Abstract
Humans have intensively sailed the Mediterranean and European Atlantic waters throughout history, from the upper Paleolithic until today and centuries of human seafaring have established complex coastal and cross-seas navigation networks. Historical literature revealed three major long-lasting maritime routes (eastern, western, northern) with four commencing locations (Alexandria, Venice, Genoa, Gibraltar) and a fourth route (circum-Italian) that connected between them. Due to oceangoing and technological constraints, most voyages were coastal, lasted weeks to months, with extended resting periods, allowing the development of fouling organisms on ship hulls. One of the abiding travellers in maritime routes is the colonial ascidian Botryllus schlosseri already known since the eighteenth century in European and Mediterranean ports. This species, was almost certainly one of the common hull fouling travellers in all trade routes for centuries. Employing COI haplotypes (1008 samples) and microsatellite alleles (995 samples) on colonies sampled from 64 pan-European sites, present-day Botryllus populations in the Mediterranean Sea/European Atlantic revealed significant segregation between all four maritime routes with a conspicuous partition of the northern route. These results reveal that past anthropogenic transports of sedentary marine species throughout millennia long seafaring have left their footprint on contemporary seascape genetics of marine organisms.
Similar content being viewed by others
Introduction
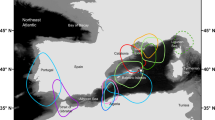
It is not really known when humans began cross sailing the Mediterranean Sea and European Atlantic, albeit the availability of documentation from the upper Paleolithic period from about 11,000 BCE1, and crossing the English Channel, as well as from the Bronze Age, about 1500 BCE2,3. Yet, it was only in the ninth century BCE that the “Phoenician commerce across the Mediterranean and beyond had taken off”1 with movement from Tyre, Cyprus, Rhodes, Crete, the Ionian Sea to Sicily, the Balearic Islands, the Straits of Gibraltar to Cadiz and even further north. More or less during the same period, the Greeks crossed the Ionian Sea and travelled to Italy1 and further west through the Straits of Gibraltar towards Atlantic Europe4,5. The traffic in the Mediterranean was augmented during the Roman Empire by thousands of ships that sailed from “anywhere to everywhere”, carrying merchandises6,7,8, people and troops. this was followed by bustling maritime traffic during the Byzantine and Middle-ages, governed by the port-hubs of Venice, Genoa and Pisa alongside with the Mamluk and Ottoman Empires1. The sea going ties between Western Europe (Atlantic coasts) and the Mediterranean intensified during the Viking age (793–1066 CE) when the Norse people of Scandinavia explored, traded with and raided Britain, Iberia, southern coasts of France and northern Italy9,10. Starting from the thirteenth century, the Italian state cities of Genoa and Venice established extensive maritime trading relations with Northern Europe1,11,12. Thus, centuries of trade routes have established complex networks of European navigation courses with prominent trajectories along the coasts and cross-seas (Fig. 1).
Due to oceanographic constraints (waves, currents, winds, storms), readiness of navigation tools, ship design, oars and sail usage, most ships travelled for centuries along the same sea routes and sailing was almost entirely coastal (Fig. 1). As a result, en route trips lasted for weeks to months, relying on crews’ experience and land identifiable topography1,7,13,14. These slow voyages were repeatedly and frequently halted for water and food refills, resting and long mooring periods, creating ideal conditions for the accumulation of fouling marine organisms and their perpetuation on submerged surfaces. Slow sailing also contributed to this phenomenon. Hull fouling is known from ancient times, already marked by the Phoenicians, the Greeks and the Romans, and centuries thereafter15,16.
One of the notorious invasive species of the hull fouling consortia is the colonial ascidian Botryllus schlosseri17,18,19,20, a common species in marinas and ports worldwide, easily spotted on submerged manmade objects like ropes, buoys and floating docks. Besides being found on boat hulls, Botryllus schlosseri’s spreading mode doesn't exhibit any sort of distance-based isolation, thus suggesting that its distribution is likely a result of human-mediated dispersal, primarily facilitated by the ongoing movement of vessels fouled by sedentary marine organisms21. This species, most likely of a European or Mediterranean Sea origin22,23, was almost certainly one of the common hull fouling travelers in all trade routes for centuries. It is thus implied that the interminable trafficking in the Mediterranean Sea and European Atlantic maritime routes may have carried its footprint forward towards the current seascape genetics of this species, reflecting long-term balance between immigration, extinction and diversification. Hence, past human-mediated dispersal activities may reveal current “historical inference from genetic data”24. Studying the present-day Botryllus schlosseri populations in the Mediterranean Sea and European Atlantic21,23,25,26,27,28,29,30, we searched here for past maritime routes’ footmarks on current portraits of COI haplotypes and microsatellite alleles repertoires.
Materials and methods
COI—animal sampling and DNA extraction
Samples were collected from Botryllus schlosseri colonies residing in 23 Mediterranean Sea and Atlantic European sites (Fig. 1; Appendix Table 1), from the underside of stones or floating docks, submerged ropes and buoys in marinas, usually 0.1–0.5 m below sea level. Sampling was restricted to specimen > 1 m away from each other’s, to avoid kin colonies30,31 or ramets of the same genotype. Tissue samples were removed from the substrates using single-edge razor blades, and placed, individually, in 1.5 ml vials containing 240 µl of lysis buffer (0.25 M Trisborat pH 8.2, 0.1 M EDTA, 2% SDS, 0.1 M NaCl and 0.5 M NaClO4), homogenized, and equal volumes of Phenol/Chloroform/isoamyl alcohol (25:24:1) were added through mixing. The vials were shipped to the laboratory at the National Institute of Oceanography, Haifa, Israel and kept at 4 °C until used. Genomic DNA was extracted according to30,32, quality was evaluated using gel electrophoresis and nanodrop, and samples were kept as stock at 4 °C.
COI amplification and analyses
DNA dilutions (1:50 or 1:100) were used in sterile double distilled water. A partial sequence of ~ 700 bp of the COI gene was amplified, using the marine invertebrates COI universal primers (HCO2198r, 5′TAAACTTCAGGG TGACCAAAAAATCA 3′ and LCO1490f., 5′GGTCAACAAATCATAAAGATATTGG 3′33). The PCR reaction was performed in 50 µl REDTaq Readymix solution (Sigma), containing 0.5 µM of each primer and 100–500 ng of template DNA. A single soak at 94 °C for 2 min was followed by 40 cycles of 94 °C for 1 min, 50 °C for 1 min and 72 °C for 1 min 30 s and a final extension step at 72 °C for 10 min. PCR products were screened on 1.25% agarose gel and successful products were sent for sequencing (Macrogen Inc., South Korea). BioEdit34 and ClustalX35) packages were used for sequence analyses and multiple alignments of the COI sequences.
GenBank COI entries
GenBank deposited COI entries pertaining to additional 33 Mediterranean and East Atlantic populations were mined (Appendix Table 1), including the information for sampling sites, articles details and authors.
Construction of putative maritime routes
Historical maritime data for shipping intensity, directions and main ports across the Mediterranean Sea and European Atlantic coasts as well as oceanographic information (winds directions and sea currents), covering > 3000 years, from the second millennium BCE until the eighteenth century CE, were mined from the literature. This maritime data, and primarily selected overviewing 16 historical sources (books, articles and websites; Appendix Text) and additional 10 sources (list of reference), enabled us to construct major pan European putative maritime routs.
COI Data analyses
Data analysis was performed by implementing four consecutive steps: (1) Fisher Exact test and Cramer’s V effect size test examined the differences between the maritime routes using SPSS-23 software; (2) genetic distances, population structures and differences between maritime routes, including principal coordinate analysis (PcoA), were employed by using GenAlEx version 6.5 software36; (3) genetic population structures (no prior assumption) were performed with NetStruct37), to construct inter-individual genetic similarity networks, clustering them into groups of genetically similar individuals; (4) cluster and geneflow analyses were performed with BAPS software38).
Microsatellites data analysis
Data pertaining to the distribution of four microsatellites in 32 sampled sites along the coasts of Scandinavia, western Europe, England and the Mediterranean21,23,25,29; Fig. 1) corresponded almost perfectly to the four chosen historical shipping routes of this study. The data was re-analyzed, using SPSS-23 software for Fisher Exact test and Cramer’s V effect size test, and GeneAlEx 6.5 was employed for the principal coordinate analysis, Fst, Nei genetic distance and Dest. Netstruct and BAPS software were used for cluster analysis and Geneflow.
Results
Historical shipping routes
The historical maritime data for shipping intensity and directions across the Mediterranean Sea and European Atlantic coasts1,4,5,7; Appendix Text) revealed that: (1) The Phoenicians sailed in the Mediterranean eastern basin starting in the fifteenth century BCE, and gradually moved westward, along the North African and European coasts towards Iberia8); (2) Starting from the first millennium BCE, the Greeks sailed in the same Phoenician routes, at first in the eastern basin and later, along European coasts towards Italy, France and Iberia; (3) Very intensive maritime traffic during the Roman Empire (27 BCE-476 CE), with ships sailing in all directions, and intensive cabotage by small vessels7. In addition, brisk traffic was also found in the eastern basin (e.g., 140–150 ships/year from Alexandria to Rome carrying wheat and barley grains6,8; Appendix text 7); 4. At the second half of the thirteenth century CE, Italian vessels (primarily from Genoa) crossed the straits of Gibraltar towards the British Iles, Scandinavia and the Baltics. Simultaneously, Venice dominated the eastward lanes; 5. During the fifteenth and sixteenth centuries, there was a substantial increase in traffic intensity (mainly cargo vessels) between the Italian city states of Genoa, Pisa and Venice and Atlantic European states in the North Sea, the Baltic states and Sweden; 6. During the eighteenth century, an intensive maritime trade existed between Marseilles and Atlantic Europe (averaging 183 ships/year39; Appendix text 6).
The above and additional detailed literature (Appendix Text) enabled us to draw three putative Mediterranean/Atlantic Europe major maritime routes (and three starting reference sites) and a fourth route that connected between two major routes (Fig. 1), as follows: (1) Eastern route-the port of Venice as the reference starting point for eastern Mediterranean trajectories, with 6 posteriorly COI sampling sites and 5 microsatellites sampling sites in Sicily, the Adriatic, Greece and the Levant; (2) Western route-the port of Genoa as the reference starting point for western Mediterranean trajectories, including the ports of Sicily, Rome, Marseilles, Barcelona and Gibraltar with 18 posteriorly COI sampling sites and 7 microsatellites sampling sites; (3) Northern route-the port of Gibraltar as the reference starting point inclusive 33 posteriorly COI sampling sites and 21 microsatellites sampling sites along the Atlantic-Iberian coasts, the English-channel, Ireland, Scotland, Holland, Germany and Scandinavia; (4) Circum-Italian route-connecting the eastern and western Mediterranean routes, revealing shipping trajectories that hugged the Italian peninsula between Venice and Genoa with vessels trafficking both westward and eastward and back since the Roman times (six COI sampling sites and four microsatellites sampling sites). Thus, some of the Italian sites included in eastern or western routes were also included in the circum-Italian route (Fig. 1; Appendix Table 1).
COI analyses
COI sequences from 1008 Botryllus schlosseri samples (419 this study, 589 GenBank deposited, alignment length: 473 bp) were assigned to 57 sites in three marine/oceanic regions: East Mediterranean, West Mediterranean and Atlantic Europe (including Spain, France, England, Ireland, Holland, Germany, Scotland, Scandinavia) along the four given routes (Table 1; Appendix Table 1). Results revealed 101 COI haplotypes. As singletons (66; appeared just once in a single sampling site) and route-private haplotypes (24; appeared in only a single route) might have biased conclusions by reinforcing route disparity through local mutations or irregular connectivity episodes. Therefore they were excluded from the analyses which resulted in 11 haplotypes (Accession numbers are found in Appendix Table 2) that appeared in at least two out of the four maritime routes and a total of 818 specimens (Table 1a, b). A second outcome was the highly diverse distributions of the COI haplotypes among the oceanic regions (Table 1b).
Fisher Exact test was then employed in two consecutive steps: the first, an overall test, resulted in highly significant differences (p = 0.000) and strong effect size (Cramer’s V = 0.57) among the routes. In the second, we analysed all six pairwise post-hoc tests between the four maritime routes, corrected for multiple testing, and revealed, again, high significant disparity of COI haplotypes between the routes (p = 0.000 Appendix Tables 3–9). The overall differentiation (PhiPT) value based on 999 permutations among the four maritime routes was 0.076 (p = 0.001) and pairwise PhiPT values ranged 0.054–0.120 (p < 0.002; Table 2). Based on Nei genetic distance the PcoA, with axes 1 and 2 capturing 93.7% of the explained variation, portrayed four distinct groups of haplotypes, fully matching the four maritime routes (Table 2; Fig. 2a). Next, a Netstruct analysis (Fig. 3a) was performed at ascending threshold levels between 0.001 to 0.017 (above which the network size of 818 samples collapsed) revealing two clusters of haplotypes for each maritime route at edge pruning threshold ranging 0.001–0.011, while at the highest edge pruning threshold of 0.017, 2,5,7, and 9 clusters appeared in the circum-Italian, the western, the eastern and the northern routes, respectively (p = 0.0001; Fig. 3a; Appendix Table 10), corresponding with haplotype numbers in the circum-Italian, and western routes (Table 1a), but not with the eastern and northern routes (9 and 11; Table 1a).
Principal coordinate analyses of the four maritime trade routes based on pairwise Nei genetic distance. (a) COI, revealing cumulative 93.7% of the explained variation in axes 1 and 2; (b) microsatellite alleles, revealing cumulative 96.9% of the explained variation in axes 1 and 2. Each of the four coloured cubes represents one of the major sailing trajectories.
Population genetics analyses on COI haplotypes (a–c) and microsatellite alleles (d–f). (a) Netstruct results showing the number of clusters for three threshold values; (b) BAPS results. The seven clusters as distributed in the four maritime routes. Cluster identities are depicted in (c). Colour widths are proportional to the number of individuals within each cluster; (c) BAPS; zero geneflow rates among the clusters. (d) Netstruct results showing the number of clusters for three threshold values. (e) BAPS results. Clusters in each of the four maritime routes. The eastern and circum-Italian routes form one cluster. Colour widths are proportional to the number of individuals within the cluster. (f) Total number of clusters and geneflow rates among and within-clusters.
BAPS analysis (Fig. 3b) revealed 2, 5, 5 and 7 clusters in the circum-Italian, eastern, western and the northern routes, respectively, corresponding with haplotype numbers of the circum-Italian and western routes but not with the eastern and northern routes (Fig. 3b; Table 1a). The analysis further enabled us to calculate the amount of geneflow among the 7 emerged clusters, revealing the absence of any geneflow (Fig. 3c). Thus, although colonies that carry the same haplotypes or share identical cluster appear in more than one route, they probably do not admix. Netstruct and BAPS analyses further revealed that the number of clusters in the northern and eastern routes is smaller than the number of COI haplotypes (Table 1a).
Microsatellites analyses
Using published results21,23,25,29, an inventory of 995 Botryllus schlosseri samples was analyzed on four microsatellite alleles: Pb29, Pb49, Pb41 and Bs811. These samples were collected from 32 sites, 5, 7, 20 in the eastern, western, and northern routes, respectively. Samples from 22 of these sites were also included in the COI analyses. In a similar way to the outcomes of COI haplotypes, the microsatellite alleles frequencies differed substantially among the routes (Appendix Table 15).
Fisher Exact test revealed that the four routes differed for each microsatellite locus (p = 0.000; Cramer’s V effect sizes range between moderate 0.33 to strong 0.60; Appendix Tables 11–14). Microsatellites basic information is summarized in Appendix Table 16. Overall population structure Fst value among the four maritime routes based on 999 permutations was 0.074 (p = 0.001) and pairwise Fst values varied between 0.013–0.114, (p = 0.001; Table 2). Overall Dest value was 0.396 (p << 0.05) and pairwise Dest ranged 0.104–0.734; p = 0.001 (Table 2). PcoA based on Nei genetic distance (Fig. 2b) portrayed four distinct groups of the microsatellite haplotypes, with axes 1 and 2 capturing 96.9% of the explained variation, fully matched with the assigned four maritime routes. Comparable to the COI outcomes, the northern route diverged by far from the others. Netstruct results showed 6 clusters at the threshold of 0.269 (Fig. 3d). BAPS results revealed 3 clusters (Fig. 3e), where two, are each restricted to northern and western routes and the third encompasses the eastern and circum-Italian together. Gene flow analyses (Fig. 3f) revealed that while there is a geneflow from cluster 2 (western route) to the other clusters, no geneflow goes to it. On the other hand, there is a geneflow between clusters 1 and 3 in both directions.
Discussion
Human mediated dispersals are millennia-old phenomena40. These occurrences, not only facilitate erosions of the species biogeographic boundaries, but in addition directly alter seascape genetics of transported species41,42, despite the difficulty of disentangling between the impacts of natural processes and contributions by human-mediated dispersal mechanisms24. Associated with functional connectivity bearings, the balance between immigration of new genetic variants, and the extinction and diversification of existing genetic repertoire, are key factors that determine population genetics characteristics for decades to come43. Within this conceptualization, one of the leading drivers for the unintentional transfer of marine organisms is the global maritime traffic44, when long-term functional connectivity trajectories, may establish a ‘genetic memory’ of historical sea voyages and ancient population genetics characteristics. These seascape genetics may persist for centuries, leaving their footprints on present-day population genetics and furthermore serving as an important proxy for understanding the genetic assemblies of contemporary populations45,46. This is primarily true for species with limited ability for self-dispersal and limited larval mobility47 such as with Botryllus schlosseri31.
Using two independent molecular markers for genetic parameters (four nuclear microsatellite loci and 11 mitochondrial COI haplotypes) we analyzed here original and archive-mined data of 64 sampling sites (Appendix Table 1; the lack of B. schlosseri population genetic data from North African coasts is further noted), pertaining to coalesce into four European maritime routes (termed here as eastern, western northern and circum-Italian). We further performed identical population genetics tests (Fisher exact test, Fst and Dest, Nei genetic distance, Principal coordinate analysis, and two cluster analyses) on both genetic markers, with outcomes revealing the same results of significant segregation and differentiation between all routes, with the most conspicuous segregation for the northern route. The outcomes from the microsatellite analyses also uncovered an unexpected absence of gene flow toward the western route (Fig. 3f). This outcome could be partially clarified by the likelihood that abiotic factors, such as water temperatures and salinity, play a role in restructuring of Botryllus schlosseri seascape genetics, together with the influence of maritime traffic. As a support, Reem et al.23 identified genetic disparities between the eastern and western Mediterranean basins, as well as within the eastern basin of the Mediterranean Sea. These disparities were proposed to potentially arise from abiotic differences between the basins, which might slow down the rate of interconnectivity.
During Medieval times, maritime traffic in the Mediterranean was networked by the Genoa/Spain and Venice/Ottomans major trajectories4,6,48. The Genoa/Spain ships travelled mostly in the western and European-Atlantic routes, while the Venetian/Ottoman ships dominated the eastern route. While shipping routes in the Mediterranean Sea and European Atlantic commenced long ago, maritime transportation expanded dramatically as from the late 1500s49. Yet, the wind dependent ship velocity was slow (maximum 4–5 knots14,50 and did not change dramatically since ancient times8,14. The dependence on wind resulted in long journeys and frequent and long mooring breaks during periods of rough seas50. In contrast, the speed of modern vessels ranges between 20–26 knots, they are less sensitive to oceanographic conditions and are moored for much shorter periods than in the past due to advanced operation procedures in ports51, 52. Hundreds to thousands of slow-moving trade ships per year crossed the Mediterranean Sea and European Atlantic for centuries, to/from several major ports of origin (Appendix texts), while no evidence whatsoever has suggested that this traffic was halted in any period during the past ca. 3000 years. There is also no evidence showing that navigation routes have ever changed in historical times as a result of climatic changes14.
Maritime activities are associated with the transport of sedentary organisms that recruit to the ship hauls, a millennia-old phenomenon38, anecdotally documented in the literature for calcifying organisms e.g.,15,53,54,55,56,57,58. Yet, very little is known about the associations between the early sea voyages and the dispersal of fouling and boring communities in ancient wooden sea-going ships. Thus, the application of molecular biology methods, as performed here, may broaden our understanding on the impact of past invasions on current population genetics characteristics. In addition to the outcomes of natural phenomena, like post glacial impacts on genetics repertoires25,59, this study reveals that historical European intensive seafaring from the beginning of the Holocene, through the Bronze and Iron ages and toward the Classical Antiquity up to the industrial revolution have left their footprint on contemporary seascape genetics (revealed from the independent analyses of both, nuclear and mitochondrial markers) of one of the most common European urochordate, Botryllus schlosseri, a model case for presumable many other occurrences.
The study of seascape genetics aims to elucidate how spatially variable structural and environmental parameters (in the present study, intensive seafaring through major navigation routes) impact population genetic properties of marine organisms60. Past anthropogenic transport of species not only may impact genetic homogenization61, primarily under weak natural connectivity (such as with Botryllus schlosseri21,30), but may even result in constructed population genetics properties, ghost of past maritime trajectories, providing keys for understanding past, current and future seascape genetics.
Data availability
The dataset of microsatellites generated during the current study appear in Appendix Table 15, The accession numbers of the COI Haplotypes that are deposited in GenBank repository are found in appendix Table 2.
References
Abulafia, D. The Great Sea: A Human History of the Mediterranean (Oxford University Press, 2011).
Samson, A. V. M. Offshore finds from the Bronze age in North-western Europe: the shipwreck scenario revisited. Oxf. J. Archeol. 25, 371–388 (2006).
Clark, P. (ed.) The Dover Bronze Age Boat (English Heritage, 2004).
Moscati, S. The Phoenicians (Tauris Publishers, 2001).
Roller, D. W. Through the Pillars of Herakles Greco-Roman Exploration of the Atlantic (Routledge, 2006).
Rickman, G. E. The grain trade under the Roman Empire. Mem. Am. Acad. Rome 36, 261–275 (1980).
Horden, P. & Purcell, N. The Corrupting Sea: A Study of Mediterranean History (Wiley, 2000).
Gal, D., Saaroni, H. & Cvikel, D. Mappings of potential sailing mobility in the Mediterranean during antiquity. J. Archaeol. Method Theory https://doi.org/10.1007/s10816-022-09567-5 (2022).
Milman, N. & Pedersen, P. Evidence that the Cys282Tyr mutation of the HFE gene originated from a population in southern Scandinavia and spread with the Vikings. Clin. Genet. 64, 36–47 (2003).
Price, N. The Vikings in Spain, North Africa and the Mediterranean. In The Viking World (eds Price, N. & Brink, S.) 462–469 (Taylor & Francis, 2008).
Munroe, J. H. The low countries’ export trade in textiles with the Mediterranean basin, 1200–1600: A cost-benefit analysis of comparative advantages in overland and maritime trade routes. Int. J. Marit. Hist. 11, 1–30 (1999).
Van Gelder, M. Trading Places: The Netherlandish Merchants in Early Modern Venice (Lib. Econ. Hist. Brill NV Leiden Boston, 2009).
Tucci, U. Sur la pratique vénitienne de la navigation au XVIe siècle [quelques remarques]. Ann. Econ. Soc. Civilis. 1, 72–86 (1958).
Pryor, J. H. The geographical conditions of galley navigation in the Mediterranean. In The Age of the Galley: Mediterranean Oared Vessels Since Pre-classical Times (ed. Gardiner, R.) (Conway Maritime Press, 1995).
Catta, J. D. Note sur quelques crustacés erratiques. Ann. Sci. Nat. Zool. 3, 1–33 (1876).
Woods Hole Oceanographic Institute. Contribution No. 580: Marine fouling and its prevention. U.S. naval Institute Annapolis Maryland (George Banta Publishing, 1952)
Godwin, L. S. Hull fouling of maritime vessels as a pathway for marine species invasions to the Hawaiian Islands. Biofouling 19, 123–131 (2003).
Carver, C. E., Mallet, A. L. & Vercaemer, B. Biological synopsis of the colonial tunicates Botryllus schlosseri and Botrylloides violaceus. Can. Man. Rep. Fish. Aquat. Sci. 2747, 256 (2006).
Farrapeira, C. M. R., Ferreira, G. F. A. & Tenório, D. O. Intra-regional transportation of a tugboat fouling community between the ports of Recife and Natal, nortrheast Brazil. Braz. J. Oceanogr. 58, 1–14 (2010).
Lacoursière-Roussel, A., Forrest, B. M., Guichard, F., Piola, R. F. & McKindsey, C. W. Modeling biofouling from boat and source characteristics: a comparative study between Canada and New Zealand. Biol. Invasions 14, 2301–2314 (2012).
Reem, E., Mohanty, I., Katzir, G. & Rinkevich, B. Population genetic structure and modes of dispersal for the colonial ascidian Botryllus schlosseri along the Scandinavian Atlantic coasts. Mar. Ecol. Prog. Ser. 485, 143–154 (2013).
Berrill, N. J. The Tunicata with an Account of the British Species (Ray Society, 1950).
Reem, E., Douek, J., Paz, G., Katzir, G. & Rinkevich, B. Phylogenetics biogeography and population genetics, of the ascidian Botryllus schlosseri in the Mediterranean Sea and beyond. Mol. Phylog. Evol. 107, 221–231 (2017).
Einfeldt, A. L., Jesson, L. K. & Addison, J. A. Historical human activities reshape evolutionary trajectories across both native and introduced ranges. Ecol. Evol. 10, 6579–6592 (2020).
Ben-Shlomo, R., Paz, G. & Rinkevich, B. Post glacial period and current invasions shape population genetics of botryllid ascidians along European Atlantic coasts. Ecosystems 9, 1118–1127 (2006).
Bock, D. G., MacIsaac, H. J. & Cristescu, M. E. Multilocus genetic analyses differentiate between widespread and spatially restricted cryptic species in a model ascidian. Proc. R. Soc. B. 279, 2377–2385 (2012).
Nydam, M. L., Giesbrecht, K. B. & Stephenson, E. E. Origin and dispersal history of two colonial ascidian clades in the Botryllus schlosseri species complex. PLoS ONE 12, e0169944 (2017).
Yund, P. O., Collins, C. & Johnson, S. L. Evidence of a native northwest Atlantic COI haplotype clade in the cryptogenic colonial ascidian Botryllus schlosseri. Biol. Bull. 228, 201–216 (2015).
Paz, G., Douek, J., Caiqing, M., Goren, M. & Rinkevich, B. Genetic structure of Botryllus schlosseri (Tunicata) populations from the Mediterranean coast of Israel. Mar. Ecol. Prog. Ser. 250, 153–162 (2003).
Rinkevich, B., Paz, G., Douek, J. & Ben-Shlomo, R. Allorecognition and microsatellite allele polymorphism of Botryllus schlosseri from the Adriatic Sea. In The Biology of Ascidians (eds Sawada, H. & Yokosawa, H.) 426–435 (Springer, 2001).
Grosberg, R. K. Limited dispersal and proximity-dependent mating success in the colonial ascidian Botryllus schlosseri. Evolution 41, 372–384 (1987).
Graham, D. E. The isolation of high molecular weight DNA from whole organisms or large tissue masses. Anal. Biochem. 85, 609–613 (1978).
Folmer, O., Black, M., Hoeh, W., Lutz, R. & Vrijenhoek, R. DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Mol. Mar. Biol. Biotechnol. 3, 294–299 (1994).
Hall, T. A. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp. Ser. 41, 95–98 (1999).
Thompson, J. D., Gibson, T. J., Plewniak, F., Jeanmougin, F. & Higgins, D. G. The Clustal X windows interface: Flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 25, 4876–4882 (1997).
Peakall, R. & Smouse, P. E. GenAlEx6: genetic analysis in Excel. Population genetics software for teaching and research. Mol. Ecol. Notes 6, 288–295 (2006).
Greenbaum, G., Templeton, A. R. & Bar-David, S. Inference and analysis of population structure using genetic data and network theory. Genetics 202, 1299–1312 (2016).
Corander J., Marttinen, P., Sirén, J. & Tang, J. BAPS: Bayesian analysis of population structure version 5.3. Department of Mathematics, Åbo Akademi University Finland. http://web.abo.fi/fak/mnf/mate/jc/software/BAPS5manual.pdf (2009).
Pourchasse, P. Trade between France and Sweden in the eighteenth century. Forum Navale 67, 92–114 (2011).
Ojaveer, H. et al. Historical baselines in marine bioinvasions: Implications for policy and management. PLoS ONE 13, e0202383. https://doi.org/10.1371/journal.pone.0202383 (2018).
Carlton, J. T. Deep invasion ecology and the assembly of communities in historical time. In Biological Invasions in Marine Ecosystems (eds Rilov, G. & Crooks, J. A.) 13–56 (Springer, 2009).
Otto, S. P. Adaptation, speciation and extinction in the anthropocene. Proc. R. Soc. B. 285, 2018–2047. https://doi.org/10.1098/rspb.2018.2047 (2018).
Roman, J. Diluting the founder effect: cryptic invasions expand a marine invader’s range. Proc. R. Soc. B 273, 2453–2459. https://doi.org/10.1098/rspb.2006.3597 (2006).
Iacarella, J. C. et al. Climate change and vessel traffic create networks of invasion in marine protected areas. J. Appl. Ecol. 57, 1793–1805 (2020).
Epps, C. W. & Keyghobadi, N. Landscape genetics in a changing world: disentangling historical and contemporary influences and inferring change. Mol. Ecol. 24, 6021–6040 (2015).
Chafin, T. K. et al. Spatial population genetics in heavily managed species: separating patterns of historical translocation from contemporary gene flow in white-tailed deer. Evol. Appl. 14, 1673–1689 (2021).
Darling, A. J., Kuenzi, A. & Reitzel, A. M. Human-mediated transport determines the non-native distribution of the anemone Nematostella vectensis, a dispersal-limited estuarine invertebrate. Mar. Ecol. Prog. Ser. 380, 137–146 (2009).
Dumrec, B. (2017). Flottes publiques, flottes privees à Venise (XII-XV siecles) in La mer dans L'histoire, le moyen age (ed. Balard, M.) 138–147 (Boydel Press, 2017).
Natkiel, R. & Preston, A. Atlas of Maritime History 1st edn. (Facts on File Inc., 1986).
Casson, L. Speed under sail of ancient ships. Trans. Proc. Am. Philol. Assoc. 82, 136–148 (1951).
Psaraftis, H. N. & Kontovas, C. A. Speed models for energy-efficient maritime transportation: a taxonomy and survey. Transp. Res. Part C 26, 331–351 (2013).
Nam, K. P. & Sang, C. S. Tendency toward mega containerships and the constraints of container terminals. J. Mar. Sci. Eng. 7, 131 (2019).
Darwin, C. A monograph on the subclass Cirripedia, with figures of all the species. The Balanidae (or sessile cirripedes); the Verrucidae (Ray Society, 1854).
Holthuis, L. B. & Heerebout, G. R. Vondsten van de zeepok Balanus tintinnabulum (Linnaeus, 1758) in Nederland. Zool. Bijdr. 13, 24–31 (1972).
Stevens, W. Progress report on the marine excavation at Red Bay, Labrador: a summary of the 1983 field season no. 240. (Min. of Supply and Services Canada 1986).
Petersen, K. S., Rasmussen, K. L., Heinemeier, J. & Rud, N. Clams before Columbus?. Nature 359, 679 (1992).
Kerckhof, F. & Cattrijsse, A. Exotic Cirripedia (Balanomorpha) from buoys off the Belgian coast. Senck. Marit. 31, 245–254 (2001).
Essink, K., Oost, A. P., Streurman, H. J. & Van der Plicht, J. Are medieval Mya arenaria (Mollusca; Bivalvia) in the Netherlands also clams before Columbus?. Neth. J. Geosci. 96, 9–16 (2017).
Panova, M. et al. Glacial history of the North Atlantic marine snail, Littorina saxatilis, inferred from distribution of mitochondrial DNA lineages. PLoS ONE 6, e17511. https://doi.org/10.1371/journal.pone.0017511 (2011).
Riginos, C. & Liggins, L. Seascape genetics: populations, individuals, and genes marooned and adrift. Geogr. Compass 7(3), 197–216 (2013).
Hudson, J., Bourne, S. D., Seebens, H., Chapman, M. A. & Rius, M. The reconstruction of invasion histories with genomic data in light of differing levels of anthropogenic transport. Philos. T. R. Soc. B. 377, 20210023. https://doi.org/10.1098/rstb.2021.0023 (2022).
Lampman, Late Medieval Land & Maritime Trade Routes map. World history encyclopedia. https://www.worldhistory.org/image/9759/late-medieval-land--maritime-trade-routes/ (2018).
Acknowledgements
We thank G. Paz for technical support and graphic design, J. Eliav, N. Helfman and D. Gal from the Department of Marine Civilisations at Haifa University for introducing us with historical literature sources and for helpful discussions and to G. Greenbaum from the Hebrew University of Jerusalem for assisting in the implementation of the NetStruct software. This study was partially supported by the Israel Science Foundation (to BR no. 172/17), by the United States—Israel Binational Science Foundation (to BR no. 2015012) and by Ocean Citizen, EC-Horizon program (to BR no. 101093910).
Author information
Authors and Affiliations
Contributions
E.R., J.D. and B.R. designed the study. B.R. and E.R. collected the Botryllus samples. E.R. and J.D. performed data collection, conducted the molecular analyses and performed the statistical tests. E.R. and B.R. co-drafted the manuscript together. All authors read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Reem, E., Douek, J. & Rinkevich, B. Historical navigation routes in European waters leave their footprint on the contemporary seascape genetics of a colonial urochordate. Sci Rep 13, 19076 (2023). https://doi.org/10.1038/s41598-023-46174-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-46174-0
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.