Abstract
Alzheimer's, Parkinson’s, and Huntington’s are the most common neurodegenerative diseases that are incurable and affect the elderly population. Discovery of effective treatments for these diseases is often difficult, expensive, and serendipitous. Previous comparative studies on different model organisms have revealed that most animals share similar cellular and molecular characteristics. The meta-SNP tool includes four different integrated tools (SIFT, PANTHER, SNAP, and PhD-SNP) was used to identify non synonymous single nucleotide polymorphism (nsSNPs). Prediction of nsSNPs was conducted on three representative proteins for Alzheimer's, Parkinson’s, and Huntington’s diseases; APPl in Drosophila melanogaster, LRRK1 in Aedes aegypti, and VCPl in Tribolium castaneum. With the possibility of using insect models to investigate neurodegenerative diseases. We conclude from the protein comparative analysis between different insect models and nsSNP analyses that D. melanogaster is the best model for Alzheimer’s representing five nsSNPs of the 21 suggested mutations in the APPl protein. Aedes aegypti is the best model for Parkinson’s representing three nsSNPs in the LRRK1 protein. Tribolium castaneum is the best model for Huntington’s disease representing 13 SNPs of 37 suggested mutations in the VCPl protein. This study aimed to improve human neural health by identifying the best insect to model Alzheimer's, Parkinson’s, and Huntington’s.
Similar content being viewed by others
Introduction
Neurodegenerative diseases (NDs) are neurological disorders caused by progressive decline in brain function resulting from gradual neuronal death1. They are incurable and mostly affect the elderly population. Their incurability refers to the neural death which is the main cause of these diseases, and the late diagnosis where most symptoms appear in late stages of the diseases. The prevalence of age-related neurodegenerative diseases is increasing with age worldwide. The most recognized NDs were Alzheimer's disease (AD), Parkinson's disease (PD), and Huntington's disease (HD) respectively2 (Fig. S1). As they are not curable, their symptoms appear in late stages and lead to death. They negatively affect the quality of life of patients and their families both socially and economically3. Most NDs result from a combination of genetic and environmental factors, such as PD and AD, whereas others are caused by inherited mutant genes, such as HD.
Insects are suggested to serve as research model organisms because of their easy handling, small in size, small rearing places, relatively low rearing cost, short life cycles, high fecundity, rapid and simple gene manipulation, and fewer ethical permissions compared to vertebrate models4,5. The genomes of different model organisms have been sequenced in parallel with the human genomes, starting with Drosophila melanogaster6. The availability of multiple insect genomes creates an outstanding potential for comparative genomics among insects and between insects and humans. These comparative studies provide an effective tool for investigating human gene function compared to model insects7,8 Table S1. Many insect genes share common ancestry and function with human genes9. Decision-making centres in the brains of insects and mammals share many similarities in physiology although they have evolved independently10. The central complex in insects and the basal ganglia in vertebrates are similar in the maintenance of behavioural actions11. The hippocampi of vertebrates and mushroom bodies of arthropods were also similar in learning and memory (Fig. S2)12. Furthermore, insects provide phenotypic characteristics representing different NDs13,14 as shown in Table S215. The dysfunctional brains of insects enable us to learn more about human brain diseases. In the AD Drosophila model for example, appearance of degenerated neurons and signs of edema in the hippocampus improve our understanding about what is happening16. In PD Bombyx mori model, The p-translucent silkworm is caused by downregulation of the DJ-1 gene, resulting in an increase in the oxidative stress response of the body, which leads to oxidative damage to the nerves and tissues17,18.
Dysfunctional gene behaviour is commonly caused by mutations that are primarily responsible for the development of illnesses. Many disease-causing mutations have been identified in the genome, around 0.5 million are SNPs19. This means that one base is replaced by one other base. Such mutations may involve synonymous and non-synonymous single nucleotide variants (SNVs) or SNPs that may fall within coding sequences of genes, non-coding regions of genes, or intergenic regions20. SNPs play a significant role and increase the susceptibility toward many diseases. Synonymous SNPs (sSNPs) in coding regions have no effect on translated proteins21. However, they can also affect mRNA stability and translation rate. Nonsynonymous SNPs (nsSNPs), which cause amino acid substitutions, have a direct impact on protein structure and function. SNPs in non-coding regions may affect gene splicing and other biological processes such as RNA degradation and transcription22.
Computational tools are used to predict the effects of mutations on protein function and structure. They are important for the analysis of SNVs and their prioritisation for experimental characterization. Using a sequence homology algorithm, computational tools can identify mutations that are significantly pathogenic based on their alignment with known pathogenic mutations as in SIFT, and PANTHER tools23,24. Other computational tools utilise artificial neural networks, and support vector machines to classify the nsSNVs into diseased or neutral substitutions as SNAP, and PhD-SNP tools25,26. Consensus-based approaches tool that integrate multiple algorithms to determine the pathogenicity of nsSNPs as Meta-SNP tool, that combine (SIFT, PANTHER, SNAP, and PhD-SNP)27.
Proving a causal link between a gene and disease is expensive and time-consuming. Therefore, the comprehensive prioritisation of candidate SNPs and determination of the best model to simulate the disease before experimental testing drastically reduces the associated costs, saves time, and accelerates the process of drug discovery as shown in (Fig. 1). Our aim is to highlight the best insect to model Alzheimer's, Parkinson's, and Huntington’s diseases; even in case of the selection of a specific protein to be deeply studied or for overall simulating one of the diseases. Based on the predicted nsSNPs in insect proteins compared to human proteins, simulating diseases’ mechanisms and pathways will be easier and will help improve drug discovery of these NDs.
Materials and methods
This research paper was approved by the research ethics committee from the Faculty of Science, Ain Shams University (ASU-SCI/ENTO/2023/8/1).
Dataset retrieval
All retrieved data are from publicly available databases
-
(a)
The information of the genes of interest was retrieved based on the most influential disease-causing genes from the literature “GeneReviews®—NCBI Bookshelf'' (https://www.ncbi.nlm.nih.gov/books/)28, KEGG DISEASE Database (https://www.kegg.jp/kegg/disease/)29,30,31 (Figs. S3, S4, and S5), and other manual searches using keywords “Alzheimer genes, Parkinson genes, Huntington genes” on Pubmed and Google Scholar. Selected genes included 10 genes for AD, 13 genes for PD, and 10 genes for HD, as shown in Table S3 (accessed: February, 2023).
-
AD: APP, COL25A1, GRN, HDAC6, MAPT, Nep2, PSN-1, PSN-2, RAC1, and SORL1.
-
PD: PRKN, Pink1, DJ-1, GAK, VPS35, UCHL1, EIF4G1, ATP13A2, GIGYF2, HTRA2, PLA2G6, FBXO7, and LRRK2.
-
HD: HTT, DMBK, GRIK2, VCP, VPS13A, ATXN1, ALS, MJD, UBQLN2, and CACNA1A.
-
-
(b)
Protein sequences were retrieved from NCBI using the Protein database (https://www.ncbi.nlm.nih.gov/protein/), Blastp tool (https://blast.ncbi.nlm.nih.gov/Blast.cgi)32, and genes with similar protein architectures were searched using NCBI's SPARCLE (https://www.ncbi.nlm.nih.gov/protfam/) as a resource of the sequential arrangement of CD domains.
-
(c)
Human nsSNPs within the coding region were selected for the APP, LRRK2, and VCP genes (each as a representative gene for Alzheimer’s, Parkinson’s, and Huntington’s diseases, respectively). Polymorphism data were retrieved from the dbSNP-SNP database of NCBI (https://www.ncbi.nlm.nih.gov/snp/) with selection criteria (pathogenic and somatic).
Model selection
The selection of insect models was based on some criteria; these eight selected insect models were taxonomized in four different insect orders (the four largest insect orders)33. Selected insect models are the most common, fully sequenced insects, and are the most representative species of their orders. According to their taxonomic classification; (1) Order Diptera: Drosophila melanogaster (D. melanogaster), Musca domestica (M. domestica), Anopheles gambiae (An. gambiae), and Aedes aegypti (A. aegypti), (2) Order Hymenoptera: Apis mellifera (A. mellifera), (3) Order Coleoptera: Tribolium castaneum (T. castaneum), (4) Order Lepidoptera: Bombyx mori (B. mori), and Galleria mellonella (G. mellonella). The insect models were confirmed based on InsectBase (http://v2.insect-genome.com/Classify/Model%20Organism)34. In addition to the Mus musculus as a transition mammalian model and a distant from insect models.
Bioinformatics analyses
-
1.
Pairwise alignment was performed to detect protein homology and identify query coverage and percentage of protein identity. Alignment was performed between each protein in H. sapiens against its homolog in the selected model organisms using BLASTP (Basic Local Alignment Search Tool for protein) with default parameters from NCBI32, except (GRN and NEP2) alignments were performed against D. melanogaster because they showed no alignment against H. sapiens.
-
2.
In Silico SNP prediction of disease-causing variants was performed using the publicly available tool Meta-SNP (meta-predictor of disease-causing variants)27,35,36. This tool permits the detection of disease-associated nsSNVs for both well-identified and predicted amino acid sequences (SNPs based on dbSNP of humans) (accessed 22 June, 2023). This approach is characterised by other methods by integrating four existing methods: PANTHER, PhD-SNP, SIFT, and SNAP with defined default threshold parameters PANTHER, PhD-SNP, and Meta-SNP: Between 0 and 1 (If > 0.5, mutation is predicted Disease), SIFT: Positive Value (If > 0.05 mutation is predicted Neutral), SNAP: Output normalised between 0 and 1 (If > 0.5, mutation is predicted Disease).
-
(a).
A local alignment search was performed between the substituted amino acid in the human protein and its homolog protein in the selected insect model using BLASTP and manual search, depending on finding the best match using 5 aa before and 5 aa after the substituted amino acid to provide a proper short sequence needed to find the accurate position of required aa.
-
(b)
The matched amino acids and protein sequence were entered into the meta-SNP analysis tool to determine the probability of causing disease for amino acid substitutions according to the human nsSNPs.
-
(a).
Results
Pairwise alignment
Pairwise alignment using blastp with default parameters was conducted for each selected insect protein against its homolog in humans, except for GRN and NEP2, where pairwise alignment was conducted against the fruit fly (Tables S4, S5, and S6). A sharp cut-off value for homology, 75% query coverage, and 30% protein identity was applied to filter the results with the more meaningful values37,38,39 .
The results showed that:
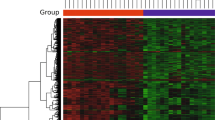
For Alzheimer’s disease Table 1: A. mellifera shows greater identity to H. sapiens than D. melanogaster for APP protein. Musca domestica has more identity with H. sapiens than D. melanogaster for COL25A1 protein. Aedes aegypti is the nearest in identity to D. melanogaster for the GRN protein. Musca domestica, and A. aegypti show greater identity to H. sapiens than D. melanogaster for HDAC6 protein. Galleria mellonella has the closest Tau/Mapt protein identity to H. sapiens beside D. melanogaster. Aedes aegypti shows greater identity to D. melanogaster for the Nep2 protein. Tribolium castaneum, and B. mori have more identical Psn1 and Psn2 to H. sapiens than D. melanogaster. For RAC1 T. castaneum, M. domestica, and G. mellonella were more identical to H. sapiens than D. melanogaster. In the absence of SORL1 in D. Melanogaster; A. mellifera, T. castaneum, and A. aegypti showed a higher identity with H. sapiens. As shown in (Fig. 2).
The heatmap shows the percentage of protein identity for AD proteins between different insect models, 1 Mus musculus, 2 Drosophila melanogaster, 3 Apis mellifera, 4 Tribolium castaneum, 5 Bombyx mori, 6 Musca domestica, 7 Galleria mellonella, 8 Anopheles gambiae, 9 Aedes aegypti. Where deep colour refers to high protein identity and light colour refers to low protein identity. The heatmap was generated using RStudio version 2022.12.0 + 353.
For Parkinson’s disease Table 2: M. domestica and A. gambiae show better protein identity to H. sapiens than D. melanogaster for DJ-1 protein. Tribolium castaneum has greater GAK, HTRA2, LRRK2, and EIF4G1 protein identity to H. sapiens than D. melanogaster. In the case of VPS35, A. mellifera showed the highest protein identity with H. sapiens. B. mori showed higher UCHL1 protein identity than D. melanogaster. For A. aegypti and G. mellonella, ATP13A2 protein showed more identity to H. sapiens than D. melanogaster. In addition, G. mellonella has a better GIGYF2 protein identity with H. sapiens than D. melanogaster. A. gambiae showed greater identity to H. sapiens than D. melanogaster for the PLA2G6 protein. As shown in Fig. 3.
The heatmap shows the percentage of protein identity for PD proteins between different insect models, 1 Mus musculus, 2 Drosophila melanogaster, 3 Apis mellifera, 4 Tribolium castaneum, 5 Bombyx mori, 6 Musca domestica, 7 Galleria mellonella, 8 Anopheles gambiae, 9 Aedes aegypti. Where deep colour refers to high protein identity and light colour refers to low protein identity. The heatmap was generated using RStudio version 2022.12.0 + 353.
For Huntington’s disease Table 3: A. mellifera has higher HTT, UBQLN2, and DMBK protein identity to H. sapiens than D. melanogaster. B. mori has better GRIK2 and ATXN1 protein identity to H. sapiens than D. melanogaster. Apis mellifera, A. gambiae, and A. aegypti were closer to H. sapiens than D. melanogaster for the VCP protein. Tribolium castaneum has greater VPS13A and ATXN3 protein identity to H. sapiens than D. melanogaster. A. gambiae showed better CACNA1A protein identity with H. sapiens than D. melanogaster. As shown in Fig. 4.
The heat map shows the percentage of protein identity for HD proteins between different insect models, 1 Mus musculus, 2 Drosophila melanogaster, 3 Apis mellifera, 4 Tribolium castaneum, 5 Bombyx mori, 6 Musca domestica, 7 Galleria mellonella, 8 Anopheles gambiae, 9 Aedes aegypti. Where deep colour refers to high protein identity and light colour refers to low protein identity. The heatmap was generated using RStudio version 2022.12.0 + 353.
In Silico nsSNPs prediction
In Silico nsSNPs prediction is performed using five integrated tools (SIFT, PANTHER, SNAP, PhD-SNP, and Meta-SNP).
Polymorphism data for APP (NP_001191231.1), LRRK2 (NP_940980.4), and VCP (NP_009057.1) proteins were retrieved from the NCBI dbSNP database as a publicly available database. Accordingly, APP was found to contain four missense SNPs in its coding regions. The LRRK2 gene was found to have one missense SNP in its coding region, and the VCP gene was found to have five missense SNPs in its coding region, but two of them rs779834525, and rs1420316004, were related to the FANCG gene and not VCP.
For Alzheimer’s disease, SNP analysis was performed on D. melanogaster App-like protein (NP_001245452.1) as a homolog of H. sapiens APP with 36.27% protein identity, using reference human SNPs rs63750264 (V > L,F,I), rs63750643 (T > A), rs63750671 (A > G), and rs193922916 (A > V,G).
-
1.
In rs63750264, Val680Phe or Val680Leu, or Val680Ile in humans matches Val at positions 94, 863, and 869 in D. melanogaster.
-
2.
In rs63750643, Thr677Ala in humans matches the Thr at position 742 in D. melanogaster.
-
3.
In rs63750671, human Ala655Gly matches Ala at positions 180, 802, and 820 in D. melanogaster.
-
4.
In rs193922916, Ala636Val or Ala636Gly in humans matches Ala at positions 138, 246, 713, and 758 in D. melanogaster.
Prediction using the SIFT, PANTHER, SNAP, PhD-SNP, and Meta-SNP tools results from 21 input-suggested mutations. Eleven mutations were predicted, Five of the 11 mutations showed deleterious or diseased points Table 4. Mutations V94F, V94L, A758G, A758V, and A820G are thought to be pathogenic in the AD D. melanogaster model according to PANTHER, Phd-SNP, and Meta-SNP while SIFT and SNAP couldn’t identify the effects of nsSNPs. In spite of the fact that A820G has the highest reliability index.
For Parkinson’s disease, SNP analysis was performed on A. aegypti LRRK1 protein (XP_021698550.1) as a homolog of H. sapiens LRRK2 with 27.47% protein identity, using the reference human SNP rs33939927 (R > S,G,C).
-
1.
In rs33939927, Arg1441Ser or Arg1441Gly, or Arg1441Cys in humans matches Arg at position 1218 in A. aegypti.
Prediction using the SIFT, PANTHER, SNAP, PhD-SNP, and Meta-SNP tools results from three input suggested mutations. Three mutations showed deleterious or diseased points Table 5. Mutations R1218C, R1218C, and R1218S are thought to be pathogenic in the PD A. aegypti model according to Phd-SNP, and Meta-SNP while PANTHER, SIFT and SNAP couldn’t identify the effects of nsSNPs. In spite of the fact that R1218C has the highest reliability index.
For Huntington’s disease, SNP analysis was performed on T. castaneum VCP-like protein (XP_008192481.1) as a homolog of H. sapiens VCP with 43.99% protein identity, using the reference human SNPs rs121909330 (R > C,G,S), rs121909334 (R > P,Q), and rs387906789 (R > C,G,S).
-
1.
In rs121909330, Arg155Cys or Arg155Gly, or Arg155Ser in humans matches Arg at positions 268, 282, and 836 in T. castaneum.
-
2.
In rs121909334, Arg191Pro or Arg191Gln in human matches 618, 639, 739, 89, 217, 743, 618, 82, 330, 411, and 462 in T. castaneum.
-
3.
In rs387906789, Arg159Cys or Arg159Gly, or Arg159Ser in humans matches Arg at positions 710, and 750 in T. castaneum.
Prediction using the SIFT, PANTHER, SNAP, PhD-SNP, and Meta-SNP tools results from 37 input-suggested mutations. Fifteen mutations were predicted, thirteen of the 15 mutations showed deleterious or diseased points Table 6. Mutations R268C, R268G, R268S, R282C, R282G, R282S, R836C, R710C, R710G, R710S, R750C, R750G, and R750S are thought to be pathogenic in the HD T. castaneum model according to PANTHER, Phd-SNP, SIFT, SNAP and Meta-SNP. In spite of the fact that R268C, R268G, R282G, R710C, R750C, R750G have the highest reliability index.
Prediction; Neutral: Neutral variants. Disease: Disease causing variants.
Outputs: Value reported under each prediction.
PANTHER, PhD-SNP, and Meta-SNP: between 0 and 1 (if > 0.5, mutation is predicted disease).
SIFT: Positive Value (If > 0.05, mutation is predicted Neutral). SNAP: Output normalised between 0 and 1 (if > 0.5, mutation is predicted disease).
RI: A Reliability Index between 0 and 10 provides a means of focusing on the most accurate predictions.
Discussion
Neurodegenerative diseases are devastating diseases which are incurable and mostly result in the death of patients. To accelerate the search for treatments and save money, effort, and time, there is a need to determine the best model that mimics human disease. In turn, this leads to improved human neural health. Pairwise alignment was applied to each protein against humans for all proteins except (GRN and NEP2) against the fruit fly because they showed no alignment against H. sapiens. We determined the best insect for studying each protein separately by selecting the highest query coverage with the highest protein identity.
In this study, a total of eight insect models were used to find out which of them is the best to model each of AD, PD and HD.
For Alzheimer’s, the best overall two models according to the average protein identity percentage for the 10 selected proteins were D. melanogaster then A. gambiae. Drosophila melanogaster is believed to have nearly 75% of human disease-causing genes functional homologs15,40,41. The fruitfly showed a high protein identity to human with reasonable query coverage in GRN, COL25A1, MAPT and RAC1. They can express different phenotypes of induced AD15. From the 10 proteins, APP was selected as a representative of AD related proteins in human. The analysis of nsSNPs related to APPl protein in the fruit fly showed predicted pathogenic nsSNPs (V94F, V94L, A758G, A758V, and A820G) that could be used for further studies on the induction of familial forms of early-onset Alzheimer's disease and cerebral amyloid angiopathy, and study the factors that increase total Aβ levels42,43. Anopheles gambiae is known to become an important model organism for the study of insect-parasite interactions and innate immune responses to protozoan parasites44. Anopheles gambiae shows better protein identity to H. sapiens than D. melanogaster for DJ-1, VCP and PLA2G6 proteins. Moreover, A. gambiae infection with Toxoplasma gondii promotes the accumulation of glutamate. Glutamate is a neurotransmitter in the brain that triggers neurodegenerative diseases such as Alzheimer’s disease and Parkinson’s disease in individuals predisposed to such conditions45. Thus in turn makes A. gambiae a potential model to study the pathology of these AD.
For Parkinson’s, the best two models according to the average protein identity percentage for the 13 selected proteins were A. aegypti then A. mellifera. A. aegypti has an advanced nervous system, with sensory organs used to locate their hosts in their environment46. On applying a sublethal dose of spinosyn insecticides to A. aegypti. Parkinson's disease-related genes were significantly enriched in spinetoram-exposed mosquitoes compared with controls47. Through our studies, it showed a high protein identity to human with reasonable query coverage for PARK6, VPS35, ATP13A2 and PLA2G6. From the 13 proteins, LRRK2 was selected as a representative of PD related proteins in human. The analysis of nsSNPs related to LRRK1 protein in the yellow fever mosquito showed predicted pathogenic nsSNPs (R1218C, R1218C, and R1218S) that could be used for induction of PD through mutations in the catalytic domains that may result in hyperactivation of the kinase domain, and show Lewy Body pathology48. Apis mellifera is more similar to vertebrates in terms of RNA (Ribonucleic acid) interference, DNA (Deoxyribonucleic Acid) methylation, and circadian rhythm49. It showed a high protein identity to human with reasonable query coverage in PARK2, VPS35 and ATP13A2. Honey bees’ ethanol exposure causes changes in their body and wing kinematics50. Mechanisms identified in the cellular stress response to ethanol, such as the oxidative stress response, are also involved in Parkinson’s disease51. Apis mellifera is a key social behavioural model that displays sophisticated cognitive abilities52. This makes it possible to analyse the changes occurring in honeybee brains during learning and remembering and increases the opportunity to be used also as a model for AD, along with the ability to identify new genome-based single-nucleotide polymorphisms (SNPs)14,53.
For Huntington’s, T. castaneum then B. mori were the best models according to the average protein identity percentage. Tribolium castaneum has more olfactory receptors and detoxification genes than D. melanogaster and other insects and may be better adapted to its environment45. It shows a higher genetic homology to humans when compared to other invertebrate models, such as D. melanogaster54. Therefore, T. castaneum is one of the most suitable genetic models for post-genomic studies such as proteomics and functional genomics. It showed a high protein identity to human with reasonable query coverage in GRIK2, VPS13A and UBQLN2. From the 10 proteins, VCP was selected as a representative of HD related proteins in human. The analysis of SNPs related to VCPl protein in the Red flour beetle revealed predicted pathogenic nsSNPs (R268C, R268G, R268S, R282C, R282G, R282S, R836C, R710C, R710G, R710S, R750C, R750G, and R750S) that could be used for further studies on the gene role in cell division, the cell apoptosis, repairing damaged DNA, and formation of abnormal proteins build up in muscle, bone and brain cells that lead to induction of HD. These protein aggregations interfere with the normal functions of the brain cells55,56. The PINK1 protein from the T. castaneum beetle (TcPINK1) exhibits catalytic activity toward ubiquitin, parkin, and generic substrates and provides a basis for further studies on human Parkinson’s disease57. Bombyx mori shares 58% of diseased human homologs genes, which are related to neurodegenerative diseases such as HD, oxidative stress, and protein degradation-associated genes58. Bombyx mori has higher identical VPS35, and UCHL1 to H. sapiens than D. melanogaster. Downregulation of the DJ-1 gene causes p-translucent silkworm as a result of increased oxidative stress response of the body, which leads to oxidative damage to the nerves and tissues17,18.
Galleria mellonella didn’t represent the best model for any of the three studied NDs, although it has a similar innate immune response to that of mammals, regardless of whether it evolved separately from mammals several thousand years ago29,30,31. Comparative studies of genomes have shown that it has numerous homologues of human genes encoding proteins involved in pathogen recognition or signal transduction59,60. According to our study, it showed a high protein identity to human with reasonable query coverage in MAPT, ATP13A2, GIGYF2 and RAC1. In addition, its larvae can cultivate Bacteria such as Borrelia burgdorferi61, Enterococcus faecalis62, and Staphylococcus aureus63, which are believed to play a role in neuroinflammation and may contribute to AD.
Musca domestica has a strong immune system and has been used as a model to investigate the presence of enhanced detoxification64. Applying its larval extract on an AD diseased mouse has therapeutic effects against memory impairment, structural damage, and oxidative stress65. According to our study, it showed a high protein identity to human with reasonable query coverage in RAC1, COL25A1, HDAC6, DJ-1, GRIK2, VPS13A, VCP and UBQLN2.
These findings will assist in the selection of the best model for further studies in simulation diseases, deep understanding for mutations and their effects and how to fix them genetically or through improving drug discovery. The average percentage of protein identity between the different insect models and the selected proteins is provided in the supplementary data, as shown in Figs. 5, and 6.
Conclusion
The increasing prevalence of neurodegenerative diseases such as Alzheimer's, Parkinson’s and Huntington's necessitates improvement in our understanding of these diseases. The research strategy for NDs is two-armed; one of them focuses on finding actual treatments that work on delaying symptoms or preventing disease development, whereas the other depends on searching for tools that can be used to detect the earliest and indirect signs of the disease and this is our point. Thus, it is crucial to simulate the disease, identify the counterparts of human diseased genes, test and apply their findings to easily handled model organisms. Comparative analysis has the potential to improve research and drug development for human diseases.
In this study, a total of 61 SNPs were checked in APPl, LRRK1 and VCPl proteins of D. melanogaster, A. aegypti and T. castaneum respectively by five prediction tools; 21 out of 29 SNPs showed a deleterious effect and 8 of the 21 showed high reliability index. For the 21 deleterious nsSNPs, most of them are located on the functional domains of the proteins.
Although mammalian models are more similar to humans, insects are often preferred because of their shorter lifespan and fewer ethical constraints. Human insect disease models provide new tools for drug discovery to overcome current limitations by using them at different stages as models that show a significant response to many drugs that act on the mammalian central nervous system (CNS) instead of differences in their brains, which allows researchers to find new therapeutic strategies.
In conclusion A. mellifera, T. castaneum, B. mori, A. aegypti besides D. melanogaster have promising future in the field of medical research and provide valuable insights into common neurodegenerative diseases as AD and PD and rare diseases as HD. This study provides comprehensive information on the available insect models on the protein-level resources and analysis of the predicted functional nsSNPs to improve human neural health by finding the best insect model to study Alzheimer’s disease, Parkinson’s disease, and Huntington’s disease, and to find answers to complex biological questions as the functional impacts of these variants. This will happen by using the findings of the predicted nsSNPs for example to enhance wet-labs experiments and detect the proper position to be knocked down and find out the pathological effects of it and on determining the possible affected genes or proteins on induction of one of the NDs in its proper models.
Recommendation
To maximise the benefits, we recommend the provision of stock centres of different insect models, mutant and transgenic strains, microarrays, or RNA interference libraries, and working on updating annotations, providing more genome sequencing and assembly of sequenced insects. Additionally, we recommend the development of tools specific to insect model organisms.
Data availability
All retrieved data (Human data or models data) are from publicly available databases. All data generated or analysed during this study are included in this published article [and its supplementary information files].
References
Checkoway, H., Lundin, J. I. & Kelada, S. N. Neurodegenerative diseases. IARC Sci. Publ. 163, 407–419 (2011).
Ali, A. M. & Kunugi, H. Royal jelly as an intelligent anti-aging agent—A focus on cognitive aging and Alzheimer’s disease: A review. Antioxidants 9(10), 1–46. https://doi.org/10.3390/antiox9100937 (2020).
Chekani, F., Bali, V. & Aparasu, R. R. Quality of life of patients with Parkinson’s disease and neurodegenerative dementia: A nationally representative study. Res. Soc. Adm. Pharm. 12(4), 604–613. https://doi.org/10.1016/j.sapharm.2015.09.007 (2016).
Denell, R. Establishment of tribolium as a genetic model system and its early contributions to evo-devo. Genetics 180(4), 1779–1786. https://doi.org/10.1534/genetics.104.98673 (2008).
Bingsohn, L., Knorr, E. & Vilcinskas, A. The model beetle Tribolium castaneum can be used as an early warning system for transgenerational epigenetic side effects caused by pharmaceuticals. Comp. Biochem. Physiol. Part C Toxicol. Pharmacol. 185, 57–64 (2016).
Nitta, Y. & Sugie, A. Studies of neurodegenerative diseases using Drosophila and the development of novel approaches for their analysis. Fly 16(1), 275–298 (2022).
Roy, S. Genomics and bioinformatics in entomology. Entomol. Ornithol. Herpetol. Curr. Res. https://doi.org/10.4172/2161-0983.1000e107 (2013).
Severson, D. W. & Behura, S. K. Mosquito genomics: Progress and challenges. Annu. Rev. Entomol. 57, 143–166 (2012).
Michels Thompson, L. & Marsh, J. L. Invertebrate models of neurologic disease: Insights into pathogenesis and therapy. Curr. Neurol. Neurosci. Rep. 3, 442–448 (2003).
Bridi, J. C. et al. Ancestral regulatory mechanisms specify conserved midbrain circuitry in arthropods and vertebrates. Proc. Natl. Acad. Sci. U.S.A. 117(32), 19544–19555. https://doi.org/10.1073/pnas.1918797117 (2020).
Strausfeld, N. J. & Hirth, F. Deep homology of arthropod central complex and vertebrate basal ganglia. Science (New York, N.Y.) 340(6129), 157–161. https://doi.org/10.1126/science.1231828 (2013).
Daniel, S. & Seil, C. (n.d.). The Strikingly Similar Brains of Flies and Men. Retrieved September 19, 2022, from http://www.sciencemag.org/content/340/6129/157.short
Brandt, A., Joop, G. & Vilcinskas, A. Tribolium castaneum as a whole-animal screening system for the detection and characterization of neuroprotective substances. Arch. Insect Biochem. Physiol. https://doi.org/10.1002/arch.21532 (2019).
Lee, H. Y., Lee, S. H. & Min, K. J. Insects as a model system for aging studies. Entomol. Res. 45(1), 1–8. https://doi.org/10.1111/1748-5967.12088 (2015).
Pandey, U. B. & Nichols, C. D. Human disease models in drosophila melanogaster and the role of the fly in therapeutic drug discovery. Pharmacol. Rev. 63(2), 411–436. https://doi.org/10.1124/pr.110.003293 (2011).
Ahmed, A., Ghallab, E. H., Shehata, M., Zekri, A. R. N. & Ahmed, O. S. Impact of nano-conjugate on Drosophila for early diagnosis of Alzheimer’s disease. Nanotechnology 31(36), 365102 (2020).
Chen, W. W., Zhang, X. I. A. & Huang, W. J. Role of neuroinflammation in neurodegenerative diseases. Mol. Med. Rep. 13(4), 3391–3396 (2016).
Meng, X., Zhu, F. & Chen, K. Silkworm: A promising model organism in life science. J. Insect Sci. 17(5), 97 (2017).
Jia, M. et al. Computational analysis of functional single nucleotide polymorphisms associated with the CYP11B2 gene. PLoS ONE 9(8), e104311 (2014).
Mooney, S. D., Krishnan, V. G. & Evani, U. S. Bioinformatic tools for identifying disease gene and SNP candidates. Methods Mol. Biol. 628, 307–319. https://doi.org/10.1007/978-1-60327-367-1_17 (2010).
Bromberg, Y. Chapter 15: Disease gene prioritization. In PLoS Computational Biology. https://doi.org/10.1371/journal.pcbi.1002902 (2013).
Tey, H. J. & Ng, C. H. Computational analysis of functional SNPs in Alzheimer’s disease-associated endocytosis genes. PeerJ https://doi.org/10.7717/peerj.7667 (2019).
Sim, N. L. et al. SIFT web server: Predicting effects of amino acid substitutions on proteins. Nucleic Acids Res. 40(W1), W452–W457 (2012).
Thomas, P. D. et al. PANTHER: A library of protein families and subfamilies indexed by function. Genome Res. 13(9), 2129–2141 (2003).
Bromberg, Y. & Rost, B. SNAP: Predict effect of non-synonymous polymorphisms on function. Nucleic Acids Res. 35(11), 3823–3835 (2007).
Capriotti, E. & Fariselli, P. PhD-SNPg: A webserver and lightweight tool for scoring single nucleotide variants. Nucleic Acids Res. 45(W1), W247–W252 (2017).
Capriotti, E., Altman, R. B. & Bromberg, Y. Collective judgment predicts disease-associated single nucleotide variants. BMC Genom. 14, 1–9 (2013).
Hoeppner, M. A. NCBI Bookshelf: Books and documents in life sciences and health care. Nucleic Acids Res. 41(D1), D1251–D1260 (2012).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28(1), 27–30. https://doi.org/10.1093/nar/28.1.27 (2000).
Kanehisa, M. Toward understanding the origin and evolution of cellular organisms. Protein Sci. Publ. Protein Soc. 28(11), 1947–1951. https://doi.org/10.1002/pro.3715 (2019).
Kanehisa, M., Furumichi, M., Sato, Y., Kawashima, M. & Ishiguro-Watanabe, M. KEGG for taxonomy-based analysis of pathways and genomes. Nucleic Acids Res. 51(D1), D587–D592. https://doi.org/10.1093/nar/gkac963 (2023).
Johnson, M. et al. NCBI BLAST: A better web interface. Nucleic Acids Res. 36(suppl_2), W5–W9 (2008).
Gaston, K. J. The magnitude of global insect species richness. Conserv. Biol. 5(3), 283–296 (1991).
Mei, Y. et al. InsectBase 2.0: A comprehensive gene resource for insects. Nucleic Acids Res. 50(D1), D1040–D1045. https://doi.org/10.1093/nar/gkab1090 (2022).
Hall, M. et al. The WEKA data mining software: An update. ACM SIGKDD Explor. Newsl. 11(1), 10–18 (2009).
Pei, J. & Grishin, N. V. AL2CO: Calculation of positional conservation in a protein sequence alignment. Bioinformatics 17(8), 700–712 (2001).
Pearson W. R. An introduction to sequence similarity (“homology”) searching. In Current Protocols in Bioinformatics, Chapter 3, 3.1.1–3.1.8. https://doi.org/10.1002/0471250953.bi0301s42 (2013).
Kilinc, M., Jia, K. & Jernigan, R. L. Improved global protein homolog detection with major gains in function identification. Proc. Natl. Acad. Sci. 120(9), e2211823120 (2023).
Novoa, E. M., Pouplana, L. R. D., Barril, X. & Orozco, M. Ensemble docking from homology models. J. Chem. Theory Comput. 6(8), 2547–2557 (2010).
Dhankhar, J., Agrawal, N. & Shrivastava, A. An interplay between immune response and neurodegenerative disease progression: An assessment using Drosophila as a model. J. Neuroimmunol. https://doi.org/10.1016/j.jneuroim.2020.577302 (2020).
Yamamoto, S. et al. A drosophila genetic resource of mutants to study mechanisms underlying human genetic diseases. Cell 159(1), 200–214 (2014).
Müller, U. C. & Zheng, H. Physiological functions of APP family proteins. Cold Spring Harbor Perspect. Med. 2(2), a006288. https://doi.org/10.1101/cshperspect.a006288 (2012).
Giri, M., Zhang, M. & Lü, Y. Genes associated with Alzheimer’s disease: An overview and current status. Clin. Interv. Aging 11, 665–681. https://doi.org/10.2147/CIA.S105769 (2016).
Sharakhova, M. V. et al. Update of the Anopheles gambiae PEST genome assembly. Genome Biol. 8(1), R5. https://doi.org/10.1186/gb-2007-8-1-r5 (2007).
Li, F. et al. Insect genomes: progress and challenges. Insect Mol. Biol. 28(6), 739–758. https://doi.org/10.1111/imb.12599 (2019).
Matthews, B. J., McBride, C. S., DeGennaro, M., Despo, O. & Vosshall, L. B. The neurotranscriptome of the Aedes aegypti mosquito. BMC Genom. 17, 32. https://doi.org/10.1186/s12864-015-2239-0 (2016).
Wang, L. et al. Sublethal exposure to spinetoram impacts life history traits and dengue virus replication in Aedes aegypti. Insect Sci. 30(2), 486–500 (2023).
Jia, F., Fellner, A. & Kumar, K. R. Monogenic Parkinson’s disease: Genotype, phenotype, pathophysiology, and genetic testing. Genes 13(3), 471. https://doi.org/10.3390/genes13030471 (2022).
Łoś, A., Bieńkowska, M. & Strachecka, A. Honey bee (Apis mellifera) as an alternative model invertebrate organism. Medycyna Weterynaryjna 75(2), 93–106 (2019).
Ahmed, I., Abramson, C. I. & Faruque, I. A. Honey bee flights near hover under ethanol-exposure show changes in body and wing kinematics. PLoS One 17(12), e0278916. https://doi.org/10.1371/journal.pone.0278916 (2022).
Peng, B. et al. Role of alcohol drinking in Alzheimer’s disease, Parkinson’s disease, and amyotrophic lateral sclerosis. Int. J. Mol. Sci. 21(7), 2316 (2020).
Shpigler, H. Y. et al. Behavioral, transcriptomic and epigenetic responses to social challenge in honey bees. Genes Brain Behav. 16(6), 579–591. https://doi.org/10.1111/gbb.12379 (2017).
Honeybee Genome Sequencing Consortium. Insights into social insects from the genome of the honeybee Apis mellifera. Nature 443(7114), 931–949 (2006).
Grünwald, S. et al. The red flour beetle Tribolium castaneum as a model to monitor food safety and functionality. In Yellow Biotechnology I: Insect Biotechnologie in Drug Discovery and Preclinical Research (ed. Vilcinskas, A.) 111–122 (Springer, 2013).
Ju, J. S., Miller, S. E., Hanson, P. I. & Weihl, C. C. Impaired protein aggregate handling and clearance underlie the pathogenesis of p97/VCP-associated disease. J. Biol. Chem. 283(44), 30289–30299. https://doi.org/10.1074/jbc.M805517200 (2008).
Meyer, H. & Weihl, C. C. The VCP/p97 system at a glance: Connecting cellular function to disease pathogenesis. J. Cell Sci. 127(Pt 18), 3877–3883. https://doi.org/10.1242/jcs.093831 (2014).
Adamski, Z. et al. Beetles as model organisms in physiological, biomedical and environmental studies—A review. Front. Physiol. https://doi.org/10.3389/fphys.2019.00319 (2019).
Tabunoki, H., Bono, H., Ito, K. & Yokoyama, T. Can the silkworm (Bombyx mori) be used as a human disease model?. Drug Discov. Ther. 10(1), 3–8. https://doi.org/10.5582/ddt.2016.01011 (2016).
Singkum, P., Suwanmanee, S., Pumeesat, P. & Luplertlop, N. A powerful in vivo alternative model in scientific research: Galleria mellonella. Acta Microbiol. Immunol. Hung. 66(1), 31–55 (2019).
Serrano, I., Verdial, C., Tavares, L. & Oliveira, M. The virtuous Galleria mellonella model for scientific experimentation. Antibiotics 12(3), 505. https://doi.org/10.3390/antibiotics12030505 (2023).
Chakravarthi, S. T. & Joshi, S. G. An association of pathogens and biofilms with Alzheimer’s disease. Microorganisms 10(1), 56 (2021).
Underly, R., Song, M. S., Dunbar, G. L. & Weaver, C. L. Expression of Alzheimer-type neurofibrillary epitopes in primary rat cortical neurons following infection with Enterococcus faecalis. Front. Aging Neurosci. 7, 259 (2016).
Zubair Alam, M. et al. Infectious agents and neurodegenerative diseases: Exploring the links. Curr. Top. Med. Chem. 17(12), 1390–1399 (2017).
Xu, Y., Tao, S., Hinkle, N., Harrison, M. & Chen, J. Salmonella, including antibiotic-resistant Salmonella, from flies captured from cattle farms in Georgia, U.S.A. Sci. Total Environ. 616–617, 90–96. https://doi.org/10.1016/j.scitotenv.2017.10.324 (2018).
Tang, Y. et al. The protective effects of protein-enriched fraction from housefly (Musca domestica) against aged-related brain aging. J. Nutr. Sci. Vitaminol. 66(5), 409–416. https://doi.org/10.3177/jnsv.66.409 (2020).
Acknowledgements
This research is financially supported and funded by the Academy of Scientific Research and Technology (ASRT), Scholarship of scientist of next generation (SNG) cycle 6, Grant no. ASRT/SNG/BGM/2018-8. I would like to express appreciation to Dr. Enas Ghalab for her academic advice and assistance during the planning and development of this research work.
Funding
Open access funding provided by The Science, Technology & Innovation Funding Authority (STDF) in cooperation with The Egyptian Knowledge Bank (EKB).
Author information
Authors and Affiliations
Contributions
M.E.-H. designed the work flow and fellow up its progress, M.G.S. revised the work and edited it, E.A.A.A. performed analyses, interpreted the results and wrote, and M.G.S. substantively revised the work. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Al-Ayari, E.A., Shehata, M.G., EL-Hadidi, M. et al. In silico SNP prediction of selected protein orthologues in insect models for Alzheimer's, Parkinson's, and Huntington’s diseases. Sci Rep 13, 18986 (2023). https://doi.org/10.1038/s41598-023-46250-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-46250-5
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.