Abstract
Neural networks are frequently employed to model species distribution through backpropagation methods, known as backpropagation neural networks (BPNN). However, the complex structure of BPNN introduces parameter settings challenges, such as the determination of connection weights, which can affect the accuracy of model simulation. In this paper, we integrated the Grey Wolf Optimizer (GWO) algorithm, renowned for its excellent global search capacity and rapid convergence, to enhance the performance of BPNN. Then we obtained a novel hybrid algorithm, the Grey Wolf Optimizer algorithm optimized backpropagation neural networks algorithm (GNNA), designed for predicting species’ potential distribution. We also compared the GNNA with four prevalent species distribution models (SDMs), namely the generalized boosting model (GBM), generalized linear model (GLM), maximum entropy (MaxEnt), and random forest (RF). These models were evaluated using three evaluation metrics: the area under the receiver operating characteristic curve, Cohen’s kappa, and the true skill statistic, across 23 varied species. Additionally, we examined the predictive accuracy concerning spatial distribution. The results showed that the predictive performance of GNNA was significantly improved compared to BPNN, was significantly better than that of GLM and GBM, and was even comparable to that of MaxEnt and RF in predicting species distributions with small sample sizes. Furthermore, the GNNA demonstrates exceptional powers in forecasting the potential non-native distribution of invasive plant species.
Similar content being viewed by others
Introduction
Species distribution models (SDMs) use known geographical occurrences of species and corresponding environmental conditions, such as bioclimatic variables and abiotic variables, to predict the potential distribution of species1,2,3. SDMs have become important tools for ecologists to study ecological issues such as species diversity4,5,6, species conservation7,8,9 and biological invasions10,11. In the last decades, a large number of SDMs have been proposed, including regression models (e.g., generalized linear model, GLM)12,13,14,15, classification models (e.g., generalized boosting model, GBM)16,17,18, complex models (e.g., random forest, RF; maximum entropy, MaxEnt)16,19,20,21, and ensemble models22,23. Notably, SDMs such as GLM, GBM, MaxEnt, and RF, are extensively applied in investigating ecological and evolutionary theories24,25, assessing climate change impacts8,26,27, managing invasive species10,11, and identifying conservation areas7,8.
Despite their widespread use, the predictive performance of SDMs can varies significantly across different algorithms2,3,28,29, posing challenges for reliable forecasts30,31,32. Most research in this filed has focused on comparing the predictive success of various SDMs, endorsing those with superior performance2,3,33,34,35. However, there are few studies on optimization of SDMs that are abandoned due to poor predictive performance36. With the development of machine learning, backpropagation neural networks (BPNN) have gained advantages in ecological research where data rarely meet parametric statistical assumptions and non-linear relationships are prevalent37,38,39. However, BPNN also have some disadvantages, such as high dependency on the initial weights, the tendency to be trapped in the local optimum, and slow convergence38,40,41, which are particularly pronounced in species distribution predictions3,28.
Swarm intelligence optimization algorithms (SIOAs), known for their simplicity, flexibility, and high efficiency, have been used as the primary technique to solve global optimization problems42,43,44. It should be mentioned that the SIOAs mainly introduce randomness in the search process to reduce the possibility of falling into the local optimum42. Therefore, it is of practical significance to use the SIOAs to obtain the optimal solution to the global optimization problem. In the past decades, the SIOAs has developed rapidly and becomes a hotspot in many fields42,43,44,45,46,47,48. So far, many different types of SIOAs have been proposed, such as the Grey Wolf Optimizer (GWO) algorithm43, the butterfly optimization algorithm (BOA)44, and the sparrow search algorithm (SSA)42, each demonstrating success across different optimization tasks41,49.
Motivated by these developments, our study introduced a novel hybrid algorithm that leverages the GWO to enhance the BPNN’s predictive performance for species distribution. We detailed the construction of this hybrid algorithm and evaluated its performance against BPNN and the prevalent SDMs (GBM, GLM, MaxEnt, and RF) using data on 23 species. Additionally, we explored the hybrid model’s ability to predict the spatial distribution of an invasive species, aiming to showcase its effectiveness in spatial distribution prediction.
Materials and methods
Backpropagation neural networks and Grey Wolf Optimizer algorithm
Backpropagation neural networks (BPNN) are capable of handling both continuous and categorical data40,50. They exhibit some attractive properties, including the ability to capture nonlinearity and tolerance noise, but they also have some drawbacks, such as being highly dependent on initial solutions and falling into the local optimum38,40,41. The Grey Wolf Optimizer (GWO) algorithm can effectively balance local optimization and global search with its adaptive convergence factor and information feedback mechanism and obtain high convergence speed and solution accuracy43.
Construction of the hybrid algorithm
In this paper, we proposed a novel hybrid algorithm for predicting the potential distribution of species, called Grey Wolf Optimizer algorithm optimized backpropagation neural networks algorithm (GNNA). Specifically, we used the BPNN to construct GNNA. GNNA is not a simple combination of GWO and BPNN but uses the good global search ability and fast convergence ability of GWO to determine the optimal threshold and optimal weight of BPNN. The specific GNNA process is as follows:
-
1.
Determine the basic structure of the BPNN. The three-layer BPNN was selected, the number of nodes in the hidden layer was determined to be 5 and the training set and test set were randomly generated according to 4:1.
-
2.
Initialize the basic parameters. The gray wolf population size was set as 20, the maximum number of iterations was 100, the upper bound of the gray wolf was 1, and the lower bound of the gray wolf was − 1. Initialize the gray wolf position and parameters A, a and C. The dimension of each gray wolf position information was calculated according to the number of layers in each layer of BPNN (dimension = input layer number × hidden layer number + hidden layer number + hidden layer number × output layer number + output layer number).
-
3.
Determine the fitness function. The activation function in the hidden layer and the output layer were adopted Sigmoid type function. The learning rate was 0.01 and the training goal was 0.00001.
-
4.
Calculate the fitness values of all search agents according to the threshold and weight and update the position information of the remaining gray wolves \(\omega\) and parameters Ai, a and Ci.
-
5.
Divide the data into test data and training data, and record the optimal search agent and its corresponding error.
-
6.
Determine whether the maximum number of iterations was met. If the condition was met, terminate the cycle; otherwise, repeat steps (4) to (6).
-
7.
Get the result. The final position of the gray wolf \(\alpha\), the minimum error of the position of the gray wolf \(\alpha\), and error between test data and training data.
Update the gray wolf position according to the following equations. First, calculate the distance vectors between the individual and the prey (Eqs. 1 and 2).
where, \(C_{i} \left( t \right)\left( {i = 1,2,3} \right)\) represents the random vectors; \(r_{i} \left( t \right)\left( {i = 1,2,3} \right)\) represents the random vectors in which every element is in [0,1]; \(D_{p} \left( t \right)(p = \alpha ,\beta ,\delta )\) represents the distance vectors between p and other individuals, \(\circ\) represents the Hadamard product, || represents the absolute value of each element in the vectors; \(X_{p} \left( t \right)(p = \alpha ,\beta ,\delta )\) represents the current position of p; \(X(t)\) represents the current position of the gray wolf.
Second, the positions of the first three wolves are updated according to the following equations:
where, \(A_{i} \left( t \right)(i = 1,2,3)\) represents the convergence vector; \(r_{i + 3} \left( t \right)(i = 1,2,3)\) represents the random vectors in which every element is in [0,1]; components of \(a(t)\) are linearly decreased from 2 to 0 during iteration; \(X_{i} \left( t \right)\left( {i = 1,2,3} \right)\) represents the updated position of the first three wolves.
Finally, adjust the position of the offspring gray wolf according to the following equations:
where, \(\omega_{i} (i = 1,2,3)\) represents respectively the learning rate of wolf \(\omega\) to wolf \(\alpha ,\beta ,\delta\); \(\left\| {X_{i} \left( t \right)} \right\|\) represents the 2-norm of position vector \(X_{i} \left( t \right)\), and \(X_{\omega } (t + 1)\) represents the position of the offspring gray wolves. The pseudo code of the GNNA is shown as follows (Algorithm 1).

Algorithm 1 Pseudo code of the GNNA.
Comparing GNNA predictive performance with BPNN and four commonly used SDMs
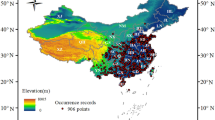
We first compared the predictive performance of GNNA with BPNN, posing the explicit hypothesis that GNNA would outperform BPNN and achieve good absolute predictive performance. To this aim, we downloaded occurrence records for 23 species after 1970 from the Global Biodiversity Information Facility (GBIF, http://www.gbif.org/) and removed duplicate records within a 5 km radius. These species have diverse characteristics in the climate, elevation, and range of their habitat (the number of records and details for each species are shown in Table S1 and Table S2 in Supporting Information). We also categorized the 23 species into three kinds of sample sizes according to the number of occurrence records (Table S1 in Supporting Information). In addition, for each species, we randomly generated pseudo-absence data according to three times the number of occurrence records. Each occurrence and pseudo-absence point is associated with a vector composed of climate values, corresponding to bioclimatic variables, which are downloaded from WorldClim 2.1 (http://www.worldclim.org/) at a raw resolution of 2.5 arc-min51 and selected by Pearson’s correlation test (r) with |r|< 0.7. Abbreviations and full names of bioclimatic variables are listed in Table S3, and the bioclimatic variables obtained for each species are shown in Table S4.
As a preliminary step, we constructed SDMs for all 23 species through BPNN and GNNA. Specifically, for each species, we first randomly split 80% of the species data into training data and the remaining 20% into testing data. We then evaluated the predictive performance of the model by computing three metrics widely used in ecological research, namely the area under the receiver operating characteristic curve (AUC, Swets52), Cohen's kappa (KAPPA, Cohen53), and the true skill statistic (TSS, Allouche et al.54). We repeated this splitting procedure 12 times and then took the median of the evaluation metrics. In this study, we used a threshold value at which the TSS is maximized to determine presences and absences.
We then applied four commonly used SDMs, namely GLM14,15, GBM16,18, RF19, and MaxEnt20, to all 23 species and compared their predictive performance with GNNA. We followed Brun et al.55 and Zhang et al.56 to set complex parameters for each of the four SDMs involved in the comparison, aiming to make them sufficiently comparable to GNNA. For GLM, the response curve was set to polynomial and the search direction for stepwise regression was set to both; for RF, the number of variables randomly sampled as candidates at each split was set to 5, the number of trees to grow was set to 1000, and the minimum size of terminal nodes was set to 5; for GBM, the maximum depth of each tree was set to 3, the total number of trees was set to 1000, and a shrinkage parameter applied to each tree in the expansion was set to 0.01; for MaxEnt, the maximum number of iterations was set to 100. We performed these SDMs in the R environment (version 4.1.1, R Core Team, 2021) using the packages ‘stats’ (version 4.0.5), ‘randomForest’ (version 4.6–14), ‘gbm’ (version 2.1.8), and ‘dismo’ (version 1.3–5). The data (i.e., species data and bioclimatic variables) and data partitioning used for the four SDMs (i.e., GLM, GBM, RF, and MaxEnt) described above are the same as GNNA and BPNN, which is to facilitate the direct comparison of the predictive performance of the four SDMs with that of GNNA.
Comparison of spatial distribution predictions—an application case of an invasive species
In addition to the comparison of predictive performance (measured by metrics), the comparison of the prediction of spatial distribution should be taken into consideration. The prediction of spatial distribution is concerned with practical application, especially that of invasive species. We provided an example for predicting the distribution of an invasive plant, Mimosa bimucronata (DC.) Kuntze (M. bimucronata), which is native in South America and has now invaded the southern coastal region of China. We applied GNNA, BPNN, and the four commonly used SDMs to predict the native and non-native distribution of the species under the current environment, respectively. We used native occurrence records to train the SDMs and predicted both native and non-native potential distributions. At the same time, non-native occurrence records were used to verify the prediction performance of the SDMs for the potential distribution. The occurrence records of M. bimucronata in South America were obtained from GBIF (http://www.gbif.org/), and the occurrence records of M. bimucronata in China were obtained from the study of Xie et al.57. The environmental variables and parameter settings of the SDMs were consistent with those described above in section “Comparing GNNA predictive performance with BPNN and four commonly used SDMs”.
Results
Comparison of predictive performance between GNNA and BPNN
Overall, the three evaluation metrics consistently showed that GNNA had better predictive performance than BPNN (Fig. 1a–c). Specifically, 20 out of 23 species performed better with GNNA based on having higher metric values for two or more metrics (Fig. 1d–f). The percentage improvement in predictive performance of GNNA over BPNN, no matter which metric was used to measure it, decreased as the sample size increased (Table 1). When the sample size was small, the predictive performance of GNNA was improved by about 2% compared with that of BPNN, while when the sample size was large (middle and big), the predictive performance of GNNA was improved by less than 0.3% compared with that of BPNN (Table 1). The predictive performance of GNNA gradually stabilized with increasing sample size, with a wide inter-quartile range (IQR) when the sample size was small and a narrower IQR when the sample size was large (middle and big) (Table 1).
Comparison of predictive performance between GNNA and BPNN under three evaluation metrics (AUC, KAPPA, and TSS). (a–c) Represent the density distribution of 23 species under AUC, KAPPA, and TSS, respectively. (d–f) Show the comparison of predictive performance of GNNA and BPNN under AUC, KAPPA, and TSS for each species, respectively.
Comparison of predictive performance between GNNA and four commonly used SDMs
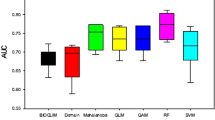
Overall, the predictive performance of GNNA was better than that of GBM and GLM, but slightly lower than that of RF and MaxEnt (Fig. 2b–d). Specifically, 14 out of 23 species (about 61% of species) showed better predictive performance of GNNA than GBM, and 12 out of 23 species (about 52% of species) showed better predictive performance of GNNA than GLM (Fig. 2a). Only about five out of 23 species (about 22% of species) showed better predictive performance for GNNA than for RF and MaxEnt (Fig. 2a). The predictive performance of GNNA was comparable to that of MaxEnt and RF in predicting the distributions of species with small sample sizes (such as S. dareiformis and C. flavum) (Fig. 2e–g).
Comparison of the predictive performance of the GNNA model with four commonly used species distribution models (SDMs, i.e., GBM, GLM, RF, and MaxEnt) under three evaluation metrics (i.e., AUC, KAPPA, and TSS). (a) Represents how many species out of the 23 species show that GNNA has better predictive performance than four commonly used SDMs. GNNA > ** means that the predictive performance of GNNA is better than that of ** under the same species, and ** means the four commonly used SDMs. (b–d) Represent the comparison of the predictive performance of GNNA and four commonly used SDMs under the three evaluation metrics, respectively. (e–g) Represent the predictive performance of the 23 species under GNNA and four commonly used SDMs under the three evaluation metrics, respectively.
Comparison of spatial distributions predicted by GNNA, BPNN, and the four commonly used SDMs
The native distribution is mainly concentrated on the southern edge of Brazil (Fig. 3), as shown by the almost identical findings from GNNA, BPNN, and the four commonly used SDMs (i.e., MaxEnt, RF, GBM, and GLM) in predicting native distribution areas. However, there are some obvious differences when predicting non-native distribution areas. In addition to the prediction results, all models consistently show that Guangxi, Guangdong, and Hainan are the main distribution areas of non-native species (Fig. 4). The prediction results of GNNA, MaxEnt, and RF also showed a high probability of invasion in Chongqing, which is consistent with the occurrence record of M. bimucronata found in Chongqing (Fig. 4a–c).
Current distribution of M. bimucronata in South America (native) based on GNNA, BPNN, and the four commonly used SDMs (i.e., MaxEnt, RF, GBM, and GLM), respectively. The black point represents the occurrence records of M. bimucronata in South America. Figures were created using R 4.1.1 (https://www.R-project.org/).
Current distribution of M. bimucronata in China (non-native) based on GNNA, BPNN, and the four commonly used SDMs (i.e., MaxEnt, RF, GBM, and GLM), respectively. The black triangle represents the occurrence records of M. bimucronata in China. The occurrence records of M. bimucronata in China are not used to predict its distribution in non-native (China), but only to test whether the predicted distribution is valid. Figures were created using R 4.1.1 (https://www.R-project.org/).
Discussion
The proposed hybrid algorithm, GNNA, demonstrates a substantial enhancement in predictive performance over the traditional BPNN, as evidenced by three distinct evaluation metrics. The advancement of predictive performance remains a primary goal in developing new methods for creating SDMs36,58, and our research provides a new idea for combining existing SDMs with SIOAs to develop SDMs. In addition, the stability of GNNA is affected by the sample size and increases with the increase in sample size. Nevertheless, certain species within our study did not exhibit this trend when applying GNNA, which may be attributed to either their widespread geographical distribution or potential inaccuracies in occurrence records which sourced from the GBIF.
Our comparative analysis reveals that the predictive performance of GNNA was better than that of GLM and GBM, and delivering predictive results on compare with MaxEnt and RF when species with small sample sizes. Despite the notable superiority of GNNA over the four commonly used SDMs in certain cases (e.g., S. dareiformis and C. flavum), relying solely on a single SDM could result in skewed interpretations within ecological research3,59. It is well-established that no single SDM can consistently deliver high predictive performance across diverse species and regions29,35,60. In ecological research, researchers often depend on the consistent results of multiple SDMs or ensemble models to fortify the credibility of their findings2,23,61,62,63. Therefore, our proposed GNNA has great potential to serve as an integral base learner within ensemble model constructions.
Furthermore, biological invasion is a global issue that ecologists have been concerned about for decades64,65,66,67. Effectively predicting the potential distribution of invasive alien plants provides is crucial for developing prevention and control strategies against their spread68,69. SDMs have been increasingly used to predict the potential distribution of invasive plants in recent years11,57,69. The GNNA proposed in this study also showed superior ability in predicting the non-native potential distribution of invasive plants.
Conclusions
This study introduces an SIOA GWO into SDMs, and constructs a hybrid algorithm GNNA to improve the predictive performance of SDMs. Specifically, compared with BPNN, the predictive performance of the hybrid algorithm GNNA proposed in this paper is significantly improved. In addition, GNNA, which has excellent predictive performance comparable to common SDMs such as MaxEnt and RF, can be used as a good base learner for ensemble models. Up to now, many different types of SIOAs have been proposed, and these SIOAs have been tested to have superior optimization capabilities. We will try to combine more SIOAs with SDMs in future work.
Data availability
The cleaned occurrence records for the 23 real plant species investigated in this study: Dryad https://datadryad.org/stash/share/XhPyzK093jJB0x3cyH4x0ujpbDTkAgmqBDDUjZcSh3o.
References
Austin, M. Species distribution models and ecological theory: A critical assessment and some possible new approaches. Ecol. Model. 200, 1–19 (2007).
Hao, T., Elith, J., Guillera-Arroita, G. & Lahoz-Monfort, J. J. A review of evidence about use and performance of species distribution modelling ensembles like BIOMOD. Divers. Distrib. 25, 839–852 (2019).
Li, X. & Wang, Y. Applying various algorithms for species distribution modelling. Integr. Zool. 8, 124–135 (2013).
Fitzpatrick, M. C. et al. Forecasting the future of biodiversity: A test of single-and multi-species models for ants in North America. Ecography 34, 836–847 (2011).
Moullec, F. et al. Using species distribution models only may underestimate climate change impacts on future marine biodiversity. Ecol. Model. 464, 109826 (2022).
Guisan, A. & Thuiller, W. Predicting species distribution: Offering more than simple habitat models. Ecol. Lett. 8, 993–1009 (2005).
Williams, J. N. et al. Using species distribution models to predict new occurrences for rare plants. Divers. Distrib. 15, 565–576 (2009).
Brunton, A. J., Conroy, G. C., Schoeman, D. S., Rossetto, M. & Ogbourne, S. M. Seeing the forest through the trees: Applications of species distribution models across an Australian biodiversity hotspot for threatened rainforest species of Fontainea. Glob. Ecol. Conserv. 2023, e02376 (2023).
Marmion, M., Parviainen, M., Luoto, M., Heikkinen, R. K. & Thuiller, W. Evaluation of consensus methods in predictive species distribution modelling. Divers. Distrib. 15, 59–69 (2009).
Aidoo, O. F. et al. A machine learning algorithm-based approach (MaxEnt) for predicting invasive potential of Trioza erytreae on a global scale. Eco. Inform. 71, 101792 (2022).
Padalia, H., Srivastava, V. & Kushwaha, S. Modeling potential invasion range of alien invasive species, Hyptis suaveolens (L.) Poit. in India: Comparison of MaxEnt and GARP. Ecol. Inf. 22, 36–43 (2014).
Friedman, J. H. Multivariate adaptive regression splines. Ann. Stat. 19, 1–67 (1991).
Hastie, T., Tibshirani, R. & Buja, A. Flexible discriminant analysis by optimal scoring. J. Am. Stat. Assoc. 89, 1255–1270 (1994).
McCullagh, P. Generalized linear models. Eur. J. Oper. Res. 16, 285–292 (1984).
Nelder, J. A. & Wedderburn, R. W. Generalized linear models. J. R. Stat. Soc. Ser. A Stat. Soc. 135, 370–384 (1972).
Elith, J., Leathwick, J. R. & Hastie, T. A working guide to boosted regression trees. J. Anim. Ecol. 77, 802–813 (2008).
Hastie, T., Tibshirani, R., Friedman, J. H. & Friedman, J. H. The Elements of Statistical Learning: Data Mining, Inference, and Prediction, vol. 2 (Springer, 2009).
Friedman, J. H. Greedy function approximation: A gradient boosting machine. Ann. Stat. 2001, 1189–1232 (2001).
Breiman, L. Random forests. Mach. Learn. 45, 5–32 (2001).
Phillips, S. J., Anderson, R. P. & Schapire, R. E. Maximum entropy modeling of species geographic distributions. Ecol. Model. 190, 231–259 (2006).
Ripley, B. D. Pattern Recognition and Neural Networks (Cambridge University Press, 2007).
Breiner, F. T., Guisan, A., Bergamini, A. & Nobis, M. P. J. M. I. E. Evolution: Overcoming limitations of modelling rare species by using ensembles of small models. Methods Ecol. Evol. 6, 1210–1218 (2015).
Araújo, M. B. & New, M. Ensemble forecasting of species distributions. Trends Ecol. Evol. 22, 42–47 (2007).
Schorr, G., Holstein, N., Pearman, P., Guisan, A. & Kadereit, J. Integrating species distribution models (SDMs) and phylogeography for two species of Alpine Primula. Ecol. Evol. 2, 1260–1277 (2012).
de Araújo, C. B., Marcondes-Machado, L. O. & Costa, G. C. The importance of biotic interactions in species distribution models: A test of the Eltonian noise hypothesis using parrots. J. Biogeogr. 41, 513–523 (2014).
Crimmins, S. M., Dobrowski, S. Z. & Mynsberge, A. R. Evaluating ensemble forecasts of plant species distributions under climate change. Ecol. Model. 266, 126–130 (2013).
Taylor, P. J., Ogony, L., Ogola, J. & Baxter, R. M. South African mouse shrews (Myosorex) feel the heat: Using species distribution models (SDMs) and IUCN Red List criteria to flag extinction risks due to climate change. Mammal Res. 62, 149–162 (2017).
Norberg, A. et al. A comprehensive evaluation of predictive performance of 33 species distribution models at species and community levels. Ecol. Monogr. 89, e01370 (2019).
Pearson, R. G. et al. Model-based uncertainty in species range prediction. J. Biogeogr. 33, 1704–1711 (2006).
Koo, K. A. et al. Potential climate change effects on tree distributions in the Korean Peninsula: Understanding model & climate uncertainties. Ecol. Model. 353, 17–27 (2017).
Thuiller, W., Guéguen, M., Renaud, J., Karger, D. N. & Zimmermann, N. E. Uncertainty in ensembles of global biodiversity scenarios. Nat. Commun. 10, 1446 (2019).
Araújo, M. B. & Guisan, A. Five (or so) challenges for species distribution modelling. J. Biogeogr. 33, 1677–1688 (2006).
Gobeyn, S. et al. Evolutionary algorithms for species distribution modelling: A review in the context of machine learning. Ecol. Model. 392, 179–195 (2019).
Kampichler, C., Wieland, R., Calmé, S., Weissenberger, H. & Arriaga-Weiss, S. Classification in conservation biology: A comparison of five machine-learning methods. Ecol. Inf. 5, 441–450 (2010).
Segurado, P. & Araujo, M. B. An evaluation of methods for modelling species distributions. J. Biogeogr. 31, 1555–1568 (2004).
Yu, H., Cooper, A. R. & Infante, D. M. Improving species distribution model predictive accuracy using species abundance: Application with boosted regression trees. Ecol. Model. 432, 109202 (2020).
Lek, S. & Guégan, J.-F. Artificial neural networks as a tool in ecological modelling, an introduction. Ecol. Model. 120, 65–73 (1999).
Özesmi, S. L., Tan, C. O. & Özesmi, U. Methodological issues in building, training, and testing artificial neural networks in ecological applications. Ecol. Model. 195, 83–93 (2006).
Chen, Y.-H. & Chang, F.-J. Evolutionary artificial neural networks for hydrological systems forecasting. J. Hydrol. 367, 125–137 (2009).
Faris, H., Aljarah, I. & Mirjalili, S. Training feedforward neural networks using multi-verse optimizer for binary classification problems. Appl. Intell. 45, 322–332 (2016).
Mirjalili, S. How effective is the Grey Wolf Optimizer in training multi-layer perceptrons. Appl. Intell. 43, 150–161 (2015).
Xue, J. & Shen, B. A novel swarm intelligence optimization approach: Sparrow search algorithm. Syst. Sci. Control Eng. 8, 22–34 (2020).
Mirjalili, S., Mirjalili, S. M. & Lewis, A. Grey wolf Optimizer. Adv. Eng. Softw. 69, 46–61 (2014).
Arora, S. & Singh, S. Butterfly optimization algorithm: A novel approach for global optimization. Soft Comput. 23, 715–734 (2019).
Kamboj, V. K., Bath, S. & Dhillon, J. Solution of non-convex economic load dispatch problem using Grey Wolf Optimizer. Neur. Comput. Appl. 27, 1301–1316 (2016).
Komaki, G. & Kayvanfar, V. Grey Wolf Optimizer algorithm for the two-stage assembly flow shop scheduling problem with release time. J. Comput. Sci. 8, 109–120 (2015).
Yang, X. S. & Hossein-Gandomi, A. Bat algorithm: A novel approach for global engineering optimization. Eng. Comput. 29, 464–483 (2012).
Karaboga, D. & Akay, B. A comparative study of artificial bee colony algorithm. Appl. Math. Comput. 214, 108–132 (2009).
Aljarah, I., Faris, H. & Mirjalili, S. Optimizing connection weights in neural networks using the whale optimization algorithm. Soft Comput. 22, 1–15 (2018).
Hancock, J. T. & Khoshgoftaar, T. M. Survey on categorical data for neural networks. J. Big Data 7, 1–41 (2020).
Fick, S. E. & Hijmans, R. J. WorldClim 2: New 1-km spatial resolution climate surfaces for global land areas. Int. J. Climatol. 37, 4302–4315 (2017).
Swets, J. A. Measuring the accuracy of diagnostic systems. Science 240, 1285–1293 (1988).
Cohen, J. A coefficient of agreement for nominal scales. Educ. Psychol. Meas. 20, 37–46 (1960).
Allouche, O., Tsoar, A. & Kadmon, R. Assessing the accuracy of species distribution models: Prevalence, kappa and the true skill statistic (TSS). J. Appl. Ecol. 43, 1223–1232 (2006).
Brun, P. et al. Model complexity affects species distribution projections under climate change. J. Biogeogr. 47, 130–142 (2020).
Zhang, H. T., Guo, W. Y. & Wang, W. T. The dimensionality reductions of environmental variables have a significant effect on the performance of species distribution models. Ecol. Evol. 13, e10747 (2023).
Xie, C., Li, M., Jim, C. Y. & Liu, D. Spatio-temporal patterns of an invasive species Mimosa bimucronata (DC.) Kuntze under different climate scenarios in China. Front. Forests Glob. Change 6, 1144829 (2023).
Stevens, B. S. & Conway, C. J. Predictive multi-scale occupancy models at range-wide extents: Effects of habitat and human disturbance on distributions of wetland birds. Divers. Distrib. 26, 34–48 (2020).
Zhang, H.-T. & Wang, W.-T. Prediction of the potential distribution of the endangered species Meconopsis punicea Maxim under future climate change based on four species distribution models. Plants 12, 1376 (2023).
Elith, J. et al. Novel methods improve prediction of species’ distributions from occurrence data. Ecography 29, 129–151 (2006).
Friedman, J. H. & Popescu, B. E. Predictive learning via rule ensembles. Ann. Appl. Stat. 2008, 916–954 (2008).
Hao, T., Elith, J., Lahoz-Monfort, J. J. & Guillera-Arroita, G. Testing whether ensemble modelling is advantageous for maximising predictive performance of species distribution models. Ecography 43, 549–558 (2020).
Seni, G. & Elder, J. F. Ensemble methods in data mining: Improving accuracy through combining predictions. Synthes. Lect. Data Min. Knowl. Discov. 2, 1–126 (2010).
Diagne, C. et al. High and rising economic costs of biological invasions worldwide. Nature 592, 571–576 (2021).
Vinogradova, Y. et al. Invasive alien plants of Russia: Insights from regional inventories. Biol. Invasions 20, 1931–1943 (2018).
Pyšek, P., Brundu, G., Brock, J., Child, L. & Wade, M. Twenty-five years of conferences on the ecology and management of alien plant invasions: The history of EMAPi 1992–2017. Biol. Invasions 21, 725–742 (2019).
Rai, P. K. & Singh, J. Invasive alien plant species: Their impact on environment, ecosystem services and human health. Ecol. Indic. 111, 106020 (2020).
Gallien, L., Münkemüller, T., Albert, C. H., Boulangeat, I. & Thuiller, W. Predicting potential distributions of invasive species: Where to go from here?. Divers. Distrib. 16, 331–342 (2010).
Panda, R. M. & Behera, M. D. Assessing harmony in distribution patterns of plant invasions: A case study of two invasive alien species in India. Biodivers. Conserv. 28, 2245–2258 (2019).
Acknowledgements
We thank the support of the Innovation Team of Intelligent Computing and Dynamical System Analysis and Application. This work was supported by the National Natural Science Foundation of China (no. 32260293), the Natural Science Foundation of Gansu Province (no. 21JR11RA023), the Scientific Research Project for Colleges and Universities of Gansu Province (no. 2022QB-017), the Foundation Research Funds for the Central Universities (no. 31920240049).
Author information
Authors and Affiliations
Contributions
H-T Z: methodology (lead); formal analysis (lead); data curation (lead); writing – original draft preparation (lead); writing – review and editing (equal); T-T Y: methodology (equal); formal analysis (equal); W-T W: Conceptualization (lead); formal analysis (equal); writing – original draft preparation (equal); writing – review and editing (lead); funding acquisition (lead).
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, HT., Yang, TT. & Wang, WT. A novel hybrid model for species distribution prediction using neural networks and Grey Wolf Optimizer algorithm. Sci Rep 14, 11505 (2024). https://doi.org/10.1038/s41598-024-62285-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-62285-8
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.