Abstract
Biomarker-based differential diagnosis of the most common forms of dementia is becoming increasingly important. Machine learning (ML) may be able to address this challenge. The aim of this study was to develop and interpret a ML algorithm capable of differentiating Alzheimer’s dementia, frontotemporal dementia, dementia with Lewy bodies and cognitively normal control subjects based on sociodemographic, clinical, and magnetic resonance imaging (MRI) variables. 506 subjects from 5 databases were included. MRI images were processed with FreeSurfer, LPA, and TRACULA to obtain brain volumes and thicknesses, white matter lesions and diffusion metrics. MRI metrics were used in conjunction with clinical and demographic data to perform differential diagnosis based on a Support Vector Machine model called MUQUBIA (Multimodal Quantification of Brain whIte matter biomArkers). Age, gender, Clinical Dementia Rating (CDR) Dementia Staging Instrument, and 19 imaging features formed the best set of discriminative features. The predictive model performed with an overall Area Under the Curve of 98%, high overall precision (88%), recall (88%), and F1 scores (88%) in the test group, and good Label Ranking Average Precision score (0.95) in a subset of neuropathologically assessed patients. The results of MUQUBIA were explained by the SHapley Additive exPlanations (SHAP) method. The MUQUBIA algorithm successfully classified various dementias with good performance using cost-effective clinical and MRI information, and with independent validation, has the potential to assist physicians in their clinical diagnosis.
Similar content being viewed by others
Introduction
Neurodegenerative dementias are a common and increasing cause of mortality and disability worldwide, particularly in older age1. The most common form of neurodegenerative dementia worldwide is Alzheimer’s dementia (AD), but recent epidemiological studies and refinement of new clinical criteria have shown that frontotemporal dementia (FTD) and dementia with Lewy bodies (DLB) are also common forms2. Specifically, DLB accounts for 5–7% of all dementias in the elderly3, FTD about 7%4, with one in four cases occurring late in life5, while AD may contribute to 60–70% of cases overall6. These neurodegenerative dementias are heterogeneous in their clinical presentation and underlying pathophysiology, although they share overlapping features7.
Biomarkers provide a powerful approach to understand the spectrum of neurological diseases by identifying them from the earliest manifestations to the final stages8. Increased diagnostic accuracy allows more precise prognostic approaches and often leads to specific treatments and optimal patient care9. In this context, it is important to determine which diagnostic markers can most reliably identify the different pathologies that lead to dementia. The main challenge for researchers and clinicians is to determine biomarkers that not only identify AD but can simultaneously distinguish between patients with FTD, DLB and cognitively normal controls (CN). Currently, imaging biomarkers assessed by magnetic resonance imaging (MRI) in conjunction with clinical examinations and neurocognitive assessments are the most commonly used tests to diagnose neurodegenerative dementias10. In recent years, several MRI-based imaging sequences or modalities have been introduced into clinical practice. The most commonly used MRI sequences are: structural T1-weighted 3D (T13D) and T2 Fluid Attenuated Inversion Recovery (FLAIR) images, which provide morphological measurements of the brain. In addition, Diffusion Tensor Imaging (DTI) is a well-established technique that is particularly useful for studying white matter (WM) integrity11.
The development of accurate image analysis pipelines combined with advanced classification methods could improve differential diagnosis12. Indeed, automated MRI segmentation tools can systematically generate brain morphometric features with minimal operator-differences, although a limitation is that some of these tools require a lot of processing time and computational power.
The best-known segmentation algorithms are FreeSurfer (FS), which can extract volume, area and thickness of many brain regions of interest (ROI) and the Lesion Prediction Algorithm (LPA), which can quantify WM hyperintensities. Both algorithms have been validated against manual raters and performed well13,14,15. As for DTI analysis, TRActs Constrained by UnderLying Anatomy (TRACULA) is one of the best validated tools for reconstructing WM pathways16.
The results of automated MRI pipelines can be used to develop machine learning (ML) tools with good classification performance. Support Vector Machines (SVM) are among the widely used supervised ML algorithms because they are easy to implement while being effective in diagnostic classification tasks17,18. In some cases, imaging variables can be used in conjunction with clinical and neuropsychological variables as input to multivariate data analyses and ML algorithms19. These models have been shown to be an effective strategy for identifying features capable of discriminating between different classes and subtypes of disease20,21, with results comparable to or better than neuropsychological tests alone22,23.
Indeed, ML in neuroscience is an ever-growing area of research based on learning relationships from large and complex data sets with the ability to apply the learned rules to other similar unseen data. Often, these tools appear to be able to detect brain patterns that are beyond human perception and can help clinicians to highlight and interpret medical findings24. To this end, tools for global and local interpretability of ML models have recently been developed25.
The present study was conducted within the framework of the Italian Network for Neuroscience and Neurorehabilitation (RIN) (https://www.reteneuroscienze.it/en/), established in 2017 by the Italian Ministry of Health. The RIN (1) promotes collaboration among the National Research Hospitals (IRCCS), (2) facilitates the dissemination of information on clinical/scientific community, and (3) promotes the use of harmonized protocols and advanced ML tools to enhance clinical practice26,27,28.
With this background, we developed and explained how our ML algorithm classified subjects into the four diagnostic classes (i.e.: AD, FTD, DLB, CN) based on sociodemographic, clinical, and imaging data. Our objectives were to: (1) discover the most informative combination of biomarkers to distinguish the different forms of dementia; (2) investigate the pathophysiological role of WM alterations multimodally; (3) provide an interpretation of how MUltimodal QUantification of Brain whIte matter biomArkers in dementia (MUQUBIA) works.
Methods
Study design
This study included the following steps: data preprocessing, selection of discriminative features, classification of subjects, SHapley Additive exPlanations (SHAP) analyses.
MRI images were processed with automated tools to extract the volume and thickness of cortical and subcortical brain regions, WM lesions, and WM diffusion metrics. All these values were used to train and test the MUQUBIA model for classification into diagnostic groups with a hold-out strategy.
Data
Subjects with a clinical diagnosis of AD, FTD, DLB, or CN were selected from 5 data sets.
The databases used for data collection were:
-
Alzheimer’s Disease Neuroimaging Initiative (ADNI)29: 84 AD, 15 DLB (from Neuropathology Data, http://adni.loni.usc.edu/methods/neuropath-methods/), 80 CN;
-
Frontotemporal Lobar Degeneration Neuroimaging Initiative (FTLDNI): 135 FTD, 10 CN;
-
National Alzheimer's Coordinating Center (NACC)30: 26 AD, 27 DLB, 18 CN;
-
NIH Parkinson's Disease Biomarkers Program (PDBP)31: 60 DLB;
-
Newcastle University, Newcastle upon Tyne32,33,34,35: 51 DLB.
The FTLDNI database contained sufficient FTD data for our purposes. All three FTD subtypes (i.e.: behavioural variant, semantic variant, and progressive non-fluent aphasia) were considered. AD and CN were selected from a larger sample to avoid size imbalance. For these three classes, only subjects with all three available sequences at the same time-point and DTI directions greater than 12 were included. Because there were no available open access databases of DLB patients with all three sequences needed for this study, we also included subjects with at least one sequence for the DLB group (Supplementary Table S2), thus improving the sample size and allowing more accurate data imputation. A sample of no less than 100 subjects was assembled for each diagnostic class. Sociodemographic, clinical, and imaging variables were collected for all subjects. Neuropsychological test scores were collected in our study but not included in the analysis because the assessment protocol for CN does not always include neuropsychological characterization. The clinical assessment used was the global score of the Clinical Dementia Rating (CDR) Dementia Staging Instrument.
Supplementary Table S1 lists the diagnostic and selection criteria for each study considered. For a complete list of subjects, diagnoses, and data sets used in this study see Supplementary Table S2.
MR imaging
Data used in the preparation of this article were obtained from the Alzheimer’s Disease Neuroimaging Initiative (ADNI) database (adni.loni.usc.edu). The ADNI was launched in 2003 as a public–private partnership, led by Principal Investigator Michael W. Weiner, MD. The primary goal of ADNI has been to test whether serial magnetic resonance imaging (MRI), positron emission tomography (PET), other biological markers, and clinical and neuropsychological assessment can be combined to measure the progression of mild cognitive impairment (MCI) and early Alzheimer’s disease (AD).
ADNI and FTLDNI data were collected from the Imaging Data Archive (IDA) web-portal of the Laboratory of NeuroImaging (LONI) (http://adni.loni.usc.edu).
NACC and PDBP data were downloaded from their respective web portals: https://naccdata.org/ and https://pdbp.ninds.nih.gov/.
The Newcastle data were provided directly by the Translational and Clinical Research Institute, Newcastle University.
Table 1 reports the imaging characteristics for each sequence and data set. Combining data from multiple hospitals is useful to build ML models that are invariant to systematic inter-scanner effects and to overcome differences in field strengths and acquisition protocols36.
Pipelines for image processing
N4 correction, from Advanced Normalization Tools (ANTs)37, was performed for all images to correct smooth intensity variations in MRI. The pipelines used for image processing in this study were FS version 6.0, LPA, and TRACULA.
FS is a pipeline for segmenting the cortical and subcortical brain structures using volumetric T13D images, where each voxel is labeled based on a probabilistic atlas13,14. The T13D MRIs were processed using the cross-sectional stream over the recon-all script using the Desikan-Killiany atlas and, when available in high quality, FLAIRs were used to improve the segmentation of the pial surfaces38. Volumes of subcortical regions in native space were normalized to FS estimated total intracranial volume (eTIV). Normalization was performed by dividing the volume of the region by the eTIV of the subject and multiplying the ratio by a reference value of 1409 ml39 to remove the effect of head size40. Cortical thickness values were not normalized19.
LPA is an algorithm for the quantification of the WM lesions that is part of the Lesion Segmentation Toolbox (LST)41. First, FLAIR images were linearly registered to T13D and each voxel was classified as cerebrospinal fluid (CSF), gray matter, or WM using the Statistical Parametric Mapping Tool v12.0 (SPM-12) tissue probability maps. Intensity distributions were calculated for each of them and weighted based on the spatial probability of belonging to WM. Finally, the map was converted to a binary lesion mask and its volume in native space (normalized to eTIV) was calculated.
TRACULA is a tool for automatic reconstruction of a set of 18 major WM pathways16. It uses prior information about the anatomy and relative positions of the WM tracts in relation to surrounding anatomical structures, obtained from a set of cognitively normal training subjects in which the tracts were manually labeled to produce tractography streamlines42. After mitigating image distortions due to eddy currents and B0 field inhomogeneities, TRACULA fits the shape of the tracts to both the subject's diffusion data and the anatomical neighborhood priors derived from the subject's T1 data. Fractional anisotropy (FA) and mean diffusivity (MD) were extracted from the diffusion data in MNI template space for each of the 18 reconstructed pathways. Then, the mean FA and mean MD of 48 ROIs were obtained from the WM John Hopkins University (JHU-ICBM-labels-1 mm) atlas43 and applied to the TRACULA maps.
Quality control of the processed outputs was performed by experienced neuroscientists (SD, AR) who inspected the images and the results of each pipeline slice by slice, and discarded those with poor quality or incorrect segmentation (Fig. 1). The influence of WM-hyperintensity load on FA and MD values in MUQUBIA selected tracts was assessed with two multivariate linear regression models44 (Supplementary Table S3). To investigate possible bias due to different image acquisition protocols in the datasets, we compared the distributions of MRI features of subjects with the same diagnosis from different datasets (inter-cohort variability), and the distributions of MRI features of subjects with different diagnoses from the same dataset (intra-cohort variability) (Supplementary Fig. S8).
MUQUBIA classification steps
Figure 2 shows the workflow for the creation of the Support Vector Machine (SVM) model.
Steps to create and test MUQUBIA. (a) Images of 506 subjects were processed to obtain the full set of features. (b) Missing values were replaced with median values. (c) The data were split into training set (70% of the subjects) and test set (30%) to avoid any bias in the selection of features and in the classification performance. (d) Values were standardized. (e) The full set of features was pruned to avoid overfitting using a bidirectional sequential feature selection approach. (f) The non-linear SVM model was built and fine-tuned on the training and validation sets, while being tested on the test set left aside. Acronyms: ft, features; MD, Mean Diffusivity; SVM, Support Vector Machine; WM, White Matter.
The imaging biomarkers, CDR scores, and sociodemographic information served as input to the SVM algorithm, which was run in Python 3.7.11. The framework we used was based on the scikit-learn library version 0.22.245.
The data set was randomly shuffled, with 70% of subjects forming the training set and 30% forming the test set. All 5 data sets were included in both the training and test data sets. None of the features resulted in more than 50% missing data. For the missing values, we employed the median as a method of imputation46. The statistical comparison between the original biomarker values of the training and test sets is presented in Supplementary Table S6 to demonstrate homogeneity between the two groups. All values were standardized by removing the mean and scaling to the variance of the feature distributions of the subjects from the whole training sample (z-scores).
To test the adequacy of the training sample size we modeled the relationship between training sample size and accuracy using the post-hoc “learning curve fitting” method47. The results are shown in Supplementary Fig. S1.
Machine learning models tend to overfit and become less generalizable when dealing with high-dimensional features, a well-known phenomenon called the “curse of dimensionality”48. A large set of features generally implies the presence of irrelevant, redundant, or correlated variables. To overcome this, our algorithm performed feature selection, considering only those features that maximized the accuracy of the classification task in the training set evaluated with a five-fold cross validation (CV) approach. This procedure allowed us to determine which variables were most informative for the diagnostic categories selected in this study. To determine the best set, a forward and backward sequential feature selection approach was followed, with each feature added to the model individually49. If accuracy increased, the feature was considered important; otherwise, it was discarded. After the selection process was completed, the surviving features were further reduced to obtain a Variance Inflation Factor (VIF) below the threshold value of 5 for each of them (see Table 4), indicating that there was no collinearity50.
To increase computational efficiency, the one-versus-rest (OVR) method was used to transform the multi-class problem into multiple binary classifications. The classification results were obtained using a non-linear SVM51. We optimized the search for the best hyperparameters using a five-fold CV splitting strategy over a grid search to find the best combination of SVM kernel, C and γ values. We also used L2 regularization.
Finally, SVM performance was evaluated using the following metrics: accuracy, precision, recall, F1 score, Area Under the Curve (AUC), Receiver Operating Characteristic curve (ROC). The global metrics, except for the accuracy, are macro-averaged, that is the arithmetic mean of the individual class performance.
In the context of ML, interpretability is necessary to explain the outcome of a model. In this study, Shapley values were calculated using the library SHAP, version 0.40.025, to better understand the contribution of each feature expression.
The clinical challenge for the MUQUBIA algorithm was to distinguish between the different types of dementia. Because CDR is a clinical score collected by clinicians during the assessment process to differentiate the healthy from the dementia state, we evaluated the performance of our model even without including this scale in the feature set (Supplementary Fig. 2) to avoid circularity and minimize potential bias in favor of CN classification.
Statistics
Differences in the variances of the feature distribution of each diagnostic class between the original data set and the data set with imputed medians were assessed using the Brown-Forsythe test. Differences in sociodemographic, clinical, neuropsychological and morphological feature distributions among diagnostic groups, and inter- intra-cohort differences were assessed using the Kruskal–Wallis test for continuous variables and the Chi-squared test for dichotomous variables. Post-hoc analyses were performed to test differences between the four diagnostic groups by pairwise comparisons of the Wilcoxon rank sum test for continuous variables and a pairwise comparison between pairs of proportions for dichotomous variables. The p-values of the post-hoc analyses were adjusted with the Benjamini–Hochberg correction. To compare the neuropathological multilabel evidence with the MUQUBIA results, the metric LRAP (Label Ranking Average Precision) was calculated. Similarity between test and train ROC curves was assessed using the DeLong’s test. All statistical analyses were performed using R version 3.6.3, and the significance level was set at 0.05 for all tests.
Pipeline availability
The single subject classification tool based on the MUQUBIA models was also made publicly available through the neuGRID platform (https://neugrid2.eu)21,52,53, an on-line high-performance computing (HPC) infrastructure that provides source code, tools, and data for image processing and ML analysis (see Supplementary Fig. S5).
Results
Subjects
The final data set included 506 subjects: 110 AD, 135 FTD, 153 DLB and 108 CN. Demographic, clinical, neuropsychological, and ApoE information are shown in Table 2. Only neuropsychological tests that followed the same protocol in all 5 data sets were considered.
Feature set and sanity check
Image processing yielded a total of 336 features: 202 from FS (including 132 volumes and 70 cortical thickness values); 2 from LPA (WM lesion volume and WM lesion number); 36 from TRACULA (18 FA, 18 MD values for WM pathways); 96 features from the application of the JHU atlas ROIs to the FA and MD maps. The full list of features is reported in Supplementary Table S4.
Table 3 reports the number of outputs deemed acceptable after visual inspection for each pipeline and diagnostic group, as well as the consistency of the success rate for each pipeline in the 4 diagnostic groups.
MUQUBIA algorithm
The training sample of MUQUBIA included 354 subjects, while 152 subjects formed the test group. The best hyper-parameters among those tested with the GridSearchCV function (i.e. kernel: linear, polynomial, sigmoid, radial basis function (RBF); C: 1, 10, 100, 1000, 10,000; γ: 0.1, 0.01, 0.001, 0.0001, 0.00001), were RBF kernel, C equal to 1000, and γ equal to 0.0001. For the entire analysis, consisting of image processing and classification of the subjects, the algorithm requires 10 h on a machine running Ubuntu Server 18.04 LTS version on a Sun Grid Engine scheduler equipped with 1300 GB RAM and 214 cores. Most of the requested time is spent for image analysis.
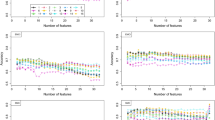
The algorithm selected 24 features, but two of them were discarded because of a VIF above 5, namely: fractional anisotropy of the left retrolenticular part of the internal capsule and left postcentral thickness. Figure 3 shows the imaging features selected by the bidirectional selection process implemented in MUQUBIA. The 22 features composing the best set are listed in Table 4. The features were ranked from highest to lowest importance in distinguishing the four diagnostic classes. The set of best features was composed by CDR, 19 MRI features, age and gender. The influence of age and gender on the MRI features was assessed and the results are reported in Supplementary Table S5. Across all diagnoses, CDR was the most important feature. The results of the Kruskal–Wallis test showed that the diagnostic groups differed significantly with respect to the selected variables. Post-hoc analyses revealed p-values below 0.05 in at least one comparison for all the features.
Representation of brain regions corresponding to imaging features selected by MUQUBIA to distinguish the different diagnostic classes (AD, DLB, FTD, CN). The color of each brain region reflects the ability of the corresponding feature to discriminate among the different classes (averaged mean Shapley value). Acronyms: L, left; R, right.
The Brown-Forsythe test always yielded a p-value greater than 0.05 (Supplementary Table S7), indicating that the original variance of the data set was not altered by median imputation.
SHAP analysis
Figure 4 shows the average influence of the features on the prediction of each diagnosis, the values of CDR have the greatest influence especially for the classification of CN and AD, whereas the FA of the left corticospinal tract, among the others, influences the classification of DLB and AD groups the most.
Contribution of each feature to the classification, represented by the mean Shapley magnitude values. The graph shows the importance of each variable for each diagnostic group. Acronyms: AD, Alzheimer’s Dementia; FTD, Frontotemporal Dementia; DLB, Dementia with Lewy Body; CN, Cognitive Normal; FA, Fractional Anisotropy; MD, Mean Diffusivity; LH, left hemisphere; RH, right hemisphere.
The global interpretability plot (Fig. 5), shows whether a feature shifts the MUQUBIA prediction toward other diagnostic classes and the relative contribution of each feature. The plot consists of all points standardized. Focusing on the CN class, low values of CDR have a very high impact on the determination of this diagnosis. High values of temporal ROIs (left hippocampal volume and left entorhinal thickness) also have a high influence, as does a low MD value of right medial lemniscus. Other MRI measures do not provide simple or practical information on how they influence MUQUBIA outcome. Atrophy of the left frontal pole, associated with the increase of MD in the right medial lemniscus and the decrease of FA in the fronto-occipital fasciculus, influences the prediction of FTD class in addition to the degeneration of the corticospinal tract. For DLB class, the corticospinal tract represented an imaging biomarker of great importance, especially with a reduced value of FA, although this tract is not a classic biomarker for DLB. Other imaging biomarkers, such as preservation of MD in the retrolenticular part of the internal capsule and preservation of left cortical thickness (entorhinal and inferior parietal), have an impact on the classification of DLB patients. For AD, lack or moderate impairment of FA for the corticospinal tract and high scores for CDR have a major impact on classification, followed by damage and shrinkage of some ROIs of the left hemisphere, such as: left superior fronto-occipital fasciculus, inferior-parietal thickness, entorhinal thickness. In general, age represents one of the most important factors for classification in all dementias.
Global interpretability plots for each diagnostic class. Each dot corresponds to a subject in the training set. The position of the dot on the x-axis shows the effect of that feature on the prediction of the model for that subject. If multiple dots land at the same x position, they piled up to show density. The features are ordered by the sum of the Shapley values. Colors are used to display the standardized value of each feature (colder colors represent lower values, warmer colors represent higher values). Acronyms: AD, Alzheimer’s Dementia; FTD, Frontotemporal Dementia; DLB, Dementia with Lewy Body; CN, Cognitive Normal; LH, left hemisphere; RH, right hemisphere; FA, Fractional Anisotropy; MD, Mean Diffusivity.
Additional information can be derived from the partial dependence plot of the main features (Fig. 6). This plot shows the marginal effect that two features have on the predicted outcome of MUQUBIA. Once the first feature was selected, the second was automatically chosen, picking out the feature with the strongest interaction with first one. Most of the plots show complex correlations between the two features and the Shapley values (Supplementary Fig. S7), which are discussed in more detail in the “Discussion” section.
SHAP partial dependence plots for each diagnostic class (AD, DLB, FTD, CN). Each subplot shows the marginal effect that two features have on the predicted diagnosis. Once the first feature is chosen, the second is selected based on the feature with which the first feature interacts most strongly. The color of a dot indicates the value for the second feature. The color of each plot changes progressively from blue to red (or vice-versa) as you move along the axes. Colder colors represent lower values, warmer colors represent higher values of the second feature. Acronyms: AD, Alzheimer’s Dementia; FTD, Frontotemporal Disease; DLB, Dementia with Lewy Body; CN, Cognitive Normal; LH, left hemisphere; RH, right hemisphere; FA, Fractional Anisotropy; MD, Mean Diffusivity.
Finally, to increase the interpretability and to understand potential problems of MUQUBIA we analyzed some correctly and incorrectly predicted subjects in Supplementary Fig. S3 and in Supplementary Fig. S4.
MUQUBIA performance on training set
The classification resulted in the following global metrics: accuracy 91.53%, macro-precision 91.62%, macro-recall 90.82%, macro-F1 score 90.92%, AUC 98.44%.
MUQUBIA performance on test set
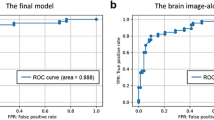
The SVM classification task for the subjects in the test set (Fig. 7) resulted in the following global metrics: accuracy 87.50%, macro-precision 88.00%, macro-recall 88.36%, macro-F1 score 87.88%, AUC 97.79%. The DeLong test revealed no significant differences (p > 0.05) between the ROC curves of the training and test sets for each class. A summary of the performance metrics is provided in Table 5.
Classification metrics obtained with MUQUBIA, trained with the same selected features but without CDR, are shown in Supplementary Fig. S2. Performance decreased slightly, especially in the case of CN. However, the classification task yielded the following global metrics: accuracy 84%, macro-precision 84%, macro-recall 84%, macro-F1 score 83%, AUC 96%.
MUQUBIA performance on neuropathological assessed subsample of the test set
Table 6 reports the LRAP value used to compare the agreement between the MUQUBIA probability estimates with the National Institute on Aging and Alzheimer’s Association protocol54 for neuropathological assessment of 9 patients in our test group. The LRAP metric is classically used in multilabel ranking problems55. It determines the percentage of higher-ranked labels that resemble the true labels for each of the given samples. The score obtained is always greater than 0, and the best score is 1.
MUQUBIA report
An example of the MUQUBIA report generated with the on-line tool on the neuGRID platform is available as supplementary material (Supplementary Fig. S6).
Discussion
In this work, we developed an automated ML algorithm based on multimodal MRI capable of discriminating the most common forms of dementia. The performance of this classifier was validated using quality metrics that resulted in high scores for accuracy, macro-precision, macro-recall, macro-F1 and AUC. The classifier was successful in discriminating between the 4 groups (AD, FTD, DLB and CN) characterized by different neuropsychological scores and ApoE expression (Table 2). The algorithm selected CDR, age, gender information, MRI-based diffusion metrics, volumetric and cortical thickness values as the best differentiating features.
SVM performance did not differ significantly between the test and training sets using 22 informative features; and performances on training set were higher than performance on the test set arguing against severe overfitting56.
In the test set group, MUQUBIA scored highest in discriminating CN from the others, with excellent discrimination performance for each diagnostic class. The lowest performance was in detecting the AD group. This could be due to the overlap with other types of dementia, especially DLB57. Neuropathological brains assessed by Montine’s criteria were also correctly classified by MUQUBIA with very good performance (LRAP = 95%).
The MRI features studied were appropriate to selectively distinguish AD, FTD, DLB and to differentiate them from cognitively normal aging. The neuroimaging features were extracted from FS and TRACULA pipelines, making mandatory only the T13D and DTI to run the MUQUBIA algorithm. Optionally, the FLAIR can be used to improve the pial segmentation and to reduce segmentation errors caused by WM hyperintensities. The WM hyperintensity information extracted from the LPA does not seem to affect MUQUBIA, as this aspect is likely already present in the DTIs as increased MD and decreased FA. It is known that WM hyperintensity may have an impact on the DTI metrics, although in the present study and in relation to the features selected by MUQUBIA, only the tract of the superior fronto-occipital fasciculus was weakly affected.
In addition to cortical/subcortical gray matter information, which has long been considered informative biomarkers, WM diffusion metrics have also been shown to be important for ML classification. These metrics appear to be useful in distinguishing AD from FTD17, and, albeit to a lesser extent, in distinguishing AD from DLB35.
The implemented data-driven MUQUBIA approach identified the best set of features, many of which were consistent with those described in the literature, while others were unexpected. For the benefit of the reader, the discussion of the results was organized according to the following 3 main macro-groups:
1. Clinical and socio-demographic features:
Among the most important features in our model there is the CDR, a well-known test for detecting and assessing the severity of dementia58; therefore, it is not surprising that it turned out to be the most informative feature. Interestingly, the SHAP partial dependence plot (Fig. 6) shows that the probability of being classified as cognitively normal by MUQUBIA is greater when the CDR score is zero and the MD value of the medial lemniscus tract is low, indicating no degeneration. Higher values of MD, may instead, progressively reduce the weight of the (non-pathological) CDR score in classifying a person as cognitively normal. This could be very promising information, especially for secondary prevention, which, by combining multimodal ad hoc biomarkers, would allow more accurate, sensitive, and earlier stratification of individuals at the pre-dementia stage than using CDR alone59. As expected, the MUQUBIA model without CDR performed worse in the classification of CN, but also in AD, DLB, and FTD confirming the importance of CDR also in the classification of dementia groups, as explained by the Shapley values (Fig. 4).
In addition, although neurological diseases are naturally assumed to affect only the elderly, this is not always the case. From the Shapley analysis, younger individuals belonging to the CN class are more likely to drop out (Fig. 5). The younger age of the FTD group must also be taken into account to explain possible brain imaging deviation and possible errors of our model.
Interestingly, according to the literature, DLB is associated with male preponderance60, and this was also observed in our DLB group. Finally, MUQUBIA seems to be strongly influenced by the degeneration of the left corticospinal tract, which is more pronounced in women than in men, when classifying AD subjects.
2. Cortical and sub-cortical features:
DLB is associated with less global atrophy than AD, whereas posterior cingulate atrophy was similar in AD and DLB. AD patients showed more atrophy of the medial temporal lobe structures compared to DLB61. Hippocampal atrophy was not limited to the AD and DLB groups, but has also been noted in FTD, although to a lesser extent than in AD62. Conversely, FTD patients showed greater atrophy of the temporal pole and orbitofrontal areas than AD patients, while AD patients showed greater atrophy of the posterior cingulate and inferior parietal regions63. In our study, no significant differences were found between DLB and CN with respect to the temporal pole, inferior parietal and orbitofrontal areas.
According to the literature, we found the putamen volume of AD is intermediate between CN and FTD, showing more atrophy in the latter64. DLB showed volumetric atrophy in the putamen65, with a moderate influence in the MUQUBIA model, or a slight influence in other basal ganglia such as the left pallidum66. Even in FTD, where there is limited and conflicting evidence in the literature regarding the volumetry of deep gray matter structures, our results tend to confirm the findings of Möller et al., with respect to the basal ganglia, and show that FTD patients are characterized by the most severe atrophy compared with other diagnostic groups as well as that atrophy of the pallidum contributes to the classification of FTD patients in MUQUBIA model. Further specific efforts will be needed to clarify this point in future studies.
Surprisingly, the volume of the left frontal pole was highest in FTD and differed significantly from all other patients examined in this study. This can be partly explained by the younger age of FTD compared with the other groups by approximately a decade. Consistent with the literature, patients with AD had smaller volumes of the frontal pole, isthmus of cingulate and left pars opercularis67 compared with CN subjects.
Cortical thickness was a sensitive and comprehensive marker to distinguish AD from other dementias. Cortical shrinkage of the left entorhinal cortex has been reported to be greater in AD than in DLB68, but similar in AD and FTD69. Left inferior parietal thickness, also greater in FTD, proved to be a robust marker to disentangle AD from FTD for MUQUBIA70.
Moreover, the SHAP partial dependence plot (Fig. 6) showed that MUQUBIA classifies patients as AD when a concomitant reduction in left inferior parietal thickness is associated with a reduction in total left cortical volume, which has been linked in previous studies to a decrease in semantic fluency71. Likewise, the SHAP partial dependence analysis (Fig. 6) revealed that MUQUBIA tends to classify patients in the DLB class when they exhibit lower total left cortical volume and a reduction in left parsopercularis thickness. This observation aligns with the existing literature, that links speech fluency impairment to these important regions in DLB72,73.
3. DTI feature
FA of the left corticospinal tract was lower in AD than in CN74. Degeneration of the corticospinal tract has also been described in FTD75. Instead, there is no clear evidence in the literature of damage of this tract in the DLB group76, although this tract had a major effect on MUQUBIA. Possible explanations may be found in the larger group size used in our study than in other efforts and the quality of the DTI pipeline and scans we used to quantify the DTI metrics.
FA of the splenium of the corpus callosum and the superior fronto-occipital fasciculus was lower in AD than in CN72, although the lowest FA values of these pathways occurred in DLB. DLB also showed lower values for FA than all other groups in many other pathways and ROIs77. According to the literature, DLB showed higher MD in brainstem areas78, such as in the pontine crossing tract, compared to CN. Other imaging biomarkers, such as the preservation of the retrolenticular part of the internal capsule, influenced MUQUBIA toward DLB classification. This is correct given that motor and sensory fibres run through this ROI79 and must be maintained integer to prevent dysphagia and swallowing dysfunction. FTD and AD were the most affected groups in the right retrolenticular part of the internal capsule80. The medial lemniscus MD proved to be the third most important feature for classifying FTD patients in MUQUBIA. As previously mentioned, FTD was characterized by the degeneration of the corticospinal tract81 similar to AD. The SHAP partial dependence plot (Fig. 6) for the FTD class also revealed that MUQUBIA finds a direct relationship between left corticospinal tract FA and right medial lemniscus MD values indicating a specific form of frontal neurodegeneration. Last but not least, the correlation between these two tracts could confirm interesting findings on the detection of subtypes of frontotemporal lobar degeneration82.
Benefits from MUQUBIA
Recently, the number of studies using ML has steadily increased because ML enables a fully data-driven and automated approach. ML is indeed flexible in discovering patterns, complex relationships, and predicting unobserved outcomes in data, starting from a sufficient number of observations83, especially with increasing complexity, where classical statistical methods may be rather ineffective84.
Research studies often address the binary classification between two clinical conditions (i.e.: AD vs. CN; FTD vs. CN; FTD vs. AD, etc.…), but this does not reflect the reality of the clinician who needs to make a diagnosis considering multiple neurodegenerative diseases at the same time. Although the field of neurodegenerative diseases has been extensively researched85, to our knowledge, few studies have implemented an MRI-based ML algorithm for the classification of AD, FTD, DLB and CN56,86,87, and to date, no study has used DTIs and multimodal analyses simultaneously. MUQUBIA is the first ML algorithm for differential diagnosis to use DTI together with T13D and FLAIR on a very robust sample size. In fact, Klöppel et al. recruited a small group of FTD and DLB, whereas Koikkalainen et al. and Tong et al. included a broader range of dementias (such as vascular dementia and subjective memory complaints), but still with fewer subjects per group and with worse performance compared with MUQUBIA (i.e.: Klöppel et al.: accuracy of 65%; Koikkolainen et al.: accuracy of 70.6%; Tong et al.: accuracy of 75.2%). Moreover, Tong et al. used CSF biomarkers that required an invasive procedure such as lumbar puncture which is difficult to obtain in a large population. This could also affect the applicability in daily routine and clinical practice in hospitals compared to the data needed as input to MUQUBIA. Many advanced research frameworks recommend the analysis of amyloid, tau, or 18F-fluorodeoxyglucose positron emission tomography (PET) scans of the brain and CSF to better classify patients88. However, these expensive procedures may limit their actual utility and are not available in the normal clinical setting. MUQUBIA requires routinely available MRIs, a clinical test, and a few demographic information, so it can be considered widely applicable without incurring excessive costs and burdening patients unnecessarily.
The online MUQUBIA tool does not require manual or “a priori” preprocessing, and the end-user does not need to have prior knowledge of the algorithm, although a quality check of the ROI segmentation is always advisable.
In addition, experienced neuroradiologists are often not available in routine clinical practice outside of a specialized memory clinic, so an automated method capable of extracting and interpreting the information with high precision would be of great clinical value.
A strength of this study is that the DTIs followed heterogeneous acquisition protocols, e.g., gradient directions vary from a minimum of 19 (low) to a maximum of 114 (high). The FLAIR and T13D parameters differed, bringing this study closer also to a real-world clinical scenario.
Limitations and future developments
We have considered various types of neurodegenerative diseases, which account for a large proportion of dementia cases, but this approach to differential diagnosis is far from complete. We did not attempt to define subtypes, such as posterior cortical atrophy in AD or the language or semantic variant in FTD or psychiatric and delirium onset in DLB. This study has limitations related to a partial influence of age and gender on certain MRI features, particularly in the FTD or in DLB. In fact, FTD group is the youngest and has an average age of onset of 56 years, while AD and DLB occurs later9. DLB group instead showed a preponderance of male. These confounders could help the classifier to identify more easily these groups and additional experiments should be performed to exclude this point. The fact that inter-cohort variability was lower than intra-cohort variability hints that the effect of etiology of dementia on MRI features is more important than potential bias induced by heterogeneous acquisition protocols, still the classifier might be further improved by trying to minimize the “center-effect” and reduce the few differences observed89.
Future efforts will aim to speed up processing times with new tools, such as FastSurferCNN, that exploit deep neural networks and graphical processor units to reduce image preprocessing in minutes.
Finally, due to difficulties finding datasets that contained multimodal and multiclass data, this study lacked a complete independent validation data set, but in the future, MUQUBIA should be validated with independent data sets given the upcoming Big-Data era.
Conclusion
The fully automated classifier developed in this study can discriminate between AD, FTD, DLB and CN with good to excellent performance. Our ML classifier can help clinicians as a second opinion tool to better diagnose the different forms of dementia based on routine and cost-effective biomarkers such as age, gender, CDR and automatically extracted MRI features. It is important to point out that the interpretability and explainability of the methods of ML provide important clues, allow to go beyond the slogan “ML is a black-box”, and lead to the discovery of new informative data-driven candidate biomarkers.
Data availability
Publicly available data sets were analyzed in this study: ADNI and FTLDNI are accessible through the Laboratory of NeuroImaging (LONI) web portal (http://adni.loni.usc.edu). NACC and PDBP data are available through the following web portals: https://naccdata.org/ and https://pdbp.ninds.nih.gov/. MUQUBIA algorithm is publicly accessible through the neuGRID platform (https://www.neugrid2.eu).
Change history
18 January 2024
A Correction to this paper has been published: https://doi.org/10.1038/s41598-024-51435-7
References
GBD 2016 Neurology Collaborators. Global, regional, and national burden of neurological disorders, 1990–2016: A systematic analysis for the Global Burden of Disease Study 2016. Lancet Neurol. 18, 459–480. https://doi.org/10.1016/S1474-4422(18)30499-X (2019).
Benussi, A. et al. Classification accuracy of transcranial magnetic stimulation for the diagnosis of neurodegenerative dementias. Ann. Neurol. 87, 394–404. https://doi.org/10.1002/ana.25677 (2020).
McKeith, I. G. et al. Research criteria for the diagnosis of prodromal dementia with Lewy bodies. Neurology 94, 743–755. https://doi.org/10.1212/WNL.0000000000009323 (2020).
Van der Flier, W. M. & Scheltens, P. Amsterdam dementia cohort: Performing research to optimize care. J. Alzheimers Dis. 62, 1091–1111. https://doi.org/10.3233/JAD-170850 (2018).
Onyike, C. U. & Diehl-Schmid, J. The epidemiology of frontotemporal dementia. Int. Rev. Psychiatry 25, 130–137. https://doi.org/10.3109/09540261.2013.776523 (2013).
Young, J. J., Lavakumar, M., Tampi, D., Balachandran, S. & Tampi, R. R. Frontotemporal dementia: Latest evidence and clinical implications. Ther. Adv. Psychopharmacol. 8, 33–48. https://doi.org/10.1177/2045125317739818 (2018).
Armstrong, R. A., Lantos, P. L. & Cairns, N. J. Overlap between neurodegenerative disorders. Neuropathology 25, 111–124. https://doi.org/10.1111/j.1440-1789.2005.00605.x (2005).
Mayeux, R. Biomarkers: Potential uses and limitations. NeuroRx 1(2), 182–188. https://doi.org/10.1602/neurorx.1.2.182 (2004).
Erkkinen, M. G., Kim, M. O. & Geschwind, M. D. Clinical neurology and epidemiology of the major neurodegenerative diseases. Cold Spring Harb. Perspect. Biol. 10, a033118. https://doi.org/10.1101/cshperspect.a033118 (2018).
Frisoni, G. B., Fox, N. C., Jack, C. R. Jr., Scheltens, P. & Thompson, P. M. The clinical use of structural MRI in Alzheimer disease. Nat. Rev. Neurol. 6, 67–77. https://doi.org/10.1038/nrneurol.2009.215 (2010).
Risacher, S. L. & Saykin, A. J. Neuroimaging in aging and neurologic diseases. Handb. Clin. Neurol. 167, 191–227. https://doi.org/10.1016/B978-0-12-804766-8.00012-1 (2019).
Amelio, L. & Amelio, A. Classification methods in image analysis with a special focus on medical analytics. In Machine Learning Paradigms. Intelligent Systems Reference Library Vol. 149 (eds Tsihrintzis, G. et al.) (Springer, Cham, 2019). https://doi.org/10.1007/978-3-319-94030-4_3.
Fischl, B. et al. Whole brain segmentation: Automated labeling of neuroanatomical structures in the human brain. Neuron 33, 341–55. https://doi.org/10.1016/s0896-6273(02)00569-x (2002).
Fischl, B. et al. Automatically parcellating the human cerebral cortex. Cereb. Cortex 14, 11–22. https://doi.org/10.1093/cercor/bhg087 (2004).
Ribaldi, F. et al. Accuracy and reproducibility of automated white matter hyperintensities segmentation with lesion segmentation tool: A European multi-site 3T study. Magn. Reson. Imaging. 76, 108–115. https://doi.org/10.1016/j.mri.2020.11.008 (2021).
Yendiki, A. et al. Automated probabilistic reconstruction of white-matter pathways in health and disease using an atlas of the underlying anatomy. Front. Neuroinform. 5, 23. https://doi.org/10.3389/fninf.2011.00023 (2011).
Bron, E. E. et al. Multiparametric computer-aided differential diagnosis of Alzheimer’s disease and frontotemporal dementia using structural and advanced MRI. Eur. Radiol. 8, 3372–3382. https://doi.org/10.1007/s00330-016-4691-x (2017).
Dukart, J. et al. Alzheimer’s disease neuroimaging initiative. Meta-analysis based SVM classification enables accurate detection of Alzheimer’s disease across different clinical centers using FDG-PET and MRI. Psychiatry Res. 212, 230–6. https://doi.org/10.1016/j.pscychresns.2012.04.007 (2013).
Westman, E., Aguilar, C., Muehlboeck, J. S. & Simmons, A. Regional magnetic resonance imaging measures for multivariate analysis in Alzheimer’s disease and mild cognitive impairment. Brain Topogr. 26, 9–23. https://doi.org/10.1007/s10548-012-0246-x (2013).
Kim, J. P. et al. Machine learning based hierarchical classification of frontotemporal dementia and Alzheimer’s disease. Neuroimage Clin. 23, 101811. https://doi.org/10.1016/j.nicl.2019.101811 (2019).
Archetti, D. et al. Inter-cohort validation of SuStaIn model for Alzheimer’s disease. Front. Big Data 4, 661110. https://doi.org/10.3389/fdata.2021.661110 (2021).
Möller, C. et al. Alzheimer disease and behavioral variant frontotemporal dementia: Automatic classification based on cortical atrophy for single-subject diagnosis. Radiology 3, 838–48. https://doi.org/10.1148/radiol.2015150220 (2016).
Klöppel, S. et al. Accuracy of dementia diagnosis: A direct comparison between radiologists and a computerized method. Brain 131, 2969–74. https://doi.org/10.1093/brain/awn239 (2008).
Erickson, B. J., Korfiatis, P., Akkus, Z. & Kline, T. L. Machine learning for medical imaging. Radiographics 37, 505–515. https://doi.org/10.1148/rg.2017160130 (2017).
Lundberg, S. M. & Lee, S.-I. A unified approach to interpreting model predictions. In Proceedings of the 31st International Conference on Neural Information Processing Systems (NIPS'17). Curran Associates Inc., Red Hook, NY, USA, 4768–4777. Preprint at 1705.07874 (2017).
Redolfi, A. et al. Medical Informatics Platform (MIP): A pilot study across clinical Italian cohorts. Front. Neurol. 11, 1021. https://doi.org/10.3389/fneur.2020.01021 (2020).
Nigri, A. et al. Quantitative MRI harmonization to maximize clinical impact: The RIN-neuroimaging network. Front. Neurol. 13, 855125. https://doi.org/10.3389/fneur.2022.855125 (2022).
Palesi, F. et al. MRI data quality assessment for the RIN: Neuroimaging Network using the ACR phantoms. Phys. Med. 104, 93–100. https://doi.org/10.1016/j.ejmp.2022.10.008 (2022).
Petersen, R. C. et al. Alzheimer’s Disease Neuroimaging Initiative (ADNI): Clinical characterization. Neurology 74, 201–9. https://doi.org/10.1212/WNL.0b013e3181cb3e25 (2010).
Beekly, D. L. et al. The National Alzheimer’s Coordinating Center (NACC) database: The uniform data set. Alzheimer Dis. Assoc. Disord. 21, 249–258. https://doi.org/10.1097/WAD.0b013e318142774e (2007).
Ofori, E., Du, G., Babcock, D., Huang, X. & Vaillancourt, D. E. Parkinson’s disease biomarkers program brain imaging repository. Neuroimage 124, 1120–1124. https://doi.org/10.1016/j.neuroimage.2015.05.005 (2016).
Firbank, M. J. et al. Diffusion tensor imaging in dementia with Lewy bodies and Alzheimer’s disease. Psychiatry Res. 155, 135–145. https://doi.org/10.1016/j.pscychresns.2007.01.001 (2007).
Firbank, M. J. et al. High resolution imaging of the medial temporal lobe in Alzheimer’s disease and dementia with Lewy bodies. J. Alzheimers Dis. 21, 1129–1140. https://doi.org/10.3233/jad-2010-100138 (2010).
Firbank, M. J. et al. Neural correlates of attention-executive dysfunction in Lewy body dementia and Alzheimer’s disease. Hum. Brain Mapp. 37, 1254–70. https://doi.org/10.1002/hbm.23100 (2016).
Donaghy, P. C. et al. Diffusion imaging in dementia with Lewy bodies: Associations with amyloid burden, atrophy, vascular factors and clinical features. Parkinsonism Relat. Disord. 78, 109–115. https://doi.org/10.1016/j.parkreldis.2020.07.025 (2020).
Archetti, D. et al. Multi-study validation of data-driven disease progression models to characterize evolution of biomarkers in Alzheimer’s disease. Neuroimage Clin. 24, 101954. https://doi.org/10.1016/j.nicl.2019.101954 (2019).
Tustison, N. J. et al. N4ITK: Improved N3 bias correction. IEEE Trans. Med. Imaging 29, 1310–20. https://doi.org/10.1109/TMI.2010.2046908 (2010).
Lindroth, H. et al. Examining the identification of age-related atrophy between T1 and T1 + T2-FLAIR cortical thickness measurements. Sci. Rep. 9, 11288. https://doi.org/10.1038/s41598-019-47294-2 (2019).
Reite, M. et al. Brain size and brain/intracranial volume ratio in major mental illness. BMC Psychiatry 10, 79. https://doi.org/10.1186/1471-244X-10-79 (2010).
De Francesco, S. et al. Norms for automatic estimation of hippocampal atrophy and a step forward for applicability to the Italian population. Front. Neurosci. 15, 656808. https://doi.org/10.3389/fnins.2021.656808 (2021).
Schmidt, P. et al. An automated tool for detection of FLAIR-hyperintense white-matter lesions in multiple sclerosis. Neuroimage 59, 3774–3783. https://doi.org/10.1016/j.neuroimage.2011.11.032 (2012).
Watanabe, A. et al. The detection of white matter alterations in obsessive-compulsive disorder revealed by TRActs Constrained by UnderLying Anatomy (TRACULA). Neuropsychiatr. Dis. Treat. 14, 1635–1643. https://doi.org/10.2147/NDT.S164058 (2018).
Mori, S. et al. Stereotaxic white matter atlas based on diffusion tensor imaging in an ICBM template. Neuroimage 40, 570–582. https://doi.org/10.1016/j.neuroimage.2007.12.035 (2008).
Svärd, D. et al. The effect of white matter hyperintensities on statistical analysis of diffusion tensor imaging in cognitively healthy elderly and prodromal Alzheimer’s disease. PLoS ONE 12, e0185239. https://doi.org/10.1371/journal.pone.0185239 (2017).
Pedregosa, F. et al. Scikit-learn: Machine learning in python. Journal of Machine Learning Research. 12, 2825–2830 (2011).
Acuña, E. & Rodriguez, C. The treatment of missing values and its effect on classifier accuracy. In Classification, Clustering, and Data Mining Applications. Studies in Classification, Data Analysis, and Knowledge Organisation (eds Banks, D. et al.) (Springer, Berlin, 2004). https://doi.org/10.1007/978-3-642-17103-1_60.
Balki, I. et al. Sample-size determination methodologies for machine learning in medical imaging research: A systematic review. Can. Assoc. Radiol. J. 70, 344–353. https://doi.org/10.1016/j.carj.2019.06.002 (2019).
Berisha, V. et al. Digital medicine and the curse of dimensionality. NPJ Digit. Med. 4, 153. https://doi.org/10.1038/s41746-021-00521-5 (2021).
Jović, A., Brkić, K. & Bogunović, N. A review of feature selection methods with applications. In 38th International Convention on Information and Communication Technology, Electronics and Microelectronics (MIPRO), 1200–1205 (IEEE, 2015). https://doi.org/10.1109/MIPRO.2015.7160458.
Craney, T. A. & Surles, J. G. Model-dependent variance inflation factor cutoff values. Qual. Eng. 14, 391–403. https://doi.org/10.1081/QEN-120001878 (2002).
Abraham, A. et al. Machine learning for neuroimaging with scikit-learn. Front. Neuroinform. 8, 14. https://doi.org/10.3389/fninf.2014.00014 (2014).
Redolfi, A., Bosco, P., Manset, D., Frisoni, G. B., neuGRID consortium. Brain investigation and brain conceptualization. Funct. Neurol. 28, 175–90. https://doi.org/10.11138/FNeur/2013.28.3.175 (2013).
Redolfi, A. et al. Italian, European, and international neuroinformatics efforts: An overview. Eur. J. Neurosci. https://doi.org/10.1111/ejn.15854 (2022).
Hyman, B. T. et al. National institute on aging-Alzheimer’s association guidelines for the neuropathologic assessment of Alzheimer’s disease. Alzheimers Dement. 8, 1–13. https://doi.org/10.1016/j.jalz.2011.10.007 (2012).
Liu, Z. et al. Listening to mental health crisis needs at Scale: Using natural language processing to understand and evaluate a mental health crisis text messaging service. Front. Digit. Health 3, 779091. https://doi.org/10.3389/fdgth.2021.779091 (2021).
Klöppel, S. et al. Applying automated MR-based diagnostic methods to the memory clinic: A prospective study. J. Alzheimers Dis. 47, 939–954. https://doi.org/10.3233/JAD-150334 (2015).
Koenig, A. M., Nobuhara, C. K., Williams, V. J. & Arnold, S. E. Biomarkers in Alzheimer’s, frontotemporal, Lewy body, and vascular dementias. Focus (Am. Psychiatr. Publ.) 16, 164–172. https://doi.org/10.1176/appi.focus.20170048 (2018).
Huang, H. C., Tseng, Y. M., Chen, Y. C., Chen, P. Y. & Chiu, H. Y. Diagnostic accuracy of the Clinical Dementia Rating Scale for detecting mild cognitive impairment and dementia: A bivariate meta-analysis. Int. J. Geriatr. Psychiatry 36, 239–251. https://doi.org/10.1002/gps.5436 (2021).
Saxton, J. et al. Functional and cognitive criteria produce different rates of mild cognitive impairment and conversion to dementia. J. Neurol. Neurosurg. Psychiatry 80, 737–743. https://doi.org/10.1136/jnnp.2008.160705 (2009).
Kane, J. P. M. et al. Clinical prevalence of Lewy body dementia. Alzheimers Res. Ther. 10, 19. https://doi.org/10.1186/s13195-018-0350-6 (2018).
Mak, E., Su, L., Williams, G. B. & O’Brien, J. T. Neuroimaging characteristics of dementia with Lewy bodies. Alzheimers Res. Ther. 6, 18. https://doi.org/10.1186/alzrt248 (2014).
Muñoz-Ruiz, M. A. et al. Structural MRI in frontotemporal dementia: Comparisons between hippocampal volumetry, tensor-based morphometry and voxel-based morphometry. PLoS ONE 7, e52531. https://doi.org/10.1371/journal.pone.0052531 (2012).
Möller, C. et al. More atrophy of deep gray matter structures in frontotemporal dementia compared to Alzheimer’s disease. J. Alzheimers Dis. 44, 635–647. https://doi.org/10.3233/JAD-141230 (2015).
Looi, J. C. et al. Differential putaminal morphology in Huntington’s disease, frontotemporal dementia and Alzheimer’s disease. Aust. N. Z. J. Psychiatry 46, 1145–58. https://doi.org/10.1177/0004867412457224 (2012).
Cousins, D. A. et al. Atrophy of the putamen in dementia with Lewy bodies but not Alzheimer’s disease: An MRI study. Neurology 61, 1191–5. https://doi.org/10.1212/01.wnl.0000091889.20347.30 (2003).
Watson, R., Colloby, S. J., Blamire, A. M. & O’Brien, J. T. Subcortical volume changes in dementia with Lewy bodies and Alzheimer’s disease. A comparison with healthy aging. Int. Psychogeriatr. 28, 529–36. https://doi.org/10.1017/S1041610215001805 (2016).
Vasconcelos, L. G. et al. The thickness of posterior cortical areas is related to executive dysfunction in Alzheimer’s disease. Clinics (Sao Paulo) 69, 28–37. https://doi.org/10.6061/clinics/2014(01)05 (2014).
Sun, X. et al. Destruction of white matter integrity in patients with mild cognitive impairment and Alzheimer disease. J. Investig. Med. 62, 927–33. https://doi.org/10.1097/JIM.0000000000000102 (2014).
Frisoni, G. B. et al. Hippocampal and entorhinal cortex atrophy in frontotemporal dementia and Alzheimer’s disease. Neurology 52, 91–100. https://doi.org/10.1212/wnl.52.1.91 (1999).
Du, A. T. et al. Different regional patterns of cortical thinning in Alzheimer’s disease and frontotemporal dementia. Brain 130, 1159–66. https://doi.org/10.1093/brain/awm016 (2007).
Vonk, J. M. J. et al. Semantic loss marks early Alzheimer’s disease-related neurodegeneration in older adults without dementia. Alzheimers Dement (Amst). 12, e12066. https://doi.org/10.1002/dad2.12066 (2020).
Blanc, F. et al. Cortical thickness in dementia with lewy bodies and Alzheimer’s disease: A comparison of prodromal and dementia stages. PLoS ONE 10, e0127396. https://doi.org/10.1371/journal.pone.0127396 (2015).
Ash, S. et al. Impairments of speech fluency in Lewy body spectrum disorder. Brain Lang. 120, 290–302. https://doi.org/10.1016/j.bandl.2011.09.004 (2012).
Lee, S. H. et al. Tract-based analysis of white matter degeneration in Alzheimer’s disease. Neuroscience 301, 79–89. https://doi.org/10.1016/j.neuroscience.2015.05.049 (2015).
Crespi, C. et al. Diffusion tensor imaging evidence of corticospinal pathway involvement in frontotemporal lobar degeneration. Cortex 125, 1–11. https://doi.org/10.1016/j.cortex.2019.11.022 (2020).
Delli Pizzi, S. et al. Structural Connectivity is differently altered in dementia with Lewy body and Alzheimer’s disease. Front. Aging Neurosci. 7, 208. https://doi.org/10.3389/fnagi.2015.00208 (2015).
Kiuchi, K. et al. White matter changes in dementia with Lewy bodies and Alzheimer’s disease: A tractography-based study. J. Psychiatr. Res. 45, 1095–100. https://doi.org/10.1016/j.jpsychires.2011.01.011 (2011).
Watson, R. et al. Characterizing dementia with Lewy bodies by means of diffusion tensor imaging. Neurology 79, 906–14. https://doi.org/10.1212/WNL.0b013e318266fc51 (2012).
Bozzali, M. et al. Brain tissue damage in dementia with Lewy bodies: An in vivo diffusion tensor MRI study. Brain 128, 1595–604. https://doi.org/10.1093/brain/awh493 (2005).
Zhang, Y. et al. White matter damage in frontotemporal dementia and Alzheimer’s disease measured by diffusion MRI. Brain 132, 2579–2592. https://doi.org/10.1093/brain/awp071 (2009).
Lillo, P. et al. Grey and white matter changes across the amyotrophic lateral sclerosis-frontotemporal dementia continuum. PLoS ONE 7, e43993. https://doi.org/10.1371/journal.pone.0043993 (2012).
Josephs, K. A. et al. Corticospinal tract degeneration associated with TDP-43 type C pathology and semantic dementia. Brain 136, 455–70. https://doi.org/10.1093/brain/aws324 (2013).
Bzdok, D., Krzywinski, M. & Altman, N. Machine learning: A primer. Nat. Methods 14, 1119–1120. https://doi.org/10.1038/nmeth.4526 (2017).
Bzdok, D., Altman, N. & Krzywinski, M. Statistics versus machine learning. Nat. Methods 15, 233–234. https://doi.org/10.1038/nmeth.4642 (2018).
Tanveer, M. et al. Machine learning techniques for the diagnosis of Alzheimer’s disease: A review. ACM Trans. Multimed. Comput. Commun. Appl. 16(1s), 1–35. https://doi.org/10.1145/3344998 (2020).
Koikkalainen, J. et al. Differential diagnosis of neurodegenerative diseases using structural MRI data. Neuroimage Clin. 11, 435–449. https://doi.org/10.1016/j.nicl.2016.02.019 (2016).
Tong, T. et al. Five-class differential diagnostics of neurodegenerative diseases using random undersampling boosting. Neuroimage Clin. 15, 613–624. https://doi.org/10.1016/j.nicl.2017.06.012 (2017).
Jack, C. R. Jr. et al. NIA-AA research framework: Toward a biological definition of Alzheimer’s disease. Alzheimers Dement. 14(535), 562. https://doi.org/10.1016/j.jalz.2018.02.018 (2018).
Garcia-Dias, R. et al. Neuroharmony: A new tool for harmonizing volumetric MRI data from unseen scanners. Neuroimage 220, 117127. https://doi.org/10.1016/j.neuroimage.2020.117127 (2020).
Acknowledgements
PDBP
Data and biospecimens used in preparation of this manuscript were obtained from the Parkinson’s Disease Biomarkers Program (PDBP) Consortium, supported by the National Institute of Neurological Disorders and Stroke at the National Institutes of Health. Investigators include: Roger Albin, Roy Alcalay, Alberto Ascherio, Thomas Beach, Sarah Berman, Bradley Boeve, F. DuBois Bowman, Shu Chen, Alice Chen-Plotkin, William Dauer, Ted Dawson, Paula Desplats, Richard Dewey, Ray Dorsey, Jori Fleisher, Kirk Frey, Douglas Galasko, James Galvin, Dwight German, Lawrence Honig, Xuemei Huang, David Irwin, Kejal Kantarci, Anumantha Kanthasamy, Daniel Kaufer, James Leverenz, Carol Lippa, Irene Litvan, Oscar Lopez, Jian Ma, Lara Mangravite, Karen Marder, Laurie Orzelius, Vladislav Petyuk, Judith Potashkin, Liana Rosenthal, Rachel Saunders-Pullman, Clemens Scherzer, Michael Schwarzschild, Tanya Simuni, Andrew Singleton, David Standaert, Debby Tsuang, David Vaillancourt, David Walt, Andrew West, Cyrus Zabetian, Jing Zhang, and Wenquan Zou. Most of the PDBP Investigators have not participated in reviewing the data analysis or content of the manuscript. Grant Numbers: U01 NS100620; U01 NS100610.
ADNI
Data collection and sharing for this project was funded by the Alzheimer's Disease Neuroimaging Initiative (ADNI) (National Institutes of Health Grant U01 AG024904) and DOD ADNI (Department of Defense award number W81XWH-12-2-0012). ADNI is funded by the National Institute on Aging, the National Institute of Biomedical Imaging and Bioengineering, and through generous contributions from the following: AbbVie, Alzheimer’s Association; Alzheimer’s Drug Discovery Foundation; Araclon Biotech; BioClinica, Inc.; Biogen; Bristol-Myers Squibb Company; CereSpir, Inc.; Cogstate; Eisai Inc.; Elan Pharmaceuticals, Inc.; Eli Lilly and Company; EuroImmun; F. Hoffmann-La Roche Ltd and its affiliated company Genentech, Inc.; Fujirebio; GE Healthcare; IXICO Ltd.; Janssen Alzheimer Immunotherapy Research & Development, LLC.; Johnson & Johnson Pharmaceutical Research & Development LLC.; Lumosity; Lundbeck; Merck & Co., Inc.; Meso Scale Diagnostics, LLC.; NeuroRx Research; Neurotrack Technologies; Novartis Pharmaceuticals Corporation; Pfizer Inc.; Piramal Imaging; Servier; Takeda Pharmaceutical Company; and Transition Therapeutics. The Canadian Institutes of Health Research is providing funds to support ADNI clinical sites in Canada. Private sector contributions are facilitated by the Foundation for the National Institutes of Health (www.fnih.org). The grantee organization is the Northern California Institute for Research and Education, and the study is coordinated by the Alzheimer’s Therapeutic Research Institute at the University of Southern California. ADNI data are disseminated by the Laboratory for Neuro Imaging at the University of Southern California.
FTLDNI
FTLDNI was funded through the National Institute of Aging, and started in 2010. The primary goals of FTLDNI were to identify neuroimaging modalities and methods of analysis for tracking frontotemporal lobar degeneration (FTLD) and to assess the value of imaging versus other biomarkers in diagnostic roles. The Principal Investigator of NIFD was Dr. Howard Rosen, MD at the University of California, San Francisco. The data are the result of collaborative efforts at three sites in North America. For up-to-date information on participation and protocol, please visit http://memory.ucsf.edu/research/studies/nifd. Grant Number: R01 AG032306.
NACC
The NACC database is funded by NIA/NIH Grant U24 AG072122. NACC data are contributed by the NIA-funded ADRCs: P30 AG019610 (PI Eric Reiman, MD), P30 AG013846 (PI Neil Kowall, MD), P50 AG008702 (PI Scott Small, MD), P50 AG025688 (PI Allan Levey, MD, PhD), P50 AG047266 (PI Todd Golde, MD, PhD), P30 AG010133 (PI Andrew Saykin, PsyD), P50 AG005146 (PI Marilyn Albert, PhD), P50 AG005134 (PI Bradley Hyman, MD, PhD), P50 AG016574 (PI Ronald Petersen, MD, PhD), P50 AG005138 (PI Mary Sano, PhD), P30 AG008051 (PI Thomas Wisniewski, MD), P30 AG013854 (PI Robert Vassar, PhD), P30 AG008017 (PI Jeffrey Kaye, MD), P30 AG010161 (PI David Bennett, MD), P50 AG047366 (PI Victor Henderson, MD, MS), P30 AG010129 (PI Charles DeCarli, MD), P50 AG016573 (PI Frank LaFerla, PhD), P50 AG005131 (PI James Brewer, MD, PhD), P50 AG023501 (PI Bruce Miller, MD), P30 AG035982 (PI Russell Swerdlow, MD), P30 AG028383 (PI Linda Van Eldik, PhD), P30 AG053760 (PI Henry Paulson, MD, PhD), P30 AG010124 (PI John Trojanowski, MD, PhD), P50 AG005133 (PI Oscar Lopez, MD), P50 AG005142 (PI Helena Chui, MD), P30 AG012300 (PI Roger Rosenberg, MD), P30 AG049638 (PI Suzanne Craft, PhD), P50 AG005136 (PI Thomas Grabowski, MD), P50 AG033514 (PI Sanjay Asthana, MD, FRCP), P50 AG005681 (PI John Morris, MD), P50 AG047270 (PI Stephen Strittmatter, MD, PhD).
Newcastle
This work was supported by the NIHR Newcastle Biomedical Research Centre (BRC) based at Newcastle upon Tyne Hospitals NHS Foundation Trust and Newcastle University. Individual studies received funding from the Wellcome Trust; Avid Radiopharmaceuticals, Inc., a wholly owned subsidiary of Eli Lilly and Company; Alzheimer's Research UK; and the Newcastle Healthcare Charity.
RIN
The present study was funded by the National funding of Italian Ministry of Health in the framework of the grant RETE IRCCS DI NEUROSCIENZE E NEURORIABILITAZIONE (Imaging Project - RRC-2016-2361095; RRC-2017-2364915; RRC-2018-2365796; RRC-2019-23669119_001, RCR-2022-23682285) and Ministry of Economy and Finance (CCR-2017-23669078). The Italian IRCCS advanced neuroimaging network is constituted by the following centers: IRCCS Istituto Auxologico Italiano (Milan); IRCCS Ospedale pediatrico Bambino Gesù (Rome); Fondazione IRCCS Istituto neurologico “Carlo Besta” (Milan); IRCCS Centro Neurolesi “Bonino Pulejo” (Messina); IRCCS Fondazione Don Carlo Gnocchi Onlus (Milan); IRCCS Istituto Centro San Giovanni di Dio Fatebenefratelli (Brescia); IRCCS Ospedale pediatrico “Giannina Gaslini” (Genoa); IRCCS Istituto Clinico Humanitas (Milan); Istituto di Ricerche Farmacologiche Mario Negri IRCCS (Milan); Istituti Clinici Scientifici Maugeri, IRCCS (Pavia); IRCCS Eugenio Medea (Bosisio Parini); Fondazione IRCCS Istituto Neurologico “Casimiro Mondino” (Pavia); IRCCS NEUROMED – Istituto Neurologico Mediterraneo (Pozzilli); IRCCS Associazione Oasi Maria SS Onlus – Troina (Enna); Fondazione IRCCS Ca’ Granda Ospedale Maggiore Policlinico (Milan); IRCCS Fondazione Ospedale San Camillo (Venice); IRCCS Ospedale San Raffaele (Milan); IRCCS Fondazione Santa Lucia (Rome); IRCCS Istituto di Scienze Neurologiche (Bologna); IRCCS SDN Istituto di ricerca diagnostica e nucleare (Naples); IRCCS Fondazione Stella Maris (Pisa); IRCCS San Martino (Genova); IRCCS Gemelli (Roma).
Author information
Authors and Affiliations
Consortia
Contributions
S.D.: formal analysis, investigation, software, data curation, visualization, validation, writing. C.C.: software, methodology, review—editing. D.A.: software, methodology, review—editing. C.M.: resources, methodology, review—editing. R.R.: methodology, review—editing. A.N.: project administration, review—editing. M.G.B.: project administration, review—editing. F.T.: project administration, review—editing. R.L.: project administration, review—editing. E.D.: project administration, review—editing. B.B.: methodology, review—editing. K.K.: resources, methodology, review—editing. M.F.: resources, methodology, review—editing. J.P.T.: resources, methodology, review—editing. P.T.: resources, methodology, review—editing. A.R.: conceptualization, methodology, resources, formal analysis, data curation, writing, supervision. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this Article was revised: In the original version of this Article the consortia names were incorrectly given as “for the ADNI, Frontotemporal Lobar Degeneration Neuroimaging; NIA Alzheimer’s Disease Centers; and the RIN – Neuroimaging Network”. The correct consortium name is “the RIN – Neuroimaging Network”.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
De Francesco, S., Crema, C., Archetti, D. et al. Differential diagnosis of neurodegenerative dementias with the explainable MRI based machine learning algorithm MUQUBIA. Sci Rep 13, 17355 (2023). https://doi.org/10.1038/s41598-023-43706-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-43706-6
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.