Abstract
The family Cervidae is the second most diverse in the infraorder Pecora and is characterized by variability in the diploid chromosome numbers among species. X chromosomes in Cervidae evolved through complex chromosomal rearrangements of conserved segments within the chromosome, changes in centromere position, heterochromatic variation, and X-autosomal translocations. The family Cervidae consists of two subfamilies: Cervinae and Capreolinae. Here we build a detailed X chromosome map with 29 cattle bacterial artificial chromosomes of representatives of both subfamilies: reindeer (Rangifer tarandus), gray brocket deer (Mazama gouazoubira), Chinese water deer (Hydropotes inermis) (Capreolinae); black muntjac (Muntiacus crinifrons), tufted deer (Elaphodus cephalophus), sika deer (Cervus nippon) and red deer (Cervus elaphus) (Cervinae). To track chromosomal rearrangements during Cervidae evolution, we summarized new data, and compared them with available X chromosomal maps and chromosome level assemblies of other species. We demonstrate the types of rearrangements that may have underlined the variability of Cervidae X chromosomes. We detected two types of cervine X chromosome—acrocentric and submetacentric. The acrocentric type is found in three independent deer lineages (subfamily Cervinae and in two Capreolinae tribes—Odocoileini and Capreolini). We show that chromosomal rearrangements on the X-chromosome in Cervidae occur at a higher frequency than in the entire Ruminantia lineage: the rate of rearrangements is 2 per 10 million years.
Similar content being viewed by others
Introduction
Order Artiodactyla1 (recent Cetartiodactyla) is a large mammalian order that includes camels, whales, pigs, hippos, and ruminants (the suborder of animals with divided stomachs). The family Cervidae is the second most diverse in the suborder Ruminantia2. The systematic relationships of ruminants remain controversial. In supplementary Fig. 1 ruminants phylogeny is present in according to deep investigation of Cervidae species3. Cervidae includes two subfamilies: Cervinae and Capreolinae3. Representatives of the family are widespread in America and Eurasia and have high economic (farm animals, hunting) and ecosystem values (food source for carnivores, impact on vegetation). For decades the systematic relationships of Cervidae have been a hotly debated topic. Recent studies, based on sequences of the complete mitochondrial genome4 and on all available data on 318 existing and extinct species5, significantly clarified not only Cervidae but also artiodactyl phylogeny. There are also studies integrating molecular, morphological, and bioinformatics approaches3. Previously the family was divided into three subfamilies Cervinae, Capreolinae, and Hydropotinae. Recent phylogenetic data place Hydropotes inermis, a monotypic Hydropotinae species, in the subfamily Capreolinae3. Now, the subfamily Capreolinae is divided into four tribes: Capreolini, Alceini, Odocoileini, Rangiferini, and the subfamily Cervinae into two tribes: Cervini, Muntiacini3. In recent phylogenic research based on whole genome sequencing analysis, the status of the Muntiacini tribe was upgraded to subfamily6.
The accumulated cytogenetic data for the Cervidae family allows us to trace the evolution of karyotypes. Cervidae karyotypes are characterized by diversity in the diploid chromosome number (2n = 6–70)7, 8 and have evolved by tandem and Robertsonian translocations of acrocentric chromosomes9, also involving sex chromosomes. Comparative chromosome painting with whole chromosome painting probes has been employed in several studies10,11,12,13,14,15,16. These studies showed that artiodactyl autosomes evolved through fissions, fusions, and inversions. Recent research has shown the undervalued contribution of intrachromosomal rearrangements and identified evolutionary breakpoint regions not only in Cervidae16 but in ruminant genome evolution16, 17.
In most eutherian orders, only autosomal syntenic segments undergo reshuffling while the X chromosome remains highly conserved, as shown by cross-species chromosome painting18 and G-banding19. Contrarily, the X chromosome in Artiodactyla and Ruminantia demonstrates a high level of evolutionary rearrangements shown by G-banding19 and molecular cytogenetics studies15, 20,21,22,23. X-chromosomal changes in Ruminantia include inversions, centromere shifts, heterochromatic variation, and X-autosomal translocations. Recently the X chromosome evolution in different representatives of artiodactyl species was studied by high-resolution BAC (Bacterial Artificial Chromosomes) mapping15, 20, 22,23,24. Cervidae X chromosomes were investigated previously by band-specific probes25, BACs15, 20, 22, 24, and oligo26 probe localization. A substantial part of the cervid X chromosome can be formed by heterochromatin which can be interspersed or present in blocks in the intercalary or pericentromeric regions. The centromeric heterochromatin is mostly composed of satellite DNAs that are also present in centromeres of autosomes and gonosomes27,28,29. In Cervidae, the increased size of gonosomes caused by intercalary and pericentromeric heterochromatin blocks has been observed in reindeer25, 27, tufted deer10, and Indian muntjac30. In addition, interspecific variation of the X chromosome provides a supplemental source of phylogenetic information in the form of evolutionary rearrangements as cytogenetic markers.
There is substantial data on chromosome-level genome assemblies of Cervidae species26, 31,32,33,34. But not all species with a chromosome-level assembly have an assembled X chromosome26. In the present study, we extend the list of four species studied by detailed X chromosome BAC mapping to include species from both subfamilies and four tribes: Capreolinae—reindeer (Rangifer tarandus, Rangiferini), gray brocket deer (Mazama gouazoubira, Odocoileini), Chinese water deer (Hydropotes inermis, Capreolini); Cervinae—black muntjac (Muntiacus crinifrons, Muntiacini), tufted deer (Elaphodus cephalophus, Muntiacini), sika deer (Cervus nippon, Cervini), and red deer (Cervus elaphus, Cervini). We analyze previously established X chromosome assemblies and compare them with the cattle genome. We reveal chromosomal rearrangements that occur on the X chromosome in the Cervidae family. Moreover, we summarize new and previously established data and describe the fine picture of rearrangements on the cervid X chromosome in an evolutionary context.
Results
BAC mapping of the X chromosome in Cervidae
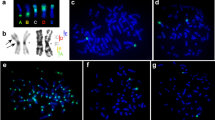
In Cervidae family, an elevated level of morphological variation of the X chromosome was revealed using GTG staining19 and confirmed by BAC-clone hybridization in several cervid species20, 22, 24. To investigate the order of conserved syntenic segments on X chromosomes in the Cervidae family, 29 bovine BAC-clones were localized using FISH (fluorescence in situ hybridization) on X chromosomes of seven species from both subfamilies: black muntjak, tufted deer, sika deer, red deer, reindeer, Chinese water deer, and gray brocket deer in a series of pairwise FISH experiments. An example of the localization of BAC clones on reindeer chromosomes is presented in Fig. 1. All FISH data are presented in supplementary materials (Suppl. 2).
The X chromosomes in Cervidae tend to accumulate the heterochromatin. For visualization of heterochromatin blocks on X-chromosomes of reindeer, Chinese water deer, gray brocket deer, Eurasian elk (Alces alces), Siberian roe deer (Capreolus pygargus), black muntjac, tufted deer, milu deer (Elaphurus davidianus), fallow deer (Dama dama), sika deer and red deer) we performed the Combined Method of Heterogeneous Heterochromatin Detection (CDAG)35 (Fig. 2). In Cervinae species we observe only pericentromeric heterochromatin, whereas in Capreolinae species we detecte interstitial heterochromatic blocks not only on the reindeer X chromosome (published previously25, 27) but on Chinese water deer and gray brocket deer X chromosomes. Remarkably, an autosome to X chromosome translocation was previously identified in tufted deer10. We compared G-banded and CDAG-stained chromosomes from this study and those published previously4. The individual studied here does not have this translocation, which may indicate an intraspecific chromosome polymorphism for this rearrangement36.
Heterochromatin on X chromosome in Cervidae species revealed by chromomycin A3-DAPI after G-banding (CDAG)35 staining (AT-enriched (blue) and GC-enriched (green)): reindeer (RTA), Chinese water deer (HIN), gray brocket deer (MGO), Eurasian elk (AAL), Siberian roe deer (CPY), black muntjac (MCR), Indian muntjac (MMU)30, tufted deer (ECE), milu (EDA), fallow deer (DDA), sika deer (CNI) and red deer (CEL). Centromere positions are designated by a white circle, blocks of interstitial heterochromatin by arrows.
In total, comparative analysis of BACs’ order reveal thee syntenic blocks: X Syntenic Block 1 (15 BACs, XSB1, pink), X Syntenic Block 2 (8 BACs, XSB2, yellow), and X Syntenic Block 3 (6 BACs, XSB3, blue)20. The order of the BACs and syntenic blocks was detected for all investigated species. We identify two types of cervid X, that correspond to the morphology of chromosome—submetacentric and acrocentric (Fig. 3). The submetacentric type is found only in Capreolinae species (Siberian roe deer20, roe deer22, Eurasian elk20, reindeer). The variation of the X chromosome of these species was conditioned by centromere shift and heterochromatin expansion. Whereas the acrocentric type is present in both subfamilies: Chinese water deer, gray brocket deer (Capreolinae), black muntjac, tufted deer, sika deer, red deer, fallow deer20, milu deer22 (Cervinae). We identified an error in BACs order in blue and yellow conservative segments on the milu deer X chromosome in our previous publication20. The analysis of the corrected order with other research22 and new data shows that, milu deer X chromosome represents the typical acrocentric type of cervid X. We also compared BACs’ order from previous research24 of gray brocket deer X chromosome with new data. The variation of the acrocentric type of X is conditioned by autosomal translocation and heterochromatin expansion.
Two types of cervid X (submetacentric and acrocentric) and the order of three syntenic blocks and cattle bacterial artificial chromosome clones (BAC, CHORI-240) on reindeer (RTA X) and Chinese water deer (HIN X) X chromosomes are shown as an example. Heterochromatin blocks in reindeer are shown according to previous research25 and new CDAG data. The order of BACs whose names are in gray is not definitive due to close positioning at the same locus.
Comparative analysis of cervid X chromosome assemblies
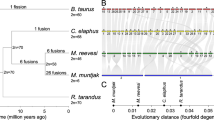
To perform analysis of chromosome-level assemblies of X, we used available material from GenBank NCBI. We performed an alignment of the whole X chromosome assemblies of cattle37 and five deer species: Chinese water deer (Hydropotes inermis)32, Chinese muntjac (Muntiacus reevesi)33, black muntjac (Muntiacus crinifrons)32, red deer (Cervus elaphus)31, Yarkand deer (Cervus hanglu yarkandensis)34 (Fig. 4).
The alignment of the X chromosome in six artiodactyl species: cattle (Bos taurus)37, Chinese water deer (Hydropotes inermis)32, Chinese muntjac (Muntiacus reevesi)33, black muntjac (Muntiacus crinifrons)32, red deer (Cervus elaphus)31, Yarkand deer (Cervus hanglu yarkandensis)34. Cattle bacterial artificial chromosome clones (BAC, CHORI-240) are designated by black dots on the cattle X chromosome. The X chromosome centromeres correspond to the leftmost position for five species. The cattle X chromosome centromere is on the right of BAC 386M8. BACs’ positions in the bovine genome are listed in table (Suppl. 3).
All five assembled cervid X chromosomes belong to the acrocentric type. The analysis of X chromosome alignments reveals several rearrangements that span large genomic areas as well as multiple micro rearrangements (Fig. 4). Some rearrangements include genomic regions corresponding to the BACs from the set mapped here by FISH to the X chromosomes. Major rearrangements that include mapped BACs are a translocation, several inversions and duplications. In particular relative to the cattle X chromosome, there is a translocation of the region encompassing BACs from 93K24 to 316D2 and inversion of the large region that includes BACs 108D16 and 54D24. The presence of these two rearrangements is in concordance with maps constructed by FISH on Chinese water deer and black muntjac X chromosomes (Fig. 3, Suppl. 2). Additionally, in the Chinese muntjac X chromosome we detected prominent inversions between 386M8/108D16 and 29N7/386M8. In Chinese muntjac translocation between 386M8 and 108D16 to pericentromeric region, whereas in black muntjac two inversions occurred between 29N7 and 386M8 BACs’ and in subtelomeric region were identified. The areas of the translocation and two inversions were not covered by the set of BACs. The quality of genome assemblies varies greatly, so the rearrangements identified here by genome comparison are not certain and need validation in further research, for example, by BAC localization.
Some smaller rearrangements are detected by comparative analysis of the alignment. We can see multiple microduplications (blue color) on the Cervidae X chromosome, and also multiple micro inversions (orange) in several species. An in-depth study is required to show whether these micro rearrangements represent real inversions and duplications. Maybe some, or all, are assembly or program artifacts that vary greatly in quality.
Discussion
Artiodactyl sex chromosomes, especially the X, demonstrate high levels of evolutionary rearrangements, including inversions, centromere shift, heterochromatic variation, and X-autosomal translocations19. Recently, a series of investigations of X chromosome evolution in different artiodactyl species, including several Cervidae species, were published employing high-resolution BAC mapping15, 20, 22, 24. Combining new data with previously obtained data15, 20, 22, 24 is possible to trace the path of transformation of the X chromosome in various branches of the Cervidae family. Figure 5 illustrates the transformation of the Cervidae X chromosome.
Evolutionary changes in the structure of the Cervidae X chromosome are depicted on the phylogenetic tree of the family (the tree topology is from5). PAX is the Pecoran ancestral X chromosome20. Major conservative segments of artiodactyl X are shown in pink (XSB 1), yellow (XSB 2), and blue (XSB 3). Centromere positions are designated by a black spot. Arrows show the orientation of the conservative segments. Chromosome changes are shown on the phylogenetic tree near their respective branches: CR—centromere reposition, Inv—inversion, Tr—translocation. The timescale is in million years (MY) of evolution. Tufted deer (ECE), black muntjac (MCR), and reindeer (RTA) X chromosomes are shown with an X-autosomal translocation (green block) or a heterochromatic expansion (gray block). Reindeer (RTA) X chromosome presented in an inverted position (with q arm on top).
The assembled data show that the X chromosome of Cervinae is more conserved than the X chromosome of Capreolinae. Interestingly, the data demonstrate a higher rate of chromosome rearrangements in the Capreolinae X chromosome, while the Cervinae X chromosome is more preserved through species radiation. In the subfamily Cervinae, we identify the acrocentric type of cervid X chromosome with the same order of BACs and pericentromeric heterochromatin. This type of X chromosome evolves from the Pecoran ancestral X chromosome (PAX) by one centromere reposition and one inversion. The PAX was reconstructed according to data obtained by X chromosome BAC mapping in species from all artiodactyl families20. Variation in X chromosome morphology was observed in Muntiacini tribe due to X-autosomal translocations10, 38 and heterochromatin expansion. On the contrary, in the subfamily Capreolinae, we find both types of cervid X chromosome—acrocentric and submetacentric. The European elk (AAL)20, reindeer (RTA), Siberian roe deer (CPY) 20 and roe deer (C. capreolus)22 have the submetacentric type of cervid X. In general, the X chromosome order of BAC corresponds to PAX in the studied capreolins, but only the X chromosome of European elk (AAL)20 has the ancestral centromere position. In two capreol tribes: Capreolini (Chinese water deer, HIN) and Odocoileini (gray mazama, MGO), we find an acrocentric type of cervid X chromosome. An accumulation of interstitial heterochromatin is also described for the three species: reindeer (RTA), Chinese water deer (HIN), and gray mazama (MGO).
In Capreolinae, the inversion between 93K24/54D24 occurred in paraphyletic groups including water deer (HIN, Capreolini) and gray brocket deer (MGO, Odoloiceini), which may indicate the presence of a common ancestor for these two species. Indeed, the relationships within the Capreolinae subfamily remain controversial, with paraphilias between Capreolini and Odocoileini39, 40. However, according to both molecular and morphological data there is no evidence of monophyly for these two species4. The occurrence of this inversion in the relatively distant species presents another case of paraphyly within the Capreolinae family. This type of rearrangement is observed in X chromosomes of the subfamily Cervinae. Differences between the acrocentric type of X chromosome in Capreolinae and Cervinae are shown in the whole chromosome alignment map (Fig. 4). In general, the inversion is between 14O10 and 25P8. Inversions in 93K24/54D24 demonstrate the presence of the previously undiscovered breakpoint, which appears in the Cervidae ancestral X chromosome. The alignment demonstrates several major rearrangements consistent with the BAC maps and several new rearrangements not covered by a set of 29 BACs. Overall, the alignment shows that the BAC’s order is well preserved among all five studied cervid X chromosomes, except for several rearrangements, and that the BAC maps are precise and robust.
For ruminants, evolutionary breakpoints41 or hot spots of karyotype evolution were identified in previous studies17. In addition to the ruminant X chromosome hotspots located between 514O22/316D2, and 108D16/214A3 (108D16/48F6) that were identified in previous studies20, rearrangements also occur between 93K24/54D24, with a break in the blue conserved block. Rearrangements at this point have been identified only for Cervidae species, and it must be assumed that this hotspot is specific to this family. In this region, in the cattle genome, there are transcription factors genes, LOC genes (genes with low homology level) and repeated sequences according to previous data17.
In rodents, intrachromosomal rearrangements of the X chromosome are associated with large clusters of intrachromosomal duplications and/or repeated DNA sequences which are present in ancestral species but have subsequently disappeared during evolution42. We searched for the presence of repeated sequences at cervid X chromosome evolutionary breakpoints and identified genomic coordinates (in Chinese water deer, Chinese muntjac, black muntjac, red deer, and Yarkand deer) of three breakpoints regions (514O22/316D2, 108D16/48F6, and 93K24/54D24) flanking the evolutionary rearrangements (Suppl. 4), and identified repeated sequences in the intervals using RepeatMasker. We identified an increase of LINEs and LTR elements and a decrease of GC-level, SINEs, and DNA element percentages relative to the whole X chromosome only in the breakpoint 108D16/48F6 in all species investigated (Suppl. 5). Further detailed analysis of breakpoint regions is warranted with an additional comparison of the repeatmasked elements in nearby non-breakpoint regions.
Conclusion
High-resolution X chromosome maps of species in the family Cervidae provide unique information about intrachromosomal evolution and rearrangements. The detailed analysis of the 29 cattle BACs across multiple species by FISH mapping, CDAG staining and bioinformatic analysis allowed us to identify major changes in the course of cervid X chromosome evolution. We detected two types of cervine X-chromosome—acrocentric and submetacentric. The acrocentric type is found in three independent deer linages (subfamily Cervinae, and Capreolinae tribes—Odocoileini and Capreolini). The relationships within Capreolinae subfamily remain controversial, with paraphilias between Capreolini and Odocoileini39, 40 tribes. The X chromosome rearrangements identified represent phylogenetic markers that may help to resolve these complicated phylogenetic relationships. According to our data an increase in the rate of X chromosome evolution is observed within the Cervidae family. In ruminants the speed of X chromosome evolution is 1 rearrangement per 15 million years14. In the family Cervidae, we find an average rate, including X-autosome translocations, of 2 rearrangements per 10 million years.
Methods
Compliance with ethical standards
The study was carried out in compliance with the ARRIVE guidelines. All experiments were approved by the Ethics Committee on Animal and Human Research at the IMCB SB RAS, Russia (No. 01/21 from January 26, 2021), following all relevant guidelines and regulations. This article does not contain any experiments on human subjects performed by any of the coauthors.
Species
The research was completed using equipment and materials of the Core Facilities Centre “Cryobank of cell cultures” Institute of Molecular and Cellular Biology SB RAS (Novosibirsk, Russia). Cell cultures of black muntjac (Muntiacus crinifrons), tufted deer (Elaphodus cephalophus) and gray brocket deer (Mazama gouazoubira) were provided by Cambridge Resource Center for Comparative Genomics, Cambridge University, UK. Cell cultures of milu deer (Elaphurus davidianus) and fallow deer (Dama dama) were provided by Laboratory of Genomic Diversity (NCI, Frederick, MD, USA). Cell cultures of sika deer (Cervus nippon), red deer (Cervus elaphus), Eurasian elk (Alces alces), Siberian roe deer (Capreolus pygargus) and reindeer (Rangifer tarandus) were prepared from ear biopsy at Institute of Molecular and Cellular Biology SB RAS (Novosibirsk, Russia).
Chromosome preparation
The procedure for establishing the fibroblast cell line from an ear biopsy was described previously43. Metaphase chromosomes were obtained from fibroblast cell lines. Briefly, cells were incubated at 37 °C in 5% CO2 in medium αMEM (Gibco), supplemented with 15% fetal bovine serum (Gibco), and antibiotics (ampicillin 100 μg/mL, penicillin 100 μg/mL, amphotericin B 2.5 μg/mL). Metaphases were obtained by adding colcemid (0.02 mg/L) and ethidium bromide (1.5 mg/mL) to actively dividing culture for 3–4 h. Hypotonic treatment was performed with 3 mM KCl, 0.7 mM sodium citrate for 20 min at 37 °C and followed by fixation with 3:1 methanol—glacial acetic acid (Carnoy`s) fixative. Metaphase chromosome preparations were made from a suspension of fixed fibroblasts, as described previously44. G-banding on metaphase chromosomes prior to fluorescence in-situ hybridization (FISH) was performed using standard procedure45. Chromomycin A3-DAPI-after G-banding (CDAG) staining procedure was performed as described earlier35.
FISH procedure
The protocol for selection and coordinates of BAC-clones was reported in previous research20. The list of BAC-clones from CHORI-240 library is shown in table (Suppl. 3). BAC clones’ DNA was isolated using the Plasmid DNA Isolation Kit (BioSilica, Novosibirsk, Russia) and amplified with GenomePlex Whole Genome Amplification kit (Sigma-Aldrich Co., St. Louis, MO, USA). Labeling of BAC clone DNA was performed using GenomePlex WGA Reamplification Kit (Sigma-Aldrich Co., St. Louis, MO, USA) by incorporating biotin-16-dUTP or digoxigenin-dUTP (Roche, Basel, Switzerland).
Dual-color FISH experiments were conducted as described by Yang and Graphodatsky46. Trypsin-treated chromosomes were immobilized in 0.5% formaldehyde in PBS followed by formamide denaturing and overnight probe hybridization at 40◦C. Digoxigenin-labeled probes were detected using anti-digoxigenin-CyTM3 (Jackson Immunoresearch), whereas biotin-labeled probes were identified with avidin-FITC (Vector Laboratories) and anti-avidin FITC (Vector Laboratories). Images were captured and processed using VideoTesT 2.0 Image Analysis System and a Baumer Optronics CCD Camera mounted on an Olympus BX53 microscope (Olympus).
Bioinformatic analysis
To perform analysis of chromosome-level assemblies of X, we used available material from GenBank NCBI. Whole X chromosome assemblies were compared with the cattle (NC_037357.1)37 X chromosome using D-GENIES47 resources by aligner Minimap2 v2.24. After alignment we selected six whole X chromosome assemblies: Hydropotes inermis CM035303.1, Muntiacus reevesi CM035268.1, Muntiacus crinifrons CM018500.1, Cervus elaphus OU343077.1, Cervus hanglu yarkandensis CM021225.131,32,33,34 and Bos taurus GK000030.2. After selection we aligned X chromosome assemblies using a whole-genome alignment tool minimap2 2.26-r117548, 49. The search for synteny and structural rearrangements between the X chromosomes was performed using SyRI 1.6.350. For final visualization of intrachromosomal rearrangements plostsr 1.1.051 software was used.
To identify breakpoint coordinates we used aligned in D-GENIES X chromosomes and fixed coordinates. We compared the obtained coordinates with the BAC clones’ coordinates and revealed genome coordinates of intervals (Suppl. 4). Intervals between 514O22/316D2, 108D16/48F6, and 93K24/54D24 were identified. Since only acrocentric chromosomes were used for bioinformatic analysis, instead of the interval 93K24/54D24, which is already broken in these species, the interval 386M2/5424 was used. To identify repeated sequences in a breakpoint interval and in the whole X chromosome of investigated species, RepeatMasker (Dfam 3.2) was used. Visualization of major classes of repeated sequences was performed in Excel (Suppl. 5).
Data availability
All data generated or analyzed during this study are included in this article. In research were used data from GenBank NCBI: Bos taurus (NC_037357.1, GK000030.2), Hydropotes inermis (CM035303.1), Muntiacus reevesi (CM035268.1), Muntiacus crinifrons (CM018500.1), Cervus elaphus (OU343077.1), Cervus hanglu yarkandensis (CM021225). Further enquiries can be directed to the corresponding authors.
References
Prothero, D. R. et al. On the unnecessary and misleading taxon “Cetartiodactyla”. J. Mamm. Evol. 29, 93–97. https://doi.org/10.1007/s10914-021-09572-7 (2022).
Webb, S. D. Evolutionary history of new world Cervidae. In Antelopes, Deer, and Relatives: Fossil Record, Behavioral Ecology, Systematics, and conservation (eds. Vrba, E. & Schaller, G.) 38–64 (Yale University Press, New Haven, London, 2000).
Heckeberg, N. S. The systematics of the Cervidae: A total evidence approach. PeerJ 8, e8114 (2020).
Hassanin, A. et al. Pattern and timing of diversification of Cetartiodactyla (Mammalia, Laurasiatheria), as revealed by a comprehensive analysis of mitochondrial genomes. C. R. Biol. 335, 32–50 (2012).
Zurano, J. P. et al. Cetartiodactyla: Updating a time-calibrated molecular phylogeny. Mol. Phylogenet. Evol. 133, 256–262 (2019).
Chen, L. et al. Large-scale ruminant genome sequencing provides insights into their evolution and distinct traits. Science 364, eaav6202 (2019).
Fontana, F. & Rubini, M. Chromosomal evolution in Cervidae. Biosystems 24, 157–174 (1990).
Duarte, J. M. B. Morphologic and cytogenetic description. Mammalia 67, 403–410 (2003).
Rubes, J. et al. Comparative molecular cytogenetics in Cetartiodactyla. Cytogenet. Genome Res. 137, 194–207 (2012).
Huang, L., Chi, J., Nie, W., Wang, J. & Yang, F. Phylogenomics of several deer species revealed by comparative chromosome painting with Chinese muntjac paints. Genetica 127, 25–33 (2006).
Kulemzina, A. I. et al. Cross-species chromosome painting in Cetartiodactyla: Reconstructing the karyotype evolution in key phylogenetic lineages. Chromosome Res. 17, 419–436 (2009).
Kulemzina, A. I. et al. Chromosome painting in Tragulidae facilitates the reconstruction of Ruminantia ancestral karyotype. Chromosome Res. 19, 531 (2011).
Kulemzina, A. I. et al. Comparative chromosome painting of pronghorn (Antilocapra americana) and saola (Pseudoryx nghetinhensis) karyotypes with human and dromedary camel probes. BMC Genet. 15, 68 (2014).
Kulemzina, A. I. et al. Comparative chromosome map and heterochromatin features of the gray whale karyotype (Cetacea). Cytogenet. Genome Res. 148, 25–34 (2016).
Proskuryakova, A. A. et al. Comparative chromosome mapping of musk Ox and the X chromosome among some bovidae species. Genes 10, 857 (2019).
Proskuryakova, A. A. et al. Comparative studies of karyotypes in the Cervidae family. Cytogenet. Genome Res. 162, 312–322 (2022).
Farré, M. et al. Evolution of gene regulation in ruminants differs between evolutionary breakpoint regions and homologous synteny blocks. Genome Res. 29, 576–589 (2019).
Ferguson-Smith, M. A. History and evolution of cytogenetics. Mol. Cytogenet. 8, 19 (2015).
Graphodatsky, A. S., Perelman, P. L. & O’Brien, S. J. Atlas of mammalian chromosomes (2nd edition). Wiley-Blackwell (2020).
Proskuryakova, A. A. et al. X chromosome evolution in Cetartiodactyla. Genes 8, 216 (2017).
Robinson, T. J., Harrison, W. R., Ponce de Leon, F. A., Davis, S. K. & Elder, F. F. B. A molecular cytogenetic analysis of X chromosome repatterning in the Bovidae: Transpositions, inversions, and phylogenetic inference. Cytogenet. Genome Res. 80, 179–184 (1998).
Frohlich, J. et al. Karyotype relationships among selected deer species and cattle revealed by bovine FISH probes. PLoS ONE 12, e0187559 (2017).
Perucatti, A. et al. Advanced comparative cytogenetic analysis of X chromosomes in river buffalo, cattle, sheep, and human. Chromosome Res. 20, 413–425 (2012).
Bernegossi, A. M. et al. Cytogenetic mapping of cattle BAC probes for the hypothetical ancestral karyotype of the family Cervidae. Cytogenet. Genome Res. 162, 140–147 (2022).
Lee, C. et al. Defining the anatomy of the Rangifer tarandus sex chromosomes. Chromosoma 107, 61–69 (1998).
Poisson, W. et al. Chromosome-level assembly of the Rangifer tarandus genome and validation of cervid and bovid evolution insights. BMC Genom. 24, 1–17 (2023).
Lee, C., Ritchie, D. B. C. & Lin, C. C. A tandemly repetitive, centromeric DNA sequence from the Canadian woodland caribou (Rangifer tarandus caribou): Its conservation and evolution in several deer species. Chromosome Res. 2, 293–306 (1994).
Vozdova, M. et al. Sequence analysis and FISH mapping of four satellite DNA families among Cervidae. Genes 11, 584 (2020).
Vozdova, M. et al. Satellite DNA in neotropical deer species. Genes 12, 123 (2021).
Biltueva, L. S. et al. Chromosomes of the Indian muntjac (Muntiacus muntjak): Comeback. Cell Tissue Biol. 14, 407–412 (2020).
Pemberton, J., Johnston, S. E., Fletcher, T. J., Darwin Tree of Life Consortium. The genome sequence of the red deer, Cervus elaphus Linnaeus 1758. Wellcome Open Res. 6, 336 (2021).
Yin, Y. et al. Molecular mechanisms and topological consequences of drastic chromosomal rearrangements of muntjac deer. Nat. Commun. 12, 6858 (2021).
Mudd, A. B., Bredeson, J. V., Baum, R., Hockemeyer, D. & Rokhsar, D. S. Analysis of muntjac deer genome and chromatin architecture reveals rapid karyotype evolution. Commun. Biol. 3, 1–10 (2020).
Ba, H. et al. Chromosome-level genome assembly of Tarim red deer, Cervus elaphus yarkandensis. Sci. Data 7, 187 (2020).
Lemskaya, N. A. et al. A combined banding method that allows the reliable identification of chromosomes as well as differentiation of AT-and GC-rich heterochromatin. Chromosome Res. 26, 307–315 (2018).
Cao, X., Jiang, H. & Zhang, X. Polymorphic karyotypes and sex chromosomes in the tufted deer (Elaphodus cephalophus): Cytogenetic studies and analyses of sex chromosome-linked genes. Cytogenet. Genome Res. 109, 512–518 (2005).
Zimin, A. V. et al. A whole-genome assembly of the domestic cow, Bos taurus. Genome Biol. 10, 1–10 (2009).
Yang, F., Carter, N. P., Shi, L. & Ferguson-Smith, M. A. A comparative study of karyotypes of muntjacs by chromosome painting. Chromosoma 103, 642–652 (1995).
Heckeberg, N. S., Erpenbeck, D., Wörheide, G. & Rössner, G. E. Systematic relationships of five newly sequenced cervid species. PeerJ 4, e2307 (2016).
Gutiérrez, E. E. et al. A gene-tree test of the traditional taxonomy of American deer: The importance of voucher specimens, geographic data, and dense sampling. ZooKeys 697, 87 (2017).
Larkin, D. M. et al. Breakpoint regions and homologous synteny blocks in chromosomes have different evolutionary histories. Genome Res. 19, 770–777 (2009).
Rubtsov, N. B. et al. Reorganization of the X chromosome in voles of the genus Microtus. Cytogenet. Genome Res. 99, 323–329 (2002).
Beklemisheva, V. R. et al. The ancestral carnivore karyotype as substantiated by comparative chromosome painting of three pinnipeds, the walrus, the Steller sea lion and the Baikal seal (Pinnipedia, Carnivora). PLoS ONE 11, e0147647 (2016).
Yang, F. et al. A complete comparative chromosome map for the dog, red fox, and human and its integration with canine genetic maps. Genomics 62, 189–202 (1999).
Seabright, M. A rapid banding technique for human chromosomes. The Lancet 298, 971–972 (1971).
Yang, F. & Graphodatsky, A. S. Animal probes and ZOO-FISH. In Fluorescence In Situ Hybridization (FISH) — Application Guide (ed. Liehr, T.) 323−346. https://doi.org/10.1007/978-3-540-70581-9_29 (Springer, Berlin, Heidelberg, 2017).
Cabanettes, F. & Klopp, C. D-GENIES: Dot plot large genomes in an interactive, efficient and simple way. PeerJ 6, e4958 (2018).
Li, H. Minimap2: Pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Li, H. New strategies to improve minimap2 alignment accuracy. Bioinformatics 37, 4572–4574 (2021).
Goel, M., Sun, H., Jiao, W.-B. & Schneeberger, K. SyRI: Finding genomic rearrangements and local sequence differences from whole-genome assemblies. Genome Biol. 20, 1–13 (2019).
Goel, M. & Schneeberger, K. plotsr: Visualizing structural similarities and rearrangements between multiple genomes. Bioinformatics 38, 2922–2926 (2022).
Acknowledgements
The work was supported by a research Grant of the Russian Science Foundation (RSF, 19-14-00034-П). We thank Dmitri Mezentsev and Vasilina Belik from Primorsky Safari-Park (https://safaripark25.ru/) for provided sika deer and reindeer ears biopsy. We would like to sincerely thank Dr. Stephen O’Brien (Laboratory of Genomic Diversity), Dr. Mitchell Bush (Conservation and Research Center, National Zoological Park, Virginia, USA), June Bellizzi and director Richard Hahn (Catoctin wildlife Zoo and Preserve, MD, USA) for kindly providing samples. We would like to thank Mary Thompson (NCI-Frederick, USA) for establishing cell lines. We thank Dr. Anna V. Kukekova and Dr. Jennifer Lynn Johnson (Animal Sciences Department, College of ACES, University of Illinois at Urbana-Champaign, USA) for providing bacterial artificial chromosome clones.
Author information
Authors and Affiliations
Contributions
Conceptualization A.A.P. and A.S.G.; FISH data curation A.A.P. and E.S.I.; cell culture preparation A.A.P. and P.L.P.; chromosome preparation P.L.P., A.A.P., E.S.I.; formal analysis A.A.P. and E.S.I.; funding acquisition A.S.G.; investigation A.A.P. and E.S.I.; methodology A.A.P.; bioinformatic analysis A.I.M.; project administration A.S.G.; resources D.M.L., M.A.F., F.Y., O.V.U.; bacterial artificial chromosomes analysis D.M.L.; supervision A.S.G. and A.A.P.; visualization A.A.P. and E.S.I.; writing original draft A.A.P. and E.S.I.; writing – review & editing P.L.P., A.I.M., D.M.L., M.A.F., F.Y., O.V.U., A.S.G.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Proskuryakova, A.A., Ivanova, E.S., Makunin, A.I. et al. Comparative studies of X chromosomes in Cervidae family. Sci Rep 13, 11992 (2023). https://doi.org/10.1038/s41598-023-39088-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-39088-4
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.