESCAT). Ann. Oncol. 30, 365–373 (2019).
Vergote, I. et al. European experts consensus: BRCA/homologous recombination deficiency testing in first-line ovarian cancer. Ann. Oncol. 33, 276–287 (2022).
Janavičius, R. Founder BRCA1/2 mutations in the Europe: implications for hereditary breast-ovarian cancer prevention and control. EPMA J. 1, 397–412 (2010).
Sekine, M., Nishino, K. & Enomoto, T. Differences in ovarian and other cancers risks by population and BRCA mutation location. Genes 12, 1050 (2021).
Claes, K., Poppe, B., Coene, I., De Paepe, A. & Messiaen, L. BRCA1 and BRCA2 germline mutation spectrum and frequencies in Belgian breast/ovarian cancer families. Br. J. Cancer 90, 1244 (2004).
Labidi-Galy, S. I. et al. Association of location of BRCA1 and BRCA2 mutations with benefit from olaparib and bevacizumab maintenance in high-grade ovarian cancer: phase III PAOLA-1/ENGOT-ov25 trial subgroup exploratory analysis. Ann. Oncol. 34, 152–162 (2023).
Target Ovarian Cancer. BOUQUET: A Trial Looking At The Effects Of Different Drugs In Platinum-Resistant Ovarian Cancer With Various Genetic Changes. https://targetovariancancer.org.uk/about-ovarian-cancer/clinical-trials/bouquet-trial-looking-effects-different-drugs-platinum
Sava, J. Phase 3 Confirmatory Study of Avutometinib/Defactinib in LGSOC Finalizes Design. https://www.targetedonc.com/view/phase-3-confirmatory-study-of-avutometinib-defactinib-in-lgsoc-finalizes-design
O’Dwyer, P. J. et al. The NCI-MATCH trial: lessons for precision oncology. Nat. Med. 29, 1349–1357 (2023).
Chae, Y. K. et al. Phase II study of AZD4547 in patients with tumors harboring aberrations in the FGFR pathway: results from the NCI-MATCH Trial (EAY131) subprotocol W. J. Clin. Oncol. 38, 2407–2417 (2020).
Hoes, L. R. et al. Patients with rare cancers in the drug rediscovery protocol (DRUP) benefit from genomics-guided treatment. Clin. Cancer Res. 28, 1402–1411 (2022).
Mangat, P. K. et al. Rationale and design of the targeted agent and profiling utilization registry (TAPUR) study. JCO Precis. Oncol. 2018, 1–14 (2018).
Chae, Y. K. et al. Molecular analysis for therapy choice (MATCH) arm W: phase II study of AZD4547 in patients with tumors with aberrations in the FGFR pathway. J. Clin. Oncol. 36, 2503–2503 (2018).
Hao, X. et al. MicroRNA-195 suppresses cell proliferation, migration and invasion in epithelial ovarian carcinoma via inhibition of the CDC42/CCND1 pathway. Int J. Mol. Med. 46, 1862–1872 (2020).
Zhong, Q. et al. Cyclin D1 silencing impairs DNA double strand break repair, sensitizes BRCA1 wildtype ovarian cancer cells to olaparib. Gynecol. Oncol. 152, 157–165 (2019).
Davila, J. I. et al. Frequent POLE-driven hypermutation in ovarian endometrioid cancer revealed by mutational signatures in RNA sequencing. BMC Med. Genomic 14, 1–6 (2021).
Xie, B. et al. RB1 is an immune-related prognostic biomarker for ovarian cancer. Front. Oncol. 12, 830908 (2022).
Wu, R. C. et al. Frequent somatic mutations of the telomerase reverse transcriptase promoter in ovarian clear cell carcinoma but not in other major types of gynecologic malignancies. J. Pathol. 232, 473 (2014).
Tominaga, E. I. et al. Amplification of GNAS may be an independent, qualitative, and reproducible biomarker to predict progression-free survival in epithelial ovarian cancer. Gynecol. Oncol. 118, 160–166 (2010).
Zyla, R. E. et al. CTNNB1 mutations and aberrant β-catenin expression in ovarian endometrioid carcinoma: correlation with patient outcome. Am. J. Surg. Pathol. 45, 68–76 (2021).
Ryland, G. L. et al. RNF43 is a tumour suppressor gene mutated in mucinous tumours of the ovary. J. Pathol. 229, 469–476 (2013).
Li, Y. et al. SPOP Regulates the biological mechanism of ovarian cancer cells through the Hh signaling pathway. Onco Targets Ther. 12, 9239 (2019).
Lapke, N. et al. Genetic alterations and their therapeutic implications in epithelial ovarian cancer. BMC Cancer 21, 499 (2021).
Wolff, L. & Kiesewetter, B. Applicability of ESMO-MCBS and ESCAT for molecular tumor boards. Memo. Mag. Eur. Med. Oncol. 15, 190–195 (2022).
Verdaguer, H. et al. ESMO scale for clinical actionability of molecular targets driving targeted treatment in patients with cholangiocarcinoma. Clin. Cancer Res. 28, 1662–1671 (2022).
Moreira, A. et al. Efficacy of molecularly targeted agents given in the randomised trial SHIVA01 according to the ESMO scale for clinical actionability of molecular targets. Eur. J. Cancer 121, 202–209 (2019).
Mulet Margalef, N. et al. Genomically matched therapy in refractory colorectal cancer according to ESMO scale for clinical actionability of molecular targets: experience of a comprehensive cancer centre network. Mol. Oncol. https://doi.org/10.1002/1878-0261.13444. (2023)
Reissig, T. M. et al. Smaller panel, similar results: genomic profiling and molecularly informed therapy in pancreatic cancer. ESMO Open 8, 101539 (2023).
Fountzilas, E., Tsimberidou, A. M., Vo, H. H. & Kurzrock, R. Clinical trial design in the era of precision medicine. Genome Med. 14, 1–27 (2022).
Park, J. J. H. et al. Systematic review of basket trials, umbrella trials, and platform trials: a landscape analysis of master protocols. Trials 20, 1–10 (2019).
Lin, F. P. et al. Criteria-based curation of a therapy-focused compendium to support treatment recommendations in precision oncology. NPJ Precis. Oncol. 5, 58 (2021).
Pujade-Lauraine, E. et al. Homologous recombination repair gene mutations to predict olaparib plus bevacizumab efficacy in the first-line ovarian cancer PAOLA-1/ENGOT-ov25 trial. JCO Precis. Oncol. 7, e2200258 (2023).
Harbin, L. M., Gallion, H. H., Allison, D. B. & Kolesar, J. M. Next generation sequencing and molecular biomarkers in ovarian cancer—an opportunity for targeted therapy. Diagnostics 12, 842 (2022).
De Leeneer, K. et al. Flexible, scalable, and efficient targeted resequencing on a benchtop sequencer for variant detection in clinical practice. Hum. Mutat. 36, 379–387 (2015).
Rebbeck, T. R. et al. Association of type and location of BRCA1 and BRCA2 mutations with risk of breast and ovarian cancer. JAMA 313, 1347 (2015).
Martin Romano, P. et al. 1930O Genomic alterations in solid tumours according to ESMO scale for clinical actionability of molecular targets (ESCAT). Ann. Oncol. 31, S1092–S1093 (2020).
Chakravarty, D. et al. OncoKB: A precision oncology knowledge base. JCO Precis. Onocol. https://doi.org/10.1200/PO.17.00011. (2017)
Sanchez-Vega, F. et al. Oncogenic signaling pathways in the cancer genome atlas. Cell 173, 321–337.e10 (2018).
Acknowledgements
This study was funded by Research Foundation—Flandres (FWO-1S73622N). The funder played no role in study design, data collection, analysis and interpretation of data, or the writing of this manuscript.
Author information
Authors and Affiliations
Contributions
C.F.: Study concept and design; methodology; interpretation of data; investigation; original draft, writing—editing; visualization; project administration; funding acquisition. J.v.d.M and S.L.: Study concept; molecular analyses; writing—review and editing. K.P.: Statistical support; visualization; writing—review and editing. E.d.J.: critical review of the manuscript; writing and editing. J.V.D. and P.T.: acquisition of patient material; H.D.: Study Concept and design; methodology; writing—review and editing. K.V.: Study concept and design; revision of tumor samples; writing—review and editing. K.C.: Study concept and design; study supervision; acquisition of data; analysis and interpretation of data; critical review of the manuscript; funding acquisition. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
H.D. reports consulting fees (advisory role) paid to her institution by Pfizer, Roche, PharmaMar, AstraZeneca, Eli Lilly, Novartis, Amgen, GSK, MSD, Seagen, Gilead and other fees paid to her institution (travel, accommodations, expenses) by Pfizer, Roche, PharmaMar, Teva, AstraZeneca, MSD, GSK, Gilead and an institutional research grant by Gilead.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fieuws, C., Van der Meulen, J., Proesmans, K. et al. Identification of potentially actionable genetic variants in epithelial ovarian cancer: a retrospective cohort study. npj Precis. Onc. 8, 71 (2024). https://doi.org/10.1038/s41698-024-00565-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41698-024-00565-2