Abstract
Acquisition duration of correlated spectroscopy in vivo can be longer due to a large number of t1 increments along the indirect (F1) dimension. Limited number of t1 increments on the other hand leads to poor spectral resolution along F1. Covariance transformation (CT) instead of Fourier transform along t1 is an alternative way of increasing the resolution of the 2D COSY spectrum. Prospectively undersampled five-dimensional echo-planar correlated spectroscopic imaging (EP-COSI) data from ten malignant patients and ten healthy women were acquired and reconstructed using compressed sensing. The COSY spectrum at each voxel location was then generated using FFT, CT and a variant of CT called Inner Product (IP). Metabolite and lipid ratios were computed with respect to water from unsuppressed one-dimensional spectrum. The effects of t1-ridging artifacts commonly seen with FFT were not observed with CT/IP. Statistically significant differences were observed in the fat cross peaks measured with CT/IP/FFT. Spectral resolution was increased ~ 8.5 times (~ 19.53 Hz in FFT, ~ 2.32 Hz in CT/IP) without affecting the spectral width along F1 was possible with CT/IP. CT and IP enabled substantially increased F1 resolution effectively with significant gain in scan time and reliable measure of unsaturation index as a biomarker for malignant breast cancer.
Similar content being viewed by others
Introduction
MR spectroscopy (MRS) is an efficient biochemical tool for quantifying metabolite and lipid concentrations non-invasively in human breast tissues1,2,3,4,5,6,7,8,9. Altered biochemical concentrations in the malignant breast tissues compared to that of healthy ones have been reported in various breast cancer studies using MRS. In addition to the lipids, water and total choline measures that are commonly reported by one dimensional (1D) in vivo MRS1,2,3,4,5,6,7,8,9, two-dimensional (2D) in vivo MRS has also reported the measures of glycine (Gly), myo-Inositol (mI), saturated and unsaturated lipids and lipid unsaturation as potential biomarkers that can be used to identify malignancy in breast tissues. 2D correlated spectroscopy (COSY) is known to provide better spectral dispersion as compared to 1D, since the J-coupled multiplet resonance peaks are dispersed over two spectral dimensions as opposed to one spectral dimension in 1D MRS. The level of lipid unsaturation as a result, can be measured as the ratio of unsaturated fatty acid cross-peaks arising from J-coupling of olefinic to methylene protons in the 2D COSY spectra.
The 2D spectra were initially recorded from single volume of interest (VOI) using localized correlated spectroscopy (L-COSY) technique10,11,12. It was later extended to MR spectroscopic imaging (MRSI) sequences that record 2D COSY spectra in multiple locations at clinically feasible times, with the help of non-uniform sampling (NUS) and Compressed Sensing (CS) reconstruction13,14. Recently, a five-dimensional (5D) echo-planar correlated spectroscopic imaging (EP-COSI) technique combining 2 spectral and 3 spatial dimensions further increased the coverage area by measuring 2D spectra from multiple voxels within multiple slices, during a single scan session, and therefore helps to better localize the malignant tissues across the breast15,16.
While the multi-voxel spectroscopic imaging covers a larger area of the breast, the spectral resolution along indirect F1 (t1) spectral dimension is generally low due to the scan time limitations. Combined with artifacts due to t1-ridging caused by factors like subject motion and instrumental fluctuations17, this can lead to loss of/corrupted cross peaks, limiting the full potential of the technique. While zero-filling can improve the resolution by interpolation to some extent18, it doesn’t improve the separation between different resonant frequencies. It can also introduce ringing artifacts along the F1 domain and does not mitigate t1-ridging as well.
Recently, covariance NMR19,20 has been applied to J-resolved spectroscopic imaging in vivo for increased F1 spectral resolution without introducing ringing artifacts21. It replaced the second Fourier transformation applied to the t1 dimension with a covariance transformation (CT). The resultant spectrum has a spectral resolution in the indirect dimension equal to that in the direct dimension. Therefore, fewer number of t1 increments were required to extract the spin correlations than what is required in a conventional FFT based spectral analysis. This facilitates reducing the scan time while achieving higher spectral resolution along F122.
Although this approach has been used in various NMR experiments like total correlation spectroscopy (TOCSY) and nuclear Overhauser effect spectroscopy (NOESY)23,24,25, the adaptation to in vivo has been limited. A variant of the covariance NMR spectroscopy called inner-product (IP) NMR spectroscopy, is yet another approach which further improves the covariance NMR by making it robust against changes in the carrier frequency26. In this work, we applied the CT and IP approaches to 5D EP-COSI in-vivo to show its advantages over the conventional FFT based COSY spectrum in terms of both improved spectral resolution and minimal influence of t1-ridging, while exploring the possibility of further acceleration in scan time. It is shown the biomarkers such as unsaturation index (UI) quantified from the COSY cross peaks may be unambiguously determined from a CT/IP spectrum in the presence of t1-ridging.
Materials and methods
Subjects
Ten malignant breast masses (n = 10, mean age 52 [range: 41–71] years; grade-3 (n = 3), grade-2 (n = 4) and grade-1 (n = 3)) and healthy (n = 10, mean age 46 (range:29–60) years) volunteers were recruited. The study was performed in accordance with the Declaration of Helsinki and all the subjects gave consent according to the on-site institutional review board guidelines.
Data acquisition
The 5D EP-COSI data was acquired on a Siemens 3 T Skyra scanner (Siemens Healthineer, Erlangen, Germany) with a dedicated “receive” 24-channel phase-array breast coil and a body “transmit” coil (FOV: 160 × 160 × 120mm3, 1.5 mL voxel volume, TR/TE were 1500/35 ms), running on VE11C software platform. 64 t1 points sampled were used along F115 with a spectral bandwidth (SW) of 1250 Hz and 512 complex t2 points, and a SW of 1190 Hz along F2. A three-pulse sequence27 was employed before the global water suppression. A non-water suppressed scan with one t1 point was acquired for eddy current phase correction and coil combination28. Two spatial and one spectral dimensions (ky,kz,t1) were non-uniformly sampled with an exponentially-weighted sampling density along t1 and gaussian sampling density along the ky-kz plane for an acceleration factor of 8. The total scan time was 28 min and 48 s.
Data reconstruction
The undersampled 5D EP-COSI data was reconstructed using a Group Sparsity (GS)-based compressed sensing (CS) algorithm15,29 to estimate the unacquired samples along the ky-kz-t1 dimensions. The dominant lipid peak around 1.3 ppm was zeroed in the Fourier transform of the non-water suppressed signal to obtain a water-dominant time domain signal for the eddy current phase correction and coil combination. The spectral peak volume integrals were computed as described in16. The quantified proton resonances along the diagonal (F1-F2), and off-diagonal are listed in Table 1.
Covariance and inner-product COSY processing
After the GS-CS reconstruction, a hybrid spectral-spatial data matrix \(\mathbf{D}\)(x, y, z, t2, t1 \()\) was outputted where x, y and z were the Fourier transform of kx, ky and kz dimensions in k-space. After Fourier transforming the direct spectral dimension (t2), we get the mixed time–frequency matrix \(\mathbf{A}\)(F2, t1) for every special location (x = 1, 2, 3…, 16; y = 1, 2, 3…, 16; z = 1, 2, 3…, 8), with a stack of 1D spectra. Fourier transforming the indirect dimension t1 of \(\mathbf{A}\) yielded the conventional Fourier transformed COSY spectrum \(\mathbf{S}\)(F2, F1 ) of size (512 × 64).
A covariance transform was instead obtained from A in the following manner25.
Step 1 Make matrix \(\mathbf{A}\) offset free by subtracting the average 1D spectrum (\({\mathbf{A}}_{\mathrm{avg}}\)) from it.
\(\widetilde{\mathbf{A}} = \mathbf{A}-{\mathbf{A}}_{\mathrm{avg}}\). This 1D spectrum is formed by averaging over the t1 dimension in \(\mathbf{A}\).
Step 2 Apply a singular value decomposition (SVD)30 to the transposed mixed time–frequency matrix \({\widetilde{\mathbf{A}}}^{\mathrm{T}}\)= \(\widetilde{\mathbf{U}}\).\(\widetilde{\mathbf{W}}\).\({\widetilde{\mathbf{V}}}^{\mathrm{T}}\), where \(\widetilde{\mathbf{U}}\) and \(\widetilde{\mathbf{V}}\) are the singular vectors and \(\widetilde{\mathbf{W}}\) is the diagonal matrix with singular values as its diagonal elements.
Step 3 The final covariance transformed spectrum is then calculated as, \({\mathbf{S}}_{{{\mathbf{cov}}}} = \left( {{\tilde{\mathbf{A}}}^{{\text{T}}} .{\tilde{\mathbf{A}}}} \right)^{1/2} = { }{\tilde{\mathbf{U}}}.{\tilde{\mathbf{W}}}.{\tilde{\mathbf{V}}}^{{\text{T}}}\), which gives a high-resolution spectrum of size (512 × 512), with the spectral width along indirect spectral dimension same as that of the direct spectral dimension.
The similarity between covariance and Fourier-transformed spectra is based on the Parseval’s theorem as described in20. It follows that the covariance spectrum \({\mathbf{C}} = {\tilde{\mathbf{A}}}^{{\text{T}}} .{\mathbf{A}}\) correspond to the 2D spectrum squared, i.e., \({\mathbf{S}}^{2}\), provided that the \({\mathbf{A}}_{{{\text{avg}}}}\) in the calculation of covariance vanishes. However, it is shown later that the condition of vanishing average might not hold near the spectral center due to the relatively slow spin precision, as well as when reducing the number of t1 points26. It is shown that this problem can be overcome by discarding the average. i.e., by choosing \({\tilde{\mathbf{A}}} = {\mathbf{A}}\) in step 1. This is called the inner-product covariance transform where the Parseval’s theorem assures the correspondence between a matrix \({\mathbf{I}} = {\mathbf{A}}^{{\text{T}}} .{\mathbf{A}}\) and \({\mathbf{S}}^{2}\), even if the average doesn’t vanish26. High resolution IP covariance matrix is therefore calculated as \({\mathbf{S}}_{{{\mathbf{IP}}}} { } = { }\left( {{\mathbf{A}}^{{\text{T}}} .{\mathbf{A}}} \right)^{1/2} { } = { }{\mathbf{U}}.{\mathbf{W}}.{\mathbf{U}}^{{\text{T}}}\), where \({\mathbf{U}}\), \({\mathbf{V}}\) and \({\mathbf{W}}\) are the singular vectors singular values of \({\mathbf{A}}^{{\text{T}}}\).
Even though high resolution can be achieved by CT while also minimizing t1-ridging, reduced number of t1 points can introduce spurious correlations in the CT/IP spectrum. However, the deterministic nature of the sampling scheme and resonant positions enable us to mask out these spurious correlations while retaining the advantage in resolution31. This may be approached in multiple ways. One option is to determine these spurious correlations based on their intensity, frequencies, and t1 sampling as shown in31. Another option is to use a map of true resonance frequencies obtained via high resolution FFT as a reference to identify spurious correlations. In this work, we used the resonance frequency locations identified from a prior knowledge based synthetic spectra to remove the spurious correlations.
Statistical analysis
Means, standard deviations, and 95% confidence intervals were calculated for each metabolite and lipid in IP, CT and FFT spectra. Analysis of variance procedures including Brown-Forsythe was used to test for the equality of means between the quantified IP, CT and FFT. Games-Howell and Tukey HSD multiple comparisons were used to determine the pair-wise significance among the three different groups based on the homogeneity of variances. Student’s t-test was used to compare the means of metabolite and lipid ratios between malignant and healthy tissues.
Ethical approval and informed consent
The study was approved by the institutional review board of the University of California, Los Angeles. The subjects in the study provided written informed consent.
Results
FFT versus CT versus IP
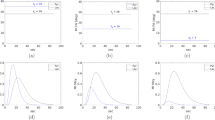
A 2D spectrum from a healthy subject generated using FFT, FFT with zero-filling along t1, covariance transform and inner product are compared in Fig. 1 as intensity plots and contour plots. FFT based spectrum shown in Fig. 1a without zero-filling had a matrix size of 512 × 64 while zero-filled FFT, CT and IP in Fig. 1b–d had matrix size of 512 × 512. Only 0–6 ppm range is shown in the figure. While FFT and zero-filled FFT had a SW of 1250 Hz along F1, both CT and IP had a SW of 1190 Hz. All four spectra had a SW of 1190 Hz along F2. Arrow 1 indicates the ringing caused by zero-padding in zero-filled FFT. Arrow 2 is pointed at the false peaks in FFT and zero-filled FFT which is in fact due to a mild t1-ridging. Arrow 3 shows the region where IP based spectra differed from the CT.
Reconstructed 2D COSY spectrum of healthy tissues from a 60-year-old woman. (a) Intensity and contour plots of FFT based spectra (0–6 ppm). Arrow 2 indicates false peak from t1-ridging. (b) Intensity and contour plots (0–6 ppm) of FFT based spectra after zero-filling t1 dimension to 512 points. Arrow 1 shows the ringing effect. (c) Intensity and contour plots of CT based spectra (0–6 ppm). (d) Intensity and contour plots of IP based spectra (0–6 ppm). Arrow 3 points out the region where IP spectrum is different from CT spectrum. (e) Intensity and contour plots of FFT based spectra (0.5–3.5 ppm). Arrow 1 indicates altered lipid cross-peak from t1-ridging. (f) Intensity and contour plots (0.5–3.5 ppm) of FFT based spectra after zero-filling along the t1 dimension to 512 points. Arrow 2 shows the ringing effect. (g) Intensity and contour plots of CT based spectra (0.5–3.5 ppm). (h) Intensity and contour plots of the IP based spectra (0.5–3.5 ppm).
Panels (e–h) show enlarged regions from figures in panels (a–d) comprising 0.5–3.5 ppm along both axes. Apart from the ringing indicated by arrow 1 in zero-filled FFT, this figure shows the effect of t1-ridging on the cross-peaks between 0.9 and 1.3 ppm, as pointed out by arrow 1 in (e, f) and the lack of this degradation in (g, h).
Another 2D spectrum from a malignant lesion identified in a 45-year-old patient is shown in Fig. 2. Sections (a)–(d) shows the 2D spectra generated using FFT, FFT with zero-filling along t1, CT and IP. T2-weighted MR image with the white box representing the VOI placement is shown in (e). Red square shows the location of the extracted 2D spectrum. Arrows 1 and 2 points out the stronger effect of t1-ridging in (a) and (b) as compared to Fig. 1. Examples of spurious correlation appearing in CT and IP are pointed out by arrows 3 and 4.
Reconstructed 2D COSY spectra from a malignant lesion identified in 45-year-old patient (grade 3 invasive ductal carcinoma, estrogen receptor positive, progesterone receptor positive, her2 positive, ki-67 = 20% and BI-RADS 5): (a) FFT based spectrum (b) FFT based spectrum after zero-filling along t1, (c) CT based spectrum, (d) IP based spectrum, (e) T2-weighted MR image with the white box representing the VOI placement. Red square shows the location of extracted spectrum. Arrows 1 and 2 point out the effect of t1-ridging in (a) and (b). Examples of spurious correlation appearing in CT and IP are pointed out by arrows 3 and 4.
Figure 3 shows the CT and IP 2D spectra of that shown in Fig. 2 after masking out the spurious correlations. Another 2D spectrum from malignant lesion identified in 41-year-old patient is shown in Fig. 4. Sections (a)–(d) shows the spectra generated using FFT, FFT with zero-filling along t1, CT and IP. T2-weighted MR image with the white box representing the VOI placement is shown in (e). Red square shows the location of extracted 2D spectrum. Arrows 1 and 2 show the effect of t1-ridging in (a) and (b).
Reconstructed 2D COSY spectra from malignant lesions identified in 41-year-old patient (grade 3 invasive ductal carcinoma and ductal carcinoma in situ, estrogen receptor positive, progesterone receptor positive, her2 positive, ki-67 = 60% and BI-RADS 5): (a) FFT based spectrum, (b) FFT based spectrum after zero-filling along t1, (c) CT based spectrum, (d) IP based spectrum, (e) T2-weighted MR image with the white box representing the VOI placement. Red square shows the location of extracted spectrum. Arrows 1 and 2 show\ the effect of t1-ridging in (a, b).
Choice of t1 points and acceleration feasibility
Figure 5 shows the effect of different schemes of t1 sampling while using FFT and CT. The 2D spectrum here is the same spectrum as shown in Fig. 1, retrospectively undersampled along t1 to study feasibility of further acceleration. Figure 5a shows the conventional FFT spectrum using the full 64 points along t1 ranging TEs from 35 to 85.4 ms at 800 μs intervals, giving 1250 Hz SW. Choosing either first 32 or last 32 points as shown in (b) and (c) reduces the spectral resolution along F1 by half and shows heavy t1-ridging effects (arrows 1 and 2). Sampling every other t1 points for 2 × acceleration as shown in (d) and (e) on the other hand halves the SW along F1 while retaining the same spectral resolution causing folding artifacts indicated by arrow 3.
Effects of t1 sampling in FFT and CT based spectra. (a) FFT spectrum using 64 points along t1 ranging TEs from 35 to 85.4 ms at 800 μs intervals, giving 1250 Hz SW. (b) FFT spectrum using first 32 t1 points. (c) FFT spectrum using last 32 t1 points. Both (b) and (c) reduce the spectral resolution along t1 by half and show heavy t1-ridging effects (arrows 1 and 2). Sampling only (d) odd t1 points or (e) even t1 points for 2× acceleration halves the SW along t1 while retaining the same spectral resolution causing folding artifacts indicated by arrow 3. (f) CT spectrum using 64 points along t1 ranging TEs from 35 to 85.4 ms at 800 μs intervals, giving 1250 Hz SW. (g) CT spectrum using first 32 t1 points (t1 SW = 1250 Hz). (h) CT spectrum using last 32 t1 points (t1 SW = 1250 Hz). Arrows 1 and 2 in (g) and (h) points out the spurious correlations appearing in the spectrum. (i) Sampling every other t1 points starting from t1 = 1. Effective t1 SW = 1250 Hz. (j) Sampling every other t1 points starting from t1 = 2. Effective t1 SW = 1250 Hz.
Panels (f)–(j) in Fig. 5 show the results of same cases as in panels (a)–(e) when CT is used instead of FFT. Arrows 1 and 2 in (g) and (h) points out the spurious correlations appearing in the spectrum. The results for IP are shown in Supplementary Figure 1 were the panels (a)–(e) correspond to the same cases as in panels (f)–(j) of Fig. 5.
Quantitation comparison
Figure 6 shows bar graphs comparing the mean (95% CI) of different metabolite and lipid ratios with respect to 1D water peak area in both malignant ((a), (c) and (e)) and healthy ((b), (d) and (f)) breasts. Plots in (a) to (d) show the ratios for different cross peaks while (e) and (f) shows the ratios for diagonal peaks in the spectrum. A statistically significant difference between the estimation of ratios from IP, CT and FFT was determined by Brown-Forsythe ANOVA for CP2 (F(2,9.99) = 5.399, p = 0.026) in malignant group, and for CP2 (F(2,9.13) = 6.477, p = 0.018), CP3 (F(2,9.23) = 6.434, p = 0.018), CP4 (F(2,9.02) = 5.569, p = 0.027) and CP5 (F(2,9.05) = 5.012, p = 0.034) in healthy group. However, the Games-Howell post hoc test showed that the difference in the estimation of the ratios between any two methods among IP, CT and FFT were not statistically significant.
Figure 7 shows bar graphs comparing the UI between IP, CT and FFT as well as across healthy and malignant groups. (a)–(b) show the results for malignant and healthy groups compared between IP, CT and FFT when the UI is computed from cross peaks above and below the diagonal. (c)–(d) show UI from cross peaks above and below the diagonal compared between healthy and malignant groups. Statistically significant differences (p < 0.05) were observed between healthy and malignant groups for UI computed from cross peaks both above and below diagonal for IP and CT, and above the diagonal for FFT. The measured values of UI from the cross peaks above and below the diagonal were very close for IP and CT with difference being < 2% for malignant and 13% for healthy, whereas they were much larger for FFT with a 25% difference for malignant and 38% difference for healthy as shown in (a) and (b).
Bar graphs comparing the unsaturation index between IP, CT and FFT as well as across healthy and malignant groups. Comparison between IP, CT and FFT when the unsaturation index is computed from cross peaks above and below the diagonal in (a) malignant group, (b) healthy group. (c) Unsaturation index from cross peaks above the diagonal compared between healthy and malignant groups. (d) Unsaturation index from cross peaks below the diagonal compared between healthy and malignant groups.
Discussion
Due to the scan time limitations, conventional 2D L-COSY spectrum is acquired with limited number of t1 points. Even with non-uniform sampling and CS based reconstruction, 64 t1 points are usually used to achieve reasonable scan time. This results in poor spectral resolution along F1. In this work, prospectively undersampled 5D EP-COSI data were reconstructed using GS-CS and 2D COSY spectra from multiple locations in malignant and healthy breast masses were analyzed using FFT, CT and IP for enhanced F1 spectral resolution, reduced t1-ridging, and acceleration feasibility for faster scan times.
Limitations of FFT based spectrum
One the main requirements of FFT based analysis is that the time increments (∆t1) for t1 points should fulfill Nyquist theorem (∆t1,Nyq = 1/SW) to avoid aliasing artifacts. Doubling the duration of t1 increment for example, results in halving the SW along F1 (625 Hz instead of 1250 Hz) while retaining the same spectral resolution (~ 19.53 Hz) if 32 t1 points are collected for 2 × acceleration. This results in aliasing artifacts along F1 due to the lack of sufficient SW (see Fig. 5d, e).
On the other hand, if SW is retained by keeping ∆t1 = 800 μs while reducing the number of t1 increments to 32, it results in poorer spectral resolution (~ 39.1 Hz) (see Fig. 5b, c). Common approach of zero-padding before FFT achieves interpolation to a larger matrix size, increasing the digital resolution, without actually increasing the spectral resolution. It can also introduce ringing in the spectrum in case of discontinuity in the time domain (see Fig. 1). The ringing can however be minimized by applying appropriate filters before zero-padding.
Advantages of CT and IP
An advantage of CT and IP over traditional 2D FFT is that the indirect dimension of S is not required to be sampled with a time increment ∆t1 that fulfills the Nyquist theorem. Furthermore, if number of t1 point needs to be reduced to accelerate the acquisition, a wider range of t1 evolution times can be used with CT and IP without sacrificing the SW, since the SW along F1 in CT and IP will be equal to that of F2 dimension. It is also reported that probing wider range of t1 evolution times allows better discrimination between true and spurious spin correlations31. Unlike zero-filled FFT, CT and IP facilitate true spectral resolution enhancement along F1 as clearly shown in the results section. Even tough uniform increments along t1 are shown in results section to demonstrate acceleration feasibility of CT and IP in comparison with FFT, non-uniform sampling along t1 is also feasible with both IP and CT. The choice of specific set of increments needs further investigation and is the subject of future work. One approach would be to use the prior knowledge simulations to identify the set of t1 increments that will maximize the sensitivity of metabolites or lipids of low concentrations.
Another advantage of the CT and IP is the lack of t1-ridging artefacts. This results in cross peaks that are much better defined in the ppm range where t1-ridges are present in the FFT spectrum, for example, the lipid cross-peaks near 1.3 ppm along F2 (see Figs. 1, 2, 3, 4). Consequently, the quantitation of these cross-peaks improves substantially. It was observed that there is a larger difference between the ratios of cross peaks on either side of the diagonal in FFT as opposed to CT and IP. This resulted in a larger difference between these cross-peaks on either side of the diagonal with FFT as opposed to CT and IP, despite the symmetric property of the COSY spectra (see Figs. 6c, d, 7). This can cause variation in the measures of UI which is one of the important potential biomarkers available in the COSY spectra depending on whether upper cross peaks or lower cross peaks are used for its computation. Ideally, the ratios should be similar on either side of the diagonal, but the lower resolution along F1, and t1 ridging effects could influence these measures. Earlier studies have reported this measure from the cross peaks either above or below the diagonal or an average of the peaks on either side16,32. Statistically significant difference among IP, CT and FFT were observed in the cross peaks CP2, CP3, CP4 and CP5, especially in healthy controls. In the ratios from malignant tissues, only the difference in CP2 was statistically significant. This may be because of the fact that the healthy breast tissues generally have dominant fat peaks compared to malignant tissues. As a result, the chances of t1 ridging is higher near the aforementioned cross peaks in healthy tissues.
Furthermore, CT and IP based approaches give substantial gain in actual spectral resolution. The CT and IP based spectra gives the same bandwidth along F1 as that of F2. Therefore, these methods gave a spectral resolution of ~ 2.32 Hz along F1, while FFT based spectra had a spectral resolution of ~ 19.53 Hz in the experiments shown in results section. The metabolite peaks between 3 and 4 ppm were therefore much better resolved with these techniques compared to FFT in spectra from malignant tissues (see Figs. 2, 3, 4). This is especially important considering the ability of 5D EP-COSI in vivo detection and quantitation of metabolites like mI and Gly, in addition to measuring lipid-based biomarkers16. Reports from ex vivo breast cancer tissues have also shown the role of these metabolites including mI, Gly and Cho in identifying malignancy5,33. Even when not as resolved as that of CT and IP, the FFT based spectra also showed higher intensities in 3 ppm-4 ppm range in malignant tissues. Hence, the results of quantitation showed that these metabolites were elevated in malignant breast tissues compared to healthy ones with all three methods (see Fig. 6e, f). Since mI and Gly are separated by only 0.006 ppm, it requires < 1 Hz spectral resolution to disentangle them at 3 T. Despite this factor, it can be argued that the combined effect of high spectral resolution (~ 2.32 Hz) and absence of t1-ridging made the identification and analysis of these potential bio markers for breast cancer in CT and IP much less ambiguous as compared to FFT.
Limitations of CT and IP
The main challenge with CT and IP is the possibility of spurious cross-peaks which has intensities above the noise level despite not representing the spin correlations. While this might arise from limited number of t1 points, we have observed that the FFT spectra with heavy t1-ridging also has tendency for stronger spurious correlations in CT and IP. Interestingly, the relationship between the number of t1 increments and spurious correlation appeared to be also dependent on the increment time used (see Fig. 5). However, the deterministic nature of the resonance positions and sampling pattern makes it easy to identify these false cross peaks31. Furthermore, since these spurious correlations generally appeared away from the true resonant positions, it was less problematic compared to effects of t1-ridging in FFT based spectra. It has also been shown that the false cross peaks can be easily identified by performing the correlation experiments with various mixing times since the artifacts should not be affected by the mixing time34. While two fold acceleration is demonstrated in the results, further acceleration introduced more artifacts in the form of spurious correlations. Even though the CT and IP based spectra should be reliable to measure known correlations, assessment of new cross-peaks should therefore need careful analysis to discard the possibility of spurious correlations.
Difference in CT and IP
While both CT and IP based spectra appeared almost identical, a notable difference was near residual water at 4.7 ppm. This is as expected since the theory of inner-product based transformation is designed to be robust against the choice of central frequency. Since our spectra were all centered at water, the frequencies near water undergo limited oscillation with increasing the evolution time which weakens the vanishing mean assumption in CT. IP on the other hand doesn’t have this requirement and hence argued to be more robust to the influence of the central frequency. However, in our application, the residual water was not important due to the water suppressed acquisition. Though the level of water itself in ratio with fat has been reported to be of importance in malignant breast cancers, this is usually calculated form the associated non-water suppressed 1D acquisition for eddy-current correction and coil combination. Therefore, both CT and IP served the purpose in a very identical manner throughout our experiments. However, it may be noted that the choice between the two should also be based on the choice of central frequency in the spectrum.
Conclusion
Reconstruction of in vivo 5D EP-COSI using CT and IP is presented in this work showing enhanced F1-resolution of COSY in comparison with FFT based spectral analysis. With CT and IP, we were able to achieve ~ 2.32 Hz spectral resolution along F1 dimension as compared to ~ 19.53 Hz resolution in FFT based spectrum. The effect of t1-ridging artifacts commonly seen in FFT spectra was not observed in CT or IP. Consequently, both CT and IP showed well defined and symmetrical cross-peaks on either side of the diagonal as evident by the quantified ratios of the lipid cross-peaks above and below the diagonal. Furthermore, CT and IP were found to permit wider range of t1 increments without affecting the actual SW along F1. This is particularly advantageous in spectral analysis of metabolite and lipid biomarkers including unsaturation index for breast cancer while making significant gains in scan time.
Data availability
The datasets generated and/or analyzed during the current study are not publicly available due to ethical and data protection restrictions, but are available from corresponding author on reasonable request and subject to an institutional data sharing agreement.
References
Aboagye, E. O. & Bhujwalla, Z. M. Malignant transformation alters membrane choline phospholipid metabolism of human mammary epithelial cells. Cancer Res. 59(1), 80–84 (1999).
Bolan, P. J. et al. MR spectroscopy of breast cancer for assessing early treatment response: Results from the ACRIN 6657 MRS trial. J. Magn. Reson. Imaging 46(1), 290–302 (2017).
Dorrius, M.D., Pijnappel, R.M., Jansen-van der Weide, M.C., Jansen, L., Kappert, P., Oudkerk, M., et al. Determination of choline concentration in breast lesions: quantitative multivoxel proton MR spectroscopy as a promising noninvasive assessment tool to exclude benign lesions. New diagnostic developments to prevent unnecessary invasive procedures in breast cancer diagnostic work-up. 2011.
Gribbestad, I., Sitter, B., Lundgren, S., Krane, J. & Axelson, D. Metabolite composition in breast tumors examined by proton nuclear magnetic resonance spectroscopy. Anticancer Res. 19(3A), 1737–1746 (1999).
Haukaas, T. H., Euceda, L. R., Giskeødegård, G. F. & Bathen, T. F. Metabolic portraits of breast cancer by HR MAS MR spectroscopy of intact tissue samples. Metabolites 7(2), 18 (2017).
Jagannathan, N., Seenu, V. & Kumar, M. Potential of in vivo proton MR spectroscopy in the assessment of breast lesions without the use of contrast agent. Radiology 223(1), 281–282 (2002).
Roebuck, J. R., Cecil, K. M., Schnall, M. D. & Lenkinski, R. E. Human breast lesions: characterization with proton MR spectroscopy. Radiology 209(1), 269–275 (1998).
Sharma, U., Mehta, A., Seenu, V. & Jagannathan, N. Biochemical characterization of metastatic lymph nodes of breast cancer patients by in vitro 1H magnetic resonance spectroscopy: a pilot study. Magn. Reson. Imaging 22(5), 697–706 (2004).
Thakur, S. B. et al. Quantitative in vivo proton MR spectroscopic assessment of lipid metabolism: Value for breast cancer diagnosis and prognosis. J. Magn. Reson. Imaging 50(1), 239–249 (2019).
Lipnick, S. et al. Combined DCE-MRI and single-voxel 2D MRS for differentiation between benign and malignant breast lesions. NMR Biomed. 23(8), 922–930 (2010).
Thomas, M. A., Binesh, N., Yue, K. & DeBruhl, N. Volume-localized two-dimensional correlated magnetic resonance spectroscopy of human breast cancer. Magn. Reson. Med. 14(2), 181–186 (2001).
Ramadan, S. et al. L-COSY of breast cancer at 3T. Eur. J. Radiol. 81Suppl1, S129-131 (2012).
Candes, E. J., Romberg, J. K. & Tao, T. Stable signal recovery from incomplete and inaccurate measurements. Commun. Pure Appl. Math. 59(8), 1207–1223 (2006).
Lustig, M., Donoho, D. & Pauly, J. M. Sparse MRI: The application of compressed sensing for rapid MR imaging. Magn. Reson. Med. 58(6), 1182–1195 (2007).
Wilson, N. E., Burns, B. L., Iqbal, Z. & Thomas, M. A. Correlated spectroscopic imaging of calf muscle in three spatial dimensions using group sparse reconstruction of undersampled single and multichannel data. Magn. Reson. Med. 74(5), 1199–1208 (2015).
Joy, A. et al. Correlated MR spectroscopic imaging of breast cancer to investigate metabolites and lipids: acceleration and compressed sensing reconstruction. BJR Open 4, 20220009 (2022).
Thomas, M. A., Hattori, N., Umeda, M., Sawada, T. & Naruse, S. Evaluation of two-dimensional L-COSY and JPRESS using a 3 T MRI scanner: from phantoms to human brain in vivo. NMR Biomed. 16(5), 245–251 (2003).
Bartholdi, E. & Ernst, R. Fourier spectroscopy and the causality principle. J. Magn. Reson. (1969) 11(1), 9–19 (1973).
Brüschweiler, R. & Zhang, F. Covariance nuclear magnetic resonance spectroscopy. J. Chem. Phys. 120(11), 5253–5260 (2004).
Brüschweiler, R. Theory of covariance nuclear magnetic resonance spectroscopy. J. Chem. Phys. 121(1), 409–414 (2004).
Iqbal, Z., Verma, G., Kumar, A. & Thomas, M. A. Covariance J-resolved spectroscopy: Theory and application in vivo. NMR Biomed. 30(8), e3732 (2017).
Snyder, D. A. Covariance NMR: Theoretical concerns, practical considerations, contemporary applications and related techniques. Progress Nuclear Magn. Reson. Spectrosc. 122, 1–10 (2021).
Zhang, F. & Brüschweiler, R. Indirect covariance NMR spectroscopy. J. Am. Chem. Soc. 126(41), 13180–13181 (2004).
Zhang, F. & Brüschweiler, R. Spectral deconvolution of chemical mixtures by covariance NMR. Chemphyschem. 5(6), 794–796 (2004).
Trbovic, N., Smirnov, S., Zhang, F. & Brüschweiler, R. Covariance NMR spectroscopy by singular value decomposition. J. Magn. Reson. 171(2), 277–283 (2004).
Takeda, K., Kobayashi, Y., Noda, Y. & Takegoshi, K. Inner-product NMR spectroscopy: A variant of covariance NMR spectroscopy. J. Magn. Reson. 297, 146–151 (2018).
Ogg, R. J., Kingsley, R. & Taylor, J. S. WET, a T1-and B1-insensitive water-suppression method for in vivo localized 1H NMR spectroscopy. J. Magn. Reson. Ser. B 104(1), 1–10 (1994).
Klose, U. In vivo proton spectroscopy in presence of eddy currents. Magn. Reson. Med. 14(1), 26–30 (1990).
Burns, B. L., Wilson, N. E. & Thomas, M. A. Group sparse reconstruction of multi-dimensional spectroscopic imaging in human brain in vivo. Algorithms 7(3), 276–294 (2014).
Stewart, G. W. On the early history of the singular value decomposition. SIAM Rev. 35(4), 551–566 (1993).
Chen, Y., Zhang, F., Bermel, W. & Brüschweiler, R. Enhanced covariance spectroscopy from minimal datasets. J. Am. Chem. Soc. 128(49), 15564–15565 (2006).
Thomas, M. A. et al. Investigation of breast cancer using two-dimensional MRS. NMR Biomed. 22(1), 77–91 (2009).
Beckonert, O., Monnerjahn, J., Bonk, U. & Leibfritz, D. Visualizing metabolic changes in breast-cancer tissue using 1H-NMR spectroscopy and self-organizing maps. NMR Biomed. 16(1), 1–11 (2003).
Masuda, Y. et al. Solid-state NMR analysis of interaction sites of curcumin and 42-residue amyloid β-protein fibrils. Bioorg. Med. Chem. 19(20), 5967–5974 (2011).
Acknowledgements
Authors acknowledge the scientific support of Andres Saucedo M.S., Dr. Zohaib Iqbal, Dr. Manoj Sarma, Dr Melissa Joines and Dr. Stephanie Lee-Felker.
Funding
This work was supported by a CDMRP grant from the US Army Breast Cancer Research Program:# W81XWH-16-1-0524.
Author information
Authors and Affiliations
Contributions
A.J.: Formal analysis, Data curation, Writing-original draft. M.A.T.: Conceptualization, Methodology, Investigation, Writing-review & editing, Project administration.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Joy, A., Thomas, M.A. Enhanced spectral resolution for correlated spectroscopic imaging using inner-product and covariance transform: a pilot analysis of metabolites and lipids in breast cancer in vivo. Sci Rep 13, 16809 (2023). https://doi.org/10.1038/s41598-023-43356-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-43356-8
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.