Abstract
Intestinal symbiotic microorganisms have a strong capacity to regulate the physiological functions of their host, and Drosophila serves as a useful model. Enterococcus faecium (E. faecium) is a member of the normal intestinal flora of animals. Lactic acid bacteria (LAB) such as E. faecium can promote the growth and development of Drosophila, but the mechanism of regulation of Drosophila is poorly understood. In this study, we found that E. faecium used a carbon source to produce probiotic acids. E. faecium is a symbiotic bacterium for Drosophila, and adult flies passed on parental flora to offspring. E. faecium promoted the growth and development of Drosophila, especially under poor nutritional conditions. E. faecium shortened the developmental process for Drosophila and accelerated the transformation from larva to pupa. Finally, E. faecium promoted the growth and development of Drosophila through TOR and insulin signalling pathways.
Similar content being viewed by others
Introduction
The human body, especially the intestinal tract, is inhabited by trillions of bacteria and other microorganisms1,2. The intestinal tract provides niches and nutrients for the microflora, which helps the host digest food, produce probiotics, and regulate a variety of host physiological activities3. Indeed, the host and the microflora establish a mutually beneficial symbiotic relationship that is essential for human health, ecology, and genetic variation4, but the complexity and diversity of the flora hinders in-depth analysis of its mechanisms5,6.
In recent years, Drosophila, has attracted increasing attention as a model for studying symbiotic relationships between hosts and bacteria due to the ease of establishing aseptic and intact systems, and its high degree of conservation with human flora and intestinal structure7. Like a mammal, the intestines of conventionally reared (CR) Drosophila are also home to numerous symbiotic microorganisms, making it a good model for asepsis (germ free, GF) or gnotobiotic analysis8. Recent studies have shown that intestinal microorganisms have important effects on the growth and development of Drosophila7,9. Firstly, Drosophila eat rotten fruit, which contains a large number of fermentative microorganisms7, indicating that these microorganisms may be beneficial to the development and growth of Drosophila10. Secondly, Drosophila ovulates in vitro, and the egg is covered with a layer of chitin, which can resist certain physical and chemical damage, making it easy to establish a sterile host via body surface disinfection. Based on a GF approach, a gnotobiotic model was established by inoculating specific bacteria7. Thirdly, the intestinal microbial structure of Drosophila is similar to that of human intestinal microorganisms11. Finally, our in-depth knowledge of genetics in Drosophila makes it convenient for us to analyse the molecular mechanism of microbial regulation of the host.
Enterococcus faecium is a Gram-positive bacterium and a member of the normal intestinal flora of animals. Because it can produce lactic acid, it belongs to the lactic acid bacteria (LAB). Studies have confirmed that there are more than 20 kinds of bacteria in Drosophila caught in the wild and raised in the laboratory, and LAB are among the most common classes of symbiotic bacteria in Drosophila12. Food spoilage caused by E. faecium facilitates host growth and reproduction, favouring the survival of semi-saprophytic insects such as fruit flies in nature, creating a new niche for them and increasing species diversity. However, a species of symbiotic E. faecium and the mechanism by which they promote the growth and development of Drosophila remain largely unknown. In this study, E. faecium was isolated from the intestine of Drosophila using selective medium, and the mechanism for promoting Drosophila growth and development was investigated.
Results
Isolation and identification of bacteria from the intestinal tract of Drosophila
The strain of bacteria was isolated from the intestines of Drosophila using selective medium13. The bacterium was confirmed to be Gram-positive and facultative anaerobic, and its morphological and biochemical characteristics were consistent with those of Enterococcus. Sequencing results showed that the 16S rRNA gene was 1103 bp, and the isolated strain was confirmed to be an E. faecium strain (GenBank accession number: KY990052) based on its closest genetic relationship with E. faecium. MEGA 6.0 software (http://www.downxia.com) was used to analyse the homology of genes and construct a phylogenetic tree (Fig. 1a). The sequences of E. faecium KY990052, Enterococcus isolated from human faeces CCFM8321 (KJ803878) and E. faecium T1 (KR909902) in cheese shared the highest homology (99%), indicating that the three strains had the closest genetic relationships. However, there was no significant similarity in the gene sequences of Acetobacter aceti and Enterobacter mori commensal to the gut of Drosophila, indicating a distant genetic relationship.
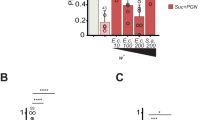
Phylogenetic tree of Enterococcus faecium and relatives and its in vitro features of culture. (a) Phylogenetic tree of Enterococcus faecium and its homologies. Bar: nucleotide divergence; number at notes present bootstrap percentages; those in parentheses are GenBank accession No. (b) Growth of Enterococcus faecium with carbon sources in medium. (c) pH curves of Enterococcus faecium growing in medium with carbon sources. Values represent mean ± SEM. (d) Determination of pH in the gut of CR Drosophila Melanogaster (n = 2000), with (DM medium) and without (no DM medium). (e) Expression of acidic results in CCR. Acidic regions indicated by pH indicator Bromophenol blue.
In vitro culture characteristics of E. faecium
E. faecium KY990052 cells grew rapidly in YCFA liquid medium with 0.25% glucose, with a logarithmic growth phase from 3 to 9 h (Fig. 1b). At 9 h, the OD600 value was 1.1 ± 0.3 near the peak, and it decreased slightly thereafter. After 3 h of culture, the pH value of the culture medium began to decrease continuously (Fig. 1c), and the lowest value was 5.1 ± 0.1. E. faecium KY990052 is a typical lactic acid-producing coccus, which can also reduce the pH value in the intestine and maintain its acidic environment. As shown in Fig. 1d, the pH of the gut of CR fly is 5.5, the pH of the single medium is 8.1, and the pH of the medium containing fly is 4.2. The latter two are statistically significant compared with the pH of the gut. These results indicated that CR Drosophila could significantly reduce the pH value of the medium. The midgut of flies contains an acidic gastric or copper cell region (CCR,14), which controls distribution and composition of the microbiota, and control the survival of ingested bacteria15. We detected that the pH of E. faecium associated intestinal CCR region was 4, and that of GF group was 7 (Fig. 1e), which was similar to that of in vitro fermentation experiment.
E. faecium colonises the Drosophila gut
In order to determine which symbiotic bacteria mediate this effect, we analysed the characteristics of bacterial communities associated with CR fly strains. As shown in Fig. 2a, the bacterial load in the intestines of 3rd instar larvae, pupae and adults was ≥ 103 and more on the 1st, 4th, 12th and 27th day after eclosion, indicating that E. faecium could effectively colonise the intestines of Drosophila. The bacterial density of the culture medium was higher, and the bacterial density per gram of medium was ≥ 105 in the larval and pupal stages, and the corresponding period at 1, 4, 12 and 27 days after adult eclosion (Fig. 2b), indicating that E. faecium could exist stably in the culture medium.
E. faecium was commensal bacteria of Drosophila. (a) The mount of internal bacterial loaded of E. faecium in various growth stages of the gut of Drosophila. (b) The mount of internal bacterial loaded of E. faecium in various growth stages of the culture medium of Drosophila. (c,d) The mount of internal bacterial loaded of the progenies of Drosophila adults mono-associated with E. faecium and CR. Values represent mean ± SEM.
Since vertical transfer is typical for microbiota acquisition, we tested whether E. faecium could be successfully transmitted from parents to offspring12. Figure 2c shows that E. faecium could be stably colonised in the 3rd instar larval, pupal and adult stages of the G1 generation of fruit flies, and the bacteria-carrying capacity in the intestines of these three stages were 104, 105 and 103, respectively, indicating that E. faecium could be vertically transmitted from generation to generation. The above results show that E. faecium is closely associated with Drosophila, and can be considered symbiotic. Similar dynamics for sustentation of the whole bacterial population and colony-forming units (CFUs) were observed during the larval, pupal and early adult stages of CR individuals (Fig. 2a,d). These results indicate that the protocol used to associate GF individuals with E. faecium accurately revealed the natural pattern of bacterial colonisation of CR during larval, pupal and early adult stages. Thus, E. faecium has a strong ability to colonise the whole larval niche of the host intestine, and maintain a population in the host digestive tract.
E. faecium association sustains larval development under nutrient scarcity
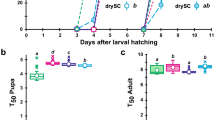
Having proved that E. faecium colonised the larval niche as a symbiotic bacterium, we tested whether E. faecium could promote the development of larvae raised on low-nutrient medium. The combination of E. faecium and nutrient-poor medium was enough to accelerate the growth of larvae, and both pupae and the adults fly eclosed earlier (Fig. 3a,b). However, this effect was not observed in medium rich in nutrients. This shows that E. faecium has a specific effect on the growth of whole body larvae, reflecting not only the nutritional effect of adding microorganisms to the fly culture medium, but also the specific biological activities of these strains. These observations showed that E. faecium accelerated the development of larvae, and led to the emergence of adults earlier under nutrient-poor conditions.
The E. faecium promoted the growth and development of Drosophila. (a) The timing of pupa formation of CR, GF or E. faecium in the different concentrations of yeast. (b) The timing of the emergence of adult of CR, GF or E. faecium in the different concentrations of yeast. (c) PTTH mRNA levels for CR, GF and E. faecium-associated Drosophila. (CR VS GF, P < 0.05, CR VS E. faecium, p > 0.05, GF VS E. faecium, p < 0.05). ns P > 0.05, *P < 0.05; values represent mean ± SEM.
In Drosophila, prothoracicotropic hormone (PTTH) secreted by the brain initiates the secretion of the steroid hormone hydroxyecdysone (20HE), which promotes the transition from 3rd instar larva to pupal stages16. Expression of the PTTH gene is regulated over time, and expression is higher in the late third instar and early pupal stages, hence expression serves as a molecular index to evaluate the development of Drosophila. The transcriptional level of PTTH in the brain of Drosophila was measured by the RT-PCR technique. Figure 3c shows that expression of PTTH peaked on the 8th day for CR, but for GF it did not reach its peak until 14 days; GF was ~ 6 days slower than CR, and expression for CR was higher than that for GF (p < 0.001). Expression of PTTH reached its peak on the 5th day for E. faecium, 4 days earlier than for GF, and levels were higher than or GF (p < 0.05). The peak was close to that for CR, which indicates that E. faecium effectively rescued the growth retardation of GF, and accelerated the transition from larval to pupal stages.
E. faecium promotes larval growth
In order to further assess the effects of E. faecium on the growth of larvae, we analysed the size of adults, which is the final parameter for the growth stage of larvae. To this end, we compared the body weight of adults raised on medium containing CR, GF or E. faecium. For the duration of the larval stage, we did not observe a difference in the weight of adults growing on nutrient-rich medium. Similarly, when larvae grew on nutrient-poor medium, no significant differences were observed between CR, GF and E. faecium-associated individuals (Fig. 4a,b). However, adults developed from GF, GF and E. faecium-associated larvae grown under poor dietary conditions weighed less than adults grown under rich conditions (Fig. 4a,b).
The E. faecium improved the growth ratio of larval, but it had little effects on weight. (a,b) The weight of male and female Drosophila on different concentrations of medium. (c,d) Larval surface of CR or GF, or E. faecium associated larvae over time when grown on rich (1.0% yeast) or poor diet (0.5% yeast). Linear regression curves are included. CR/Poor Diet, Y = 693,900X – 219,500; GF/Poor Diet, Y = 410,400X – 349,700; E. faecium/Poor Diet, Y = 740,600X – 260,100; CR/Rich Diet, Y = 855,100X − 182,500; GF/Rich Diet, Y = 516,000X − 156,100; E. faecium/Rich Diet, Y = 806,500X – 627,300. Values represent mean ± SEM (ns p > 0.05; *p < 0.05). A Under the condition of poor medium, the pupation time of CR and E. faecium-associated larvae was twofold short than GF.
Because E. faecium shortens its growth stage without impacting the size of adults, we assume that E. faecium increases the growth rate of larvae. To confirm this, we compared the size of individuals associated with CR, GF and E. faecium from L1 larval to pupal stages. There was no difference in pupation rate of CR, GF and E. faecium-associated larvae fed a rich diet (Fig. 4c), but the pupation rate of CR-related larvae fed a poor diet was two-fold higher than that of the GF group, and the results for E. faecium-associated larvae were similar to those of the CR group (Fig. 4d), confirming the results for Fig. 3c. These findings demonstrated that under conditions of nutrient deficiency, E. faecium promoted individual growth by increasing the pupation rate of larvae and shortening the growth cycle, which significantly shortened the pupation time of GF larvae.
Effect of E. faecium on intestinal cell proliferation
It has been reported that the main symbiotic bacteria in the intestinal tract of Drosophila can control the development rate, size, energy metabolism and intestinal stem cell activity of Drosophila17. Adult stem cells play a crucial role in organ homeostasis and palingenesis through their ability to maintain self-renewal and generate differentiated cells. The adult midgut of Drosophila is a frequently-used model system for stem cell biology18,19,20,21. This tissue undergoes rapid cell renewal by intestinal stem cells (ISCs), the sole mitotic cells in the gut. They generate diploid enteroblasts (EBs), which differentiate to polyploid enterocytes or diploid EEs22. We used anti-phosphorylated histone 3 (PH3) antibodies to evaluate the number of mitotic cells to determine mitotic activity23. The results in Fig. 5 show eight cells in the metaphase of midgut division for CR, compared with two cells for GF (p < 0.001), and the number of E. faecium cells was significantly higher than for GF (p < 0.001), which significantly rescued the defective intestinal cell division and proliferation of GF.
E. faecium promoted the proliferation of Drosophila intestinal cells. (a–c) The representative imaging of mitotic cells in guts of CR, GF and E. faecium flies (red as PH3 marker, blue as DAPI marker, bar 50 μm). (d) The quantification of mitosis in gut region of CR, GF and E. faecium flies. The anterior gut, midgut and posterior are parts of gut regions, n = 10. Values represent mean ± SEM. ns p > 0.05, **p < 0.01.
Effects of E. faecium on the steroid hormone ecdysone and regulation of the insulin signal pathway
In Drosophila, the larval period and the larval growth rate are controlled by two crucial hormones: the steroid hormone ecdysone (Ecd) and Drosophila insulin-like peptides (dILPs), respectively24. Transcription factor E74B is one of the early genes to act in response to increased Ecd concentrations, and it serves as a molecular marker of Ecd activity25. To determine whether E. faecium directly affects these growth signals, we compared the strength of these signals in CR, GF and E. faecium-associated larvae. Figure 6a shows that for CR, the highest mRNA levels for E74B were reached at day 14 after oviposition, while for GF, peak expression of this gene was significantly delayed until day 20 after oviposition, and the difference was statistically significant (p < 0.001). Transcriptional levels peaked on day 16 in E. faecium-associated larvae, hence the delayed expression of the gene in GF was clearly rescued. However, levels of E74B mRNA peaked sharply in CR and E. faecium-associated larvae, while GF larvae showed a lower peak that was delayed. Interestingly, in GF larvae, levels of E74B mRNA were already increased on day 6 (although this was not statistically significant), and there was no pupariation at this time (pupariations first observed on day 23; Fig. 3b). These results showed that E. faecium-associated 3rd instar larvae showed early and stronger Ecd peaks, which rescued the delayed Ecd expression of GF.
The E. faecium promoted the secretion of hormone and was required to regulate the insulin signal pathway. (a–c) E74B, InR, dILP3 and dILP5 mRNA levels for CR, GF and E. faecium-associated Drosophila. (d) Glucose levels for CR, GF and E. faecium-associated Drosophila. Values represent mean ± SEM. ns p > 0.05; *p < 0.05, ***p < 0.001.
We next measured InR gene expression levels as a marker for intracorporal dILP activity. InR gene transcription reflects negative regulation of the InR signalling pathway through the activity of the fork head transcription factor (dFOXO). The InR gene is a negative regulator of intracorporal dILP activity; lower InR expression is associated with greater dILP activity26. As shown in Fig. 6b, InR expression was consistently lower in CR larvae than in GF larvae. The E. faecium-associated larvae were effectively rescued by over-expression of the InR gene in GF, suggesting that E. faecium bacteria are equivalent to CR associated with increased intracorporal InR signalling during the period of larval growth. We then measured dILP3 and dILP5 gene expression as a positive marker for insulin signal pathway activity. As shown in Fig. 6c, dILP3 and dILP5 expression levels were higher in CR larvae than in GF larvae. E. faecium-associated larvae were effectively rescued from low expression of dILP3 and dILP5 genes in GF. These results showed that E. faecium is associated with increased dILP activity during larval growth, and this is similar to the effect of CR.
It is known that insulin/insulin-like growth factor signalling (IIS) mutant animals display diabetic phenotypes, with increased levels of circulating sugars27,28,29,30,31. As shown in Fig. 6b,c, GF showed abnormal expression of dILP3, dILP5 and InR genes. In the Fig. 6d, we also measured levels of sugars in CR, GF and E. faecium-associated larvae, and detected elevated levels of total sugars (glucose) in GF compared with CR larvae (p < 0.001), and glucose was the major disaccharide in insects. E. faecium effectively reduced the higher levels of glucose in GF, and there was no difference compared with CR. Taken together, our results support the notion that although differing in terms of kinetics, E. faecium-associated larvae strengthen the systemic generation of two hormonal growth signals.
Discussion
Due to the relative simplicity of the microbial community and facile establishment of the bacterial host, Drosophila provides a useful model that avoids complexity and is suitable for revealing the molecular mechanisms by which microflora modulate host physiology. Our current results showed that intestinal microorganisms had an important effect on the development of Drosophila. The bacterial community in the digestive tract of Drosophila is relatively simple; the dominant bacteria are Firmicutes (Acetobacter and Lactobacillus). Given the great differences in the growth environments of fruit flies, it is possible that their symbiotic microbes are much more complex than expected. Herein, E. faecium KY990052 was isolated from the intestinal tract of Drosophila for the first time, revealing greater diversity and complexity of the intestinal bacteria of Drosophila (Fig. 1). This strain can effectively colonise the intestinal tract of Drosophila and be passed from parents to offspring, suggesting that E. faecium is a symbiotic organism of Drosophila (Fig. 2). At the organism level, E. faecium increased the growth rate of Drosophila, promoted Drosophila growth, and accelerated the eclosion rates of pupae and adults (Fig. 3). At the molecular level, E. faecium enhanced the production of two key growth hormones (Ecd and dILPs) in the larval stage, thus promoting the growth rate of the larval stage, and ultimately the growth and development of Drosophila (Fig. 6).
LAB are Gram-positive bacteria commonly found in the gastrointestinal tracts of animals, fermented food, dairy products, and natural environments including soil and water32. LAB are also common symbiotic microorganisms of Drosophila, including Lactobacillus and Enterococcus. Enterococcus is a large genus of LAB. In Drosophila, many species have been identified as commensal bacteria, including E. faecium33. LAB convert carbohydrates from raw materials into lactic acid, and they are often used in food fermentation to obtain a variety of high-quality end products34. Studies have shown that Enterococcus participates in and acts on a variety of physiological processes in the host12,35,36,37, but there are few reports on how it affects the development of the host. E. faecium decomposes nutrients in fruit using its own enzyme system, and the decay caused by E. faecium is not only beneficial to its own reproduction, but also provides more available nutrients for fruit flies, and creates a new niche for the host37. This is beneficial for the survival and reproduction of fruit flies. Drosophila are semi-saprophytic insects that mainly depend on eating rotten fruit, and they form a mutually beneficial symbiotic relationship with E. faecium in the natural environment.
This symbiotic relationship is reflected in the fact that E. faecium promoted the growth and development of Drosophila. Ours and other studies showed that Drosophila cannot survive without microorganisms, even when given adequate nutrients such as protein and glucose, indicating that microorganisms are necessary for the growth and development of Drosophila5. Members of the Drosophila intestinal flora can affect the genetic factors of some animals and the composition of important nutrients in the body, such as triglyceride (fat) content; high diversity among intestinal microorganisms can improve the digestion and absorption of nutrients and increase nutritional reserves in the host body38. Many animals have an intestinal region with a low pH to aid protein digestion, absorb nutrients such as calcium, iron and vitamin B12, and kill enteric pathogens and parasites acquired orally39. Our current study showed that E. faecium can utilise carbohydrates (glucose) and lower the pH to 5.1 (Fig. 1). Furthermore, E. faecium is resistant to bile salts and poor gastrointestinal conditions. Its main function is automatic aggregation and adhesion, and it synthesises a variety of bacteriocins called enteroglobulins; these small, low-molecular-weight, heat-resistant, ribosomal antibacterial peptides possess antibacterial activity40,41. Microorganisms in the intestinal tract of Drosophila promote immune activity by stimulating innate immune signals of intestinal epithelial cells42. Herein, we explored the signalling mechanism by which E. faecium promotes the growth and development of Drosophila.
The insulin signalling pathway dynamically regulates the transcription and translation of many proteins involved in glucose uptake, promotes intestinal cell division and proliferation, and regulates the whole process of body development, growth and metabolism7. It has been reported that the main symbiotic bacteria in the intestinal tract of Drosophila can regulate insulin signalling and the target of rapamycin (TOR) signal pathway, the host self-balance program, and control development rate, size, energy metabolism and intestinal stem cell activity9,12,17,43. In Drosophila, the TOR pathway regulates the hormone signal of larval growth in a tissue-specific manner, and the production of Ecd is controlled by the prothoracic gland in the middle stage of the 3rd instar larvae42. E. faecium-associated larvae can also express the E74B gene in advance, which rescues the expression defect of GF (Fig. 6a). Systemic InR signalling is regulated by remote control of dILP secretion by neurons, which is regulated by TOR activity in the fat body44,45. We found that the LAB E. faecium is a major symbiotic organism in the intestinal tract of Drosophila. Figure 6b,c show that E. faecium rescued defective expression of insulin signalling pathway genes (InR, dILP3 and dILP5) in GF, which controlled levels of sugars in the body (Fig. 6d), and had a similar effect to CR. The results showed that E. faecium could promote the development of Drosophila through the insulin signal pathway alone. This TOR signalling activity in the fat body triggered whole body InR signal transduction and increased the growth rate. In the prothoracic gland, TOR signalling enhanced the production of Ecd in the latter stages of larval development, and shortened the length of the growth stage. The combined effects of increased TOR activity and hormone signals led to optimal whole body growth. We also found that E. faecium increased the number of mitotic cells in the midgut of Drosophila (Fig. 5), which helped to maintain homeostasis and functioning in the host intestine, and created the necessary conditions for the absorption and utilisation of nutrients.
The above results showed that E. faecium promoted the growth and development of Drosophila, and increased the species diversity of intestinal microorganisms. This diverse intestinal microflora participates in the digestion, absorption and synthesis of nutrients. Drosophila provides a good model for intestinal microorganisms in humans and other animals. Using Drosophila as a model to study the role of symbiotic bacteria can provide important information pertinent to human diseases.
Materials and methods
Strain culture and Drosophila melanogaster breeding
All Drosophila were reared at 25 °C in 50–60% humidity under a 12 h/12 h light/dark cycle. The Oregon R strain was used for wild-type flies. Drosophila were raised on standard cornmeal sugar agar medium (Agar 7.5 g, grape juice 58 ml, sucrose 8.2 g, yeast 58.8 g, corn meal 58.8 g, distilled water 1000 ml)46.
Isolation and identification of bacteria
Fifteen CR-associated adults were anesthetised with carbon dioxide, placed in tubes, disinfected twice with 75% ethanol, washed twice with phosphate-buffered saline (PBS), ground, and cultured in a 100 μL volume at 37 °C for 24 h on plates. Single colonies were selected and cultured in liquid medium for 24 h. DNA was extracted by pyrolysis and PCR was carried out on the 16S rRNA gene using universal primers. The 16SrRNA gene sequence was submitted to GenBank to obtain the registration number of the strain. The 16S rRNA gene sequences of nine strains were downloaded from the GenBank database. Based on the homology of the 16SrRNA gene, MEGA 6.0 software was employed to generate a phylogenetic tree using the neighbour-joining method.
Bacteria culture and cell counting
Bacteria were isolated from Drosophila using specific media from the China General Microbiological Culture Collection Center, and identified from 16 SrRNA sequences with a PCR primer set47. To culture commensal bacteria, selective media were used to assay the bacterial population of E. faecium in 200 ml of liquid YCFA medium with 0.25% glucose48.
In vitro culture of E. faecium
Isolated E. faecium cells (2.5%) were injected into YCFA liquid medium containing 0.25% glucose (1L medium contains 10 g tryptone, 2.5 g yeast extract, 4 g NaHCO3, 1 g cysteine, 1 g inulin, 0.45 g K2HPO4, 0.9 g NaCl, 0.09 g MgSO4·7H2O, 0.09 g CaCl2, 1 mg resazurin, 10 mg haemin, 10 μg biotin, 10 μg cobalamin, 30 μg p-aminobenzoic acid, 50 μg folic acid, 150 μg pyridoxamine, 50 μg thiamine, 50 μg vibioflain, 13.8 mM glucose) with H2/CO2 in the gas phase47, and its cultured in a 37 °C incubator, and sampled at 0, 3, 6, 9,12 and 24 h for OD600 and pH measurement.
pH indicator, blue dye
Putting 150ul of 2% Bromophenol blue sodium (pH indicator, Sigma) was added to a food vial, and stir the blue fuel with a sterile rod. Flies were fed for 10 h. For E. faecium feeding, flies were fed 200 ul of 1 OD bacteria (in 5% sucrose) mixed with 100ul 2% pH indicator in filter paper for 24 h.
Establishment of germ-free (GF) and gnotobiotic populations
The GF model has been described previously in the literature9,49. Based on GF, 1 OD of E. faecium (take 100 μl of the bacterial solution, about 105 density) solution was added to medium and a gnotobiotic model of Drosophila (100 μl embryo) was established49. At least five tubules were included in each group.
Colonisation and generation transmission of E. faecium in Drosophila
Larvae at 2nd to 3rd instar stages were taken from 0 to 6 days after oviposition, along with pupae from 6 to 8 days and adults emerging at 1, 4 and 12 days. Drosophila were disinfected twice with 75% ethanol, washed twice with PBS, and ground. A step dilute release coating plate (YCFA) was placed in a 37 °C incubator for 48 h to observe and count colony growth (CFU/gut). In order to avoid the effects of yeast meal containing microbial factors on Drosophila development, casein hydrolysate was used to replace yeast meal9. At the corresponding growth stage of Drosophila, 1 g casein culture medium was added to the coating plate, which was diluted and cultured at 37 °C for 24 h. Colony growth was observed and counted (CFU/g).
CR or E. faecium-associated adults were transferred to casein medium after strict bacteria-free handling, and discarded after oviposition. After pupation from eggs, they were disinfected with 75% ethanol twice, cleaned with phosphate buffer twice, and transferred to sterile 4% yeast medium. After emergence and oviposition, the adult G0 generation was discarded. The intestinal bacteria content of 3rd larvae, pupae and adults of G1 generation was detected, using the same detection method as the colonization experiment (CFU/gut).
Determination of developmental duration and growth rate of Drosophila
The number of CR, GF and E. faecium-associated pupae and adults in 0.25%, 0.5% and 1% (casein%) casein medium was recorded, and the time of pupae formation and adult emergence was determined. There were at least 20 flies per tubule. The following equation was used:
where T is the development period of the corresponding stage, Tm is the days from spawning to the corresponding stage, and Nm is the number of Drosophila in the corresponding stage on day Tm. Casein medium at three concentrations comprised 0.5 g/L sucrose, 0.7 g/L corn flour, 0.1 g/L agar, and 0.025 g/L, 0.05 g/L or 0.1 g/L casein.
On the 3rd day after adult emergence, 40 fruit flies in the three groups (CR, GF and E. faecium) were weighed. Ten fruit flies (0 days larvae to pupae) were placed on a glass slide, frozen at − 20 °C for 1 h, images were taken under a DM4000 microscope (Leica, Wetzlar, Germany), and ImageJ software (https://imagej.en.softonic.com/) was used to measure the area.
Immunofluorescence staining
Ten CR, GF and E. faecium-associated adults (emergence 3 days) were selected, their intestines were dissected in PBS, and fixed in 4% paraformaldehyde solution at room temperature for 30 min. 0.3% PBST (Triton X-100) containing 0.2% goat serum and 0.1% fetal bovine serum was added and incubated for 30 min, rabbit primary antibody PH3 (Millipore, H0412; 1:1000) was added and incubated overnight at 4 °C, and anti-rabbit secondary antibody (Invitrogen, WP20007, 1:1000) was added and incubated for 2 h. PBST was used to wash samples. DAPI (Invitrogen, 1:1000) staining for 10 min, PBST washing, tablet sealing, microscopy image collection (Leica DM4000), and statistical analysis were then performed. Finally, the average value was taken.
Real-time PCR
Ten CR, GF and E. faecium-associated Drosophila were selected from 0.5% casein medium (3-day larvae to 3-day pupa) were selected and total RNA was extracted by the TRIzol method. A 5 μg sample of RNA was used as template, and oligo-DT primer was used for reverse transcription of mRNA to synthesise cDNA using a Prime Script RT reagent Kit (TaKaRa). Primer sequences are provided in previous work16,25,26. Primer sequence: PTTH (F:AACAGTGGCGGATTCGGTT, R:TACTCGGAGCATTGGAGGCAT), E74B ( F: GAATCCGTAGCCTCCGACTGT, R:AGGAGGGAGAGTGGTGGTGTT), InR (F: AACAGTGGCGGATTCGGTT, R: TACTCGGAGCATTGGAGGCAT), rp49 (F: GACGCTTCAAGGGACAGTATCTG, R: AAACGCGGTTCAGCATGA), dILP3 (F: AAGCTCTGTGTGTATGGCTT, R: AGCACAATATCTCAGCACCT), dILP5 (F: AGT3TCTCCTGTTCCTGATCC, R: CAGTGAGTTCATGTGGTGAG). Each group did 10 technical replicates and 3 biological replicates. Results were calculated using the formula ΔCt = Ct (target gene) − Ct (reference gene), and relative expression levels were determined using the 2–ΔΔCt method.
Sugar measurement
Fifty CR, GF and E. faecium- associated 3rd instar larvae were collected, rinsed three times in sterile water, ground, centrifuged at 4000g for 2 min, and 2 μL of the supernatant was treated with a glucose kit (Sigma, St. Louis, MO, USA) to determine the absorbance value (wavelength 505 nm).
Data analysis
The experiment was repeated three times or more, and the average value for each group was calculated and subjected to Student’s t-test. Results are presented as mean ± standard error of the mean (S.E.M.). *p < 0.05, **p < 0.01, ***p < 0.001.
Data availability
The datasets generated during the current study are available in the [Genbank] repository, The Genbank accession number: KY990052 and the Website: [https://www.ncbi.nlm.nih.gov/nuccore/KY990052].
References
Fernandez-Millan, E. & Guillen, C. Multi-organ crosstalk with endocrine pancreas: A focus on how gut microbiota shapes pancreatic beta-cells. Biomolecules. 12, (2022).
Round, J. L. & Mazmanian, S. K. The gut microbiota shapes intestinal immune responses during health and disease. Nat. Rev. Immunol. 9, 313–323 (2009).
Marchesi, J. R. et al. The gut microbiota and host health: A new clinical frontier. Gut. 65, 330–339 (2016).
Dethlefsen, L., McFall-Ngai, M. & Relman, D. A. An ecological and evolutionary perspective on human-microbe mutualism and disease. Nature. 449, 811–818 (2007).
Cao, Y., Wang, Y., Zheng, X., Li, F. & Bo, X. Revecor: An R package for the reverse ecology analysis of microbiomes. BMC Bioinf. 17, 294 (2016).
Scanlan, P. D., Knight, R., Song, S. J., Ackermann, G. & Cotter, P. D. Prevalence and genetic diversity of blastocystis in family units living in the United States. Infect. Genet. Evol. 45, 95–97 (2016).
Lee, W. J. & Brey, P. T. How microbiomes influence metazoan development: insights from history and drosophila modeling of gut-microbe interactions. Annu. Rev. Cell Dev. Biol. 29, 571–592 (2013).
Ryu, J. H. et al. Innate immune homeostasis by the homeobox gene caudal and commensal-gut mutualism in drosophila. Science. 319, 777–782 (2008).
Shin, S. C. et al. Drosophila microbiome modulates host developmental and metabolic homeostasis via insulin signaling. Science. 334, 670–674 (2011).
Erkosar, B., Storelli, G., Defaye, A. & Leulier, F. Host-intestinal microbiota mutualism: “learning on the fly”. Cell Host Microbe. 13, 8–14 (2013).
Capo, F., Wilson, A. & Di Cara, F. The intestine of drosophila melanogaster: An emerging versatile model system to study intestinal epithelial homeostasis and host-microbial interactions in humans. Microorganisms. 7, (2019).
Storelli, G. et al. Lactobacillus plantarum promotes drosophila systemic growth by modulating hormonal signals through tor-dependent nutrient sensing. Cell Metab. 14, 403–414 (2011).
Lopez-Siles, M. et al. Cultured representatives of two major phylogroups of human colonic Faecalibacterium Prausnitzii can utilize pectin, uronic acids, and host-derived substrates for growth. Appl. Environ. Microbiol. 78, 420–428 (2012).
Dubreuil, R. R. Copper cells and stomach acid secretion in the drosophila midgut. Int. J. Biochem. Cell Biol. 36, 745–752 (2004).
Li, H., Qi, Y. & Jasper, H. Preventing age-related decline of gut compartmentalization limits microbiota dysbiosis and extends lifespan. Cell Host Microbe. 19, 240–253 (2016).
Garelli, A., Gontijo, A. M., Miguela, V., Caparros, E. & Dominguez, M. Imaginal discs secrete insulin-like peptide 8 to mediate plasticity of growth and maturation. Science. 336, 579–582 (2012).
Blum, J. E., Fischer, C. N., Miles, J. & Handelsman, J. Frequent replenishment sustains the beneficial microbiome of drosophila melanogaster. Mbio. 4, e813–e860 (2013).
Lucchetta, E. M. & Ohlstein, B. The drosophila midgut: A model for stem cell driven tissue regeneration. Wiley Interdiscip. Rev. Dev. Biol. 1, 781–788 (2012).
Ohlstein, B. & Spradling, A. Multipotent drosophila intestinal stem cells specify daughter cell fates by differential notch signaling. Science. 315, 988–992 (2007).
Micchelli, C. A. & Perrimon, N. Evidence that stem cells reside in the adult drosophila midgut epithelium. Nature. 439, 475–479 (2006).
Ohlstein, B. & Spradling, A. The adult drosophila posterior midgut is maintained by pluripotent stem cells. Nature. 439, 470–474 (2006).
Park, J. S. et al. Increased centrosome amplification in aged stem cells of the drosophila midgut. Biochem. Biophys. Res. Commun. 450, 961–965 (2014).
Li, Y. et al. Capsaicin functions as drosophila ovipositional repellent and causes intestinal dysplasia. Sci. Rep. 10, 9963 (2020).
Hietakangas, V. & Cohen, S. M. Regulation of tissue growth through nutrient sensing. Annu. Rev. Genet. 43, 389–410 (2009).
Karim, F. D. & Thummel, C. S. Ecdysone coordinates the timing and amounts of E74a and E74B transcription in drosophila. Genes Dev. 5, 1067–1079 (1991).
Puig, O. & Tjian, R. Transcriptional feedback control of insulin receptor by Dfoxo/Foxo1. Genes Dev. 19, 2435–2446 (2005).
Baker, K. D. & Thummel, C. S. Diabetic larvae and obese flies-emerging studies of metabolism in drosophila. Cell Metab. 6, 257–266 (2007).
Edgar, B. A. How flies get their size: Genetics meets physiology. Nat. Rev. Genet. 7, 907–916 (2006).
Rulifson, E. J., Kim, S. K. & Nusse, R. Ablation of insulin-producing neurons in flies: Growth and diabetic phenotypes. Science. 296, 1118–1120 (2002).
Brogiolo, W. et al. An evolutionarily conserved function of the drosophila insulin receptor and insulin-like peptides in growth control. Curr. Biol. 11, 213–221 (2001).
Tatar, M. et al. A mutant drosophila insulin receptor homolog that extends life-span and impairs neuroendocrine function. Science. 292, 107–110 (2001).
Garcia-Solache, M. & Rice, L. B. The enterococcus: A model of adaptability to its environment. Clin. Microbiol. Rev. 32, 1 (2019).
Wan, K. H. et al. Complete genome sequence of enterococcus durans oregon-R-modencode strain Bdgp3, a lactic acid bacterium found in the drosophila melanogaster gut. Genome Announc. 5, (2017).
De Filippis, F., Pasolli, E. & Ercolini, D. The food-gut axis: Lactic acid bacteria and their link to food, the gut microbiome and human health. Fems Microbiol. Rev. 44, 454–489 (2020).
Wong, A. C., Chaston, J. M. & Douglas, A. E. The inconstant gut microbiota of drosophila species revealed by 16S Rrna gene analysis. Isme J. 7, 1922–1932 (2013).
Chandler, J. A., Lang, J. M., Bhatnagar, S., Eisen, J. A. & Kopp, A. Bacterial communities of diverse drosophila species: Ecological context of a host-microbe model system. Plos Genet. 7, e1002272 (2011).
Cox, C. R. & Gilmore, M. S. Native microbial colonization of drosophila melanogaster and its use as a model of enterococcus faecalis pathogenesis. Infect. Immun. 75, 1565–1576 (2007).
Chaston, J. M., Dobson, A. J., Newell, P. D. & Douglas, A. E. Host genetic control of the microbiota mediates the drosophila nutritional phenotype. Appl. Environ. Microbiol. 82, 671–679 (2016).
Du, Y., Luo, S., & Zhou, X. Enterococcus faecium regulates honey bee developmental genes. Int. J. Mol. Sci. 22, (2021).
Zommiti, M. et al. Evaluation of probiotic properties and safety of enterococcus faecium isolated from artisanal Tunisian meat “Dried Ossban”. Front. Microbiol. 9, 1685 (2018).
Izquierdo, E., Marchioni, E., Aoude-Werner, D., Hasselmann, C. & Ennahar, S. Smearing of soft Cheese with Enterococcus Faecium Whe 81, a multi-bacteriocin producer against Listeria monocytogenes. Food Microbiol. 26, 16–20 (2009).
Lhocine, N. et al. Pims modulates immune tolerance by negatively regulating drosophila innate immune signaling. Cell Host Microbe. 4, 147–158 (2008).
Douglas, A. E. Is the regulation of insulin signaling multi-organismal?. Sci. Signal. 4, e46 (2011).
Geminard, C., Rulifson, E. J. & Leopold, P. Remote control of insulin secretion by fat cells in drosophila. Cell Metab. 10, 199–207 (2009).
Colombani, J. et al. A nutrient sensor mechanism controls drosophila growth. Cell. 114, 739–749 (2003).
Guo, L., Karpac, J., Tran, S. L. & Jasper, H. Pgrp-Sc2 promotes gut immune homeostasis to limit commensal dysbiosis and extend lifespan. Cell. 156, 109–122 (2014).
Liu, W. et al. Enterococci mediate the oviposition preference of drosophila melanogaster through sucrose catabolism. Sci. Rep. 7, 13420 (2017).
Liu, W., Jiang, F., Bi, X. & Zhang, Y. Q. Drosophila Fmrp participates in the Dna damage response by regulating G2/M cell cycle checkpoint and apoptosis. Hum. Mol. Genet. 21, 4655–4668 (2012).
Koyle, M. L., et al. Rearing the Fruit fly drosophila melanogaster under axenic and gnotobiotic conditions. J. Vis. Exp. (2016).
Acknowledgements
We would like to thank all members of Jin faguang’s laboratory for helpful discussions. This work was supported by Key R & D Program of Shaanxi Province (2022ZDLSF01-10). We thank International Science Editing (http://www.internationalscienceediting.com) for editing this manuscript.
Author information
Authors and Affiliations
Contributions
F.J., Y.G. designed all experiments. Y.L., L.P., Y.G., L.W., J.C., F.G., P. L. and Z.L. performed experiments. Y.L. and L.P. wrote the paper. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, Y., Pan, L., Li, P. et al. Isolation of Enterococcus faecium and determination of its mechanism for promoting the growth and development of Drosophila. Sci Rep 13, 18726 (2023). https://doi.org/10.1038/s41598-023-43727-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-43727-1
This article is cited by
-
Feeding Drosophila gut microbiomes from young and old flies modifies the microbiome
Scientific Reports (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.