Abstract
The assignment of an individual to the true population of origin using a low-panel of discriminant SNP markers is one of the most important applications of genomic data for practical use. The aim of this study was to evaluate the potential of different Artificial Neural Networks (ANNs) approaches consisting Deep Neural Networks (DNN), Garson and Olden methods for feature selection of informative SNP markers from high-throughput genotyping data, that would be able to trace the true breed of unknown samples. The total of 795 animals from 37 breeds, genotyped by using the Illumina SNP 50k Bead chip were used in the current study and principal component analysis (PCA), log-likelihood ratios (LLR) and Neighbor-Joining (NJ) were applied to assess the performance of different assignment methods. The results revealed that the DNN, Garson, and Olden methods are able to assign individuals to true populations with 4270, 4937, and 7999 SNP markers, respectively. The PCA was used to determine how the animals allocated to the groups using all genotyped markers available on 50k Bead chip and the subset of SNP markers identified with different methods. The results indicated that all SNP panels are able to assign individuals into their true breeds. The success percentage of genetic assignment for different methods assessed by different levels of LLR showed that the success rate of 70% in the analysis was obtained by three methods with the number of markers of 110, 208, and 178 tags for DNN, Garson, and Olden methods, respectively. Also the results showed that DNN performed better than other two approaches by achieving 93% accuracy at the most stringent threshold. Finally, the identified SNPs were successfully used in independent out-group breeds consisting 120 individuals from eight breeds and the results indicated that these markers are able to correctly allocate all unknown samples to true population of origin. Furthermore, the NJ tree of allele-sharing distances on the validation dataset showed that the DNN has a high potential for feature selection. In general, the results of this study indicated that the DNN technique represents an efficient strategy for selecting a reduced pool of highly discriminant markers for assigning individuals to the true population of origin.
Similar content being viewed by others
Introduction
DNA probes and sequences are two important indices in gaining a deep understanding of the evolution process, and the amount of DNA sequence data is rapidly increasing1. Single nucleotide polymorphism (SNP) is a new type of marker that includes many important characteristics for evaluating animals2, crops3, and human population structure4. At present, genomic data plays a critical role in a variety of biological contexts due to its numerous advantages. However, the curse of dimensionality (small n and large p) is a major limitation to their ability for practical applications. The lack of complete pedigrees and misidentification of parents affects the accuracy of genetic evaluations, and consequently, the efficiency of breeding programs. Identification of the discriminant SNP(s) process is one of the most appealing opportunities to exploit genomic data, for practical use, including determining the population of origin for unknown individuals2. Many researchers have widely investigated discriminant SNP(s) and genetic diversity5,6,7,8. Researchers can use such SNP markers for developing a cheap customized panel to trace the breeds. Furthermore, the SNP(s) can provide a reliable solution for the traceability of breed-specific branded products9.
In feature selection, researchers seek to identify key variables and eliminate annoying (or noisy) variables10. The same condition is true for biological data11, especially SNP markers. In various areas of breeding, we are always looking for SNP markers with enormous effects. Now, we import the issue to machine learning, especially the neural network approach. In genetics, this process is also known as Tag SNP Selection Problem (TSSP)12.
Mimicking the behavior of the biological brain in the nerve system is the base of Artificial Neural Networks (ANNs), which are the information processing tools13. Researchers have argued the shortcomings of ANN, including the complexity of analysis, computational cost, and time consumption. However, we must mention that ANN’s high prediction accuracy compensates its drawbacks to a great extent. Deep Neural Networks (DNN) have been employed to analyze biological data14,15. They have many applications in feature abstraction and selection16,17. DNNs were able to construct many biological prediction models18, but their power of feature selection had been ignored for individual discrimination.
The ANNs have recently been applied as a powerful statistical modeling technique for many areas of different biological data, especially in the animal sciences19,20. Fernández, et al.21 have indicated that ANNs were suitable to be used in fields of time series data for weekly milk prediction and clustering individuals in goat flocks. Ince and Sofu22 modeled data with ANN for the prediction of the sheep milk yield by using the back-propagation algorithm.
For feature selection (FS) based on ANN, a comparison was made in this study to discriminate among different horse breeds as well as to assign new individuals to their breed. Statistically, in the analysis of GWAS, all SNPs act separately and conduct the research with significant results. The consequence of this analysis obtains the identification of significant SNP markers, but the relationships between them are ignored. While the network approach is more reliable and logical monitoring all SNPs simultaneously leads to better results efficiently.
To obtain the best results, allele dosage has been applied to ANNs, which is a completely unbiased estimation. The Garson (weights) algorithm illustrates behavioral instability in the analysis, which can be considered a weakness23. Unlike most studies, Olden, et al.24 examined the performance of the Garson algorithm in the variable selection on simulated data, and have found that it has the lowest efficiency compared with other studied algorithms. Ibrahim25 showed that the Olden and Garson methods had the weakest results. The results of Fischer26 revealed that the Garson algorithm has a higher degree of stability in modeling non-linear relationships. Additionally, other studies have used the Garson and Olden algorithms, which are only applicable to ANN with a single hidden layer.
To the best of our knowledge, researchers had not investigated the potential of feature selection by ANN approaches for assigning individuals in horse breeds. We have analyzed the ANN’s potential to characterise, whether ANNs can be used as a tool for tackling the curse of dimensionality of SNP(s) data. We attempted to compare the DNN alongside a brief description of Garson and Olden methods to gain the relative importance of variables (SNP markers). While the DNN is a multiple hidden layer ANN, the two mentioned methods are compatible with a single hidden layer. This paper is one of the first studies to determine the discriminant SNP(s) on a large scale by using the sophisticated methods of ANN approaches. We have conducted this study intending to find distinct SNP markers to reduce the dimensions of the SNP panels as well as comparing different variable selection methods such as Garson and Olden through the ANN approach.
Results and discussion
Feature selection: comparison between three approaches
In the current research, we have used the three feature selection (FS) methods namely Olden, Garson, and DNN. Neural networks are commonly referred to as powerful and efficient statistical modeling techniques by various researchers25. Many studies have compared different FS methods26,27,28,29. The selection criteria for the variables in the DNN structure were the absolute value of the first hidden layer connection weights that they assumed as the regression coefficient. According to the DNN procedure, 4270 SNP markers had been selected for the rest of the analysis. The Garson and Olden algorithms led to a selection of 4937 and 7999 SNP markers for further analysis, respectively. The reason for choosing a more significant number of SNP tags for the Olden algorithm is the low transparency of the PCA plot. We must have mentioned that increasing the number of tags did not increase transparency anymore, this could be due to no linear relationship between SNPs number and PCA plot transparency. Moreover, the absolute increase of markers did not include a useful index for improvement unless the marker allele frequencies were different across subpopulations.
After the selection process of SNP markers, all SNP markers were sorted based on the calculated coefficient. The 460 top-rank SNP of each approach was selected, and all sub-SNP sets were compared to each other to find the common markers (Table 1). Table 1 represents the common SNP(s) in the prime 460 SNP markers. It indicates that all three methods had at least a 34% overlap (the average number of common SNPs is 158).
Regarding Table 1, we have found the lowest number of SNP markers between the DNN and Garson approaches. This phenomenon could be owing to the weights of the first layer in the two approaches. We have obtained the most significant number of SNP markers between Garson and Olden. This evidence shows that Garson and Olden had similar mechanisms for feature selection by using NN’s weights in the input-hidden and hidden-output layers. The Spearman correlation for coefficients of common markers indicated a strong relationship between Garson and Olden methods (98.10%). Also, the association obtained between DNN and Garson methods is 43.1%, which is confirmed by the number of common SNP markers.
In general, most of the studies have widely used the Olden and Garson approaches. The results of Olden, et al.24 revealed that the Olden method was the best overall methodology for processing and identifying the variable importance in the neural network, especially when the inputs had a weak or strong correlation with output. Fischer26 compared the Olden and Garson methods and reported that the results obtained by the Garson method are preferable and more stable than those obtained by the Olden method for nonlinear relationships. Findings from his study have shown that ranks obtained by the Garson approach may be more reliable than the Olden method, especially when those ranks are used for modeling nonlinear data such as positive and negative quadratics and interactive data. The results of these studies indicated that the Olden (Connection weights) method had an excellent performance for different assumptions and, Garson (Weights), as the ancestor of the weighted methods, had a various behavior in these studies.
All mentioned studies used the simulated or ecological data in which the maximum input variables were less than 20 variables. At first glance, both Olden and Garson’s algorithms used the input-hidden and hidden-output connection weights for calculating the importance of variables. The linear regression modeling habe been used as a control method on the real datasets for evaluating the input's significance in some studies23,25, and some others have used simulated data where the data have mostly contained the linear24,28 or semi-linear relationship27. However, the DNN approach could raise the performance and efficiency of the artificial neural network in circumstances where a large number of input variables (for example, genomic data of the globally equine breeds) have confronted the system.
Feature selection: a comparison based on PCA analysis
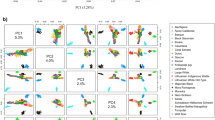
In the first place to assess the degree of divergence among samples, the principal component analysis (PCA) was applied to determine how the animals were allocated to the groups30. The actual coefficients of SNP markers have been obtained step by step according to the original PCA plot, which is according to the numerical analysis in mathematics. In other words, after choosing a new coefficient, the PCA plot was drawn, and the breed distinction was compared with the main PCA plot created by 50K SNP markers panel (Fig. 1). After marker selection and discovering the subsets of markers, PCA analysis was performed using all three sub-SNP(s) and total 50k SNP(s) available on SNP chip (Figures S1 (DNN), S2 (Garson), and S3 (Olden)).
The results indicated an excellent performance of PCA in distinct individuals into separated groups. PCA analysis has identified two subpopulations of Thoroughbred, (TB_UK & TB_US), as one breed, and a similar result was obtained for Standardbred (STBDNor & STBDUS) too. In Fig. 1, some breeds overlapped, but according to the symbols of each breed, we can say that these breeds are properly distinct from each other. Some breeds like Clyd, Shire, Shet, Ice, Mini, and TB (UK-US), were located in corners of the PCA plot, and this fact is due to the geographic boundaries of their countries (Table 5). In other words, these breeds belong to countries that have common borders. As a result, they might have more genetic resource exchanges with each other. Although STBD (including Nor and US) overlapped with Paint and Quarter breeds, they were completely separated by likelihood assessment. Asian breeds (AKTK, ARR, and CSP) were located near the center of the PCA plot and overlapped with Central European Breeds (CEB). It is highlighting this point that Asian breeds have a lot of common characteristics with CEB. The PCA analysis was performed for each method by selected SNP markers (Figs. S1 (DNN), S2 (Garson), and S3 (Olden)). The breed distinction is in good agreement with the main PCA plot created by 50K SNP markers (Fig. 1).
Assessment of different methods and the number of SNP(s) to assignment
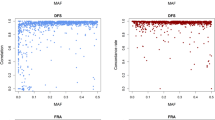
We have estimated the likelihood of assigning 795 individual genotypes to their known origins (or breeds) by the Paetkau, et al.31 approach. Although one particular breed (Shire) had at least one failure assignment by each method. In general, all three feature selection methods assigned most of the individuals to the right population. It resulted in a 9% reduction in the potential of the assignment procedure. Two individuals in the Shire breed failed in all subsets. Red arrows indicate these individuals in Fig. 2.
With the analysis of assignment and concerning values of LLR, obtained results showed that one failure was recognized as Belgian breed by three methods, and the other one was known as different breeds like Paint, Quarter, Swiss warmblood, and Thoroughbred-US. By using three methods, the first individual has 97.30% accuracy to be assigned to the correct race (Shire). By DNN, and Olden approaches, the second individual also had 91.89% accuracy for being appointed into the right breed. For further explanation, these failures might be due to hybrid or crossbreeding parentage. There were two Shire individuals in the center of the PCA plot (Fig. 2); the assignment method indicated that they belong to their breed (Green arrows). In Fig. 3, we have demonstrated the correctness plots for three feature selection algorithms at various strict levels.
The success percentage of genetic assignment for DNN (a), Garson (b), and Olden (c) methods in the three stringency levels. The levels LLR > 2, 3 and 4 show that the individuals 100, 1000 and 10000 times are more likely to be assigned to the right population than the other one. The success rate of 70% in the analysis was obtained by three methods with the number of markers of 110, 208, and 178 tags for DNN, Garson, and Olden methods, respectively.
As shown in Fig. 3, all three methods revealed different behavior for the success percentage of genetic assignment. In the DNN, the success rate in selecting the correct animal breed was more than in the other methods. The sufficient number of SNP markers required to correctly assign an unknown animal to its exact breed/origin at different threshold levels (90%, 95%, and 98%) have been shown for DNN, Garson, and Olden methods in Table 2.
We have accurately calculated the percentages of individuals and correct assignments for different numbers of SNP markers. Testing the performance of each approach has been done at four different levels of LLR analysis. We found that DNN performed better than the other two approaches by achieving 93% accuracy at the most stringent threshold (LLR > 4) (Table 3). In this section, the Garson method did not perform well.
The results revealed that the DNN outperformed other methods with fewer SNP markers. Generally, about 500 discriminant SNP markers enabled us to assign new individuals to the right groups using different ways. There are some issues related to the comparison of results in this study with other ones. First, many previous studies used another type of marker with only a limited number of tags32,33,34,35. Second, there were different methods in several studies36. Maudet, et al.32 found out that, by using 23 microsatellite loci, they could be assigned more than 90% of individuals to their breed. Negrini, et al.37 used the limited set of available SNP markers for an individual assignment. Aiming to determine the range of the minimum number of SNP markers (from 60 to 140), Wilkinson, et al.38 worked for assigning individuals in 17 Bovine breeds.
Model validation
PCA and LLR analysis for validation data
We have used a separate dataset to test the model. Firstly, we have applied the PCA analysis to find the relationship among the breeds like the training dataset (Fig. 4).
In Fig. 4, the Quarter and Warmblood have a small overlap. We identified and extracted the selective SNP markers of 3 feature selection methods (from panel 50K) in the evaluation dataset. Common extracted SNP markers were maintained for later analysis. We have isolated and extracted 839 (Fig. S4—DNN) from 4270 for DNN, 370 (Fig. S5—Garson) from 4940 for Garson, and 1718 (Fig. S6—Olden) from 7999 for Olden approaches in the validation data-set, respectively. Then, we have found the 85 (DNN), 15 (Garson), and 49 (Olden) SNP markers in the evaluation set based on the 460 top-rank SNP markers in the training set, respectively. The LLR analysis was performed for two series of data extracted from the test data and the results have been presented in Table 4.
The results of this section revealed that all three artificial neural networks had an excellent performance. The Garson method with a minimum number of markers (fifteen) had a 60% accuracy, which may be due to the low number of animals and the distinction between the source in the test data, because there are significant differences between the countries of Switzerland, France, and England (the continent of Europe) and the countries of the Middle East and the Americas (Asian and American continents).
By using one dataset, there is a possibility to observe a negligible amount of kinship relationships. Because all individuals are sampled from one herd, kinship relationships are practically inevitable in the research. Therefore, using new data from other sources reduces the probability of kinship among individuals. If unknown or novel information is introduced to the desired network, the least errors will get. Previously obtained results of the network were reliable enough for DNN to infer the right class of novel information precisely. In this case (DNN), the system undoubtedly possesses much power and much success in correctly determining the essential features.
Neighbor-Joining tree of allele-sharing distances for validation data
For a better understanding, we have used the Neighbor-Joining tree of allele-sharing distances on the validation dataset. Neighbor-Joining analysis performs better than PCA analysis on topics such as breed-level differentiation, the intermingling of breeds, outliers, genetic isolation, etc. First, we have analyzed whole genomic data (32419 SNP markers, 120 horses) to show the breed-level differentiation in validation data (Fig. S7).
Then, the Neighbor-Joining analysis was done for each obtained dataset (Fig. S8 (DNN), S9 (Garson), S10 (Olden)) to demonstrate the breed distinction in comparison to the whole data. In Fig. S7, except for two groups (Quarter Horse and Warmblood) and despite the low amount of SNP markers, the rest of the breeds were in their real groups. It is critical to consider that two breeds (Quarter Horse and Warmblood), may have an unusual overlap due to the low number of markers.
We have drawn Fig. S8 by using the markers selected by the DNN. It is noteworthy that the classification of individuals is mostly successful, and there is no significant overlap between breeds. The Neighbor-Joining plot (Fig. S9) drawn by the selected markers of the Garson method did not have a good quality in terms of the classification of individuals. In Fig. S9, there was a great deal of unusual overlap between the breeds, and only the Thoroughbred was identified as a pure breed due to the small number of individuals. The number of outsiders in the results of this dataset was very high (red arrows).
The Olden method had the same performance similar to the DNN and whole data (Fig. S10). In a way, its plot was promising. Perhaps the only disadvantage of the Olden method compared to the other two is that despite the high number of SNP markers, two individuals (Arabian-3 and QuarterHorse-1) still have been identified as outsiders.
Conclusion
We have used the weights of the first hidden layer of the DNN, for selecting and ranking variables (SNPs). Artificial neural networks (ANNs) will receive a great deal of attention in the various scientific fields, given that they are powerful statistical modeling techniques. However, in an attempt to provide useful insights into the contributions of the input (independent) variables in the prediction process, they have been labeled as the “black box” technique. As mentioned earlier, many published studies had been conducted to clarify the interpretation of the connection between the neurons in ANN.
By comparing the results, the Garson and Olden procedures only work with a single hidden layer and single output unit, while multiple layer networks (DNN) do not suffer these limitations. Regarding log-likelihood ratio (LLR) for the individual assignment, the obtained results by this research revealed that ANN’s feature selection methods could be used for genomic data, especially for dimension reduction by DNNs. This finding solves the most critical issue for genetics researchers in dealing with the considerable dimension of data. Researchers can use DNN in the field of animal sciences because of the high performance of breed discriminants. Researchers in the field of genetics and breeding are seeking to reduce the number of biomarkers to find a link between the observed phenotype and these markers.
The result of this study showed that the DNN has a high potential for feature selection in genomic data along with more flexibility in the application of ANNs in the field of animal sciences. Results also showed that using the connection weight of the first hidden layer in a DN Network provides the possibility to reach a high optimum level of accuracy for ranking and selecting the variables (SNP(s)). Another conclusion of this research is that the most critical weights for output values of every variable in a DN Network are the weights in the first hidden layer because all connected loads of the next layers are functions of the first layer's connected load. If three analyzes of PCA, LLR, and Neighbor-Joining achieve the desirable results, we will get the real discriminative features.
It is necessary to point out that the results of this study shed some lights on the using of DN Networks (especially pattern recognition) in genetics and breeding. Feature selection in the genetic field particularly on SNP markers is in the infancy period. The computation time will be reduced significantly. It should also be noted that the DNN network is increasing computing time but it was decreasing the error rate significantly. It can open a new opportunity to extend human insights.
Finally, we think that this will be a fruitful approach to the study of existing domestic populations, such as inferior local breeds and strains in developing countries. In general, the present paper highlighted the importance of variable selection from the varying point of view, including the socio-economic perspective (for developing a low-cost customized assay for assigning the breeds or tracing the origin of animal products derived from diverse species).
Materials and methods
The data for training ANN
A total of 795 animals from 37 breeds of horse populations were genotyped by using the Illumina SNP 50k Bead chip (Illumina, San Diego, CA, USA). Petersen et al.7 have already described the comprehensive description and necessary details of data mining. In summary, Table 5 has given the breed names, the ID of breeds, the geographic origin, minor allele frequency (MAF), Heterozygosity, and the number of animals. Genotype data are coded as the number of reference SNP allele carries, that is, 0 (for AA), 1 (for AB), and 2 (for BB). In the present study, a further filtration for the call rate (the proportion of SNP genotypes) less than 99% was used to discard the missing genotypes39,40.
Moreover, raw predictor variable data (SNP matrix) is used as the input variable in ANN. It is assumed that each of these markers represents a mathematical variable that can only hold 3 inputs (0, 1, and 2).
The data for testing and validation methods
To assess the performance of the ANN methods, learning and evaluation were performed using two separate datasets, respectively. The testing dataset contains 120 individuals from eight breeds (Table 6 includes the sample information). You can find all the details and information about the validation data in the article by Schaefer, et al.41. Data preprocessing included extracting common SNP markers between panels of 50K and 2M. This process resulted in the identification of 32K markers, and 14K of these markers remained after quality control (call rate 99%) for further analysis.
ANN model and construction
Artificial neural networks represent complex structures that are generated by fundamental units (elements) called neurons22. Neurons and their connections create a specific network architecture such as multilayer perceptron (MLP), self-organizing map (SOM), etc.13. In terms of genomic data analysis, we used two types of ANN architecture. The first one is a feed-forward multilayer perceptron (DNN) with two hidden layers, and the second one is a standard single hidden layer (ANN) with a back-propagation algorithm for the weight adjustments42,43. In Figure 5, The architecture of a single hidden layer ANN has been shown for better understanding. Neural net44 and Neural Net Tools45 packages were applied by R software (version 3.4.0)46 to select informative and unique SNP markers that are within each breed. The mentioned algorithms (Garson and Olden) have been utilized by ANN to detect the relative importance of variables for the breed diversity characterization.
The large dimension of the SNP-panel leads to a stack overflow error in the computing process. De Oña and Garrido29 have proposed the usage of a set of neural networks instead of a single one. In contrast to29 in the present work, the high-density SNP chip was partitioned into the sub-datasets with the same dimension and were used as input to identify the discriminant SNP(s).
Feature selection: Garson and Olden
Weights (Garson approach), had been described by Garson47 and has also been modified by Goh48. It was used to identify the relative importance of input variables by the calculated weights within connections in a supervised neural network. The Garson approach indicates relative importance values as the absolute magnitude ranging from zero to one (0-1). Olden and Jackson49 had proposed connection weights, also known as the Olden approach that has been used in this research.
Feature selection: DNN approach and its architecture
For the DNN approach, the ANN with two hidden layers was used to identify the discriminant SNP(s) within breeds. Many combinations exist for selecting the number of nodes in the hidden layer50. The optimal number of nodes in the first and second hidden layers detected 40 and 38 nodes after testing a range of combinations. Finally, ANN with Garson and Olden algorithms contained 40 nodes in the hidden layer.
We have used the final fitted weights of the neural network for selecting the genetic markers. In the DNN approach, we assumed there was a linear relationship between the variable and the response12. We considered the SNP markers to retain a direct relationship with the horse breeds. (Eq. 1).
where Y is the matrix of observed values for the desired breeds, g is a vector of weights of SNP markers, and e is the vector of residual terms. X is known as the design matrix that relates the elements of g to its corresponding element in Y. Assuming that higher coefficient values in this (regression) equation have a significant effect on the output variable, the absolute maximum weight obtained by DNN led to the selection of SNP markers that caused the diversity of the breeds.

Figure 6 shows the whole analysis process. The researchers must determine the features according to Eq. (2), after the convergence of the neural network (Fig. 6). Feature selection is based on the absolute value of the weights of the first hidden layer. It should be noted that 40 weights have been calculated for each variable. In this step, the maximum value is obtained for each variable. If the obtained value was greater than the coefficient of Eq. (2), then that variable was selected as the effective SNP marker.
By considering Eq. (2), it is assumed that all variables are doing their job with maximum potential. Then, a selection threshold was defined to choose a small set of variables. As previously described, in this status, the effects of all variables are not estimated equally and we see the minimum and maximum values among them. The reason for assuming maximum potential is that we do not know what is the actual effect of each variable in biological data. Therefore, we considered every marker on the same level and allowed them to make their inferences and results. Regarding stages Turn 1 and Turn 2, it can be explained that sometimes the result of feature selection in subsequent analyzes is not desirable. Finally, further analysis to evaluate the individual assignment accuracy and qualify all three sub-SNP sets was done by a manual script in R software version (3.4.0).
Individual assignment analysis
There are several available approaches for genetic assignment31,51,52. The method of Paetkau, et al.31 has been used for the assignment analysis (as had been described by38), and it had high effectiveness on individual assignment when high levels of genetic differentiation between reference populations existed52. It is noteworthy that the SNP markers were applied instead of the microsatellites. We have calculated the log-likelihood ratios (LLR) to accurately assess the performance of the assignment procedure. The log-likelihood ratios (LLR) will be calculated by comparing the probability of an individual assigned to its real population to the probability of it assigned to another population (Eqs. 3 and 4).
where,
Different stringency thresholds are applied as confidence levels of assignment precision. Four stringency levels were used: LLR > 1, 2, 3 & 4, which means a multi-locus genotype should be 10, 100, 1000 & 10000 times more similar to the true population rather than the other one. If a calculated LLR value was lower than the selected stringency levels, the individual genotype would fail to assign to its unique origin. In other words, it would assign to the pseudo reference population. The correct assignment of an individual genotype to its known origin occurred when the calculated LLR was greater than the selected stringency levels.
The aim of evaluating a classification model is to evaluate and understand its flexibility, behavior, and prediction ability in dealing with new or unknown samples.
Ethics statement
Training Data-set: DNA sampling was limited to the collection of blood by jugular venipuncture performed by a licensed veterinarian or from hairs pulled from the mane or tail by the horse owner or researcher. All animal work was conducted in accordance with and approval from the international and national governing bodies at the institutions in which samples were collected (the University of Minnesota Institutional Animal Care and Use Committee (IACUC); the University of Kentucky IACUC; the University College Dublin, Animal Research Ethics Committee; Swiss Law on Animal Protection and Welfare; the Ethical Board of the University of Helsinki; the Animal Health Trust Clinical Research Ethics Committee; Norwegian Animal Research Authority; UK Home Office License; and the Lower Saxon state veterinary office).
Testing Data-set: DNA samples were previously collected with approval from the Animal Care and Use Committees at the respective institutions. All animal work was performed in accordance and with approval from international and national governing bodies at the institutions where the samples were collected (University of Minnesota Institutional Animal Care and Use Committee (IACUC); University of California, Davis Institutional Animal Care and Use Committee (protocol #17491); University of Kentucky Institutional Animal Care and Use Committee (IACUC); Ethics Committee for Animal Experiments in Uppsala, Sweden (Number C121/14); Institutional animal care and use committee at Cornell University (protocol 2008-0121); University of California, Davis IACUC 19205; Hebrew University’s approval number AG-23476-07; Institutional Animal Care and Use Committee (IACUC), the Lower Saxony state veterinary office- registration number 11A 160/7221.3-2.1-015/11, 8.84-02.05.20.12.066; University of Sydney Animal Ethics Committee: AEC APPROVAL NUMBER: N00/9-2009/3/5109; permit no. BE75/16, veterinary service of the Canton of Bern; Institutional ethics committee of the University of Veterinary Medicine Vienna Good Scientific Practice guidelines and national legislation; Italian Ministry of Agricultural, Food and Forestry Policies (Mipaaf); Ethical Committee of the Canton of Bern (BE33/07, BE58/10 and BE10/13)) No commercial animals were used in this study. Written informed client consent describing the purpose and duration of the study, procedures, potential risks and benefits and containing study contact information were obtained from private owners.
Data availability
Training Data-set: All SNP genotype data are available at the NAGPR Community Data Repository (animalgenome.org) for the purpose of reconstructing the analyses. The only exception is the data collected from the Tennessee Walking Horse, which, under agreement from the granting agency (to the University of Minnesota from the Foundation for the Advancement of the Tennessee Walking Show Horse (FAST) and the Tennessee Walking Horse Foundation (TWHF)), is only available under a Material Transfer Agreement (MTA) between interested individuals and the University of Minnesota. Testing Data-set: Whole genome sequences are available in the following NCBI BioProjects: PRJEB14779, PRJNA273402, and PRJEB10098. Additional sequences are restricted in availability due to pre-existing material transfer agreements and can be requested by contacting the contributing investigator in Additional file 1: Table S1. Genotypes for horses on the MNec2M array will be released upon publication. Genome positions for all 23 million discovered SNPs have been submitted to dbSNP as well as the European Variation Archive.
References
Heather, J. M. & Chain, B. The sequence of sequencers: The history of sequencing DNA. Genomics 107, 1–8. https://doi.org/10.1016/j.ygeno.2015.11.003 (2016).
Dimauro, C. et al. Selection of discriminant SNP markers for breed and geographic assignment of Italian sheep. Small Ruminant Res. 128, 27–33. https://doi.org/10.1016/j.smallrumres.2015.05.001 (2015).
Ganal, M. W., Altmann, T. & Röder, M. S. SNP identification in crop plants. Curr. Opin. Plant Biol. 12, 211–217. https://doi.org/10.1016/j.pbi.2008.12.009 (2009).
Paschou, P. et al. PCA-Correlated SNPs for Structure Identification in Worldwide Human Populations. PLoS Genetics 3, e160. https://doi.org/10.1371/journal.pgen.0030160 (2007).
Gautier, M. et al. A whole genome Bayesian scan for adaptive genetic divergence in West African cattle. BMC Genom. 10, 550. https://doi.org/10.1186/1471-2164-10-550 (2009).
Dimauro, C. et al. Use of the canonical discriminant analysis to select SNP markers for bovine breed assignment and traceability purposes. Anim. Genet. 44, 377–382. https://doi.org/10.1111/age.12021 (2013).
Petersen, J. L. et al. Genetic Diversity in the modern horse illustrated from genome-wide SNP data. PLoS ONE 8, e54997. https://doi.org/10.1371/journal.pone.0054997 (2013).
Boutorh, A. & Guessoum, A. Complex diseases SNP selection and classification by hybrid association rule mining and artificial neural network-based evolutionary algorithms. Eng. Appl. Artif. Intell. 51, 58–70. https://doi.org/10.1016/j.engappai.2016.01.004 (2016).
Lewis, J. et al. Tracing cattle breeds with principal components analysis ancestry informative SNPs. PLoS ONE 6, e18007. https://doi.org/10.1371/journal.pone.0018007 (2011).
Meenachi, L. & Ramakrishnan, S. Metaheuristic search based feature selection methods for classification of cancer. Pattern Recogn. 119, 108079. https://doi.org/10.1016/j.patcog.2021.108079 (2021).
Paul, D., Saha, S. & Mathew, J. Fusion of evolvable genome structure and multi-objective optimization for subspace clustering. Pattern Recogn. 95, 58–71. https://doi.org/10.1016/j.patcog.2019.05.033 (2019).
He, J. & Zelikovsky, A. In The 26th Annual International Conference of the IEEE Engineering in Medicine and Biology Society 2840–2843 (IEEE).
Arbib, M. A. The Handbook of Brain Theory and Neural Networks (MIT press, 2003).
Dean, J. et al. Large scale distributed deep networks. Advances in Neural Information Processing Systems 25 (2012).
Min, S., Lee, B. & Yoon, S. Deep learning in bioinformatics. Brief. Bioinform. 18, 851–869 (2017).
Li, Y. et al. DEEPre: Sequence-based enzyme EC number prediction by deep learning. Bioinformatics 34, 760–769 (2018).
Luo, F., Wang, M., Liu, Y., Zhao, X.-M. & Li, A. DeepPhos: Prediction of protein phosphorylation sites with deep learning. Bioinformatics 35, 2766–2773. https://doi.org/10.1093/bioinformatics/bty1051 (2019).
Di Lena, P., Nagata, K. & Baldi, P. Deep architectures for protein contact map prediction. Bioinformatics 28, 2449–2457 (2012).
Sanzogni, L. & Kerr, D. Milk production estimates using feed forward artificial neural networks. Comput. Electron. Agric. 32, 21–30. https://doi.org/10.1016/S0168-1699(01)00151-X (2001).
Torres, M., Hervás, C. & Amador, F. Approximating the sheep milk production curve through the use of artificial neural networks and genetic algorithms. Comput. Oper. Res. 32, 2653–2670. https://doi.org/10.1016/j.cor.2004.06.025 (2005).
Fernández, C., Soria, E., Martin, J. & Serrano, A. J. Neural networks for animal science applications: Two case studies. Expert Syst. Appl. 31, 444–450 (2006).
Ince, D. & Sofu, A. Estimation of lactation milk yield of Awassi sheep with artificial neural network modeling. Small Ruminant Res. 113, 15–19 (2013).
Gevrey, M., Dimopoulos, I. & Lek, S. Review and comparison of methods to study the contribution of variables in artificial neural network models. Ecol. Model. 160, 249–264. https://doi.org/10.1016/S0304-3800(02)00257-0 (2003).
Olden, J. D., Joy, M. K. & Death, R. G. An accurate comparison of methods for quantifying variable importance in artificial neural networks using simulated data. Ecol. Model. 178, 389–397 (2004).
Ibrahim, O. A comparison of methods for assessing the relative importance of input variables in artificial neural networks. J. Appl. Sci. Res. 9, 5692–5700 (2013).
Fischer, A. How to determine the unique contributions of input-variables to the nonlinear regression function of a multilayer perceptron. Ecol. Model. 309, 60–63. https://doi.org/10.1016/j.ecolmodel.2015.04.015 (2015).
Kemp, S. J., Zaradic, P. & Hansen, F. An approach for determining relative input parameter importance and significance in artificial neural networks. Ecol. Model. 204, 326–334 (2007).
Paliwal, M. & Kumar, U. A. Assessing the contribution of variables in feed forward neural network. Appl. Soft Comput. 11, 3690–3696 (2011).
De Oña, J. & Garrido, C. Extracting the contribution of independent variables in neural network models: A new approach to handle instability. Neural Comput. Appl. 25, 859–869. https://doi.org/10.1007/s00521-014-1573-5 (2014).
Ringnér, M. What is principal component analysis?. Nat. Biotechnol. 26, 303. https://doi.org/10.1038/nbt0308-303 (2008).
Paetkau, D., Calvert, W., Stirling, I. & Strobeck, C. Microsatellite analysis of population structure in Canadian polar bears. Mol. Ecol. 4, 347–354 (1995).
Maudet, C., Luikart, G. & tarberlet, P. Genetic diversity and assignment tests among seven French cattle breeds based on microsatellite DNA analysis. J. Anim. Sci. 80, 942–950 (2002).
Ciampolini, R. et al. Statistical analysis of individual assignment tests among four cattle breeds using fifteen STR loci. J. Annim. Sci. 84, 11–19 (2006).
Negrini, R. et al. Differentiation of European cattle by AFLP fingerprinting. Anim. Genet. 38, 60–66. https://doi.org/10.1111/j.1365-2052.2007.01554.x (2007).
Negrini, R. et al. Breed assignment of Italian cattle using biallelic AFLP® markers. Anim. Genet. 38, 147–153. https://doi.org/10.1111/j.1365-2052.2007.01573.x (2007).
McKay, S. D. et al. An assessment of population structure in eight breeds of cattle using a whole genome SNP panel. BMC Genet. 9, 37. https://doi.org/10.1186/1471-2156-9-37 (2008).
Negrini, R. et al. Assessing SNP markers for assigning individuals to cattle populations. Anim. Genet. 40, 18–26 (2009).
Wilkinson, S. et al. Evaluation of approaches for identifying population informative markers from high density SNP Chips. BMC Genet. 12, 45. https://doi.org/10.1186/1471-2156-12-45 (2011).
Milne, L. In AI-Conference 571–571 (World Scientific Publishing).
Li, B. et al. Genomic prediction of breeding values using a subset of SNPs identified by three machine learning methods. Front. Genet. 9, 237 (2018).
Schaefer, R. J. et al. Developing a 670k genotyping array to tag ~2M SNPs across 24 horse breeds. BMC Genom. 18, 565. https://doi.org/10.1186/s12864-017-3943-8 (2017).
Rumelhart, D. E., Hinton, G. E. & Williams, R. J. Learning representation by back-propagation errors. Nature https://doi.org/10.1038/323533a0 (1986).
Cilimkovic, M. Neural networks and back propagation algorithm. Institute of Technology Blanchardstown, Blanchardstown Road North Dublin 15 (2015).
Stefan Fritsch & Guenther, F. neuralnet: Training of Neural Networks. https://journal.r-project.org/archive/2010/RJ-2010-006/index.html (2016).
Beck, M. NeuralNetTools: Visualization and Analysis Tools for Neural Networks. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6262849/ (2016).
R. Core, T. R: A Language and Environment for Statistical Computing. https://www.R-project.org/ (2017).
Garson, G. D. Interpreting neural-network connection weights. AI Expert 6, 46–51 (1991).
Goh, A. T. C. Back-propagation neural networks for modeling complex systems. Artif. Intell. Eng. 9, 143–151. https://doi.org/10.1016/0954-1810(94)00011-S (1995).
Olden, J. D. & Jackson, D. A. Illuminating the “black box”: A randomization approach for understanding variable contributions in artificial neural networks. Ecol. Model. 154, 135–150 (2002).
Sheela, K. G. & Deepa, S. N. Review on methods to fix number of hidden neurons in neural networks. Math. Probl. Eng. 2013, 11. https://doi.org/10.1155/2013/425740 (2013).
Rannala, B. & Mountain, J. L. Detecting immigration by using multilocus genotypes. Proc. Natl. Acad. Sci. USA 94, 9197–9201 (1997).
Cornuet, J. M., Piry, S., Luikart, G., Estoup, A. & Solignac, M. New methods employing multilocus genotypes to select or exclude populations as origins of individuals. Genetics 153, 1989–2000 (1999).
Author information
Authors and Affiliations
Contributions
S.M. analyzed the data (training phase), interpretation of results, and prepared the draft article. A.H.K.F. obtained data-sets, analysis (validation phase), and interpretation of the supplementary results. M.H.M. made the illustration of images in result section. M.H.M and M.K.B. contributed in the editing of the draft article.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Manzoori, S., Farahani, A.H.K., Moradi, M.H. et al. Detecting SNP markers discriminating horse breeds by deep learning. Sci Rep 13, 11592 (2023). https://doi.org/10.1038/s41598-023-38601-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-38601-z
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.