Abstract
Dengue virus (DENV) infection remains a challenging health threat worldwide. Ubiquitin-specific protease 18 (USP18), which preserves the anti-interferon (IFN) effect, is an ideal target through which DENV mediates its own immune evasion. However, much of the function and mechanism of USP18 in regulating DENV replication remains incompletely understood. In addition, whether USP18 regulates DENV replication merely by causing IFN hyporesponsiveness is not clear. In the present study, by using several different approaches to block IFN signaling, including IFN neutralizing antibodies (Abs), anti-IFN receptor Abs, Janus kinase inhibitors and IFN alpha and beta receptor subunit 1 (IFNAR1)knockout cells, we showed that USP18 may regulate DENV replication in IFN-associated and IFN-unassociated manners. Localized in mitochondria, USP18 regulated the release of mitochondrial DNA (mtDNA) to the cytosol to affect viral replication, and mechanisms such as mitochondrial reactive oxygen species (mtROS) production, changes in mitochondrial membrane potential, mobilization of calcium into mitochondria, 8-oxoguanine DNA glycosylase 1 (OGG1) expression, oxidation and fragmentation of mtDNA, and opening of the mitochondrial permeability transition pore (mPTP) were involved in USP18-regulated mtDNA release to the cytosol. We therefore identify mitochondrial machineries that are regulated by USP18 to affect DENV replication and its association with IFN effects.
Similar content being viewed by others
Introduction
Dengue virus (DENV) infection is a critical public health concern worldwide1. The incidence of potentially fatal consequences of DENV infection, such as dengue hemorrhagic fever (DHF) and dengue shock syndrome (DSS), has not notably changed in recent decades2, 3. It is estimated that 1 in 20 DENV-infected patients will suffer severe consequences, such as shock, internal bleeding and death4. The aggressive development of anti-dengue vaccines is currently underway, but these efforts still face great challenges4, 5.
Ubiquitin-specific protease 18 (USP18), also known as UBP43, is an IFN-inducible molecule preserving isopeptidase activity that deconjugates proteins by targeting interferon-stimulated gene 15 (ISG15); this process is called deISGylation6. The deISGylation process helps to protect ISG15-ISGylated target proteins from degradation via an autophagy-dependent pathway7. After recruitment by signal transducer and activator of transcription (STAT) 2, USP18 interacts with interferon alpha and beta receptor subunit 2 (IFNAR2) through its C-terminal domain and competes with Janus kinase (JAK)1 for binding and transducing signals to inhibit type I IFN (IFN-I)-mediated effects8, 9. Thus, USP18 serves as the potential target that is used by DENV to antagonize the antiviral effects of IFNs and to achieve immune evasion10. Studies have shown that the negative effects of USP18 on IFN-I may be beneficial for preventing the overactivation of immune responses in autoimmune diseases11. In a model of vesicular stomatitis virus (VSV) infection, Honke et al.8 demonstrated that by lowering IFN-I responsiveness, USP18 sustained viral replication in the local milieu to ensure the production of sufficient viral antigen for the effective induction of adaptive immunity. However, whether the reported effects of USP18 occur simply through anti-IFN signaling remains incompletely understood.
Mitochondria function as pivotal organelles for triggering antiviral immunity and, at times, may be targeted by viruses to evade immune surveillance12. As a molecule that is localized to mitochondria, how USP18 may regulate mitochondrial machinery to affect DENV replication has not been investigated. In here, we addressed (1) whether USP18 regulates DENV replication in a manner that is completely dependent on IFN signaling inhibition and (2) how USP18 may regulate DENV replication by affecting the release of mtDNA, a mechanism driving toll-like receptor 9 (TLR9) activation as reported by our group13. The results suggest that USP18 regulates DENV replication through both IFN-associated and IFN-unassociated mechanisms. By modulating mitochondrial machineries, USP18 regulates mtDNA release to the cytosol to affect DENV replication.
Results
DENV infection induced USP18 which regulated DENV replication through in part IFN-independent manner
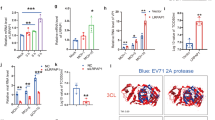
Microarray analysis revealed that USP18 was one of the genes that were upregulated in hDCs infected with DENV (Supplementary Fig. 1A). All required messages about the microarray data were included in our recently published report14. USP18 induction also occurred in primary BMDCs infected by DENV (Fig. 1A). DENV infection-induced USP18 expression and protein ISGylation depended on IFNAR and STAT1 (Supplementary Fig. 1B,C). To investigate the mechanism of USP18-regulated DENV replication and its association with IFN, USP18 expression was attenuated by introducing USP18 small interference RNA (siUSP18) into cells. The Fig. 1B showed that DENV infection induced the accumulation of USP18 in mitochondria that was dramatically reduced by siUSP18 treatment. USP18 deficiency inhibited DENV replication in different tissue cells (Fig. 1C). We used several approaches to determine whether USP18 could regulate DENV replication independently of its anti-IFN effects. The JAK1/2 inhibitor ruxolitinib, which completely blocked IFN-α-, IFN-β-, IFN-λ1- and IFN-γ-induced STAT1 phosphorylation, enhanced DENV replication, but had no effects on USP18 deficiency-mediated reduction of DENV replication (Fig. 1D). To more specifically target IFN, the effects of both anti-IFN-α neutralizing Abs and anti-IFNAR1 Abs were examined. While anti-IFN-α Abs completely blocked the IFN-α-mediated inhibition of viral production, it had limited effects on the USP18 deficiency-mediated inhibition of viral production (Fig. 1E, compare lane 11 with 5 and lane 12 with 6); TNF-α treatment served as a control in this experiment. To show the adequacy of the anti-IFN effects by different treatments, we added IFN-α into cell cultures in all experiments that were conducted to study the effects of USP18 deficiency. We also observed that anti-IFNAR1 Ab treatment could not reverse the USP18 deficiency-mediated suppression of DENV replication (Fig. 1F). Furthermore, USP18 deficiency significantly attenuated the expression of DENV NS2B protein, DENV RNA and DENV viral particles in BMDCs derived from mice lacking the ifnar1 (Fig. 2A–C) and stat1 genes (Fig. 2D–F).
DENV infection induced USP18 that regulated DENV replication through in part IFN-independent manner. Human dendritic cells (hDCs) and mouse bone marrow-derived DCs (BMDCs) (5 × 106/ml) were infected with DENV (MOI = 5 for hDCs and MOI = 0.5 for BMDCs) or mock for 24 h, and the expression of USP18 mRNA was determined by qPCR (A). A549 cells (2 × 105/ml) were introduced with siUSP18 or control siRNA (siCtrl) via lipofectamine 3000 (3 μl/ml) according to the description by the manufacturer. After this, cells were infected with DENV (MOI = 0.5) or mock for 24 h, and the expression of USP18, Tom20 and α-tubulin in cytosolic and mitochondrial fractions was determined by Western blotting (B). Similar to (B), the effects of USP18 deficiency (siUSP18 treatment) on DENV NS2B and DENV RNA levels measured by Western blots and qPCR in A549 cells and HepG2 cells were determined (C). A549 cells (2 × 105/ml) were treated with 5 μM ruxolitinib for 2 h and then stimulated with various IFNs (IFN-α (100 U/ml), IFN-β (50 U/ml), IFN-γ (10 ng/ml), and IFN-λ1 (40 ng/ml)), and the expression of USP18 and phosphorylated STAT1 was determined by Western blotting (D, upper). Similarly, effects of ruxolitinib on USP18 deficiency-regulated DENV RNA levels measured by qPCR were evaluated (D, lower). A549 cells (2 × 105/ml) treated with siCtrl or siUSP18 were pretreated with anti-IFN-α neutralizing Abs (5 μg/ml) or control Abs for 2 h and then stimulated with IFN-α (100 U/ml) or TNF-α (40 ng.ml), followed by infection with mock or DENV (MOI = 0.5) for 24 h, and the expression of mRNA of USP18 and DENV NS2B was measured (E). The effects of anti-IFNAR1 neuralizing Abs (5 μg/ml) on DENV-induced USP18 and NS2B expression with or without USP18 deficiency were measured (F). More than 3 independent experiments were analyzed for each condition. Statistical analysis was done using Student’s t test (A, C) and two-way ANOVA with Holm-Sidak (D, F) to compare differences among different treatments. *P < 0.05; **P < 0.01; ***P < 0.001 and ****P < 0.0001. Original gels are presented in Supplementary Fig. 9.
USP18 deficiency inhibited DENV replication in BMDCs from ifnar1-KO and stat1-KO mice. BMDCs were prepared from ifnar1-KO mice as described in the Materials and Methods. After introduction of siCtrl or siUSP18 via electroporation as described in the Materials and Methods, the cells (1 × 106/ml) were infected with mock or DENV (MOI = 1) for 24 h and then collected for further analysis. The expression of USP18 and NS2B, and protein ISGylation in total cell lysates was determined by Western blotting (A). The levels of DENV RNA were measured by qPCR (B). In addition, the DENV titers were evaluated by plaque assays as described in the Materials and Methods (C). Similar experiments were carried out in BMDCs from stat1-KO mice and the results were presented in (D, E and F), respectively. Values represent the mean of the individual measurements in each sample ± SEM. Statistical analysis was done using Student’s t test (A–F) to compare differences among different treatments. *P < 0.05, **P < 0.01. Original gels are presented in Supplementary Fig. 10.
USP18 regulated mtDNA release to the cytosol
The mitochondrial localization of USP18 suggests that USP18 may exert its effects by regulating the mitochondrial machinery in DENV infection. We focused on studying DENV-induced mtDNA release which activated TLR913. As shown in Fig. 3A and Supplementary Fig. 2, DENV infection reduced total mtDNA levels but increased cytosolic mtDNA levels in A549 cells and HepG2 cells, which were compatible with our observations in DENV-infected hDCs13. Notably, USP18 deficiency further enhanced the DENV-induced mtDNA release to the cytosol. The USP18-mediated effects on DENV-induced mtDNA release to the cytosol remained when IFNAR1 was deleted (Fig. 3B). Moreover, USP18 deficiency significantly enhanced DENV infection-induced oxidization of total and cytosolic mtDNA, as determined by measuring the 8-OHdG intensity (Fig. 3C). This conclusion was supported by studies using confocal microscopy demonstrating the colocalization of 8-OHdG with mtDNA and cytosolic DNA (Supplementary Fig. 3). Interestingly, the results in Supplementary Fig. 3 also suggest that siRNA mediated silencing of USP18 notably increased 8-OHdG signal in nuclear DNA in DENV-infected cells. We used an alternative approach to analyze mtDNA oxidation, namely, employing formamidopyrimidine DNA glycosylase (Fpg)-sensitive real-time PCR analysis15. As treatment of mtDNA with Fpg removes oxidized purines from the DNA and creates single-strand breaks that block PCR amplification of these sites, the difference in the qPCR amplification between Fpg-treated and Fpg-untreated DNA reflects the presence of oxidative base damage and the percentage of intact DNA; recognition and cleavage by Fpg results in a decrease in the percentage and suggests an increase in the number of sequences harboring oxidized base products15, 16. By this approach, we confirmed the enhancement of DENV infection-mediated mtDNA oxidation by USP18 deficiency (Supplementary Fig. 4). The mechanism was likely due to the suppression of OGG1, an enzyme that recognizes and removes 8-OHdG from DNA, in response to USP18 deficiency (Fig. 3D). Because full-length mtDNA cannot be released from mitochondria through mitochondrial pores, we showed that DENV infection increased fragmented mtDNA (size less than 700 bp) in the cytosol and the effect was further enhanced under USP18 deficiency (Fig. 3E and Supplementary Fig. 5).
USP18 regulated mtDNA release, oxidation and fragmentation. A549 cells (2 × 105/ml) treated with siCtrl or siUSP18 via lipofectamine 3000 were infected with mock or DENV (MOI = 0.5) for 24 h. Both total and cytosolic DNA were extracted from collected cells according to the Materials and Methods and quantified using qPCR with specific primers to measure mtDNA levels. The total mtDNA content was normalized to nuclear DNA TERT, and the relative abundance of mtDNA in cytosolic fraction was normalized with exogenously added plasmid encoding the FLAG gene (PCR3.1-flag) as described in the Materials and Methods (A). BMDCs were prepared from ifnar1-KO mice as described in the Materials and Methods and cells (1 × 106/ml) were treated similarly to (A) and mtDNA release was measured (B). A549 cells (2 × 105/ml) treated with siCtrl or siUSP18 via lipofectamine 3000 were infected by DENV (MOI = 0.5) for 24 h and the expression of 8OHdG was measured by intracellular immunostaining with anti-8OHdG Abs (1:500) and then analyzed by flow cytometry (C, upper and left lower). In addition, the 8OHdG levels in the cytosolic fractions were determined by ELISA (C, right lower). A549 cells were treated similarly to (C), collected and the expression of OGG1 in total cell lysates was determined by Western blotting (D). A549 cells (2 × 105/ml) were infected by DENV (MOI = 0.5) for 24 h and DNA was prepared from total, mitochondrial, or cytosolic fractions of treated cells as described in the Materials and Methods and run in agarose gels and analyzed by staining with Midori Green Advance Safe DNA/RNA staining kit (E, left). The signals of 6069 bp and 637 bp fragments of mtDNA and TERT in individual fractions were amplified by PCR using designated primers (Table 1), analyzed in gels, and followed by ethidium bromide staining (E, right). Statistical analysis was done using two-way ANOVA with Holm-Sidak’s multiple comparisons (A–E) to compare differences among different treatments. *P < 0.05; **P < 0.01; ***P < 0.001 and ****P < 0.0001. Original gels are presented in Supplementary Fig. 11.
Mechanisms underlying the USP18-regulated release of mtDNA
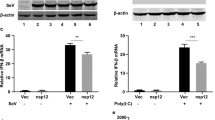
We subsequently investigated the mechanisms responsible for the effects of USP18 deficiency on mtDNA release. Using calcein-quenching assays to measure mPTP opening, we showed that USP18 deficiency significantly enhanced DENV infection-induced mPTP opening, a reflection of reduced fluorescence signal (Fig. 4A). In addition, USP18 deficiency further increased mitochondrial ROS (mtROS) production compared to DENV infection alone by both flow cytometry (Fig. 4B) and confocal microscopy (Fig. 4C) analysis. USP18 deficiency also enhanced DENV infection-mediated decrease of the mitochondrial membrane potential (Fig. 4D,E) given that the mitochondrial membrane potential reduced by siUSP18 treatment. These findings were also supported by the results examining BMDCs from IFNAR1-KO mice (Supplementary Fig. 6A–C). We then used different specific inhibitors to confirm these events. Treatment with the inositol 1,4,5-trisphosphate receptor (IP3R) antagonist xestospongin C (XeC) and the mitochondrial calcium uniporter (MCU) inhibitor ruthenium red (RuR) to block the mobilization of calcium from the endoplasmic reticulum into mitochondria increased DENV RNA levels and significantly reversed the USP18 deficiency-mediated inhibition of DENV replication (Fig. 5A). These results were further supported by the finding that RuR treatment inhibited the USP18-mediated opening of mPTP (Supplementary Fig. 7A) and the release of mtDNA to the cytosol (Supplementary Fig. 7B). Furthermore, treatment with the ROS inhibitor Mitotempo efficiently inhibited the USP18 deficiency-mediated opening of mPTP (Fig. 5B). Although we observed an increase in voltage-dependent anion-selective channel 1 (VDAC1) oligomerization in USP18-deficient cells, there were no synergistic effects between USP18 deficiency-induced and DENV infection-induced VDAC1 oligomerization (Fig. 5C). Moreover, inhibition of mtROS production and various mitochondrial pore inhibitors reversed the USP18 deficiency-mediated suppression of DENV replication and mtDNA release to the cytosol (Fig. 5D). Finally, the results demonstrated that the cause of mitochondrial dysfunction with 2′,3′ dideoxycytidine (ddC), a nucleoside analog, abolished the USP18 deficiency-mediated suppression of DENV replication, confirming the significance of mitochondrial machineries in USP18-mediated effects (Fig. 5E).
USP18 regulated mtROS production, mPTP opening, and mitochondrial membrane potential changes in DENV infection. A549 cells (2 × 105/ml) treated with siCtrl or siUSP18 were infected with mock or DENV (MOI = 0.5) for 24 h. In the calcein-quenching assay, the treated cells were mixed with 10 nM calcein and 400 μM CoCl2, and the fluorescence intensity was determined by flow cytometry (A). A549 cells (2 × 105/ml) were infected with mock or DENV (MOI = 0.5) for 24 h, and then 5 μM MitoSOXTMRe were added into the culture. After incubation for 0.5 h, the intensity of MitoSOX fluorescence observed via flow cytometry was measured and used as an indicator of mitochondrial ROS levels (B). A549 cells treated as described in (B) were stained with 1 μg/ml Hoechst 33,258 solution for cell nuclei indicator and MitoSOX Red fluorescence dye for MitoROS and examined with a confocal laser-scanning microscope as described in the Materials and Methods (C). Treated A549 cells were stained with JC-1 red/green dye and analyzed by flow cytometry to measure mitochondrial membrane potential as described in the Materials and Methods (D). Similar to (C), the intensity of mitochondrial membrane potential was analyzed by confocal microscopy after cell staining with MT-1 dye and Hoechst 33,258 (E). More than 3 independent experiments were carried out and analyzed for each condition. Statistical analysis was done using two-way ANOVA with Holm-Sidak’s multiple comparisons (A, B and D) to compare differences among different treatments. *P < 0.05; **P < 0.01; ***P < 0.001 and ****P < 0.0001.
Mechanisms of USP18-regulated mtDNA release examined by specific inhibitors. A549 cells (2 × 105/ml) delivered with siCtrl or siUSP18 were treated with indicated concentrations of RUR and Xec for 2 h and then infected with mock or DENV (MOI = 0.5) for 24 h. The levels of DENV RNA were measured (A). A549 cells (2 × 105/ml) delivered with siCtrl or siUSP18 were treated with Mitotempo (100 μM) for 2 h and then infected by DENV (MOI = 0.5) for 24 h. The collected cells were mixed with 10 nM calcein and 400 μM CoCl2 in calcein-quenching assays, and the fluorescence intensity was determined by flow cytometry to measure DENV-induced opening of mPTP (B). A549 cells (2 × 105/ml) delivered with siCtrl or siUSP18 were infected with mock or DENV (MOI = 0.5) for 24 h. The total cell lysates were incubated with 200 μM EGS at 30 °C for 15 min to stabilize the VDAC oligomer and then examined by Western blot (C). H2O2 (1 mM) treatment for 6 h served as a positive control. A549 cells (2 × 105/ml) delivered with siCtrl or siUSP18 were treated or not with various chemical compounds, including CsA (5 μM), VBIT-4 (10 μM), and Mitotempo (100 μM) for 2 h and then infected by DENV (MOI = 1) for 24 h. The DENV RNA and mtDNA levels were determined. A549 cells (5 × 104/ml) were treated or not with 100 μM ddc for 7 days to deplete mtDNA. Cells were passaged every 2–3 days. The mtDNA-depleted cells (2 × 105/ml) delivered with siCtrl or siUSP18 were infected with mock or DENV (MOI = 0.5) for 24 h. After collecting total cell lysates, the expression of USP18 and DENV NS2B was determined by Western blotting (E). More than 3 independent experiments were carried out and analyzed for each condition. Statistical analysis was done using two-way ANOVA with Holm-Sidak’s multiple comparisons (A, B, D, and E) to compare differences among different treatments. *P < 0.05; **P < 0.01; ***P < 0.001 and ****P < 0.0001. Original gels are presented in Supplementary Fig. 12.
Overexpression of USP18 enhanced DENV replication and inhibited mtDNA release
To further address the mechanisms by which USP18 regulated DENV replication, we generated usp18-knockout (KO) A549 cells with the CRISPR technique, which eliminated USP18 in both the cytosol and mitochondrial fractions Fig. 6A,B. Transfection and induced overexpression of USP18-flag upregulated DENV NS2B expression in usp18-KO cells in a dose-dependent manner (Fig. 6C). While usp18-KO increased the release of mtDNA to the cytosol, overexpression of USP18 effectively inhibited DENV infection-induced mtDNA release (Fig. 6D). Concurrently, overexpression of USP18 enhanced DENV RNA levels (Fig. 6E) and OGG1 expression (Fig. 6F). Furthermore, while usp18-KO enhanced mPTP opening in DENV-infected cells, overexpression of USP18 inhibited mPTP opening (Supplementary Fig. 8). The schematic shown in Fig. 7 illustrates how USP18 deficiency regulated mtDNA release to the cytosol in DENV infection.
Overexpression of USP18 increased DENV replication in USP18-KO cells. A549 cells were treated with puromycin-selectable lenti-CRISPR vector to deliver control sgRNAs or USP18 sgRNAs to generate USP18-knockout (KO) clone as described in the Materials and Methods. The amino acid sequences of the resulting products were shown (A). Expression of USP18, Tom20 and α-tubulin in cytosolic and mitochondrial fractions in wild-type and usp18-KO cells was determined by Western blotting (B). usp18-KO A549 cells (2 × 105/ml) were transfected with different indicated doses of usp18-flag plasmid by lipofectamine 3000 (3 μl/ml) to overexpress USP18 and then infected or not with DENV (MOI = 0.5) for 24 h. Expression of USP18 and NS2B was determined and analyzed (C). In the wild-type and usp18-KO A549 cells (2 × 105/ml) with or without overexpressing USP18, both total and cytosolic fractions were prepared, and mtDNA levels were measured as described in above figures (D). The wild-type and USP18-KO A549 cells were transfected with USP18 or control plasmid by lipofectamine 3000. The treated cells were infected by mock or DENV (MOI = 1) for 24 h and the DENV RNA was measured (E). The wild-type or USP18-KO A549 cells (2 × 105/ml) were transfected with USP18-flag plasmid (2 μg) or control empty vector and then infected by mock or DENV (MOI = 0.5) for 24 h and the expression of USP18 and OGG1 in the whole cell lysates was determined by Western blotting (F). Statistical analysis was done using one-way ANOVA Bonferroni (C) and two-way ANOVA with Holm-Sidak’s multiple comparisons (D–F) to compare differences among different treatments. Original gels are presented in Supplementary Fig. 13.
Schematic illustrates how USP18 deficiency regulated mtDNA release in DENV infection. After DENV infection, sequential events occurred in the mitochondria, and some events were enhanced in response to USP18 deficiency. USP18 deficiency resulted in several events, including the enhancement of production of mtROS, calcium mobilization to mitochondria, suppression of OGG1 expression, generation of oxidized mtDNA, formation of mtDNA fragments and opening of mPTP-VDAC pores in DENV infection. These events resulted in the release of oxidized and fragmented mtDNA from mitochondria to the cytosol to activate the downstream signaling pathways. WT, wild-type; ROS, reactive oxygen species; mPTP, mitochondrial permeability transition pore; mtDNA, mitochondrial DNA; OGG1, 8-oxoguanine DNA glycosylase 1; CsA, cyclosporin A; VBIT-4, a voltage-dependent anion channel (VDAC) oligomerization inhibitor. The figure was generated using BioRender.
Discussion
Reduced virus production in different organs was consistently observed in USP18-deficient or selective USP18 isopeptidase-deficient mice compared to wild-type mice8, 17, 18. The increased responsiveness to IFN-I signaling thus has been generally used to explain the effects of USP18 deficiency17,18,19,20,21,22. However, other evidence suggests the existence of wider ranges of USP18 activities beyond its anti-IFN effects. First, USP18 deficiency impairs antiviral immune responses during virus infection8. Second, in addition to interfering with the interaction between IFNAR and JAK to block IFN-mediated downstream effects, USP18 can interact with mitochondrial antiviral signaling (MAVS) and the tripartite motif (TRIM) to regulate virus production11. Third, USP18 deficiency resulted in a nearly 100-fold reduction in IFN-α2a-mediated downstream effects, but there was only a threefold increase in IFN-α2a-mediated antiviral activity23. Fourth, USP18 could promote bacterial replication, and USP18 deficiency inhibited bacterial production in many organs and increased animal survival; these mechanisms were mediated via the suppression of antibacterial TNF signaling24. Fifth, depending on the models examined, USP18 can play advantageous25,26,27 or disadvantageous28, 29 roles in disease pathogenesis in different animal studies. Finally, in a model of hepatitis C virus infection, Laidlaw et al.30 showed that TNF preserved antiviral effects independently of IFNs. Altogether, these results suggest that the effects of USP18 may not completely rely on attenuating the anti-IFN effects. Through different approaches, including the use of JAKs inhibitor, IFN neutralizing Abs, and anti-IFNAR Abs, we showed that USP18 could partly regulate DENV replication in an IFN-unassociated manner. This claim was further supported by studies examining BMDCs from IFNAR1-KO mice and STAT1-KO mice.
mtDNA resembles bacterial nucleic acids and serves as an important damage-associated molecular pattern (DAMP) that is recognized by nucleic acid sensors31. Several mechanisms such as changes of mitochondria permeability transition, altered mitophagy and mitochondrial dynamics occurring in conditions such as infection, cell death and inflammation are likely associated with mtDNA escape to the cytosol or the extracellular compartments32,33,34. As 16.3 kb mtDNA is compacted into a nucleoid, it cannot pass through mPTP that have diameters of 2–3 nm35, 36. Studies on isolated mitochondria or liposomes revealed that an opened mPTP can only allow the transport of fragmented mtDNA that is less than 700 bp in size and in a single or double helix conformation or linear form35. The exact composition of the mitochondrial pores remains unclear; however, some pores responsible for extrusion of mtDNA have been identified that include at least oligomerize pores of N-terminal cleavage product of gasdermin D, BAK/BAX pores, VDAC oligomers-forming pores and the non-specific mPTP37,38,39,40,41,42,43. In addition to via mitochondrial pores, mtDNA can also be released to the cytosol through the formation of mitochondrial‐derived vesicles44, 45. To trigger mtDNA fragmentation and facilitate mtDNA release, oxidation process of mtDNA appeared to be a critical step32, 41 where cytidine monophosphate kinase 2 (CMPK2) played roles14, 46. Given that many events could potentially be involved in inducing or exaggerating mtDNA oxidation, in the present study examining DENV infection, we demonstrated that the activation by Ca2 + influx, the generation of mtROS, the regulation of OGG1 expression, the oxidization and fragmentation of mtDNA and the opening of the nonspecific mPTP were participating in USP18-regulated mtDNA release to the cytosol. The conclusion was supported by a series of experiments investigating the effects of different specific inhibitors of these events.
There is evidence suggesting that USP18 can interact with MAVS and that this interaction promotes MAVS aggregation, resulting in enhancement of antiviral activity11. However, the interaction with MAVS could not explain the observed effects of USP18 in this study, where USP18 deficiency enhanced the release of mtDNA to the cytosol and suppressed viral replication. Recent studies demonstrated that the fragmentation of mtDNA appeared to be upstream of RIG-I–MAVS system activation, and mtRNA serves as a retrograde second messenger of mtDNA stress to stimulate RIG-I–MAVS signaling47, 48. This study suggests that the release of mtDNA may occur upstream of MAVS signaling. A recently published report demonstrating that oxidized mtDNA could induce gasdermin D oligomerization and pore formation to facilitate extracellular mtDNA release49 suggests possible feedback mechanisms from released oxidized mtDNA to regulate mitochondrial machineries.
Limitations of the study
The evident limitation of this study is the lack of animal data to support the significance of the observed findings in our cellular system. Nevertheless, differences were observed in USP18-deficient animals that were infected with different viruses via different infection routes. While USP18-deficient mice exhibited better protection from intracerebral challenge with lethal lymphocytic choriomeningitis virus (LCMV) or VSV infection than wild-type mice17, intravenous administration of VSV led to virus-mediated paralysis and early death in USP18-deleted mice8. In addition, there were no images showing the movement of mtDNA and the opening of mPTP, which was regulated by USP18. Which molecule is targeted by and interacts with USP18 to mediate its effects is also unclear in this study. We are currently planning more studies to address these questions.
Materials and methods
Cell culture
Human lung epithelial A549 cells and liver HepG2 cells (Bioresource Collection and Research Center, Taiwan) were maintained in F12K medium and MEM, respectively, supplemented with 10% fetal bovine serum (FBS) in a humidified atmosphere containing 5% CO2 at 37 °C. Liver Huh7 cells (Gift from Dr. Cai-yuan Xie, Tri-Service General Hospital) were maintained in DMEM, respectively, supplemented with 10% fetal bovine serum (FBS) in a humidified atmosphere containing 5% CO2 at 37 °C.
Reagents
Antibodies (Abs) against murine USP18 (A16739), and OGG1 (A4997) were purchased from ABclonal (Woburn, MA, USA). Anti-human USP18 Abs (#4813) were purchased from Cell Signaling Technology (Beverly, MA, USA). Anti-ISG15 Abs (PA5-17461) were obtained from Invitrogen (Carlsbad, CA, USA). Anti-DENV NS2B (GTX124246), anti-DENV NS3 (GTX629477), β-actin (GTX109639), anti-Flag (GTX115043) and goat IgG isotype control Abs (GTX35039) were purchased from GeneTex Inc. (Irvine, CA, USA). Anti-STAT1 Abs (SC-592) were purchased from Santa Cruz Biotechnology (Dallas, TX, USA). Anti-phosphorylated STAT1 Abs (2825-1) were purchased from Epitomics (Burlingame, CA, USA). Anti-Tom20 Abs (ab186734) were purchased from Abcam (Cambridge, UK). Anti-α-tubulin Abs (NB100-690) were purchased from Novus (Briarwood Avenue, CO, USA). Anti-8-OHdG Abs (AB5830) were from Millipore (Burlington, MA, USA). Anti-8-OHdG Abs for ELISA (SKT-120-96S) were purchased from StressMarq Biosciences Inc (Victoria, BC V8N 4G0, Canada). Donkey anti-Rabbit IgG Abs (H + L), Alexa Fluor 488 (A-21206), Donkey anti-Goat IgG (H + L), Alexa Fluor 594 (A-11058), MitoSOX (M36008), JC-1 (M34152), ProLong Diamond Antifade Mountant (P36970), Lipofectamine 3000 (L3000001) and Calcein AM (C1430) were purchased from Thermo Fisher Scientific (Waltham, MA, USA). Anti-IFN-α Abs (21100-1) (used at 5 ug/ml), Anti-IFN-α receptor 1 (21370-1) (used at 5 ug/ml), recombinant human IFN‐α (11100-1) (used at 100 U/ml), recombinant mouse IFN‐α (12100-1) (used at 100 U/ml), and recombinant human IFN‐β (11514-1) (used at 50 U/ml) were purchanced from PBL Assay Science (Piscataway, NJ, USA). MitoView Fix 640 (70,082), used similar to mitotracker, was purchanced from Biotium, Inc (Landing Parkway Fremont, CA, USA). Human GMCSF (215-GM) (used at 800 U/ml), human recombinant IL-4 (204-IL) (used at 500 U/ml), IFN‐γ (285-IF) (used at 10 ng/ml), and TNF-α (210-TA) (used at 40 ng/ml) were purchanced from R&D, Inc (New York, NY, USA). Mouse GMCSF (31503) (used at 20 ng/ml) and recombinant human IFN‐λ1 (300-02L) (used at 40 ng/ml) were purchanced from PeproTech, Inc (Rehovot, Israel). VBIT-4 (HY129122) was purchased from MedChemExpress (Monmouth Junction, NJ, USA). Ruxolitinib (tlrl-rux) was purchased from Invivogen (San Diego, CA, USA). Human CD14 + microbead (130-050-201) was purchased from Miltenyi Biotec (Bergisch Gladbach, North Rhine-Westphalia, Germany). Xestospongin C (64,950) was purchased from Cayman Chemical (Michigan 48,108 USA). Fpg enzyme (M0240S) was purchased from New England Biolabs (Ipswich, MA, USA). Midori Green Advance Safe DNA/RNA stain (MG04) was purchased from NIPPON Genetics Europe (Mariaweilerstraße Düren, Germany). PfuUltra II Fusion High-fidelity DNA Polymerase (600,670) was purchased from Agilent Technologies (Santa Clara, CA, United States). MT-1 MitoMP Detection Kit (MT13) was purchased from Dojindo (Kumamoto, Japan). Unless otherwise specified, all other reagents were purchased from Sigma Aldrich.
DENV preparation and infection
Preparation of DENV has been previously described50. The DENV2 strain New Guinea C was propagated in C6/36 mosquito cells in RPMI medium containing 5% heat-inactivated FBS and maintained at 28 °C for 7 days. Preparation of mock conditioned medium was performed using the same procedures, except buffered saline was substituted for virus inoculation. Unless specified, DENV at multiplicity of infection (MOI) 0.5 or 1 was consistently used to infect cells.
Preparation of human monocyte-derived dendritic cells and mouse bone marrow-derived dendritic cells (BMDCs)
Human monocyte-derived dendritic cells (hDCs) were prepared as described in our previous report13. After collection of peripheral blood mononuclear cells (PBMCs) from buffy coat (purchased from Blood Donation Center, Taipei, Taiwan), the CD14+ monocytes were isolated using a magnetic-activated cell isolation column (Miltenyi Biotec). The purified monocytes were cultured in RPMI 1640 medium containing 10% FBS, 800 U/ml granulocyte macrophage-colony stimulating factor (GM-CSF) and 500 U/ml interleukin (IL)-4 at a cell density of 1 × 106 cells/ml. The culture medium was replaced every other day with fresh medium containing GM-CSF and IL-4, and hDCs with a purity greater than 95% after 5–7 days of culture were used in the experiments50. The preparation of mouse BMDCs was performed according to our previous report13. In brief, male C57BL/6 mice (6–12 weeks) were purchased from the National Laboratory Animal Breeding and Research Center (Taipei, Taiwan). The Ifnar1-/- mice were from Dr. Guann-Yi Yu (National Health Research Institute (NHRI), Taiwan) and Stat1-/- mice were provided by Dr. Chien-Kuo Lee (National Taiwan University, Taiwan). All the animal studies were conducted in accordance with a protocol approved by the Institutional Animal Care and Use Committee of the National Health Research Institute (NHRI; NHRI-IACUC-107159-A-S01) and were also carried out in compliance with the ARRIVE guidelines (https://arriveguidelines.org). Bone marrow was flushed from the tibias and femurs of mouse hind legs using a needle syringe loaded with DMEM. After washing and filtering through a 40-mm nylon cell strainer, bone marrow cells were cultured in RPMI with 10 ng/ml of mGM-CSF (PeproTech Inc., New Jersey, USA) for 6 days with the medium refreshed every 2–3 days for BMDCs.
Determination of virus titers
To determine virus titers, the culture supernatants were harvested for plaque-forming assays50. Various virus dilutions were added to 80% confluent baby hamster kidney (BHK-21) cells and incubated at 37 °C for 1 h. After adsorption, cells were washed and overlaid with 1% agarose (SeaPlaque; FMC BioProducts) containing RPMI 1640 and 1% FCS. After incubation for 7 days, cells were fixed with 10% formaldehyde and stained with 0.5% crystal violet. The numbers of plaques were counted, and the results were shown as plaque forming unit (PFU) per milliliter.
Western blotting
Enhanced chemiluminescence Western blotting (Amersham, GE Healthcare Life Science, Uppsala, Sweden) was performed to study protein levels. Briefly, proteins were separated on a SDS-PAGE gel and transferred to a nitrocellulose membrane. For immunoblotting, the nitrocellulose membrane was incubated with TBS-T containing 5% nonfat milk for 1 h at room temperature. Then blotted with Abs against a specific protein for 2 h at room temperature or overnight at 4 °C. After washing, the membrane was incubated with secondary Abs conjugated to horseradish peroxidase for 1 h at room temperature. The membrane was then incubated with a substrate and exposed to X-ray film.
siRNA transfection
The sequence for USP18 siRNA were GGAUCUACGGAGUCUUCUA (Homo sapiens) and GUGGAUGGAAAGUGGUUCU (Mus musculus). Primary cells were collected and resuspended at 1 × 107 ml in modified Eagle's minimum essential medium (opti‐MEM, Invitrogen) containing 300 nM designated siRNA (Stealth RNAi™ siRNA, Invitrogen). Electroporation was performed using a BTX electroporator (San Diego, CA) with a profile of one pulse at 300 V for 3 ms. The cells (2 × 106) were then seeded in culture medium (Invitrogen, Carlsbad, CA, USA) containing 10% FBS for 24 h before subsequent treatment. For cell lines, cells were transfected with 50 nM siRNA by using Lipofectamine 3000 (3 μl/ml, Invitrogen). After transfection for 4 h, the culture medium was replaced with fresh complete medium for further experiments.
Preparation of cytosolic and mitochondrial fractions
A Mitochondria/Cytosol Fractionation Kit from Abcam (Cambridge, UK) was used to extract mitochondrial and cytosolic fractions as described by the manufacturer. In brief, 2 × 107 cells were resuspended in 0.3 ml of 1X Cytosol Extraction Buffer Mix supplemented with dithiothreitol (DTT) and protease inhibitors. After incubation on ice for 10 min, the cells were homogenized in an ice-cold Dounce tissue homogenizer (150–200 passes with the grinder). The homogenate was centrifuged at 700 × g in a microcentrifuge for 10 min at 4 °C, and the supernatant was then centrifuged at 10,000 × g in a microcentrifuge for 30 min at 4 °C. Then, the supernatant was collected (cytosolic fraction), and the pellet (intact mitochondria) was resuspended in 50 μl of the Mitochondrial Extraction Buffer Mix supplemented with DTT and protease inhibitors (mitochondrial fraction).
Measurement of mtROS levels
For mtROS level measurement, the cells were incubated with 5 μM MitoSOXTMRed (Invitrogen) in cultured medium for 30 min at 37 °C. After washing with PBS, the cells were suspended with trypsin–EDTA and were analyzed with flow cytometry13.
USP18-knockout A549 cells
USP18 sgRNA was designed and cloned into a CRISPR‒Cas9 system by the National RNAi Core Facility (RNA technology platform and gene manipulation core, Academia Sinica, Taiwan). The sequence for USP18 sgRNA were GTTCCCCTTATAGGCCTGGT (Homo sapiens). A549 cells were transfected with the plasmid carrying USP18 sgRNA for 48 h. Then, puromycin (10 μg/ml) was added to the culture medium, and the cells were incubated for 7 days; the medium was regularly replaced to eliminate untransfected cells. Subsequently, a single CRISPR‒Cas9 USP18-knockout (KO) A549 clone was selected, picked and transferred to 12-well plates for further culture. Successful KO was confirmed by Western blotting and DNA sequencing.
Overexpression of USP18 in A549 cells
A construct of human USP18 retrieved from the A549 cDNA library was cloned into pcDNA3.1 + and pcDNA3.1 + /C − (K) − DYK (GenScript Inc. Piscataway, NJ). The accuracy of the subcloning was confirmed by sequencing. For overexpression of USP18 in A549 cells, the indicated concentrations of human USP18 were transfected into A549 cells with Lipofectamine 3000 reagent according to the manufacturer’s instructions.
Quantitative RT‒qPCR
Total RNA was isolated from treated cells with NucleoZOL reagent (Macherey-Nagel, Duren, Germany) according to our previous report14. RNA concentrations were measured using a NanoDrop spectrophotometer (ND 1000 V.3.1.0; Thermo Fisher Scientific, Waltham, MA, USA). Reverse transcription was performed with a 20 μl mixture containing 2 μg of total RNA, random hexamers (Invitrogen), and a mixture containing 10X reverse transcription buffer, dNTPs, magnesium chloride, DTT, and Moloney Murine Leukemia Virus Reverse Transcriptase (MMLV RTase, Invitrogen). cDNA was prepared for further evaluation using qPCR. Briefly, 20 ng of cDNA was amplified in a total mixture volume of 20 μL consisting of 1 × KAPA SYBR FAST qPCR Master Mix (KAPA Biosystems, Boston, MA, USA) and the appropriate gene-specific primers, which were added at a final concentration of 200 nM. The primers used are shown in Table 1. The reaction conditions were 40 cycles comprising steps at 95 °C for denaturing and 60 °C for annealing and extension on a LightCycler 480 (Roche). The changes in gene expression induced by DENV infection in the presence or absence of inhibitors or siRNA were calculated with the following formula: fold change = 2−Δ(ΔCt), where ΔCt = Ct of target gene − Ct of GAPDH, and Δ(ΔCt) = ΔCt infected − ΔCt mock control.
Extraction of total, cytosolic and mitochondrial DNA
Total and cytosolic DNA extraction was performed according to our previous report13. Cells were divided into two equal aliquots. One aliquot, the control for normalization, was used to extract total DNA using a NucleoSpin Tissue kit (Macherey-Nagel, Duren, Germany). The other aliquot was resuspended in 400 ml of buffer containing 150 mM NaCl, 50 mM HEPES pH 7.4, and 25 mg/ml digitonin (EMD Chemicals, Gibbstown, NJ, USA). After rotation end-over-end for 15–20 min at 4 °C, the samples were then centrifuged at 980 × g for 3 min three times to remove cellular debris. The supernatants were collected, transferred to fresh tubes, and spun at 17,000 × g for 10 min to collect lysates (the mitochondrial fraction) and supernatants (the cytosolic fraction). The DNA in the mitochondria, cytosol and total fraction were then isolated by running the sample through a NucleoSpin Tissue column (Macherey-Nagel, Duren, Germany), subsequently eluted with buffer, and analyzed on 1% agarose gel that was stained with Midori Green Advance Safe DNA/RNA stain (NIPPON Genetics Europe, Duren, Germany).
Determination of mtDNA
To measure the levels of mtDNA, 20 ng of isolated DNA was subjected to qPCR with KAPA SYBR FAST qPCR Master Mix (KAPA Biosystems) with 200 nM mtDNA primers (ND1, ND5, and RNR2 for A549 and HepG2 cells; 12S and D-loop for BMDCs) or nuclear DNA (nDNA) primers (TERT for A549, and HepG2 cells; albumin for BMDCs). To quantify mtDNA in the cytosolic fraction, 20 ng of a purified plasmid encoding the FLAG gene (PCR3.1-flag) was added to the eluted solution according to our previous report13. The specific primers for both endogenous mtDNA and the FLAG plasmid were used to measure the relative content of cytosolic mtDNA expression normalized to that of FLAG (the primers for PCR are given in Table 1). The relative mtDNA abundance indicates the relative mtDNA content in DENV-infected cells normalized to that in mock-infected cells.
Measurement of mtDNA fragments
To amplify 637 or 6069 bp mtDNA fragment and telomerase reverse transcriptase (TERT) from the mitochondrial, cytosolic or total DNA pool of treated cells, we followed the method described by other researchers40 with some modifications. In brief, 10 ng of the extracted cytosolic and mitochondrial DNA was used as the template. In addition to the template, the reaction mixture also contained 200 nM of each primer and PfuUltra II Fusion HS DNA Polymerase (Agilent Technologies, Santa Clara, CA, USA). The PCR conditions were as follows: 95 °C for 10 min, melting temperature of 95 °C for 60 s, annealing temperature of 55 °C (for 6069 bp and 637 bp) for 1 min 30 s or 59 °C (for TERT) for 1 min and extension temperature of 72 °C for 30 s (637 bp and TERT) and 3 min (6069 bp). Then, the reaction was run for 24 (637 bp and TERT) or 40 (6069 bp) cycles, followed by incubation at 72 °C for 20 min, and the samples were maintained at 4 °C. The amplified products were analyzed on 1% agarose gels that were stained with ethidium bromide.
Measurement of mitochondrial membrane potential with JC-1 probe
The fluorescent probe, JC-1, purchased from Life Technologies was used to study mitochondrial membrane potential according to the manufacturer’s instructions and our published report13. In brief, cells (2.5 × 105 cells per each condition) were incubated with JC-1 staining solution for 30 min at 37 °C. After washes with PBS, the fluorescence intensity of JC-1 was measured by a flow cytometer (Becton Dickinson) and the results were analyzed with a FlowJo software (Becton Dickinson).
Immunofluorescence staining
Cells were collected and washed two times with PBS before being fixed with 4% paraformaldehyde on ice for 20 min13. The fixed cells were washed with PBS and permeabilized with 0.2% Triton X-100 in PBS for 10 min. Cells were washed again with PBS and blocked with PBS containing 10% BSA for 30 min. Primary Abs were added and incubated with the cells for 2 h at room temperature with occasional mixing. After removing unbound Abs by washing, secondary Abs conjugated with fluorescent dye were added and incubated in the dark for 1 h at room temperature with occasional mixing. Cellular nuclei were counterstained with 1 μg/ml DAPI or Hoechst 33,258 solution. For detecting the colocalization of DNA and mitochondria, MitoView Fix 640 fluorescence dye was used for staining mitochondria. For detecting oxidative DNA and DENV, anti-8-OHdG and anti-NS2B Abs were used. Finally, cells were mounted with mounting reagent (ProLong Diamond Antifade Mountant, Thermo Fisher Scientific) for subsequent confocal microscopy analysis.
Confocal microscopy analysis
Samples were examined with a Leica TCS SP5II and Stellaris 8 confocal laser-scanning microscope (Leica Microsystems, Wetzlar, Germany) equipped with HCX PL APO 63 × /1.4–0.6 oil and HC PL APO 100x/1.40 oil objective (Leica) in core facilities of NHRI according to our previous publication13. Image processing and colocalization analysis were performed with Leica LAS AF and LAS × life science microscope software.
Determination of 8‐OHdG levels with flow cytometry and ELISA
To determine oxidized DNA levels in the mitochondrial and cytosolic fractions51, 500 ng of DNA was added to poly-L-lysine-coated (0.01%) 96-well EIA/RIA plates (Costar, Washington, DC, USA) and incubated at 4 °C overnight. The wells were washed with PBS and blocked with PBS supplemented with 1% BSA for 2 h at room temperature. A horseradish peroxidase-conjugated mouse anti‐human 8‐OHdG Abs (StressMarq Biosciences Inc., Columbia, Canada) that recognizes oxidized nucleotides was added at 0.5 μg/ml and incubated for 2 h at room temperature. After being washed with PBS, the samples were incubated with substrate for 30 min, the reaction was stopped by adding stop solution, and the absorbance was read at 450 nm with an ELISA reader.
For intracellular 8-OHdG detection, cells were washed twice with PBS and detached with trypsin. After fixation with 4% paraformaldehyde, the cells were permeabilized with 0.05% Triton X-100 and then blocked with PBS supplemented with 1% BSA. Goat anti-human 8‐OHdG Abs (Millipore) was added and incubated for 1 h at room temperature. After washing twice with PBS, donkey anti-goat Alexa Fluor™ 488 was added and incubated for 0.5 h at room temperature. After washing twice with PBS, the samples were analyzed with flow cytometry.
Mitochondrial permeability transition pore (mPTP) opening assay
To analyze mPTP opening by calcein-quenching assay, treated cells were detached from the culture plates by trypsin–EDTA. The cells were mixed with 10 nM calcein and 400 μM CoCl2 for 10 min at 37 °C according to the manufacturer’s instructions (Thermo Fisher Scientific, Waltham, MA USA). After washing twice, the fluorescence intensity was determined by flow cytometry. To analyze mPTP opening in BMDCs, we followed the method described by other researchers52. BMDCs were mixed with 1 μM calcein-AM and 1 mM CoCl2 for 10 min at 37 °C. After washing twice, the fluorescence intensity was determined by flow cytometry.
Fpg-sensitive qPCR
The measurement of mtDNA oxidative damage with formamidopyrimidine DNA glycosylase (Fpg)-sensitive qPCR analysis was performed according to the report40. In brief, 250 ng of purified mtDNA was incubated with 8 units of Fpg in 1X NEBuffer 1 and 100 mg/mL BSA in 50 mL at 37 °C for 1 h. After inactivating the Fpg enzyme by incubation at 60 °C for 5 min, 10 ng of DNA was used for qPCR to detect Fpg-sensitive cleavage sites. Data are calculated as the quotient of signal intensities in Fpg-treated DNA relative to Fpg-untreated DNA and reflect the fraction of intact DNA.
VDAC1 cross-linking assay
Treated cells were washed with PBS and incubated with the cross-linking reagent EGS (200 μM) for 15 min at 30 °C40. The samples were analyzed with SDS‒PAGE, and the membranes were blotted with anti-VDAC1 Abs.
Statistical analysis
Statistical comparisons were performed using Student’s t-test (paired t test for primary cells and unpaired t for cell lines) or one-way analysis of variance (ANOVA). When ANOVA showed significant differences between groups, Bonferroni’s post hoc test was used to determine the specific pairs of groups that significantly differed. For multiple comparisons, two-way ANOVA with Holm-Sidak’s multiple comparisons was applied. A P value of < 0.05 was considered to indicate statistical significance. Asterisks indicate values that are significantly different from the relevant control (*P < 0.05, **P < 0.01, ***P < 0.001 and ****P < 0.0001).
Data availability
The data sets used in this study are available from the corresponding author on reasonable request.
References
Castro, M. C., Wilson, M. E. & Bloom, D. E. Disease and economic burdens of dengue. Lancet Infect. Dis. 17, e70–e78. https://doi.org/10.1016/S1473-3099(16)30545-X (2017).
Diamond, M. S. & Pierson, T. C. Molecular insight into dengue virus pathogenesis and its implications for disease control. Cell 162, 488–492. https://doi.org/10.1016/j.cell.2015.07.005 (2015).
Yong, Y. K. et al. Dengue infection—recent advances in disease pathogenesis in the era of COVID-19. Front. Immunol. 13, 889196. https://doi.org/10.3389/fimmu.2022.889196 (2022).
Pintado Silva, J. & Fernandez-Sesma, A. Challenges on the development of a dengue vaccine: A comprehensive review of the state of the art. J Gen Virol https://doi.org/10.1099/jgv.0.001831 (2023).
Thomas, S. J. Is new dengue vaccine efficacy data a relief or cause for concern?. NPJ. Vaccin. 8, 55. https://doi.org/10.1038/s41541-023-00658-2 (2023).
Malakhov, M. P., Malakhova, O. A., Kim, K. I., Ritchie, K. J. & Zhang, D. E. UBP43 (USP18) specifically removes ISG15 from conjugated proteins. J. Biol. Chem. 277, 9976–9981. https://doi.org/10.1074/jbc.M109078200 (2002).
Nakashima, H., Nguyen, T., Goins, W. F. & Chiocca, E. A. Interferon-stimulated gene 15 (ISG15) and ISG15-linked proteins can associate with members of the selective autophagic process, histone deacetylase 6 (HDAC6) and SQSTM1/p62. J. Biol. Chem. 290, 1485–1495. https://doi.org/10.1074/jbc.M114.593871 (2015).
Honke, N. et al. Enforced viral replication activates adaptive immunity and is essential for the control of a cytopathic virus. Nat. Immunol. 13, 51–57. https://doi.org/10.1038/ni.2169 (2011).
Arimoto, K. I. et al. STAT2 is an essential adaptor in USP18-mediated suppression of type I interferon signaling. Nat. Struct. Mol. Biol. 24, 279–289. https://doi.org/10.1038/nsmb.3378 (2017).
Ye, H. et al. USP18 mediates interferon resistance of dengue virus infection. Front. Microbiol. 12, 682380. https://doi.org/10.3389/fmicb.2021.682380 (2021).
Hou, J. et al. USP18 positively regulates innate antiviral immunity by promoting K63-linked polyubiquitination of MAVS. Nat. Commun. 12, 2970. https://doi.org/10.1038/s41467-021-23219-4 (2021).
Chatel-Chaix, L. et al. Dengue virus perturbs mitochondrial morphodynamics to dampen innate immune responses. Cell Host Microbe 20, 342–356. https://doi.org/10.1016/j.chom.2016.07.008 (2016).
Lai, J. H. et al. Infection with the dengue RNA virus activates TLR9 signaling in human dendritic cells. EMBO Rep. https://doi.org/10.15252/embr.201846182 (2018).
Lai, J. H. et al. Mitochondrial CMPK2 mediates immunomodulatory and antiviral activities through IFN-dependent and IFN-independent pathways. iScience 24, 102498. https://doi.org/10.1016/j.isci.2021.102498 (2021).
Al-Mehdi, A. B. et al. Perinuclear mitochondrial clustering creates an oxidant-rich nuclear domain required for hypoxia-induced transcription. Sci. Signal 5, ra47. https://doi.org/10.1126/scisignal.2002712 (2012).
Pastukh, V. M., Gorodnya, O. M., Gillespie, M. N. & Ruchko, M. V. Regulation of mitochondrial genome replication by hypoxia: The role of DNA oxidation in D-loop region. Free Radic. Biol. Med. 96, 78–88. https://doi.org/10.1016/j.freeradbiomed.2016.04.011 (2016).
Ritchie, K. J. et al. Role of ISG15 protease UBP43 (USP18) in innate immunity to viral infection. Nat. Med. 10, 1374–1378. https://doi.org/10.1038/nm1133 (2004).
Ketscher, L. et al. Selective inactivation of USP18 isopeptidase activity in vivo enhances ISG15 conjugation and viral resistance. Proc. Natl. Acad. Sci. U. S. A. 112, 1577–1582. https://doi.org/10.1073/pnas.1412881112 (2015).
Randall, G. et al. Silencing of USP18 potentiates the antiviral activity of interferon against hepatitis C virus infection. Gastroenterology 131, 1584–1591. https://doi.org/10.1053/j.gastro.2006.08.043 (2006).
Sugawara, S. et al. People with HIV-1 demonstrate type 1 interferon refractoriness associated with upregulated USP18. J. Virol. https://doi.org/10.1128/JVI.01777-20 (2021).
Taylor, J. P. et al. CRISPR/Cas9 knockout of USP18 enhances type I IFN responsiveness and restricts HIV-1 infection in macrophages. J. Leukoc. Biol. https://doi.org/10.1002/JLB.3MIA0917-352R (2018).
Kim, J. H., Luo, J. K. & Zhang, D. E. The level of hepatitis B virus replication is not affected by protein ISG15 modification but is reduced by inhibition of UBP43 (USP18) expression. J. Immunol. 181, 6467–6472. https://doi.org/10.4049/jimmunol.181.9.6467 (2008).
Murray, E. J., Burden, F., Horscroft, N., Smith-Burchnell, C. & Westby, M. Knockdown of USP18 increases alpha 2a interferon signaling and induction of interferon-stimulating genes but does not increase antiviral activity in Huh7 cells. Antimicrob. Agents Chemother. 55, 4311–4319. https://doi.org/10.1128/AAC.00644-11 (2011).
Shaabani, N. et al. The probacterial effect of type I interferon signaling requires its own negative regulator USP18. Sci. Immunol. https://doi.org/10.1126/sciimmunol.aau2125 (2018).
Goldmann, T. et al. USP18 lack in microglia causes destructive interferonopathy of the mouse brain. EMBO J. 34, 1612–1629. https://doi.org/10.15252/embj.201490791 (2015).
An, S. et al. USP18 protects against hepatic steatosis and insulin resistance through its deubiquitinating activity. Hepatology 66, 1866–1884. https://doi.org/10.1002/hep.29375 (2017).
Zhang, M. et al. USP18 recruits USP20 to promote innate antiviral response through deubiquitinating STING/MITA. Cell Res. 26, 1302–1319. https://doi.org/10.1038/cr.2016.125 (2016).
Honke, N. et al. Usp18 driven enforced viral replication in dendritic cells contributes to break of immunological tolerance in autoimmune diabetes. PLoS Pathog. 9, e1003650. https://doi.org/10.1371/journal.ppat.1003650 (2013).
Liu, X. et al. USP18 inhibits NF-kappaB and NFAT activation during Th17 differentiation by deubiquitinating the TAK1-TAB1 complex. J. Exp. Med. 210, 1575–1590. https://doi.org/10.1084/jem.20122327 (2013).
Laidlaw, S. M. et al. Tumor necrosis factor inhibits spread of hepatitis C virus among liver cells independent from interferons. Gastroenterology 153, 566–578. https://doi.org/10.1053/j.gastro.2017.04.021 (2017).
Zhong, F., Liang, S. & Zhong, Z. Emerging role of mitochondrial DNA as a major driver of inflammation and disease progression. Trends Immunol. 40, 1120–1133. https://doi.org/10.1016/j.it.2019.10.008 (2019).
Perez-Trevino, P., Velasquez, M. & Garcia, N. Mechanisms of mitochondrial DNA escape and its relationship with different metabolic diseases. Biochim. Biophys. Acta Mol. Basis. Dis. 1866, 165761. https://doi.org/10.1016/j.bbadis.2020.165761 (2020).
Xian, H. & Karin, M. Oxidized mitochondrial DNA: A protective signal gone awry. Trends Immunol. 44, 188–200. https://doi.org/10.1016/j.it.2023.01.006 (2023).
Heilig, R., Lee, J. & Tait, S. W. G. Mitochondrial DNA in cell death and inflammation. Biochem. Soc. Trans. 51, 457–472. https://doi.org/10.1042/BST20221525 (2023).
Garcia, N. & Chavez, E. Mitochondrial DNA fragments released through the permeability transition pore correspond to specific gene size. Life Sci. 81, 1160–1166. https://doi.org/10.1016/j.lfs.2007.08.019 (2007).
Wu, Z., Sainz, A. G. & Shadel, G. S. Mitochondrial DNA: Cellular genotoxic stress sentinel. Trends Biochem. Sci. 46, 812–821. https://doi.org/10.1016/j.tibs.2021.05.004 (2021).
Patrushev, M. et al. Mitochondrial permeability transition triggers the release of mtDNA fragments. Cell Mol Life Sci 61, 3100–3103. https://doi.org/10.1007/s00018-004-4424-1 (2004).
Liu, X. et al. Inflammasome-activated gasdermin D causes pyroptosis by forming membrane pores. Nature 535, 153–158. https://doi.org/10.1038/nature18629 (2016).
McArthur, K. et al. BAK/BAX macropores facilitate mitochondrial herniation and mtDNA efflux during apoptosis. Science https://doi.org/10.1126/science.aao6047 (2018).
Xian, H. et al. Oxidized DNA fragments exit mitochondria via mPTP- and VDAC-dependent channels to activate NLRP3 inflammasome and interferon signaling. Immunity 55, 1370–1385. https://doi.org/10.1016/j.immuni.2022.06.007 (2022).
Kim, J. et al. VDAC oligomers form mitochondrial pores to release mtDNA fragments and promote lupus-like disease. Science 366, 1531–1536. https://doi.org/10.1126/science.aav4011 (2019).
Yu, C. H. et al. TDP-43 triggers mitochondrial DNA release via mPTP to activate cGAS/STING in ALS. Cell 183, 636–649. https://doi.org/10.1016/j.cell.2020.09.020 (2020).
Riley, J. S. & Tait, S. W. Mitochondrial DNA in inflammation and immunity. EMBO Rep 21, e49799. https://doi.org/10.15252/embr.201949799 (2020).
Sugiura, A., McLelland, G. L., Fon, E. A. & McBride, H. M. A new pathway for mitochondrial quality control: Mitochondrial-derived vesicles. EMBO J. 33, 2142–2156. https://doi.org/10.15252/embj.201488104 (2014).
Zecchini, V. et al. Fumarate induces vesicular release of mtDNA to drive innate immunity. Nature 615, 499–506. https://doi.org/10.1038/s41586-023-05770-w (2023).
Zhong, Z. et al. New mitochondrial DNA synthesis enables NLRP3 inflammasome activation. Nature 560, 198–203. https://doi.org/10.1038/s41586-018-0372-z (2018).
Tigano, M., Vargas, D. C., Tremblay-Belzile, S., Fu, Y. & Sfeir, A. Nuclear sensing of breaks in mitochondrial DNA enhances immune surveillance. Nature 591, 477–481. https://doi.org/10.1038/s41586-021-03269-w (2021).
Tadepalle, N. & Shadel, G. S. RNA reports breaking news from mitochondria. Mol. Cell. 81, 1863–1865. https://doi.org/10.1016/j.molcel.2021.04.005 (2021).
Miao, N. et al. Oxidized mitochondrial DNA induces gasdermin D oligomerization in systemic lupus erythematosus. Nat. Commun. 14, 872. https://doi.org/10.1038/s41467-023-36522-z (2023).
Ho, L. J. et al. Dengue virus type 2 antagonizes IFN-alpha but not IFN-gamma antiviral effect via down-regulating Tyk2-STAT signaling in the human dendritic cell. J. Immunol. 174, 8163–8172 (2005).
Lood, C. et al. Neutrophil extracellular traps enriched in oxidized mitochondrial DNA are interferogenic and contribute to lupus-like disease. Nat. Med. 22, 146–153. https://doi.org/10.1038/nm.4027 (2016).
Booth, D. M., Varnai, P., Joseph, S. K. & Hajnoczky, G. Oxidative bursts of single mitochondria mediate retrograde signaling toward the ER. Mol. Cell 81, 3866–3876. https://doi.org/10.1016/j.molcel.2021.07.014 (2021).
Acknowledgements
We thank Dr. Guann-Yi Yu for providing Ifnar1-/- mice and Dr. Chien-Kuo Lee for Stat1-/- mice. This work was supported by grants from the Ministry of Science and Technology (MOST 110-2314-B-182A-127-MY3 to Dr. Lai and 109-2314-B-400-035 to Dr. Ho) and Chang Gung Memorial Hospital (CMRPG1M0021), Taiwan, R.O.C.
Funding
The studies were supported by Ministry of Science and Technology (MOST 107-2314-B-182A-132-MY3 to Dr. Lai and 109-2314-B-400-035 to Dr. Ho) and Chang Gung Memorial Hospital (CMRPG1H0101), Taiwan, R.O.C.
Author information
Authors and Affiliations
Contributions
J.-H.L. and L.-J.H. participated in the study design. D.-W. W, C.-H.W., L.-F.H., and C.-Y.H. performed the experiments and analyzed the data. S.-M. K, A.C., and Z.-F.C. provided various tools and reagents for the study and participated in data discussion. L.-J.H. and J.-H.L. analyzed the data and wrote the report. All the authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lai, JH., Wu, DW., Wu, CH. et al. USP18 enhances dengue virus replication by regulating mitochondrial DNA release. Sci Rep 13, 20126 (2023). https://doi.org/10.1038/s41598-023-47584-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-47584-w
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.