Abstract
Immune checkpoint inhibitors (ICIs) are most commonly used for melanoma and non-small cell lung cancer (NSCLC) patients. FAT atypical cadherin 1 (FAT1), which frequently mutates in melanoma and NSCLC. In this study, we aim to investigate the association of FAT1 mutations with ICI response and outcome. We collected somatic mutation profiles and clinical information from ICI-treated 631 melanoma and 109 NSCLC samples, respectively. For validation, a pan-cancer cohort with 1661 patients in an immunotherapy setting was also used. Melanoma and NSCLC samples from the Cancer Genome Atlas were used to evaluate the potential immunologic mechanisms of FAT1 mutations. In melanoma, patients with FAT1 mutations had a significantly improved survival outcome than those wild-type patients (HR: 0.67, 95% CI: 0.46–0.97, P = 0.033). An elevated ICI response rate also appeared in FAT1-mutated patients (43.2% vs. 29.2%, P = 0.032). Associations of FAT1 mutations with improved prognosis and ICI response were confirmed in NSCLC patients. In the pan-cancer cohort, the association between FAT1 mutations and favorable ICI outcome was further validated (HR: 0.74, 95% CI: 0.58–0.96, P = 0.022). Genomic and immunologic analysis showed that a high mutational burden, increased infiltration of immune-response cells, decreased infiltration of immune-suppressive cells, interferon and cell cycle-related pathways were enriched in patients with FAT1 mutations. Our study revealed that FAT1 mutations were associated with better immunogenicity and ICI efficacy, which may be considered as a biomarker for selecting patients to receive immunotherapy.
Similar content being viewed by others
Introduction
Worldwide, melanoma causes ~56,000 deaths each year1. Because of distinct access to early diagnosis and timely treatment, the morbidity and mortality of melanoma differ widely by country. For a long time, few therapy strategies were used for melanoma clinical practice owing to the unsuccess of relevant clinical trials2. In recent years, the development of genomic sequencing technologies and deep exploration of biological mechanisms have changed melanoma into a novel treatment model. Targeted therapy (e.g., BRAF inhibitors) has been demonstrated to markedly improve patients’ response and clinical outcomes3. Moreover, immune checkpoint inhibitors (ICIs), which have a capacity to prolong survival outcomes of advanced or metastatic patients, have been become the routine clinical treatment pattern for melanoma4,5.
Non-small cell lung cancer (NSCLC) is the main histologic subtype of lung cancer. Several treatment strategies were reported for NSCLC. Metastatic NSCLC patients with EGFR mutations treated with EGFR tyrosine kinase inhibitors (TKIs) have exhibited an improved survival outcome6 and the EGFR-TKIs are standard drugs in first-line treatment7. Besides, the combination of VEGF inhibitors or other chemotherapies with EGFR-TKIs was also considered as a helpful therapeutic path for NSCLC8. The emergence of ICI therapies has dramatically lengthened the survival of NSCLC patients in advanced stage or patients who produced a treatment resistance during conventional chemotherapy9.
Although melanoma and NSCLC patients who received ICI agents have revealed a preferable prognosis; however, the fact is that in clinical practice only a minority of patients could obtain a treatment response to ICIs10. Recently multiple molecular markers were determined to select patients who are responsive or resistant to ICIs, for example, tumor mutation burden (TMB)11,12, neoantigen burden (NB)13, PD-L1 protein expression14, mutations in POLE15, TP5316,17, MUC1618,19, PBRM120, and B2M21.
FAT atypical cadherin 1 (FAT1), which is a well-known tumor suppressor, plays tumor inhibition roles via the regulation of WNT/β-catenin signaling22, Hippo signaling23, and MAPK/ERK-signaling activities24. Loss of function of FAT1 contributed to tumor progression and impacted clinical outcomes. A recent study reported that FAT1 deletion generated a hybrid epithelial to mesenchymal transition status and thus promoted squamous cell carcinoma stemness and metastasis25. Consistent findings were observed in refs. 26,27 studies, that are, FAT1 mutation was associated with an inferior survival outcome in head and neck squamous cell carcinoma (HNSCC). However, a previous research revealed an inverse conclusion that human papillomavirus (HPV)-negative HNSCC patients with FAT1 mutation exhibited a better prognosis28. FAT1 loss was also demonstrated to be linked with the CDk4/6 inhibitor treatment resistance in breast cancer29,30. Besides, FAT1 or its alterations were involved in inflammatory regulation in glioma31 and clinically influenced T-cell lymphoma outcome32, suggesting that FAT1 may be an immune response regulator and participate in distinct inflammatory processes.
In this study, we retrospectively collected melanoma and NSCLC patients treated by ICIs to explore the association between FAT1 mutations and immunotherapy efficacy. Furthermore, we conducted a series of genomic and immunologic analyses to illuminate the potential mechanisms behind FAT1 mutations. Results from our work may provide helpful clues for enrolling cancer patients to receive immune treatments.
Results
FAT1 mutations in melanoma
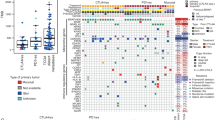
Of the 631 integrated melanoma samples, 193 (30.6%) exhibited the ICI status of complete response (CR) or partial response (PR), 430 (68.1%) were status of stable disease (SD) or progressive disease (PD), and the rest (1.3%) were unavailable. C>T was the main base substitution type in this melanoma cohort (Supplementary Fig. 1). Mutational patterns of FAT1 and the most common significantly mutated genes were shown in Supplementary Fig. 1. We observed that FAT1 is frequently mutated, accounting for 82 of 631 patients (12.9%). Amino acid transformations generated by FAT1 mutations were illustrated using a lollipop plot (Supplementary Fig. 2). Detailed protein and function changes induced by FAT1 mutations for each melanoma patient were shown in Supplementary Data 1. The workflow of this research was shown in Fig. 1.
FAT1 mutations linked with improved ICI efficacy in melanoma
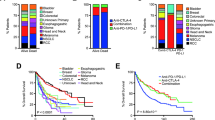
In the pooled melanoma cohort, univariate survival analysis revealed that patients with FAT1 mutations had a significantly improved ICI prognosis than those wild-type patients (median survival time: 41.9 vs. 25.6 months, Log-rank test P = 0.013; Fig. 2a). Multivariate Cox regression model with clinical confounding factors (e.g., age, sex, stage, and therapy type) taken into consideration still demonstrated a significant result (HR: 0.67, 95% CI: 0.46–0.97, P = 0.033; Fig. 2b). Associations between FAT1 mutations and ICI prognosis in individual cohorts and distinct treatment types were, respectively, shown in Supplementary Figs. 3 and 4. Further analysis showed that FAT1 mutated patients also exhibited an elevated ICI response rate (43.2% vs. 29.2%, Fisher exact test P = 0.014; Fig. 2c). And this result was still significant after adjusting for other confounding factors (OR: 0.58, 95% CI: 0.35–0.96, P = 0.032; Fig. 2d).
a Kaplan–Meier survival curves stratified by FAT1 mutational status. b Multivariate Cox regression model between FAT1 mutations and ICI outcome with age, sex, stage, and therapy type taken into account. Black box: hazard ratio; black scale bar, 95% confidence interval of hazard ratio. c Distinct ICI response rates in FAT1 mutated and wild-type subgroups. d Multivariate logistic regression model between FAT1 mutations and ICI response with age, sex, stage, and therapy type taken into account.
Relationship between FAT1 mutations and mutation burden in melanoma
We explored the relationship of FAT1 mutations with TMB owing to its important roles in predicting cancer immune response and clinical outcome. We found that a significantly enhanced TMB was enriched in melanoma patients with FAT1 mutations (Wilcoxon rank-sum test P < 0.001; Fig. 3a). Mutational signatures operative in the genome largely influence the genomic stability and mutation rates. We therefore extracted 4 mutational signatures from melanoma patients with the NMF method; they are signature 1 (age-relevant), signature 4 (smoking-relevant), signature 7 (ultraviolet light exposure-induced), and signature 11 (alkylating agent-induced). Activities of extracted signatures across all patients were illustrated in Supplementary Data 2. To eliminate the possibility that the association of FAT1 mutations with TMB was impacted by other confounders, we combined clinical variables, extracted mutational signatures, and alterations in BRCA1/2, TP53, and POLE into the multivariate logistic regression. Association between FAT1 mutations and higher TMB was still existed (OR: 9.08, 95% CI: 4.05–23.63, P < 0.001; Fig. 3b). Besides, FAT1 mutations were also associated with a higher NB (Wilcoxon rank-sum test P < 0.001; Fig. 3c). Similar results of FAT1 mutations with enhanced TMB and NB were also detected by using the melanoma somatic mutation data in the TCGA cohort (both P < 0.001; Fig. 3d, e).
a Distinct distribution of TMB in FAT1 mutated versus wild-type patients in the integrated melanoma cohort. Blue dot, TMB for wild-type patients; orange dot, TMB for mutated patients. b Multivariate logistic regression model between FAT1 mutations and TMB with clinical confounding factors, extracted mutational signatures, and mutations in DNA repair genes taken into consideration. Black box: odds ratio; black scale bar, 95% confidence interval of odds ratio. c Distinct distribution of NB in FAT1-mutated versus wild-type patients in the integrated melanoma cohort. Distinct distribution of (d) TMB and (e) NB in FAT1-mutated versus wild-type melanoma patients from TCGA cohort.
Validation in NSCLC
Of the 109 ICI-treated NSCLC patients, 36 (33.0%) harbored the status of CR or PR. FAT1 mutated in 8 (7.3%) of 109 patients. Mutational patterns of FAT1 and the most common NSCLC significantly mutated genes were shown in Supplementary Fig. 5. Detailed protein and function changes induced by FAT1 mutations for each NSCLC patient were shown in Supplementary Data 3. Kaplan–Meier survival analysis showed that NSCLC patients with FAT1 mutations had a significantly better survival outcome than wild-type patients (median survival time: 23.0 vs. 6.5 months, Log-rank test P = 0.048; Fig. 4a). Multivariate Cox regression model with confounding factors adjusted still revealed a consistent result, although it did not obtain a statistical significance (HR: 0.48, 95% CI: 0.17–1.40, P = 0.086; Fig. 4b). ICI prognostic capacity of FAT1 mutations in distinct therapy types was illustrated in Supplementary Fig. 6. Furthermore, an elevated proportion of NSCLC patients with CR or PR was also observed in FAT1 mutated subgroup (75.0% vs. 31.9%, Fisher exact test P = 0.022; Fig. 4c). This association was significant after controlling for multiple confounders (OR: 0.14, 95% CI: 0.02–0.76, P = 0.029; Fig. 4d).
a Kaplan–Meier survival curves stratified by FAT1 mutational status. b Multivariate Cox regression model between FAT1 mutations and ICI outcome with age, sex, histology subtype, smoking status, PD-L1 expression, and therapy type taken into account. Black box: hazard ratio; black scale bar, 95% confidence interval of hazard ratio. c Distinct ICI response rates in FAT1 mutated and wild-type subgroups. d Multivariate logistic regression model between FAT1 mutations and ICI response with age, sex, histology subtype, smoking status, PD-L1 expression, and therapy type taken into account.
Higher TMB and NB were observed in NSCLC patients with FAT1 mutations (Wilcoxon rank-sum test P = 0.005 and 0.006, respectively; Fig. 5a, b). Three mutational signatures were extracted based on the mutational profile of NSCLC (Supplementary Data 4). Multivariate logistic analysis was conducted with age, sex, histology subtype, smoking status, PD-L1 expression, mutational signatures, and genome repair gene mutations taken into consideration; and the association of FAT1 mutations with increased TMB was still obtained (OR: 19.49, 95% CI: 2.15–465.40, P = 0.021; Fig. 5c). We also noticed increased TMB and NB in FAT1-mutated subgroup in NSCLC patients from the TCGA (Wilcoxon rank-sum test, both P < 0.001; Fig. 5d, e).
a Distinct distribution of TMB in FAT1 mutated versus wild-type patients in the integrated NSCLC cohort. Blue dot, TMB for wild-type patients; orange dot, TMB for mutated patients. b Multivariate logistic regression model between FAT1 mutations and TMB with clinical confounding factors, extracted mutational signatures, and mutations in DNA repair genes taken into consideration. c Distinct distribution of NB in FAT1 mutated versus wild-type patients in the integrated NSCLC cohort. Black box: odds ratio; black scale bar, 95% confidence interval of odds ratio. Distinct distribution of (d) TMB and (e) NB in FAT1 mutated versus wild-type NSCLC patients from TCGA cohort.
Further corroboration in a pan-cancer ICI cohort
By using a pan-cancer cohort with distinct cancer types from MSKCC, we further investigated the ICI predictive roles of FAT1 mutations. Survival analysis showed that a significantly improved ICI prognosis was observed in FAT1 mutated patients (median survival time: 36 vs. 17 months, Log-rank test P = 0.008; Fig. 6a). And multivariate Cox regression model suggested that FAT1 mutation is an independent prognostic biomarker (HR: 0.74, 95% CI: 0.58–0.96, P = 0.022; Fig. 6b). In this cohort, an enhanced TMB was also noticed in patients with FAT1 mutations (Wilcoxon rank-sum test P < 0.001; Fig. 6c).
a Kaplan–Meier survival curves stratified by FAT1 mutational status. b Multivariate Cox regression model between FAT1 mutations and ICI outcome with clinical confounding factors taken into account. Black box: hazard ratio; black scale bar, 95% confidence interval of hazard ratio. c Association of FAT1 mutations with TMB in the pan-cancer cohort.
Immune infiltration and signaling pathways associated with FAT1 mutations
We conducted multiple immunologic analyses and pathways exploration to illuminate the potential mechanisms behind FAT1 mutations in melanoma. CIBERSORT algorithm revealed that increased infiltration of M1 macrophage and decreased infiltration of T-regulatory cells were enriched in FAT1-mutated subgroup (both P < 0.05; Fig. 7a). Consistently, by using Angelova et al. method, we observed FAT1 mutations were positively associated with pro-inflammatory immunocyte (e.g., activated CD4/CD8 cells and effector memory CD4 cells) infiltration abundance; however, negatively associated with immune-suppressive T-regulatory cell infiltration (all P < 0.05; Fig. 7b).
Distinct immunocyte infiltration abundance in FAT1 mutated versus wild-type subgroups was assessed with a CIBERSORT and b Angelova et al. methods, respectively. Blue box, FAT1 wild-type patients; orange box, FAT1-mutated patients; black scale bar, range of each immunocyte infiltration abundance. Significantly differentially infiltrating immunocytes between two subgroups were highlighted with green. c Heatmap representation of immune-related signatures enrichment in distinct FAT1 mutational subgroups. Significantly differentially enriched signature between two subgroups was highlighted with red. d Distinct proportion of activated-stroma subtype in distinct FAT1 mutational subgroups. e–g Significantly enriched signaling pathways in FAT1-mutated melanoma patients. *P < 0.05, **P < 0.01.
We further composed a heatmap with distinct enrichment of immune-related signatures in FAT1 mutated versus wild-type subgroups. Differential analysis demonstrated that the stromal cell signature exhibited a significantly reduced enrichment in FAT1 mutated group (P < 0.001; Fig. 7c). Consistent with this finding, a lower proportion of patients with activated stroma was also found in this group (36.8% vs. 50.1%, Fisher exact test P < 0.05; Fig. 7d). GSEA analysis showed that immune response-related pathways (e.g., interferon α and γ responses) and cell cycle pathways (e.g., G2M checkpoint) were markedly enriched in melanoma patients with FAT1 mutations (all FDR < 0.001; Fig. 7e–g, and Supplementary Fig. 7).
We also performed immune infiltration and pathway analyses in NSCLC patients. Results indicated that elevated infiltration of M1 macrophage and activated/effector memory CD4 cells, and decreased infiltration of M2 macrophage and T-regulatory cells were observed in FAT1 mutation group (all P < 0.05; Supplementary Fig. 8a, b). Inflammatory and interferon response-related pathways were also noticed in this mutated group; however, the pathway of epithelial–mesenchymal-transition was enriched in FAT1 wild-type subgroup (all FDR < 0.01; Supplementary Fig. 8c).
Discussion
By using melanoma and NSCLC patients treated with ICIs, we retrospectively investigated the immunotherapy predictive roles of FAT1 mutations. In addition, the association of FAT1 mutations with favorable ICI outcome was observed in a pan-cancer cohort. Genomic and multiple immunological explorations further elucidated potential biological mechanisms underlying FAT1 mutations. Findings obtained from this study suggest that FAT1 mutation may be a potential indicator for assessing ICI efficacy.
In this work, we found that FAT1 mutations were associated with an improved ICI prognosis. Further immune infiltration analysis showed an increased infiltrating abundance of CD4/CD8 T cells and a decreased abundance of T-regulatory cells in patients with FAT1 mutations. CD4 and CD8 T cells are two well-known lymphocytes, which play a vital role in cancer immune response33,34. T-regulatory cells, which are a subtype of T cells, have been demonstrated to play an immune-suppressive function in anti-tumor immunity35,36. The dynamic interaction between tumor cells and their surrounding stroma influences the progression, metastasis, and drug resistance of cancer patients37. Recent studies have revealed that stromal cells inhibit the anti-tumor response and sensitivity to immunotherapy38,39. Yoshihara et al. developed the ESTIMATE method by using gene expression data to infer the stromal cell infiltration level in mixed tumor tissue40. Moffitt et al. determined a stroma-specific subtype via the utilization of stroma-related feature genes41. Based on the above two stroma evaluation methods, we observed FAT1-mutated patients had a reduced infiltration abundance of stromal cells and a reduced proportion of activated-stroma subtype. Interferon response pathways, which were enriched in FAT1-mutated subgroup, are positive immune response regulators42,43. Higher TMB and NB have been demonstrated as promising biomarkers for predicting prognosis and immunotherapy response in several cancers11,12,44, although there is still a controversy on TMB in some clinical settings45,46. In the present study, a markedly elevated TMB and MB was observed in FAT1 mutated melanoma and NSCLC patients, which supports the potential ICI predictive roles of FAT1 mutations. Further functional experiments and clinical trials are necessary to validate these findings.
FAT1 is a typical tumor suppressor and its determined mechanisms are involved in the WNT/β-catenin pathway22, Hippo pathway23, and MAPK/ERK pathways24. Mutations in FAT1 always generate an inferior survival outcome in several cancers, such as NSCLC25 and HNSCC26,27. Recently, the immune regulation roles of FAT1 have been reported in multiple studies31,32,47. Dikshit et al. observed that FAT1-mediated glioma inflammation response via regulating the activity of PDCD4 and transcription factor AP-131. In peripheral T-cell lymphoma, FAT1 was frequently mutated and accounted for a significant proportion (39%) of patients, which provided both prognostic and therapeutic implications32. Grandi et al. reported that vaccination with a combination of FAT1-derived B cell epitope with tumor-specific B and T cell epitopes conferred a robust protective role in cancer mouse models47. The above findings further confirm the crucial roles of FAT1 in immune regulation and provide evidence for the present study.
Association of FAT1 mutations with drug sensitivity was recently reported29, that is, FAT1 loss-of-function mutations were associated with a resistance to CDK4/6 inhibitors in estrogen receptor-positive breast cancer. We also evaluated the prognostic roles of FAT1 mutations in melanoma and NSCLC patients treated with distinct chemotherapies from TCGA cohorts; and no significant survival differences were observed between FAT1 mutated and wild-type subgroups in both tumors (Log-rank test, both P > 0.05; Supplementary Fig. 9). In this study, we observed that patients with FAT1 mutations exhibited a favorable prognosis in melanoma, NSCLC, and pan-cancer cohorts with immunotherapy settings. These results indicate that FAT1 mutations play distinct roles in distinct treatment environments and it may be a potential predictive biomarker in the settings of cancer immunotherapy.
We further explored whether FAT1 mutations affect ICI treatment efficacy by regulating FAT1 expression. By using somatic mutation and transcriptomic data from TCGA melanoma and NSCLC cohorts, we analyzed the distinct FAT1 expressions in patients with distinct FAT1 mutational types (Supplementary Figs. 10 and 11). Results showed no significant expression differences were observed in all comparison subgroups (Wilcoxon test, all P > 0.05), which suggests that FAT1 mutation may not influence its own expression levels in melanoma and NSCLC. Further in-depth studies regarding the association of FAT1 mutations with FAT1 expression are warranted.
In this study, we curated eight independent melanoma datasets into a pooled cohort and observed that FAT1 mutations were associated with a favorable ICI response and outcome. Nevertheless, in individual cohorts and distinct ICI treatment types, FAT1 mutations sometimes lacked the predictive ability of immunotherapy efficacy (Supplementary Figs. 3 and 4), the relatively smaller sample size for each cohort may be a possible reason. Therefore, clinically expanded cohorts with data integration are necessary to robustly determine immunotherapy efficacy-related indicators48,49.
Limitations exist in this study. First, the association between FAT1 mutations and ICI efficacy is derived from previously published datasets, which is a retrospective study. Therefore, melanoma and NSCLC cohorts with both somatic mutation data and immunotherapy information in a prospective design are necessary. Second, this integrated study comprises several distinct cohorts, which may produce some biases in data analysis. Moreover, the lack of experimental validation is another limitation.
In summary, by integrating genomic profiles and clinical ICI data, we identified that FAT1 mutations were predictive of ICI response and outcome in melanoma, NSCLC, and pan-cancers. Further in-depth studies are needed, but FAT1 mutations may be a novel selection for enrolling cancer patients to receive immunotherapies.
Methods
Sample collection
A total of 631 melanoma samples with both somatic mutation profiles (generated by whole-exome sequencing [WES]) and clinical information were collected from previous 8 studies48,50,51,52,53,54,55,56. Besides, 109 NSCLC samples were collected from recent two studies13,57. In the above included studies, melanoma and NSCLC patients were treated with anti-PD1/PD-L1, anti-CTLA-4, or combination treatments. The Oncotator was employed to uniformly annotate somatic mutations for all patients58. Non-synonymous mutations were taken into consideration to conduct relevant analyses. Detailed treatment response information, sequencing platforms, and demographic features were shown in Supplementary Data 5 for melanoma and Supplementary Data 6 for NSCLC.
A pan-cancer ICI cohort of 1661 patients with 9 diverse cancer types in the Memorial Sloan Kettering Cancer Center (MSKCC) was also curated for further verification11. These patients underwent a 468-gene targeted sequencing and their clinical information was illustrated in Supplementary Data 7.
From Genome Data Commons (https://gdc.cancer.gov), we downloaded transcriptomic data, somatic mutational data, and clinical features of 457 melanoma and 995 NSCLC samples in the Cancer Genome Atlas (TCGA) cohort. The potential mechanistic analyses were performed based on the transcriptomic data of TCGA. All samples included in this study were acquired from previously published studies and the corresponding Institutional Ethics Committees have approved the studies.
Mutational signature extraction
We used the method proposed by Kim et al. 59 to extract mutational signatures from aggregated melanoma and NSCLC samples. In this method, Bayesian variant nonnegative matrix factorization (NMF) was applied to decompose mutation portrait matrix A with 96 base substitution categories into two nonnegative matrices W and H (i.e., A ≈ W × H), where W indicating the extracted mutational signatures and H representing the mutational activities of each mutational signature. All determined mutational signatures were then compared with the 30 well-annotated signatures in the COSMIC database (version 2) based on cosine similarity.
Tumor infiltrating immunocyte estimation
CIBERSORT method was used to evaluate the tumor abundance of 22 immunocyte subtypes with 547 feature genes from the LM22 signature60. Angelova et al. developed an 812-immune-metagene model to assess the infiltrating abundance of 31 immune cells61, gene panels for each immunocyte type were shown in Supplementary Data 8. In this study, we employed both methods to obtain a comprehensive result.
Collection of immune-related signatures
Recently revealed representative immune-related signatures were curated as follows: (1) immune and stromal cells signatures, which suggest the total immune and stromal cell infiltration abundance in the microenvironment40; (2) immune cell subsets, evaluation of T cells, B cells, and NK cells infiltration62; (3) T/NK, B/P, and M/D metagene, which, respectively, suggests the enrichment of T/NK cells, B/plasma cells, and monocytes/dendritic cells63; (4) Type 1/2 IFN response, which are two interferon responses featured by interferon α and γ, respectively64; (5) IFNγ signature, which is a well-known indicator for immune response and ICI outcome16; (6) T cell-inflamed signature, an indicator derived from the IFNγ signature65; (7) immune cytolytic activity64; (8) immune signaling molecules62; (9) cytokines and chemokines62; (10) TLS, which is tertiary lymphoid structures and links with immune response66. Detailed feature genes for each immune signature were illustrated in Supplementary Data 9.
Activated-stroma signature
A stroma-relevant signature41, which was characterized by two features (i.e., activated-stroma and normal-stroma) was proposed by Moffitt et al. study. By using the nearest template prediction (NTP) method67 with distinct feature genes of two stroma subgroups, we could obtain an activated stromal subtype.
GSVA and GSEA
Single sample gene set enrichment analysis (ssGSEA) method in the GSVA package36 was employed to calculate enrichment scores of collected immune signatures for each sample with distinct gene sets. Gene expression differential analysis according to FAT1 status was performed with R package DESeq237. The obtained t values were subsequently regarded as the input to conducting gene set enrichment analysis (GSEA) in the fgsea package (https://github.com/ctlab/fgsea). Signaling pathways from the Hallmark and KEGG databases were utilized as the background circuits.
Acquisition of mutational burden
Tumor mutation burden (TMB) was defined as the log2 transformation of total non-synonymous mutations per megabase in both WES and TCGA cohorts; for the MSKCC cohort, TMB was obtained from the supplementary information. The neoantigen burden (NB) of 224 melanoma and 109 NSCLC WES samples was estimated based on the method reported by Balachandran et al. 68. The neoantigen data of 340 melanoma and 656 NSCLC samples in the TCGA cohort was acquired from the Cancer Immunome Atlas (TCIA, https://www.tcia.at/home) project.
Statistical analyses
R software (version 4.0.5) was employed to achieve related analyses. Gene mutational patterns were shown with maftools package69. Heatmap illustration of two subgroups was completed with pheatmap package. Kaplan–Meier method was used to produce survival curves and the Log-rank test to compare the differences. Multivariate regression models within forestmodel package were utilized to adjust confounding factors. Relationship of continuous and categorical variables with FAT1 status was calculated with Wilcoxon rank-sum test and Fisher exact test, respectively.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
All data used in this study were acquired from publicly available cohorts, which are described in the “Methods” section.
Code availability
All codes supported the results of this study are available from the corresponding author upon reasonable request.
References
Bray, F. et al. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 68, 394–424 (2018).
Schadendorf, D. et al. Melanoma. Lancet 392, 971–984 (2018).
Luke, J. J., Flaherty, K. T., Ribas, A. & Long, G. V. Targeted agents and immunotherapies: optimizing outcomes in melanoma. Nat. Rev. Clin. Oncol. 14, 463–482 (2017).
Wolchok, J. D. et al. Long-term outcomes with nivolumab plus ipilimumab or nivolumab alone versus ipilimumab in patients with advanced melanoma. J. Clin. Oncol 40, 127–137 (2021).
Hu-Lieskovan, S. et al. Improved antitumor activity of immunotherapy with BRAF and MEK inhibitors in BRAF(V600E) melanoma. Sci. Transl. Med. 7, 279ra241 (2015).
Castellanos, E., Feld, E. & Horn, L. Driven by mutations: the predictive value of mutation subtype in EGFR-mutated non-small cell lung cancer. J. Thorac. Oncol. 12, 612–623 (2017).
Wu, S. G. & Shih, J. Y. Management of acquired resistance to EGFR TKI-targeted therapy in advanced non-small cell lung cancer. Mol. Cancer 17, 38 (2018).
Le, X. et al. Dual EGFR-VEGF pathway inhibition: a promising strategy for patients with EGFR-mutant NSCLC. J. Thorac. Oncol. 16, 205–215 (2021).
Doroshow, D. B. et al. Immunotherapy in non-small cell lung cancer: facts and hopes. Clin. Cancer Res. 25, 4592–4602 (2019).
Shi, F. et al. Sex disparities of genomic determinants in response to immune checkpoint inhibitors in melanoma. Front. Immunol. 12, 721409 (2021).
Samstein, R. M. et al. Tumor mutational load predicts survival after immunotherapy across multiple cancer types. Nat. Genet. 51, 202–206 (2019).
Chan, T. A. et al. Development of tumor mutation burden as an immunotherapy biomarker: utility for the oncology clinic. Ann. Oncol. 30, 44–56 (2019).
Rizvi, N. A. et al. Cancer immunology. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science 348, 124–128 (2015).
Davis, A. A. & Patel, V. G. The role of PD-L1 expression as a predictive biomarker: an analysis of all US Food and Drug Administration (FDA) approvals of immune checkpoint inhibitors. J. Immunother. Cancer 7, 278 (2019).
Wang, F. et al. Evaluation of POLE and POLD1 mutations as biomarkers for immunotherapy outcomes across multiple cancer types. JAMA Oncol. 5, 1504–1506 (2019).
Dong, Z. Y. et al. Potential predictive value of TP53 and KRAS mutation status for response to PD-1 blockade immunotherapy in lung adenocarcinoma. Clin. Cancer Res. 23, 3012–3024 (2017).
Sun, H. et al. Specific TP53 subtype as biomarker for immune checkpoint inhibitors in lung adenocarcinoma. EBioMedicine 60, 102990 (2020).
Zhang, L., Han, X. & Shi, Y. Association of MUC16 mutation with response to immune checkpoint inhibitors in solid tumors. JAMA Netw. Open 3, e2013201 (2020).
Wang, Q., Yang, Y., Yang, M., Li, X. & Chen, K. High mutation load, immune-activated microenvironment, favorable outcome, and better immunotherapeutic efficacy in melanoma patients harboring MUC16/CA125 mutations. Aging 12, 10827–10843 (2020).
Braun, D. A. et al. Clinical validation of PBRM1 alterations as a marker of immune checkpoint inhibitor response in renal cell carcinoma. JAMA Oncol. 5, 1631–1633 (2019).
Sade-Feldman, M. et al. Resistance to checkpoint blockade therapy through inactivation of antigen presentation. Nat. Commun. 8, 1136 (2017).
The tumor suppressor FAT1 modulates WNT activity in multiple cancers. Cancer Discov. 3, OF19, https://doi.org/10.1038/ng.2538 (2013).
Martin, D. et al. Assembly and activation of the Hippo signalome by FAT1 tumor suppressor. Nat. Commun. 9, 2372 (2018).
Hu, X. et al. FAT1 prevents epithelial mesenchymal transition (EMT) via MAPK/ERK signaling pathway in esophageal squamous cell cancer. Cancer Lett. 397, 83–93 (2017).
Pastushenko, I. et al. Fat1 deletion promotes hybrid EMT state, tumour stemness and metastasis. Nature 589, 448–455 (2021).
Chen, Z. et al. The proteomic landscape of growth factor signaling networks associated with FAT1 mutations in head and neck cancers. Cancer Res. 81, 4402–4416 (2021).
Lin, S. C. et al. FAT1 somatic mutations in head and neck carcinoma are associated with tumor progression and survival. Carcinogenesis 39, 1320–1330 (2018).
Kim, K. T., Kim, B. S. & Kim, J. H. Association between FAT1 mutation and overall survival in patients with human papillomavirus-negative head and neck squamous cell carcinoma. Head Neck 38, E2021–2029 (2016).
Li, Z. et al. Loss of the FAT1 tumor suppressor promotes resistance to CDK4/6 inhibitors via the Hippo pathway. Cancer Cell 34, 893–905 e898 (2018).
Loss of FAT1 Drives Resistance to CDK4/6 Inhibitors in Breast Cancer. Cancer Discov. 9, 161, https://doi.org/10.1016/j.ccell.2018.11.006 (2019).
Dikshit, B. et al. FAT1 acts as an upstream regulator of oncogenic and inflammatory pathways, via PDCD4, in glioma cells. Oncogene 32, 3798–3808 (2013).
Laginestra, M. A. et al. Whole exome sequencing reveals mutations in FAT1 tumor suppressor gene clinically impacting on peripheral T-cell lymphoma not otherwise specified. Mod. Pathol. 33, 179–187 (2020).
Ostroumov, D., Fekete-Drimusz, N., Saborowski, M., Kuhnel, F. & Woller, N. CD4 and CD8 T lymphocyte interplay in controlling tumor growth. Cell. Mol. Life Sci. 75, 689–713 (2018).
Zander, R. et al. CD4(+) T cell help is required for the formation of a cytolytic CD8(+) T cell subset that protects against chronic infection and cancer. Immunity 51, 1028–1042 e1024 (2019).
Buchan, S. L. et al. Antibodies to costimulatory receptor 4-1BB enhance anti-tumor immunity via T regulatory cell depletion and promotion of CD8 T cell effector function. Immunity 49, 958–970 e957 (2018).
Tanaka, A. & Sakaguchi, S. Regulatory T cells in cancer immunotherapy. Cell Res. 27, 109–118 (2017).
Turley, S. J., Cremasco, V. & Astarita, J. L. Immunological hallmarks of stromal cells in the tumour microenvironment. Nat. Rev. Immunol. 15, 669–682 (2015).
Wu, S. Z. et al. Stromal cell diversity associated with immune evasion in human triple-negative breast cancer. EMBO J. 39, e104063 (2020).
Lambrechts, D. et al. Phenotype molding of stromal cells in the lung tumor microenvironment. Nat. Med. 24, 1277–1289 (2018).
Yoshihara, K. et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat. Commun. 4, 2612 (2013).
Moffitt, R. A. et al. Virtual microdissection identifies distinct tumor- and stroma-specific subtypes of pancreatic ductal adenocarcinoma. Nat. Genet. 47, 1168–1178 (2015).
Borden, E. C. Interferons alpha and beta in cancer: therapeutic opportunities from new insights. Nat. Rev. Drug Discov. 18, 219–234 (2019).
Ni, L. & Lu, J. Interferon gamma in cancer immunotherapy. Cancer Med. 7, 4509–4516 (2018).
Hu, C. et al. Genomic profiles and their associations with TMB, PD-L1 expression, and immune cell infiltration landscapes in synchronous multiple primary lung cancers. J. Immunother. Cancer 9, e003773 (2021).
Goto, Y. Tumor mutation burden: is it ready for the clinic? J. Clin. Oncol. 36, 2978–2979 (2018).
Wood, M. A., Nellore, A. & Thompson, R. F. Tumor mutation burden-from doubts to concerns. JAMA Oncol. 5, 1808–1809 (2019).
Grandi, A. et al. Vaccination with a FAT1-derived B cell epitope combined with tumor-specific B and T cell epitopes elicits additive protection in cancer mouse models. Front. Oncol. 8, 481 (2018).
Miao, D. et al. Genomic correlates of response to immune checkpoint blockade in microsatellite-stable solid tumors. Nat. Genet. 50, 1271–1281 (2018).
Jia, Q., Wang, J., He, N., He, J. & Zhu, B. Titin mutation associated with responsiveness to checkpoint blockades in solid tumors. JCI Insight 4, e127901 (2019).
Snyder, A. et al. Genetic basis for clinical response to CTLA-4 blockade in melanoma. N. Engl. J. Med. 371, 2189–2199 (2014).
Van Allen, E. M. et al. Genomic correlates of response to CTLA-4 blockade in metastatic melanoma. Science 350, 207–211 (2015).
Hugo, W. et al. Genomic and transcriptomic features of response to anti-PD-1 therapy in metastatic melanoma. Cell 165, 35–44 (2016).
Zaretsky, J. M. et al. Mutations associated with acquired resistance to PD-1 blockade in melanoma. N. Engl. J. Med. 375, 819–829 (2016).
Riaz, N. et al. Tumor and microenvironment evolution during immunotherapy with nivolumab. Cell 171, 934–949 e916 (2017).
Roh, W. et al. Integrated molecular analysis of tumor biopsies on sequential CTLA-4 and PD-1 blockade reveals markers of response and resistance. Sci. Transl. Med. 9, eaah3560 (2017).
Liu, D. et al. Integrative molecular and clinical modeling of clinical outcomes to PD1 blockade in patients with metastatic melanoma. Nat. Med. 25, 1916–1927 (2019).
Hellmann, M. D. et al. Genomic features of response to combination immunotherapy in patients with advanced non-small-cell lung cancer. Cancer Cell. 33, 843–852 e844 (2018).
Ramos, A. H. et al. Oncotator: cancer variant annotation tool. Hum. Mutat. 36, E2423–2429 (2015).
Kim, J. et al. Somatic ERCC2 mutations are associated with a distinct genomic signature in urothelial tumors. Nat. Genet. 48, 600–606 (2016).
Newman, A. M. et al. Robust enumeration of cell subsets from tissue expression profiles. Nat. Methods 12, 453–457 (2015).
Angelova, M. et al. Characterization of the immunophenotypes and antigenomes of colorectal cancers reveals distinct tumor escape mechanisms and novel targets for immunotherapy. Genome Biol. 16, 64 (2015).
Cancer Genome Atlas Network. Genomic classification of cutaneous melanoma. Cell 161, 1681–1696 (2015).
Nagalla, S. et al. Interactions between immunity, proliferation and molecular subtype in breast cancer prognosis. Genome Biol. 14, R34 (2013).
Rooney, M. S., Shukla, S. A., Wu, C. J., Getz, G. & Hacohen, N. Molecular and genetic properties of tumors associated with local immune cytolytic activity. Cell 160, 48–61 (2015).
Ayers, M. et al. IFN-gamma-related mRNA profile predicts clinical response to PD-1 blockade. J. Clin. Investig. 127, 2930–2940 (2017).
Finkin, S. et al. Ectopic lymphoid structures function as microniches for tumor progenitor cells in hepatocellular carcinoma. Nat. Immunol. 16, 1235–1244 (2015).
Hoshida, Y. Nearest template prediction: a single-sample-based flexible class prediction with confidence assessment. PLoS ONE 5, e15543 (2010).
Balachandran, V. P. et al. Identification of unique neoantigen qualities in long-term survivors of pancreatic cancer. Nature 551, 512–516 (2017).
Mayakonda, A., Lin, D. C., Assenov, Y., Plass, C. & Koeffler, H. P. Maftools: efficient and comprehensive analysis of somatic variants in cancer. Genome Res. 28, 1747–1756 (2018).
Acknowledgements
This study was supported by the Shandong Provincial Youth Innovation Team Development Plan of Colleges and Universities (No. 2019-6-156, Lu-Jiao), Medicine and Health Science and Technology Development Plan Project of Shandong Province (No. 202112050480), and National Natural Science Foundation of China (No. 81872719).
Author information
Authors and Affiliations
Contributions
Q.W. conceived this study; Q.W., W.Z., Y.T., Y.G., Y.K., F.S., and C.S. collected the related data and performed main data analysis; W.Z., Y.T., Y.G., F.S., Y.K., and C.S. conducted data analysis and interpretation; W.Z., Y.T., Y.G., S.W., and Q.W. drafted and corrected the paper; Q.W. supervised this study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, W., Tang, Y., Guo, Y. et al. Favorable immune checkpoint inhibitor outcome of patients with melanoma and NSCLC harboring FAT1 mutations. npj Precis. Onc. 6, 46 (2022). https://doi.org/10.1038/s41698-022-00292-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41698-022-00292-6
This article is cited by
-
Identification of hub genes distinguishing subtypes in endometrial stromal sarcoma through comprehensive bioinformatics analysis
Scientific Reports (2024)
-
Association of ERCC family mutations with prognosis and immune checkpoint inhibitors response in multiple cancers
Scientific Reports (2023)
-
The diverse functions of FAT1 in cancer progression: good, bad, or ugly?
Journal of Experimental & Clinical Cancer Research (2022)