Abstract
DNA damage response (DDR) pathways are responsible for repairing endogenous or exogenous DNA damage to maintain the stability of the cellular genome, including homologous recombination repair (HRR) pathway, mismatch repair (MMR) pathway, etc. In ovarian cancer, current studies are focused on HRR genes, especially BRCA1/2, and the results show regional and population differences. To characterize germline mutations in DDR genes in ovarian cancer in Southwest China, 432 unselected ovarian cancer patients underwent multi-gene panel testing from October 2016 to October 2020. Overall, deleterious germline mutations in DDR genes were detected in 346 patients (80.1%), and in BRCA1/2 were detected in 126 patients (29.2%). The prevalence of deleterious germline mutations in BRCA2 is higher than in other studies (patients are mainly from Eastern China), and so is the mismatch repair genes. We identified three novel BRCA1/2 mutations, two of which probably deleterious (BRCA1 p.K1622* and BRCA2 p.L2987P). Furthermore, we pointed out that deleterious mutations of FNACD2 and RECQL4 are potential ovarian cancer susceptibility genes and may predispose carriers to ovarian cancer. In conclusion, our study highlights the necessity of comprehensive germline mutation detection of DNA damage response genes in ovarian cancer patients, which is conducive to patient management and genetic counseling.
Similar content being viewed by others
Introduction
Ovarian cancer is the third most common gynecological malignancy, accounting for 3.4% of all malignancies and 4.7% cancer deaths among women worldwide in 20201. In China, ovarian cancer disproportionately affects women's health, with approximately 57,090 newly diagnosed cases and 39,306 cancer-related deaths in 2022, showing an upward trend2. Despite a gradual decline in the incidence and mortality of ovarian cancer in the United States, the mortality rate still ranks highest among malignancies of the female reproductive system, posing a significant threat to women's health3. As for now, most ovarian cancer patients have local or distant dissemination at the time of diagnosis due to the lack of effective screening and early diagnosis methods. The prognosis of ovarian cancer is poor, with limited improvement in the long term. In recent years, the in-depth understanding of the genomic characteristics of ovarian cancer has brought breakthroughs to the treatment of patients, especially the development of anti-angiogenesis drugs and poly (ADP-ribose) polymerase (PARP) inhibitors.
Part of ovarian cancer is known to be heritable, accompany with inherited deleterious gene mutations. Genetic alteration in HRR genes may confer increased risk of multiple genetically-related malignancies, such as ovarian cancer, breast cancer, pancreatic cancers, especially BRCA1 and BRCA24,5. HRR is an error-free pathway for DNA double strands break repairing, and BRCA1/2 are the core genes of this pathway. Thus, germline mutations in HRR genes may lead to errors accumulation and predispose to malignancies such as ovarian cancer. By the age of 70, the average cumulative risks of ovarian cancer is estimated to be 60% for women with deleterious germline BRCA1 mutations, and 16.5% for women with deleterious germline BRCA2 mutations, both significantly higher than normal people6. Among Chinese ovarian cancer patients, the prevalence of deleterious germline BRCA1/2 mutations is about 16.7–28.5% in different studies7,8,9. Germline mutations in other HRR genes may also elevate the risk of ovarian cancer, such as RAD51C, RAD51D and BRIP110. The life time risk of ovarian cancer associated with these genes mutations is approximately 6–9%11,12. Meanwhile, the deficient in MMR pathway genes predispose carriers to a variety of cancers, including ovarian cancer. The estimated cumulative risks of ovarian cancer for MLH1 mutation carriers and MSH2 carriers at age 70 is 20% and 24% respectively13. In addition, attention should also be paid to germline mutations in other susceptibility genes associated with hereditary cancer syndromes, such as Li-Fraumeni syndrome (TP53), Cowden syndrome (PTEN) and Peutz-Jeghers syndrome (STK11)10,14,15,16.
Ovarian cancer patients carrying mutations in BRAC1/2 or other HRR genes may benefit from platinum-based chemotherapy and PARP inhibitors. Patients with detected BRCA1/2 mutations have a higher response rate to platinum-based chemotherapy and improved median overall survival17. Meanwhile, the presence of mutations in other HRR genes (RAD51C, RAD51D, BRIP1, ATM, etc.) may also predict favorable prognosis when receiving platinum-based chemotherapy18. In addition, patients carrying mutations in BRAC1/2 or certain HRR genes have improved progression-free survival and overall survival with PARP inhibitors therapy19,20,21,22. A positive test result may also have important implications for family members, including future generations. Once a BRCA1 or BRCA2 germline pathogenic variant has been identified in a family, testing of at-risk relatives can identify those family members who also have the familial pathogenic variant and thus need increased surveillance and specific treatments when a cancer is identified.
Although research in cancer genomics and cancer genetics over the past decade had promoted our understanding of pathogenesis and revolutionized therapies of ovarian cancer, more aspects are yet to be defined. Here, we retrospectively analyzed the germline gene mutations of consecutive ovarian cancer patients in our center (The largest gynecological cancer center in Western China has treated 450–500 newly diagnosed ovarian cancer patients in recent five years) by multi-gene panel detection in BGI genomics, containing HHR, MMR, Fanconi anemia pathway and other DNA damage response genes. The results presented here will expand our knowledge of the impact of germline mutations on heredity and treatment of ovarian cancer.
Results
Patient characteristics
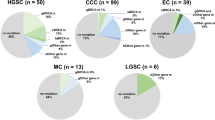
A total of 432 patients were tested for germline gene mutations using multi-gene panel. The characteristics of all patients are shown in Table 1 and Supplementary Table S1. The enrolled patients spanned an age range of 20 to 78 years, with a mean age of 52 years (Table 1). BRCA deleterious mutations (defined as likely pathogenic/pathogenic mutations) were detected in 126 patients, and among them 41 patients (32.5%) had family history of cancer, including ovarian cancer, breast cancer, colorectal cancer, gastric cancer, etc. In contrast, 71 patients (23.2%) had family history without BRCA deleterious mutations (defined as wild type/benign/likely benign mutations). At diagnosis, the median age of all patients was 52 years, with a range of from 20 to 78 years (Table 1 and Supplementary Fig. S1). Among the patients studied, 63% of them were identified as high-grade serous carcinoma (HGSC), while 37% were classified as non-HGSC (Table 1).
Overall genetic mutation landscape
Overall, deleterious germline mutations were identified in 346 patients (80.1%), suggesting that deleterious germline mutations in DDR genes are common in ovarian cancer. Moreover, at least one deleterious germline HRR gene mutation was identified in 240 patients (55.6%) and more than one in 80 patients. The most deleterious mutations observed were in BRCA1 (18.5%), BRCA2 (12.5%), ERCC5 (6%), MLH1 (5.8%), PALB2 (3.7%) and ERCC4 (3.7%) 0.155 patients (35.9%) had deleterious mutations in germline HRR genes other than BRCA1/2. Germline MMR genes deleterious mutations were identified in some patients, including MLH1 mutations in 25 patients (5.8%), MSH6 mutations in 6 patients (1.4%), MSH2 mutations in 4 patients (0.9%) and PMS2 mutations in 3 patients (0.7%). Notably, HGSC demonstrated a significantly higher incidence of pathogenic BRCA1 mutations (P-value < 0.005), while non-HGSC was characterized by a higher frequency of pathogenic NCOA4 mutations (P-value < 0.006).
When considering all the mutations, not only deleterious, FANCD2 (47%), BRCA1 (27%), BRCA2 (27%), ERCC5 (18%) and RECQL4 (17%) were the most frequently mutated genes (Fig. 1). The frequency of missense mutation was higher than them of other mutation types, except for a few genes. FANCD2 had high frequency of splice site mutation, and TSC2 and PHOX2B had high frequency of in frame deletion. High frequent mutations were enriched in DNA damage repair signal pathway, such as Fanconi anemia, HRR, nucleotide excision repair (NER), MMR and cell cycle pathway (Supplementary Table S2). And 88.2% patients were implicated by at least one Fanconi anemia pathway gene mutation.
Using the pair-wise Fisher's Exact test to explore the co-occurrence and mutually exclusive interaction of mutations between genes, it was found that the mutations in ALK and BARD1 were significantly exclusive each other in the same patient, as well as KIF1B and MSH6; whereas, the mutations in the FH and MLH1, along with BARD1 and PMS2, were significantly co-occurring in the same patient (Supplementary Fig. S2).
BRCA1/2 mutation sites and novel mutations
After comprehensively analyzing the mutation sites of BRCA1 and BRCA2, we found that the incidence of BRCA1 c.2566 T > C was significantly higher than that of other mutations, converting tyrosine at 856 site to histidine (Fig. 2A). And when analyzed in population mutation frequency database, BRCA1 c.2566 T > C was significantly enriched in Asian population (Fig. 3A). In the Chinese population, BRCA1 c.2566 T > C was more frequent in ovarian cancer patients than healthy controls (Fig. 3B). Meanwhile, BRCA2 c.8187G > T and c.10234A > G had remarkably higher frequency than other mutated sites, converting lysine at 2729 site to asparagine and isoleucine at 3412 site to valine (Fig. 2B). The frequency of BRCA2 c.8187G > T was specifically higher in Asian population than other mutated sites and moderately higher in Chinese ovarian cancer patients than healthy people (Fig. 3C,D). BRCA2 c.10234A > G was a high-frequency mutation in African and American populations (Fig. 3E). The frequency of BRCA2 c.10234A > G observed in Chinese ovarian cancer patients was 1.64 times higher than that of in normal people (Fig. 3F).
BRCA1/2 founder mutations in in different population. (A) Frequency of BRCA1 c.2566 T > C in different population. (B) Frequency of BRCA1 c.2566 T > C in Chinese ovarian cancer patients and normal people. (C) Frequency of BRCA2 c.8187G > T in different population. (D) Frequency of BRCA2 c.8187G > T in Chinese ovarian cancer patients and normal people. (E) Frequency of BRCA2 c.10234A > G in different population. (F) Frequency of BRCA2 c.10234A > G in Chinese ovarian cancer patients and normal people. AFR African/African American, AMR Latino/Admixed American, ASJ Ashkenazi Jewish, EAS East Asian, FIN European (Finnish), NFE European (non-Finnish), OTH Other.
In our study cohort, we confirmed three novel germline mutations in BRCA1/2 in ten patients that are not reported in public databases and literature (Supplementary Table S3). Two of ten patients had a family history of cancers. BRCA1 p.M1649T, p.K1622* and BRCA2 p.L2987P was identified in five patients, three patients and two patients, respectively. BRCA1 p.M1649T and BRCA2 p.L2987P were missense mutations, while BRCA1 p.K1622* was a nonsense mutation. We predicted whether these novel germline mutations were deleterious using SIFT23 and PolyPhen224. BRCA1 p.M1649T was not a deleterious mutation, but BRCA2 p.L2987P was. However, the software-predicted pathogenicity of BRCA1 p.K1622* was unknown. Considering BRCA1 p.K1622*truncated the protein, we think it is probably a deleterious mutation.
High frequency FANCD2 mutation
FANCD2 was the most mutated gene in our data, with 73.3% of patients carried four adjacent cis mutations c.1275C > T, c.1278 + 1delG, c.1278 + 3_1278 + 5delAAG and c.1278 + 15C > T, and all of these carries were heterozygous. FANCD2 c.1275C > T was a silent mutation didn’t alter FANCD2 protein sequence, and the other three mutations affected intron 16 of FANCD2. In ClinVar database, FANCD2 c.1275C > T, c.1278 + 3_1278 + 5delAAG and c.1278 + 15C > T, were labeled as benign mutations, while c.1278 + 1delG had no clinically reported significance. As GT dinucleotides were the most common 5’ splice site of intron, deletion of the first guanine would lead to splicing at wrong site. Thus, c.1278 + 1delG may dramatically impact the final transcript and function of FANCD2. Although these adjacent mutations alone were detected and reported in population database, cis mutation type of the four mutations were not reported and they may have additive effect to the function of FANCD2. We combined them as FANCD2 span mutation c.1275_1278 + 15delCTTAGTAAGTGTCAGAGACinsTTTATTGTCAGAGAT, which spanned both the exon and intron. According to the ACMG guidelines for the interpretation of sequence variants, this FANCD2 span mutation was likely pathogenic. We applied fit Chi-square calculation to evaluate the mutation with ovarian cancer risk and found the mutation of FANCD2 that significantly fails to conform Hardy–Weinberg equilibrium (P-value 2.76e − 06).
Mutation characteristic of subgroups with different ages at diagnosis
Furthermore, we investigated the association between germline mutations and age at diagnosis. In all patients, BRCA1 and CHEK2 mutations were more common in younger patients, whereas ATM, MEN1, FANCE and NTRK1 were mutated more in older patients (Fig. 4A). Germline mutations in BRCA1 and CHEK2 may lead to homologous recombination repair deficiency and cell cycle disorder, eliciting cancer at an early age. Then, we focused on deleterious mutations of BRCA1 and BRCA2, the most crucial genes increasing the risk of ovarian and breast cancer. Among the 336 patients carrying deleterious mutations, BRCA1 was with higher frequency in younger patients, while BRCA2 was moderately higher in older patients (Fig. 4B,C). We also observed that patients with FANCD2 span mutation were slightly younger than those without FANCD2 mutations, although not significantly, suggesting that FANCD2 span mutation may be associated with early onset of ovarian cancer (Fig. 4D). When analysis based on different signal pathways, patients with HRR deleterious mutations were younger than them with MMR deleterious mutations, meanwhile younger than them without HRR and MMR deleterious mutations (also named “wild type”) (Fig. 4E).
Mutation characteristic different age groups. (A) Significant mutant genes in ovarian cancer patients with different ages (separated by 52 years old). (B) Frequency of BRCA1/2 deleterious mutations in ovarian cancer patients with different ages (separated by 52 years old). (C) Age distribution of patients detected BRCA1/2 deleterious mutations. (D) Age distribution of patients detected FANCD2 mutations. (E) Age distribution of patients detected DNA damage response genes. *Indicates that the P-value is less than 0.05, **Indicates that the P-value is less than 0.01.
Mutation characteristic of subgroups with or without a family history
There are 112 patients with a family history of cancer. In all patients, FANCD2, NQO2 and PRM1D were commonly mutated in patients with a family history, while TSC1 and MSH6 mutations were occurred frequently in patients without a family history (Fig. 5A). When we focused on patients with deleterious mutations, BRCA1 was significantly associated with family history, and BRCA2 was slightly more common in patients without a family history but not significantly (Fig. 5B,C). Moreover, ERCC5, an endonuclease participating the excision repair pathway, was observed more frequently in patients without a family history (Fig. 5B).
Different gene mutations in patients with or without family history. (A) Significant mutated genes. (B) Significant mutated genes in patients carrying deleterious mutations. (C) Significant mutated genes in patients carrying BRCA1/2 deleterious mutations. (D) Significant mutated genes in patients not detected BRCA1/2 deleterious mutations. (E) RECQL4 deleterious mutations. *Indicates that the P-value is less than 0.05, **Indicates that the P-value is less than 0.01.
In patients without BRCA deleterious mutations, RECQL4 deleterious mutations were significantly accompanied with family history (Fig. 5D). In our cohort, RECQL4 deleterious mutations were observed in 5.6% of patients, of which 45.8% was c.212A > G (p.E71G) (Fig. 5E). Thus, RECQL4 deleterious mutations, especially the mutation c.212A > G may predispose carriers to ovarian cancer.
Discussion
The prevalence and spectrum of germline mutations in ovarian cancer vary across population and studies. BRCA1 and BRCA2 are the most studied ovarian cancer susceptibility genes, and are associated with hereditary breast and ovarian cancer syndrome. The reported germline BRCA1/2 mutation prevalence in ovarian cancer is ranged from 5.6% to 29.3%, with the highest frequency in Ashkenazi Jews25. And except for Poland, the frequency of germline BRCA1/2 mutation is relatively low in Northern European countries (Finland, Sweden, Denmark and Iceland), which are less than 10%25. The frequency of germline BRCA1/2 mutation in Chinese ovarian cancer patients is 16.7–28.5%, similar to that in other Eastern Asia countries (14.6% in Japanese and 19.5% in Korean)7,8,9,25. Lynch syndrome is another hereditary cancer syndrome associated with gynecologic cancers, which is owing to the deleterious germline mutations in MMR genes, and underlying approximately 5% of endometrial cancers and 1% of ovarian cancers26,27. In our cohort, at least one deleterious mutation was detected in 346 (80.1%) patients using multi-gene panel. The most observed germline deleterious mutation was BRCA1, with a frequency of 18.5%, which is similar to 13.1–20.8% in other studies7,8,9. However, the frequency of germline BRCA2 deleterious mutation was 12.5%, higher than 3.9–7.6% in other studies, which is probably due to the different geographical distribution of patients7,8,9. The patients in our cohort were mainly from Southwest China, while in other studies they were primarily from Eastern China. BRCA1 pathogenic mutations have been reported to be significantly more prevalent in HGSC7,9, which is consistent with our analysis. Meanwhile, our research revealed a significantly higher prevalence of pathogenic NCOA4 mutations in non-HGSC. The relationship between NCOA4 mutations and ovarian cancer has not yet been thoroughly investigated. Previous study has reported endometrial cancer populations harboring NCOA4 mutations are associated with a more favorable prognosis compared to wild-type28. We also observed higher germline deleterious MMR gene mutations in our study, indicating that lynch syndrome may be more frequently associated with ovarian cancer patients of Southwest China. Thus, we should be concerned more about their personal and family history of lynch syndrome associated cancers, such as endometrial cancer and colorectal cancer.
When analyzing specific mutation sites in BRCA1 and BRCA2, we identified high frequency mutation sites, BRCA1 c.2566 T > C, BRCA2 c.8187G > T and BRCA2 c.10234A > G. They were missense mutations and were not currently considered as deleterious mutations. BRCA1 c.2566 T > C converted tyrosine to histidine. Tyrosine was a non-essential amino acid and required energy to synthesize, while histidine was an essential amino acid mainly acquired from food. Therefore, the BRCA1 mutation, c.2566 T > C, reduced the energy consumption of cancer cells and promoted to proliferation. We identified three novel BRCA1/2 mutations in ten patients, each of which was observed in at least two unrelated patients. Taking into account software prediction, effects on protein function and family history, BRCA1 p.K1622* and BRCA2 p.L2987P may be germline deleterious mutations. Further detailed studies are needed to clarify the impact of these mutations on carriers and ovarian cancer patients.
In addition to the known ovarian cancer high-risk genes, we found that germline mutations in FANCD2 and RECQL4 may be associated with hereditary and early onset of ovarian cancer. FANCD2 is a key protein in Fanconi anemia signal pathway and plays an important role in many aspects of cell life, especially in DNA damage response29. Upon activation, FANCD2 is monoubiquitinated and forms heterodimer with FANCI, leading to signal amplification and downstream repair protein recruitment30. We identified a germline likely pathogenic FANCD2 span mutation, comprised of our adjacent cis mutations. 47% of patients carried FANCD2 mutations, of which 73.3% had the FANCD2 span mutation. FANCD2 span mutation located on a 23 nucleotides fragment across exon and intron, may affect protein function together. And the FANCD2 span mutation may greatly influence protein function due to wrong splice site. Our data also indicated that FANCD2 mutations, especially the FANCD2 span mutation, were related to family history and early onset of ovarian cancer. Moreover, Fanconi anemia signaling pathway is involved in the repair of inter strand crosslink, which may be induced by exposure to environmental mutagens or commonly used chemotherapeutic agents of cancers, such as platinum and furocoumarins31,32. Therefore, FANCD2 mutations potentially affect patients’ prognosis. Over expression of FANCD2 was reported as a strong negative prognostic factor in ovarian cancer, particularly in patients treated with taxane-platinum33.
RECQL4 belongs to RecQ helicase family, which plays important roles in multiple cell life processes, including DNA replication, transcription, DNA repair, and telomere maintenance34,35. RECQL4 mutations are associated with three rare autosomal-recessive syndromes, namely Rothmund–Thomson syndrome, RAPADILINO and Baller–Gerold syndrome35,36. All these three syndromes are correlated to cancer, particularly Rothmund–Thomson syndrome patients with RECQL4 mutations shown an increased risk of osteosarcoma35,36. For Rothmund–Thomson syndrome patients with RECQL4 mutations, the leading cause of death mutations is cancer, whereas in cancer-free patients the life expectancy is normal34. Recent studies also reported that germline deleterious mutations in RECQL4 are associated with predisposition to breast cancer and prostate cancer37,38,39,40,41. Besides, RECQL4 plays a role in HRR of DNA double strand breaks42. Abnormal RECQL4 may affect the prognosis of patients treated with platinum or PARP inhibitors43,44. In our study, germline deleterious mutations in RECQL4 were significantly associated with a family history in patients without BRCA1/2 mutations, suggesting that it may be also related to ovarian cancer susceptibility. In our cohort, germline deleterious mutations in RECQL4 were detected in 24 patients (5.6%), nearly half of which were c.212A > G. RECQL4 c.212A > G mutation was not a known mutation in the three autosomal-recessive syndromes, and its roles in ovarian carcinogenesis requires further investigation45,46.
In conclusion, the results presented here showed the germline DDR gene mutation spectrum of ovarian cancer patients in Southwest China, which is genetically different from the rest of China. We also identified novel BRCA mutations worthy of our further attention. Finally, we found that germline deleterious mutations in FANCD2 and RECQL4 were likely susceptibility genes for ovarian cancer, and we should be more cautious to these two genes.
The observed regional and population disparities in the prevalence of deleterious germline mutations, particularly in BRCA2 and mismatch repair genes, emphasize the significance of regional genetic variations in ovarian cancer (OC) susceptibility. This study highlights the necessity for comprehensive germline mutation testing in DNA damage response (DDR) genes for OC patients, enhancing patient management and genetic counseling. It advocates for healthcare providers to extend their genetic assessments to a broader spectrum of DDR genes, not limited to BRCA1/2. Future research should explore the functional implications of newly identified mutations and their influence on OC risk. Moreover, expansive studies across diverse populations are essential to validate the observed regional and population variances in DDR gene mutations, potentially guiding personalized treatments and better outcomes for OC patients.
Methods
Participants
We retrospectively and unselectively enrolled a cohort of patients diagnosed with advanced epithelial ovarian cancer from October 2016 to October 2020. All enrolled patients were required to meet the following criteria: (1) A pathologically confirmed diagnosis via frozen section or paraffin section or ascites cells by two expert pathologists, meeting the pathological diagnosis criteria for EOC (including serous carcinoma, endometrioid carcinoma, clear cell carcinoma, mucinous carcinoma, undifferentiated carcinoma, and carcinosarcoma); (2) Advanced radiographic imaging (CT, MRI, PET) results met primary EOC characteristics; (3) Sufficient medical history and demographic data; (4) Not undergoing comprehensive staging surgery/tumor cell debulking surgery, chemotherapy, radiotherapy, targeted therapy. Exclusion criteria included: (1) Concurrent other malignancies; (2) Incomplete clinical and pathological data; (3) Other pathological types of ovarian cancer (or non-EOC). Clinical information (age, family history, and personal history), pathology information (pathological diagnosis, tumor FIGO stage), were captured and summarized. All Eligible patients had signed informed consent for gene testing. In total, peripheral blood samples from 432 patients were collected for evaluating the specific germline alterations. Based on the current sample size, a sample power of 1 was confirmed by the PWR package in R. Blood samples were collected before any surgical or chemotherapy treatment. Each patient used an EDTA anticoagulant tube to collect 5 ml of peripheral blood, mixed it by inversion, and sent it to the testing laboratory within 5 days under conditions of 2 to 8℃. This study was approved by the Medical Ethics Committee of West China Second University Hospital, Sichuan University (Ethical Lot Number 20200076). The research was carried out according to guidelines and regulations of the ethic committee.
Next-generation sequencing
Blood samples were collected at West China Second University Hospital and sent to BGI Shenzhen Clinical Diagnostic Laboratory (the contract central testing laboratory for this study), where DNA extraction, targeted DNA sequencing, variant calling, and interpretation were performed. Briefly, genomic DNA (gDNA) was extracted from participants’ peripheral blood using the Qiagen Blood Midi Kit (Qiagen, Hilden, Germany) according to the manufacturer’s standard protocol. DNA concentration and quality were assessed by Qubit (Life Technologies, Carlsbad, USA) and agarose gel electrophoresis. The gDNA (250 ng) was randomly fragmented by the Covaris LE220 sonicator (Covaris, Woburn, USA) to generate gDNA fragments with a peak of 250 bp and then subjected to three enzymatic steps: end-repair, A-tailing, and sequencer (MGI, Shenzhen, China) adapter ligation. DNA libraries were purified with Agencourt Ampure XP beads (Beckman-Coulter, Indiana, USA), and PCR was carried out to form a pre-PCR library or pre-hybridization library, during which a unique 8 bp barcode was added to label each sample. Five to ten pre-PCR libraries were pooled equally and hybridized to a custom hereditary cancer panel (BGI, Shenzhen, China). After purification, the enriched DNA was specifically captured and amplified by PCR to obtain a post-PCR library. The post-PCR library were subjected to single-strand separation, circularization and rolling circle replication to generate DNA nano balls (DNB) and sequencing was performed with 2 × 101 bp paired-end reads on a BGISEQ-500 or MGISEQ-2000 platform (MGI, Shenzhen, China) following the manufacturer’s protocols. SNVs and INDELs in all coding exons and intron–exon boundaries (± 20 base pairs) of detected genes were identified from NGS data.
Sequencing data analysis
Raw fastq data generated by the sequencer was first filtered by SOAPnuke 1.5.0 to exclude low quality reads47. The clean reads were then aligned to the reference human genome (UCSC hg19) using the Burrows-Wheeler Aligner (BWA 0.7.12) MEM algorithm48. PCR deduplication was performed using Picard 1.87. The average depth was over 100X and the coverage at 30X exceeded 95% for each sample. Single-nucleotide variants (SNVs), small insertions and deletions (INDELs) were detected by Genome Analysis Toolkit (GATK 4.0.8.1) HaplotypeCaller49. SNVs and INDELs in all coding exons and intron–exon boundaries (± 20 base pairs) of detected genes were identified from NGS data. All of above variants were further filtered by quality depth, strand bias, mapping quality and read position. Finally, each variant was annotated by Bcfanno for gene location and predicted function in Human Genome Variation Society (HGVS) nomenclature and was ready for interpretation.
Data interpretation
Variants were classified into the following five categories according to the American College of Medical Genetics (ACMG) recommendations: class 1, benign; class 2, likely benign; class 3, variant of unknown significance (VUS); class 4, likely pathogenic; and class 5, pathogenic50. Likely pathogenic and pathogenic variants will be verified by Sanger sequencing. Individuals with likely pathogenic or pathogenic variants were defined as having deleterious variants.
Statistical analysis
All statistical analysis and plots were conducted using R version 3.6.3. Pearson's χ2-test was used as a statistical significance test in the analysis of categorical variables. We performed Student’s t-test to compare continuous variables between two groups, such as age at diagnosis. Patients with specific missing clinical data were excluded from the relevant analysis of specific clinical characteristic. Each reported P value was two-sided and P < 0.05 was considered to be statistically significant.
Ethics statement
The studies involving human participants were reviewed and approved by Medical Ethics Committee of West China Second University Hospital (Ethical Lot Number 20200076). The patients/participants provided their written informed consent to participate in this study.
Data availability
The data that support the findings of our study are available in the Supplementary material. The sequencing data generated and analysed during the current study are available in the ClinVar, accession numbers SCV003843260—SCV003843860.
References
Sung, H. et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. https://doi.org/10.3322/caac.21660 (2021).
Chen, W. et al. Cancer statistics in China, 2015. CA Cancer J. Clin. 66, 115–132. https://doi.org/10.3322/caac.21338 (2016).
Siegel, R. L., Miller, K. D., Wagle, N. S. & Jemal, A. Cancer statistics, 2023. CA Cancer J. Clin. 73, 17–48. https://doi.org/10.3322/caac.21763 (2023).
Sowter, H. M. & Ashworth, A. BRCA1 and BRCA2 as ovarian cancer susceptibility genes. Carcinogenesis 26, 1651–1656. https://doi.org/10.1093/carcin/bgi136 (2005).
Lord, C. J. & Ashworth, A. BRCAness revisited. Nat. Rev. Cancer 16, 110–120. https://doi.org/10.1038/nrc.2015.21 (2016).
Mavaddat, N. et al. Cancer risks for BRCA1 and BRCA2 mutation carriers: results from prospective analysis of EMBRACE. J. Natl. Cancer Inst. 105, 812–822. https://doi.org/10.1093/jnci/djt095 (2013).
Li, A. et al. BRCA germline mutations in an unselected nationwide cohort of Chinese patients with ovarian cancer and healthy controls. Gynecol. Oncol. 151, 145–152. https://doi.org/10.1016/j.ygyno.2018.07.024 (2018).
Wu, X. et al. The first nationwide multicenter prevalence study of germline BRCA1 and BRCA2 mutations in Chinese ovarian cancer patients. Int. J. Gynecol. Cancer 27, 1650–1657. https://doi.org/10.1097/IGC.0000000000001065 (2017).
Shi, T. et al. BRCA1 and BRCA2 mutations in ovarian cancer patients from China: Ethnic-related mutations in BRCA1 associated with an increased risk of ovarian cancer. Int. J. Cancer 140, 2051–2059. https://doi.org/10.1002/ijc.30633 (2017).
Hall, M. J. et al. Genetic testing for hereditary cancer predisposition: BRCA1/2, Lynch syndrome, and beyond. Gynecol. Oncol. 140, 565–574. https://doi.org/10.1016/j.ygyno.2016.01.019 (2016).
Sopik, V., Akbari, M. R. & Narod, S. A. Genetic testing for RAD51C mutations: In the clinic and community. Clin. Genet. 88, 303–312. https://doi.org/10.1111/cge.12548 (2015).
Meindl, A. et al. Germline mutations in breast and ovarian cancer pedigrees establish RAD51C as a human cancer susceptibility gene. Nat. Genet. 42, 410–414. https://doi.org/10.1038/ng.569 (2010).
Bonadona, V. et al. Cancer risks associated with germline mutations in MLH1, MSH2, and MSH6 genes in Lynch syndrome. JAMA 305, 2304–2310. https://doi.org/10.1001/jama.2011.743 (2011).
Gonzalez, K. D. et al. Beyond Li Fraumeni syndrome: Clinical characteristics of families with p53 germline mutations. J. Clin. Oncol. 27, 1250–1256. https://doi.org/10.1200/JCO.2008.16.6959 (2009).
Tan, M. H. et al. Lifetime cancer risks in individuals with germline PTEN mutations. Clin. Cancer Res. 18, 400–407. https://doi.org/10.1158/1078-0432.CCR-11-2283 (2012).
Hearle, N. et al. Frequency and spectrum of cancers in the Peutz-Jeghers syndrome. Clin. Cancer Res. 12, 3209–3215. https://doi.org/10.1158/1078-0432.CCR-06-0083 (2006).
Tan, D. S. P. et al. “BRCAness” syndrome in ovarian cancer: A case-control study describing the clinical features and outcome of patients with epithelial ovarian cancer associated with BRCA1 and BRCA2 mutations. J. Clin. Oncol. 26, 5530–5536. https://doi.org/10.1200/JCO.2008.16.1703 (2008).
Pennington, K. P. et al. Germline and somatic mutations in homologous recombination genes predict platinum response and survival in ovarian, fallopian tube, and peritoneal carcinomas. Clin. Cancer Res. 20, 764–775. https://doi.org/10.1158/1078-0432.CCR-13-2287 (2014).
Moore, K. et al. Maintenance olaparib in patients with newly diagnosed advanced ovarian cancer. N. Engl. J. Med. 379, 2495–2505. https://doi.org/10.1056/NEJMoa1810858 (2018).
Gonzalez-Martin, A. et al. Niraparib in patients with newly diagnosed advanced ovarian cancer. N. Engl. J. Med. 381, 2391–2402. https://doi.org/10.1056/NEJMoa1910962 (2019).
Swisher, E. M. et al. Rucaparib in relapsed, platinum-sensitive high-grade ovarian carcinoma (ARIEL2 Part 1): An international, multicentre, open-label, phase 2 trial. Lancet Oncol. 18, 75–87. https://doi.org/10.1016/s1470-2045(16)30559-9 (2017).
Ledermann, J. et al. Olaparib maintenance therapy in platinum-sensitive relapsed ovarian cancer. N. Engl. J. Med. 366, 1382–1392. https://doi.org/10.1056/NEJMoa1105535 (2012).
Sim, N. L. et al. SIFT web server: Predicting effects of amino acid substitutions on proteins. Nucleic Acids Res. 40, W452-457. https://doi.org/10.1093/nar/gks539 (2012).
Adzhubei, I. A. et al. A method and server for predicting damaging missense mutations. Nat. Methods 7, 248–249. https://doi.org/10.1038/nmeth0410-248 (2010).
Sekine, M., Nishino, K. & Enomoto, T. Differences in ovarian and other cancers risks by population and BRCA mutation location. Genes (Basel) 12, 1050. https://doi.org/10.3390/genes12071050 (2021).
Mills, A. M. & Longacre, T. A. Lynch syndrome screening in the gynecologic tract: Current state of the art. Am. J. Surg. Pathol. 40, e35-44. https://doi.org/10.1097/PAS.0000000000000608 (2016).
Kunnackal John, G., Das Villgran, V., Caufield-Noll, C. & Giardiello, F. M. Comparison of universal screening in major lynch-associated tumors: A systematic review of literature. Fam. Cancer https://doi.org/10.1007/s10689-020-00226-w (2021).
Gu, C. et al. NCOA4: An immunomodulation-related prognostic biomarker in colon adenocarcinoma and pan-cancer. J. Oncol. 2022, 5242437. https://doi.org/10.1155/2022/5242437 (2022).
Nepal, M., Che, R., Ma, C., Zhang, J. & Fei, P. FANCD2 and DNA damage. Int. J. Mol. Sci. 18, 1804. https://doi.org/10.3390/ijms18081804 (2017).
Walden, H. & Deans, A. J. The Fanconi anemia DNA repair pathway: Structural and functional insights into a complex disorder. Annu. Rev. Biophys. 43, 257–278. https://doi.org/10.1146/annurev-biophys-051013-022737 (2014).
Deans, A. J. & West, S. C. DNA interstrand crosslink repair and cancer. Nat. Rev. Cancer 11, 467–480. https://doi.org/10.1038/nrc3088 (2011).
Nalepa, G. & Clapp, D. W. Fanconi anaemia and cancer: An intricate relationship. Nat. Rev. Cancer 18, 168–185. https://doi.org/10.1038/nrc.2017.116 (2018).
Moes-Sosnowska, J. et al. Clinical importance of FANCD2, BRIP1, BRCA1, BRCA2 and FANCF expression in ovarian carcinomas. Cancer Biol. Ther. 20, 843–854. https://doi.org/10.1080/15384047.2019.1579955 (2019).
Croteau, D. L., Popuri, V., Opresko, P. L. & Bohr, V. A. Human RecQ helicases in DNA repair, recombination, and replication. Annu. Rev. Biochem. 83, 519–552. https://doi.org/10.1146/annurev-biochem-060713-035428 (2014).
Lu, H. & Davis, A. J. Human RecQ helicases in DNA double-strand break repair. Front. Cell Dev. Biol. 9, 640755. https://doi.org/10.3389/fcell.2021.640755 (2021).
Siitonen, H. A. et al. The mutation spectrum in RECQL4 diseases. Eur. J. Hum. Genet. 17, 151–158. https://doi.org/10.1038/ejhg.2008.154 (2009).
Cybulski, C. et al. Germline RECQL mutations are associated with breast cancer susceptibility. Nat. Genet. 47, 643–646. https://doi.org/10.1038/ng.3284 (2015).
Sun, J. et al. Mutations in RECQL gene are associated with predisposition to breast cancer. PLoS Genet. 11, e1005228. https://doi.org/10.1371/journal.pgen.1005228 (2015).
Paulo, P. et al. Targeted next generation sequencing identifies functionally deleterious germline mutations in novel genes in early-onset/familial prostate cancer. PLoS Genet. 14, e1007355. https://doi.org/10.1371/journal.pgen.1007355 (2018).
Fang, H. et al. RecQL4 helicase amplification is involved in human breast tumorigenesis. PLoS One 8, e69600. https://doi.org/10.1371/journal.pone.0069600 (2013).
Su, Y. et al. Human RecQL4 helicase plays critical roles in prostate carcinogenesis. Cancer Res. 70, 9207–9217. https://doi.org/10.1158/0008-5472.CAN-10-1743 (2010).
Lu, H. et al. RECQL4 promotes DNA end resection in repair of DNA double-strand breaks. Cell Rep. 16, 161–173. https://doi.org/10.1016/j.celrep.2016.05.079 (2016).
Mo, D. et al. Human helicase RECQL4 drives cisplatin resistance in gastric cancer by activating an AKT-YB1-MDR1 signaling pathway. Cancer Res. 76, 3057–3066. https://doi.org/10.1158/0008-5472.CAN-15-2361 (2016).
Veith, S. & Mangerich, A. RecQ helicases and PARP1 team up in maintaining genome integrity. Ageing Res. Rev. 23, 12–28. https://doi.org/10.1016/j.arr.2014.12.006 (2015).
Larizza, L., Roversi, G. & Volpi, L. Rothmund-Thomson syndrome. Orphanet J. Rare Dis. 5, 2. https://doi.org/10.1186/1750-1172-5-2 (2010).
Mo, D., Zhao, Y. & Balajee, A. S. Human RecQL4 helicase plays multifaceted roles in the genomic stability of normal and cancer cells. Cancer Lett. 413, 1–10. https://doi.org/10.1016/j.canlet.2017.10.021 (2018).
Chen, Y. et al. SOAPnuke: A MapReduce acceleration-supported software for integrated quality control and preprocessing of high-throughput sequencing data. Gigascience 7, 1–6. https://doi.org/10.1093/gigascience/gix120 (2018).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760. https://doi.org/10.1093/bioinformatics/btp324 (2009).
Van der Auwera, G. A. et al. From FastQ data to high confidence variant calls: The Genome Analysis Toolkit best practices pipeline. Curr. Protoc. Bioinform. 43, 11. https://doi.org/10.1002/0471250953.bi1110s43 (2013).
Richards, S. et al. Standards and guidelines for the interpretation of sequence variants: A joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 17, 405–424. https://doi.org/10.1038/gim.2015.30 (2015).
Acknowledgements
This study is supported by the Key Project of Sichuan Provincial Department of Science and Technology (19YFS0532): “Study on the key factors affecting the diagnosis and treatment of major diseases in obstetrics and gynecology”, the project of Chengdu Science and Technology administration (2021-YF05-01725-SN): “Study on the mechanism of platinum-resistance chemotherapy in ovarian cancer”, and the Science and Technology Department of Sichuan Province, Key R8D Program, 2022YFS0080. We would like to thank the patients for their participation and support of our study.
Author information
Authors and Affiliations
Contributions
Conceptualization, R.T.Y. and S.D.Z.; software, J.W.; formal analysis, K.Y.F., Q.L.L. and J.W.; investigation, K.Y.F., Q.L.L., L.S., L.Z., Y.M., J.H.C., J.Z. and D.Q.W.; resources, M.P.Z., X.Y.Y., K.M.L. and L.S.; data curation, K.Y.F., Q.L.L. and J.W.; writing—original draft preparation, K.Y.F., Q.L.L., J.W., X.Y.Y and D.S.; writing—review and editing, R.T.Y. and S.D.Z.; visualization, K.Y.F., Q.L.L., J.W., X.Y.Y and D.S.; funding acquisition, R.T.Y. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
Xinyu Yan, Jie Wang, Di Shao and Shida Zhu are employees of BGI Genomics that produces the panel test used in this study. The other authors declared no potential conflicts of interest.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fu, K., Li, Q., Wang, J. et al. Characteristics of germline DNA damage response gene mutations in ovarian cancer in Southwest China. Sci Rep 14, 6702 (2024). https://doi.org/10.1038/s41598-024-52707-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-52707-y
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.