Abstract
Temperature sensitivity of abdominal pigmentation in Drosophila melanogaster females allows to investigate the mechanisms underlying phenotypic plasticity. Thermal plasticity of pigmentation is due to modulation of tan and yellow expression, encoding pigmentation enzymes. Furthermore, modulation of tan expression by temperature is correlated to the variation of the active histone mark H3K4me3 on its promoter. Here, we test the role of the DotCom complex, which methylates H3K79, another active mark, in establishment and plasticity of pigmentation. We show that several components of the DotCom complex are involved in the establishment of abdominal pigmentation. In particular, Grappa, the catalytic unit of this complex, plays opposite roles on pigmentation at distinct developmental stages. Indeed, its down-regulation from larval L2 to L3 stages increases female adult pigmentation, whereas its down-regulation during the second half of the pupal stage decreases adult pigmentation. These opposite effects are correlated to the regulation of distinct pigmentation genes by Grappa: yellow repression for the early role and tan activation for the late one. Lastly, reaction norms measuring pigmentation along temperature in mutants for subunits of the DotCom complex reveal that this complex is not only involved in the establishment of female abdominal pigmentation but also in its plasticity.
Similar content being viewed by others
Introduction
Phenotypic plasticity is the ability of a given genotype to produce different phenotypes in response to distinct environmental conditions1. It is widely observed in the wild and is often an adaptation to predictable environmental fluctuations such as seasonal variations. Furthermore, phenotypic plasticity is thought to facilitate evolution, as a phenotype initially induced by the environment can become subsequently fixed by genetic assimilation. A few empirical studies give support to such models of evolution called “Flexible stem hypothesis” or “Plasticity first evolution”2,3,4,5,6,7. Despite these important roles in adaptation and evolution, the molecular mechanisms underlying phenotypic plasticity are only beginning to be unraveled. Here, we use the abdominal pigmentation of Drosophila melanogaster females as a model to study the mechanisms involved in phenotypic plasticity. Abdominal pigmentation is sensitive to temperature, as females grown at 18 °C are darker than those grown at 29°C8. Indeed, the expression of several pigmentation enzyme coding genes (called hereafter pigmentation genes), such as tan and yellow, is regulated by temperature9,10. In particular, tan, encoding an enzyme involved in production of black and brown melanin11, is expressed 7 times more at 18 °C than 29 °C in young females9. Environmental conditions can affect gene expression through modifications of chromatin structure, mainly via histone modifications, nucleosome remodeling or DNA methylation12,13,14. We have previously shown that the H3K4 methyl-transferase Trithorax, that deposits the H3K4me3 active histone mark, is involved in the establishment and plasticity of Drosophila pigmentation by activating tan expression through H3K4 methylation of its promoter9. Furthermore, the quantity of H3K4me3 on the tan promoter is modulated by temperature. Here, we investigate the role of the DotCom complex, involved in the deposition of the active histone mark H3K79me3, on pigmentation gene expression and thermal plasticity of abdominal pigmentation.
The mono-, di- and trimethylation of H3K79 is catalyzed by Dot1 (Disruptor of telomeric silencing-1), a non-SET domain-containing histone lysine methyl-transferase first identified in yeast15. This enzyme, encoded by grappa (gpp) in Drosophila16, is highly conserved from yeast to vertebrates and seems to have a dual role. In yeast and Drosophila, telomeric silencing is disrupted in both Dot1/gpp loss- and gain- of function mutants16,17,18. Moreover, in Drosophila, gpp mutants show phenotypes indicative of disruption of both Polycomb-Group silencing complexes (PcG) and Trithorax-Group activating complexes (TrxG)16. In mammals, Dot1 is involved in embryonic development and heterochromatin organization19. Altogether, these data reveal an important role of H3K79 methylation in the regulation of developmental genes.
Genome-wide profiling studies in different mammalian cell lines show that H3K79me2 and H3K79me3 marks localize to the promoter-proximal regions of actively transcribed genes, with a good correlation with high transcriptional levels20. In yeast, nearly 90% of the genome is methylated on H3K79 residues18. In human, these methylations are associated with activation at single-copy genes21. H3K79me2 is enriched in euchromatin whereas H3K79me3 is enriched in repeated sequences and chromocenter22. In Drosophila, the level of H3K79me3 is positively correlated with gene activity23, which is consistent with some of the phenotypes previously reported for gpp mutants such as anterior transformations of the posterior abdominal segments16.
Interestingly, other histone modifications facilitate H3K79 methylation by Dot1. In yeast, the mono-ubiquitination of histone H2B on lysine 123 (H2BK123ub) is required for an efficient deposition of methylations on H3K79 (H3K79me2/3) by Dot124,25. Structural studies showed that the interaction between Dot1 and H2BK123ub, by reducing the distance between Dot1 and the nucleosome, allows a higher methylation efficiency26. Furthermore, the acetylation of histone H4 on lysine 16 (H4K16ac) is also involved in the regulation of Dot1 activity by structuring and stabilizing the Dot1-nucleosome interaction27.
The Dot1/Gpp enzyme belongs to a large multi-subunits complex called DotCom. In mammals, this complex is composed of AF10, AF17, AF9, ENL, Skp1 and TRRAP. In Drosophila, the complex comprises the corresponding homologues: Alhambra (Alh, homologue of both AF10 and AF17), EAR (homologue of AF9 and ENL), NippedA (homologue of TRRAP) and SkpB (homologue of Skp1)28. We show here that components of the Drosophila complex DotCom (Alhambra, Ear and Gpp) are involved in the establishment and thermal plasticity of abdominal pigmentation.
Results
The DotCom complex is involved in pigmentation establishment
In order to test the role of the DotCom complex on the establishment of pigmentation in Drosophila females, we first down-regulated gpp, encoding the histone methyl-transferase, by using two UAS-RNAi transgenes (RNAi-gpp1 and RNAi-gpp2) and two drivers, pnr-Gal4 and y-Gal4. pnr-Gal4 is an « early » driver, as pnr encodes a transcription factor expressed from the embryo to the adult, along the dorso-longitudinal line29 (Supplementary Fig. 1). y-Gal4 is a « late » driver, as yellow (y) encodes a pigmentation gene expressed in the abdominal epidermis during the second half of the pupal stage30 (Supplementary Fig. 1). We observed a significant effect of both RNAi transgenes on pigmentation but, surprisingly, these effects were opposite depending on the driver (Fig. 1, Supplementary table 1, Supplementary file 1). gpp down-regulation with the pnr-Gal4 driver induced an increase in pigmentation in A5 and A6 abdominal segments, whereas with the y-Gal4 driver pigmentation decreased in A6 and A7. As the main difference between the two drivers is their stage of expression, this result suggests that Gpp plays different roles on pigmentation depending on the developmental stage. The UAS-RNAi-gpp1 line (hereafter called UAS-RNAi-gpp) was used for the following experiments, since the two gpp RNAi transgenes gave very close results.
The methyl-transferase Gpp is involved in the establishment of pigmentation. (A) Abdominal cuticles of females either control (pnr-Gal4/ + or y-Gal4/ +) or RNAi against gpp (UAS-RNAi-gpp1 and UAS-RNAi-gpp2) driven by pnr-Gal4 and y-Gal4. Cuticles were cut just beyond the dorsal midline. Hemi-abdomens are shown. Dashed white lines mark the right borders of the pnr driver expression domain. Abdominal tergites A5 to A7 are indicated. A: anterior; P: posterior; V: ventral; D: dorsal. B, C) Quantification of A5, A6 and A7 pigmentation of UAS-RNAi-gpp1 and UAS-RNAi-gpp2 females driven by pnr-Gal4 (B) or y-Gal4 (C), compared to pnr-Gal4/ + and y-Gal4/ + controls (30 females per genotype, error bars correspond to standard deviations, Tukey post-hoc test following ANOVA on aligned rank transformed data; ***: p < 0.001, **: p < 0.01).
To further confirm the role of the DotCom complex on the establishment of pigmentation, UAS-RNAi transgenes against the three other subunits of the complex were used (Fig. 2, Supplementary table 1). Using the pnr-Gal4 driver, only ear down-regulation gave interpretable results and induced a decrease of pigmentation. For NippedA, the flies presented cuticles incorrectly fused along the dorsal midline that precluded the analysis of pigmentation, whereas for alh no larvae hatched. With the y-Gal4 driver, ear and alh down-regulation induced a decrease of pigmentation, similar to the effect of gpp down-regulation. NippedA down-regulation was lethal after the second larval stage.
Subunits of the DotCom complex are involved in the establishment of pigmentation. (A) Abdominal cuticles of control females (pnr-Gal4/ + or y-Gal4/ +) and females in which genes encoding subunits of the DotCom complex have been inactivated using RNAi transgenes driven by pnr-Gal4 and y-Gal4. Cuticles were cut just beyond the dorsal midline. Hemi-abdomens are shown. Dashed white lines mark the right borders of pnr driver expression domain. Abdominal tergites A5 to A7 are indicated. A: anterior; P: posterior; V: ventral; D: dorsal. The crosses indicate lethality. (B,C) Quantification of A5, A6 and A7 pigmentation of RNAi females driven either by pnr-Gal4 (B) or y-Gal4 (C), compared to pnr-Gal4/ + and y-Gal4/ + controls (n = 30 per genotype, error bars correspond to standard deviations, t-tests; ***: p < 0.001).
In conclusion, these results showed that the DotCom complex and notably the Grappa sub-unit that encodes a H3K79 histone methyl-transferase are involved in the establishment of abdominal pigmentation but also in some essential developmental mechanisms.
Gpp plays opposite roles on pigmentation depending on the developmental stage
The major difference between the two drivers used to down-regulate gpp is the developmental stage at which they are expressed. Therefore, the opposite effects of gpp down-regulation on pigmentation revealed by these drivers could reflect the distinct roles played by gpp at different developmental stages. To test this hypothesis, we used a temperature-sensitive Gal80 repressor, Gal80ts, in combination with the pnr-Gal4 driver. This system allowed to repress gpp at different developmental stages. The UAS-RNAi-gpp was activated by shifting tubes containing developing flies from 18 to 29 °C between day 10 of development (L2 stage) and day 21 (young adult stage), and quantifying the A6 segment pigmentation in adult females. As temperature itself affects pigmentation, control flies were treated identically. Between days 10 and 12, gpp down-regulation induced an increase of pigmentation in A6, similarly to the phenotype observed with pnr-Gal4. By contrast, between days 17 and 19, a decrease of pigmentation was observed, similar to the phenotype induced by y-Gal4 (Fig. 3, Supplementary table 2). These results show that gpp is involved in different ways in pigmentation establishment during development, regulating the gene network involved in melanin production negatively during larval development and positively during the second half of the pupal life.
Gpp plays opposite roles on pigmentation depending on the developmental stage. (A) Quantification of A6 pigmentation in UAS-RNAi-gpp flies driven by Gal80ts; pnr-Gal4. The RNAi transgene was activated by shifting a pool of developing flies from 18 to 29 °C each day, from day 10 (L2 stage) to day 21 (adult stage). n = 30 per condition, error bars correspond to standard deviations, t-test; ***: p < 0.001. (B) Abdominal cuticles of Gal80ts; pnr-Gal4 control and RNAi-gpp females with the corresponding day of activation of the RNAi transgene.
Gpp regulates different pigmentation genes depending on the developmental stage
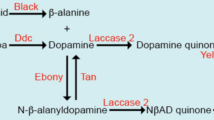
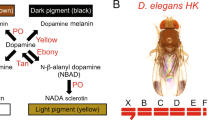
As Gpp regulates melanin production, we aimed to identify the pigmentation genes targeted, directly or indirectly, by Gpp. The pigmentation genes encode the enzymes involved in the cuticular pigment synthesis pathway, which is very dynamic. These enzymes act sequentially to produce the different pigments of the cuticle (brown and black melanin and yellow NßAD sclerotin) (Fig. 4A)9. Modification in melanin production could result from a change in expression of genes acting early in the pathway, such as TH or DDC. Alternatively, it could be caused by a deregulation of genes acting downstream, involved in the balance between yellow pigments and melanins, such as tan, ebony and yellow. We thus performed RT-qPCR experiments on A5, A6 and A7 epidermes of young RNAi-gpp females driven by y-Gal4 or pnr-Gal4 to measure the expression of the genes encoding the enzymes from the pigment synthesis pathway (Fig. 4B,C, Supplementary File 2, Supplementary Table 3). With the pnr-Gal4 driver, no significant difference in pigmentation gene expression was observed (Fig. 4B). This could be due to the fact that, pnr being expressed only around the dorso-longitudinal line, the tissue expressing pnr was too diluted in the total abdominal epidermis. To bypass this issue, we observed the expression of nEGFP transgenes reporting the expression of the pigmentation genes yellow31, ebony32 and tan33 in UAS-RNAi-gpp flies driven by pnr-Gal4 (Fig. 5). No change of GFP expression was observed for tan and ebony reporters (Fig. 5A,B). In contrast, for the yellow reporter, GFP expression increased in the expression domain of pnr compared to controls (Fig. 5C). The ratio of GFP intensity in the pnr domain to that in the lateral region (used as an internal control) was 1.76 times higher in pnr-Gal4 > RNAi gpp flies than in pnr-Gal4 control flies (Fig. 5D; Supplementary Table 4). yellow being involved in production of black melanin34, this observation is consistent with the pigmentation phenotype of pnr-Gal4 > RNAi-gpp flies. Therefore, Gpp represses yellow expression.
Gpp activates the expression of the pigmentation gene tan. (A) Synthesis pathway of cuticular pigments9. The enzymes are highlighted in red. (B,C) RT-qPCR quantification of pigmentation genes, encoding these enzymes, and bab1 expression in posterior abdomen epidermes (A5, A6 and A7 segments) of young RNAi-gpp females driven by pnr-Gal4 (B) or y-Gal4 (C) (n = 3, error bars correspond to standard deviations, t-test; ***: p < 0.001). The geometric mean of RP49 and Spt6 reference gene expression was used for normalization.
Gpp represses the expression of the pigmentation gene yellow. In (A–C), the dashed white lines represent the right hemi-segments. The dashed red line marks the limit between the dorsal pnr-Gal4 expression domain (to the left of the line) and the lateral region (to the right of the line) used as an internal control. (A,B) Effect of gpp down-regulation on tan (A) and ebony (B) nEGFP reporters. Compared to the control pnr-Gal4 (left), gpp down-regulation (right) has no effect on the expression of nEGFP driven neither by tan nor by ebony. (C) Effect of gpp down-regulation on the yellow nEGFP reporter transgene. Compared to the control pnr-Gal4 (left), gpp down-regulation (right) increases the expression of nEGFP, which extends more anteriorly in the pnr-Gal4 expression domain. (D) Computation of the ratio of the mean intensity of yellow nEGFP in the pnr domain over that in the lateral region (used as internal control) for pnr-Gal4 and pnr-Gal4 > RNAi-gpp. Error bars correspond to standard deviations. t-test; ****: p < 0.001.
With the y-Gal4 driver, the expression of tan, involved in production of brown and dark melanin, decreased, which is consistent with the pigmentation phenotype of y-Gal4 > RNAi-gpp flies (Fig. 4C). Therefore, Gpp activates tan expression. We have shown previously that the Bric-à-Brac transcription factors (Bab1 and Bab2) participate in the establishment and plasticity of pigmentation by repressing tan35. In order to test whether Gpp could activate tan through bab regulation, we quantified bab1 expression (Fig. 4B,C). No significant change in bab1 expression was observed, suggesting that Gpp regulates tan independently of Bab.
The DotCom complex is involved in thermal plasticity of abdominal pigmentation
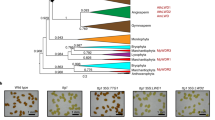
The DotCom complex being involved in pigmentation establishment, we then wondered whether it also participates in the regulatory network mediating thermal plasticity of abdominal pigmentation. To answer to this question, as thermal sensitivity of the UAS/Gal4 system prevented the use of RNAi lines, we studied a double heterozygous loss-of-function mutant for gpp (gppBG00006) and alh (alhJ8C8) to strongly destabilize the DotCom complex. Flies were grown at 18 °C, 25 °C and 29 °C. Their pigmentation was quantified in A5, A6 and A7 and compared to those of control flies (Fig. 6, Supplementary Table 5, Supplementary File 3). Pigmentation of both mutant and control females were sensitive to temperature (“T effect”). The mutant females were significantly lighter than the control females (“G effect”) for A5 but not for A6 and A7. A very strong interaction, visible as the curves of the two genotypes crossed each other, was observed between genotype and temperature (“T x G effect”) for A6 and A7 and to a lesser extent for A5. This significant interaction between genotype and temperature revealed that thermal plasticity was modified in gppBG0000/alhJ8C8 females as compared to control females. These results show that the DotCom complex participates in thermal plasticity of abdominal pigmentation.
The DotCom complex plays a role in the thermal plasticity of abdominal pigmentation. Quantification of A5, A6 and A7 pigmentation of gppBG00006/alhJ8Cmutant and control females raised at 18 °C, 25 °C or 29 °C (n = 30 per condition, error bars correspond to standard deviations). G: effect of genotype; T: effect of temperature; T x G: effect of the interaction between genotype and temperature. ANOVA on aligned rank transformed data was performed for A5. ANOVA were performed for A6 and A7 segments. * p < 0.05; ***: p < 0.001. NS: non significant.
Discussion
We show here that the DotCom complex is involved in the establishment and thermal plasticity of the abdominal pigmentation of Drosophila melanogaster females. Interestingly, Gpp, the histone methyl-transferase of the complex, participates in opposite ways in the establishment of pigmentation depending on the developmental stage. Indeed, when gpp is down-regulated from the L2 to the L3 stages, melanin production is higher, whereas when it is down-regulated later, during the second half of the pupal life, melanin production is lower. These two effects on pigmentation are mediated by the regulation of two pigmentation genes involved in melanin production, i.e. yellow for the early effect and tan for the late effect. Thus, Gpp regulates different targets during the establishment of female D. melanogaster abdominal pigmentation. It is likely that yellow is not a direct target of Gpp. Indeed, yellow starts to be expressed in the late pupal stage. However, overexpression of the yellow GFP reporter is observed when gpp RNAi is induced with the early driver pnr-Gal4, i.e. prior to expression of the endogenous gene. In addition, temperature shift experiments showed that the effect on pigmentation of gpp down-regulation driven by pnr-gal4 is determined during the larval life. Concerning tan, we observed no significant reduction of the H3K79me3 level on its promoter, second exon and enhancer in y-Gal4 > RNAi-gpp females (Supplementary Fig. 2, Supplementary File 4, Supplementary table 3), suggesting that, like yellow, tan is not directly regulated by Gpp. In contrast, we have shown previously that the histone methyl-transferase Trx participates in the deposition of the H3K4me3 active mark on tan9. The opposite roles of Gpp through the regulation of distinct pigmentation genes could explain its involvement in the thermal plasticity of abdominal pigmentation. Indeed, the expression of yellow and tan is modulated by temperature9,10. Temperature might therefore modulate the balance between the repression of yellow and the activation of tan by Grappa. This is reminiscent of Abdominal-B who plays also opposite roles on cuticle pigmentation depending on the temperature36.
In conclusion, our results clearly show that the DotCom complex and its histone methyl-transferase Gpp are involved in the establishment and the plasticity of abdominal pigmentation, through the regulation of yellow and tan pigmentation genes. However, the direct targets of this complex mediating this effect in the abdominal epidermis remain unknown. They could be identified by performing ChIP-seq experiments to study the level of H3K79 methylation in the abdominal epidermis of control and gpp RNAi flies genome-wide. Moreover, we cannot formally exclude the possibility that the DotCom complex acts independently from its histone methyl-transferase activity in this process. To address this question, we could study mutant flies that lack the catalytic activity of Gpp.
In addition, mechanistic insights could be gained by the study of Bre1 and MOF, depositing H2BK123ub and H4K16ac respectively, that facilitate H3K79 methylation by Gpp. Furthermore, a H3K79 demethylase, KDM2B, has recently been characterized in human cells37. It would be interesting to identify the Drosophila homologue of KDM2B and test its role on the establishment and plasticity of abdominal pigmentation.
Material and methods
Fly stocks
A w1118 inbred line was used as control. The pnr-Gal4 (BL-3039)29, y-Gal4 (BL-44267), UAS-RNAi-alh (BL-39057), UAS-RNAi-gpp1 (BL-42919), UAS-RNAi-gpp2 (BL-34842), UAS-RNAi-ear (BL-28068), UAS-RNAi-NippedA (BL-34849), alhJ8C8 (BL-12118) and gppBG00006 (BL-12802) lines were from the Bloomington Stock Center. In order to be tested in the same genetic background than the w1118 control, alhJ8C8 and gppBG00006 alleles were introgressed in w1118 for 10 generations by following eye colour, as these two alleles are marked by mini-white. The Gal80ts-Gal4 transgene was previously described9. The efficiency of the UAS-RNAi-gpp1 transgene was validated by RT-qPCR (Supplementary Fig. 3, Supplementary File 5, Supplementary table 3). The t_MSE-nEGFP, ebony-nEGFP (ebony ABC + intron) and yellow-nEGFP (yellow-wing-body-nEGFP) reporter transgenes were previously described31,32,33.
Quantification of gene expression by RT-qPCR
Total RNA were extracted from pools of 30 dissected posterior abdominal epidermes of females (A5, A6 and A7) with the RNAeasy Mini kit (Qiagen). Three replicates were performed in all experiments. cDNAs were synthesized with the LunaScript RT SuperMix kit (New England Biolabs) using random primers. RT-qPCR experiments were performed in a CFX96 system using SsoFast EvaGreen SuperMix (Biorad). Expression was quantified following the Pfaffl method38 using the geometric mean of the expression of RP49 and Spt6 reference genes for normalization39. All primers are listed in Supplementary table 3.
Cuticle and epidermis preparations
5 days old adult females were stored in ethanol 70% during ten days. Abdominal cuticles were cut just beyond the dorsal midline and dehydrated in ethanol 100% during 5 min. After dehydration, cuticles were mounted in Euparal (Roth). For nEGFP observation, pupal and adult abdomens were dissected in PBS, fixed 20 min in 3.7% paraformaldehyde in PBS, washed three times 10 min in PBS, and mounted in Mowiol®.
Image acquisition and GFP quantification
Cuticles were imaged as described in35. Abdominal epidermes of females expressing nEGFP under the control of t_MSE, yellow and ebony regulatory sequences were imaged using a macro-apotome (Zeiss). Young adult females were imaged for t_MSE-nEGFP and ebony-nEGFP and female pupae for yellow-nEGFP as these genes are not expressed at the same developmental stage. The mean intensity of yellow-nEGFP was measured using ImageJ. Larvae showing H2B-GFP under the control of pnr-Gal4 and yellow-Gal4 were imaged with a binocular equipped with a Leica DC480 digital camera using the Leica IM50 Image Manager software. Pharates showing H2B-GFP under the control of pnr-Gal4 and yellow-Gal4 were imaged using an Olympus BX41 fluorescence microscope (objective 4 X) equipped with a Yokogawa spinning disc and a CoolSnapHQ2 camera controlled by Metaview software (Universal Imaging).
Chromatin immunoprecipitation experiments
Chromatin immunoprecipitation (ChIP) experiments were performed as previously described9, with 3 µg of each antibody. Rabbit IgGs (Diagenode) as negative control and anti-H3K79me3 (C15410068, Diagenode) were used. Three replicates of 50 abdominal epidermes (A5, A6 and A7) of young females were analyzed. qPCR were performed on a CFX96 system using the Luna Universal qPCR Master Mix (BioLabs). Primers are listed in Supplementary table 3. Data were normalized against input chromatin.
Statistical analyses
In Figs. 2, 3, 4 and 5, pigmentation intensity, gene expression quantifications and GFP intensity ratios were compared by t-tests using the appropriate parameters depending on homogeneity of variances as assessed with Levene tests. In Fig. 1, as two RNAi lines fo gpp were used, we performed ANOVAs on aligned rank transformed data followed by Tukey post-hoc tests (Supplementary File 1). In Fig. 6, to analyse the effect of alh/gpp double mutants on pigmentation plasticity, a two-way ANOVA with genotype and temperature as factors was used. Variance homogeneity was checked using Levene tests and normality of residuals using Shapiro–Wilk tests. When variances were not homogeneous or residues not normally distributed, a non-parametric ANOVA on aligned rank transformed data was performed. All ANOVA tests were performed using RStudio (Supplementary File 1; Supplementary File 3).
Data availability
All data generated or analysed during this study are included in this published article [and its supplementary information files].
References
Pigliucci, M. Phenotypic Plasticity (JHU Press, Beyond Nature and Nurture, 2001).
Gunter, H. M., Schneider, R. F., Karner, I., Sturmbauer, C. & Meyer, A. Molecular investigation of genetic assimilation during the rapid adaptive radiations of East African cichlid fishes. Mol. Ecol. 26, 6634–6653 (2017).
Aubret, F. & Shine, R. Genetic assimilation and the postcolonization erosion of phenotypic plasticity in island tiger snakes. Curr. Biol. 19, 1932–1936 (2009).
Heil, M. et al. Evolutionary change from induced to constitutive expression of an indirect plant resistance. Nature 430, 205–208 (2004).
Corl, A. et al. The genetic basis of adaptation following plastic changes in coloration in a novel environment. Curr. Biol. CB 28, 2970-2977.e7 (2018).
Gibert, J.-M. The flexible stem hypothesis: Evidence from genetic data. Dev. Genes Evol. 227, 297–307 (2017).
West-Eberhard, M. J. Developmental Plasticity and Evolution (Oxford University Press, 2003).
Gibert, P., Moreteau, B. & David, J. R. Developmental constraints on an adaptive plasticity: Reaction norms of pigmentation in adult segments of Drosophila melanogaster. Evol. Dev. 2, 249–260 (2000).
Gibert, J.-M., Mouchel-Vielh, E., De Castro, S. & Peronnet, F. Phenotypic plasticity through transcriptional regulation of the evolutionary hotspot gene tan in Drosophila melanogaster. PLoS Genet. 12, e1006218 (2016).
Gibert, J.-M., Mouchel-Vielh, E. & Peronnet, F. Modulation of yellow expression contributes to thermal plasticity of female abdominal pigmentation in Drosophila melanogaster. Sci. Rep. 7, 43370 (2017).
True, J. R. et al. Drosophila tan Encodes a Novel Hydrolase Required in Pigmentation and Vision. PLoS Genet 1, e63 (2005).
Simola, D. F. et al. Epigenetic (re)programming of caste-specific behavior in the ant Camponotus floridanus. Science 351, aac6633 (2016).
Bonasio, R. et al. Genome-wide and caste-specific DNA methylomes of the ants Camponotus floridanus and Harpegnathos saltator. Curr. Biol. 22, 1755–1764 (2012).
Leung, A. et al. Open chromatin profiling in mice livers reveals unique chromatin variations induced by high fat diet. J. Biol. Chem. https://doi.org/10.1074/jbc.M114.581439 (2014).
Lacoste, N., Utley, R. T., Hunter, J. M., Poirier, G. G. & Côte, J. Disruptor of telomeric silencing-1 is a chromatin-specific histone H3 methyltransferase. J. Biol. Chem. 277, 30421–30424 (2002).
Shanower, G. A. et al. Characterization of the grappa gene, the Drosophila histone H3 lysine 79 methyltransferase. Genetics 169, 173–184 (2005).
Ng, H. H. et al. Lysine methylation within the globular domain of histone H3 by Dot1 is important for telomeric silencing and Sir protein association. Genes Dev. 16, 1518–1527 (2002).
van Leeuwen, F., Gafken, P. R. & Gottschling, D. E. Dot1p modulates silencing in yeast by methylation of the nucleosome core. Cell 109, 745–756 (2002).
Jones, B. et al. The histone H3K79 methyltransferase Dot1L is essential for mammalian development and heterochromatin structure. PLoS Genet. 4, e1000190 (2008).
Steger, D. J. et al. DOT1L/KMT4 recruitment and H3K79 methylation are ubiquitously coupled with gene transcription in mammalian cells. Mol. Cell. Biol. 28, 2825–2839 (2008).
Wood, K., Tellier, M. & Murphy, S. DOT1L and H3K79 methylation in transcription and genomic stability. Biomolecules 8, 11 (2018).
Malla, A. B. et al. DOT1L bridges transcription and heterochromatin formation at mammalian pericentromeres. EMBO Rep. 24, e56492 (2023).
Schübeler, D. et al. The histone modification pattern of active genes revealed through genome-wide chromatin analysis of a higher eukaryote. Genes Dev. 18, 1263–1271 (2004).
Nakanishi, S. et al. Histone H2BK123 monoubiquitination is the critical determinant for H3K4 and H3K79 trimethylation by COMPASS and Dot1. J. Cell Biol. 186, 371–377 (2009).
Valencia-Sánchez, M. I. et al. Structural basis of Dot1L stimulation by histone H2B lysine 120 ubiquitination. Mol. Cell 74, 1010-1019.e6 (2019).
Anderson, C. J. et al. Structural basis for recognition of ubiquitylated nucleosome by Dot1L methyltransferase. Cell Rep. 26, 1681-1690.e5 (2019).
Valencia-Sánchez, M. I. et al. Regulation of the Dot1 histone H3K79 methyltransferase by histone H4K16 acetylation. Science 371, eabc6663 (2021).
Mohan, M. et al. Linking H3K79 trimethylation to Wnt signaling through a novel Dot1-containing complex (DotCom). Genes Dev. 24, 574–589 (2010).
Calleja, M. et al. Generation of medial and lateral dorsal body domains by the pannier gene of Drosophila. Development 127, 3971–3980 (2000).
Wittkopp, P. J., True, J. R. & Carroll, S. B. Reciprocal functions of the Drosophila yellow and ebony proteins in the development and evolution of pigment patterns. Development 129, 1849–1858 (2002).
Jeong, S., Rokas, A. & Carroll, S. B. Regulation of body pigmentation by the Abdominal-B Hox protein and its gain and loss in Drosophila evolution. Cell 125, 1387–1399 (2006).
Rebeiz, M., Pool, J. E., Kassner, V. A., Aquadro, C. F. & Carroll, S. B. Stepwise modification of a modular enhancer underlies adaptation in a Drosophila population. Science 326, 1663–1667 (2009).
Jeong, S. et al. The evolution of gene regulation underlies a morphological difference between two Drosophila sister species. Cell 132, 783–793 (2008).
Riedel, F., Vorkel, D. & Eaton, S. Megalin-dependent yellow endocytosis restricts melanization in the Drosophila cuticle. Development 138, 149–158 (2011).
De Castro, S., Peronnet, F., Gilles, J.-F., Mouchel-Vielh, E. & Gibert, J.-M. bric à brac (bab), a central player in the gene regulatory network that mediates thermal plasticity of pigmentation in Drosophila melanogaster. PLoS Genet. 14, e1007573 (2018).
Gibert, J. M., Peronnet, F. & Schlötterer, C. Phenotypic plasticity in Drosophila pigmentation caused by temperature sensitivity of a chromatin regulator network. PLoS Genet. 3, e30 (2007).
Kang, J.-Y. et al. KDM2B is a histone H3K79 demethylase and induces transcriptional repression via sirtuin-1-mediated chromatin silencing. FASEB J. Off. Publ. Fed. Am. Soc. Exp. Biol. 32, 5737–5750 (2018).
Pfaffl, M. W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 29, e45 (2001).
Vandesompele, J. et al. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 3, RESEARCH0034 (2002).
Acknowledgements
We thank all members of the Gibert/Peronnet team for stimulating discussions and Frédérique Peronnet for critical reading of the manuscript. We thank Agnes Audibert for her help in acquiring the images of pharates expressing H2B-GFP under the control of pnr-Gal4 and y-Gal4.
Funding
The project was funded by CNRS and Sorbonne Université. RN was funded by a PhD fellowship from the Ministère de l’Enseignement Supérieur et de la Recherche. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
E.M.V. and J.M.G. jointly supervised the project. R.N. performed experiments and acquired data. R.N., E.M.V. and J.M.G. analysed and interpreted data. R.N., E.M.V. and J.M.G. wrote the article.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interest.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Narbey, R., Mouchel-Vielh, E. & Gibert, JM. The H3K79me3 methyl-transferase Grappa is involved in the establishment and thermal plasticity of abdominal pigmentation in Drosophila melanogaster females. Sci Rep 14, 9547 (2024). https://doi.org/10.1038/s41598-024-60184-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-60184-6
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.