Abstract
Head and neck squamous carcinoma (HNSC) induces high cancer-related death worldwide. The biomarker screening on diagnosis and prognosis is of great importance. This research is aimed to explore the specific diagnostic and prognostic biomarkers for HNSC through bioinformatics analysis. The mutation and dysregulation data were acquired from UCSC Xena and TCGA databases. The top ten genes with mutation frequency in HNSC were TP53 (66%), TTN (35%), FAT1 (21%), CDKN2A (20%), MUC16 (17%), CSMD3 (16%), PIK3CA (16%), NOTCH1 (16%), SYNE1 (15%), LRP1B (14%). A total of 1,060 DEGs were identified, with 396 up-regulated and 665 downregulated in HNSC patients. Patients with lower expression of ACTN2 (P = 0.039, HR = 1.3), MYH1 (P = 0.005, HR = 1.5), MYH2 (P = 0.035, HR = 1.3), MYH7 (P = 0.053, HR = 1.3), and NEB (P = 0.0043, HR = 1.5) exhibit longer overall survival time in HNSC patients. The main DEGs were further analyzed by pan-cancer expression and immune cell infiltration analyses. MYH1, MYH2, and MYH7 were dysregulated in the cancers. Compared with HNSC, their expression levels are lower in the other types of cancers. MYH1, MYH2, and MYH7 were expected to be the specific diagnostic and prognostic molecular biomarkers of HNSC. All five DEGs have a significant positive correlation with CD4+T cells and macrophages.
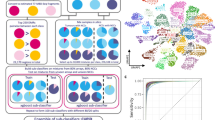
Similar content being viewed by others
Introduction
Head and neck squamous carcinoma (HNSC) accounts for 95% of head and neck cancer (HNC) and is the sixth most common malignancy worldwide1. It contains neck, ear, nose, throat, and oral and maxillofacial tumors2,3. Human papillomavirus (HPV) infection is the most common causative factor for HNSC4,5,6. Although advanced treatments have been employed in HNSC treatment, the 5-year survival rate is less than 50% in stage III and IV HNSC7. Due to the difficulty in detecting HNSC at an early stage, the delay in detection and follow-up raises the risk of mortality. To control the mortality rate, biomarker identification in diagnosis, prognosis, and metastasis is very important, which might provide essential evidence for clinical treatment.
Machine learning based on public databases provides enormous value for somatic mutation and transcriptomes data mining8. The Cancer Genome Atlas (TCGA) datasets were generally used as databases containing various mutation and expression data of cancers. In recent years, machine learning and data mining have contributed to biomarker screening for early detection, diagnosis, drug application, metastases, and prognosis of various cancers9,10,11. For example, by employing four machine learning models, Leitheiser et al.12 predicted the primary sites of HNSC metastases based on DNA methylation. Rendleman et al.13 developed a machine learning method to improve the usability of the HNSC dataset for enhancing future oncological decisions and clinical applications. Jiang et al.14 provided potential predictive value for HNSC by employing public datasets. However, the data mining on the prognosis biomarker of HNSC is still inadequate.
Realizing the excellent application prospect of machine learning and the urgent need for reliable prognosticators for HNSC, we tried to identify the biomarkers related to diagnosis and prognosis in HNSC based on gene mutation and dysregulation. Moreover, we attempted to investigate the immune infiltration of DEGs in HNSC. This study may provide a specific set of molecular biomarkers for HNSC.
Results
Mutations in HNSC patients
The mutation profile of HNSC patients from the TCGA database was obtained. Figure 1A showed that the frequency distribution of different variant types was divided into seven according to the impact on protein-coding. We found that missense mutation accounts for the majority of the variant types. In these variant types, the number of SNPs is significantly more than that of INS and DEL (Fig. 1B). Besides, C > T transversion was the primary type of single nucleotide variants (SNV) in HNSC (Fig. 1C). We show genes with a mutation frequency of top25 in Fig. 1D. The top ten genes with a mutation frequency were TP53 (66%), TTN (35%), FAT1 (21%), CDKN2A (20%), MUC16 (17%), CSMD3 (16%), PIK3CA (16%), NOTCH1 (16%), SYNE1 (15%), LRP1B (14%).
The landscape of mutation profiles in HNSC patients. (A,B) classification of mutation types according to different categories. (C) the frequency distribution of SNV mutation types. Categorizing SNPs as Transitions_vs_Transversions, the aggregated data can also be displayed as a boxplot showing the overall distribution of the six different transitions, and as a stacked bar chart showing the proportion of transitions in each sample. (D) showed Waterfall plot of the top 25 mutated genes in TCGA HNSCC cohort. The first part is the heatmap in the middle, where each row represents a gene and each column represents a sample, showing the distribution of different mutation types for each sample, and the second part is a stacked bar chart on the right, representing the different mutation types on each gene The frequency distribution of the loci, the third part is the stacked histogram above, which represents the frequency distribution of the loci of different mutation types in each sample.
Differentially expression genes calculation
In these patients, the expression levels in two groups were compared, and a total of 1061 DEGs were identified, with 396 up-regulated and 665 downregulated (Fig. 2A,B). Then, we retained the DEGs with more than 1% mutation frequency. After the screening, 187 common DEGs were obtained (Fig. 2C), including 66 up-regulated and 121 down-regulated.
PPI and key module analyses on DEGs
PPI analysis was performed on the common DEGs with more mutation frequency. The PPI network showed 144 nodes and 489 interaction pairs (Fig. 3). The DEGs with more than one connection with other DEGs were retained (Supplementary File 1). In addition, two sub-network modules were screened, with 15 nodes and 102 interaction pairs in module A (score = 14.571), and 14 nodes and 75 interaction pairs in module B (score = 11.538).
In addition, we listed the key DEGs in sub-network modules in Table 1.
Function analysis on DEGs in sub-network modules
We further analyzed the function of DEGs by GO-BP and KEGG. We found that the DEGs in module A are mainly involved in GO-BP terms, including extracellular matrix organization, extracellular structure organization, collagen fibril organization, endodermal cell differentiation, endoderm formation, endoderm development, formation of the primary germ layer, cellular response to amino acid stimulus, collagen-activated tyrosine kinase receptor signaling pathway, and collagen-activated signaling pathway (Fig. 4A). They are also involved in KEGG pathways of protein digestion and absorption, ECM-receptor interaction, AGE-RAGE signaling pathway in diabetic complications, amoebiasis, focal adhesion, human papillomavirus infection, PI3K-Akt signaling pathway, Relaxin signaling pathway, small cell lung cancer, and platelet activation (Fig. 4B).
The DEGs in module B mainly participated in muscle contraction, muscle system process, myofibril assembly, cellular component assembly involved in morphogenesis, striated muscle cell development, cardiac muscle cell development, cardiac cell development, muscle filament sliding, actin-myosin filament sliding, and cardiac muscle fiber development (Fig. 4C). In addition, KEGG pathways of hypertrophic cardiomyopathy, dilated cardiomyopathy, cardiac muscle contraction, thyroid hormone signaling pathway, adrenergic signaling in cardiomyocytes, cGMP-PKG signaling pathway, viral myocarditis, and arrhythmogenic right ventricular cardiomyopathy were enriched (Fig. 4D).
Key DEG expression validation, survival curve, and UALCAN Pan-Cancer analysis
Some of the key DEGs were validated by GEPIA. ACTN2, MYH1, MYH2, MYH7, and NEB were significantly downregulated in HNSC patients (P < 0.05; Fig. 5). Patients with lower expression of ACTN2 (P = 0.039, HR = 1.3), MYH1 (P = 0.005, HR = 1.5), MYH2 (P = 0.035, HR = 1.3), MYH7 (P = 0.053, HR = 1.3), and NEB (P = 0.0043, HR = 1.5) exhibit longer overall survival time. These results demonstrated that these DEGs might be essential for predicting the overall survival of HNSC.
Then, we evaluated their expression in other cancers (Fig. 6A–E). ACTN2 was significantly downregulated in bladder urothelial carcinoma (BLCA), kidney chromophobe (KICH), kidney renal papillary cell carcinoma (KIRP), lung adenocarcinoma (LUAD), lung squamous cell carcinoma (LUSC), prostate adenocarcinoma (PRAD), stomach adenocarcinoma (STAD), thyroid carcinoma (THCA), and uterine corpus endometrial carcinoma (UCEC) (P < 0.001); up-regulated in invasive breast carcinoma (BRCA), cholangiocarcinoma (CHOL), and colon adenocarcinoma (COAD). We can observe that although MYH1, MYH2, and MYH7 were dysregulated in some of the cancers, their expression levels are very low, except for HNSC. Therefore, MYH1, MYH2, and MYH7 can be considered specific HNSC biomarkers. As for NEB, it was significantly downregulated in BRCA, HNSC, KICH, KIRP, and THCA; up-regulated in BLCA, CHOL, COAD, esophageal carcinoma (ESCA), liver hepatocellular carcinoma (LIHC), LUAD, LUSC, rectum adenocarcinoma (READ), stomach adenocarcinoma (STAD), and UCEC (P < 0.05; P < 0.01; P < 0.001). Moreover, we evaluated the expression levels of ACTN2, MYH1, MYH2, MYH7, and NEB in HPV-positive and -negative HNSC samples. Their expression levels showed no significant differences in HPV-positive and negative HNSC samples (Fig. 6F).
The expression of ACTN2, MYH1, MYH2, MYH7, and NEB in Pan-cancers. (A–E) represented the expression levels of ACTN2, MYH1, MYH2, MYH7, and NEB in Pan-cancers. (F) represented the expression levels of ACTN2, MYH1, MYH2, MYH7, and NEB in HPV-positive and -negative HNSC samples. ACC, Adrenocortical carcinoma; BLCA, Bladder Urothelial Carcinoma; BRCA, Breast invasive carcinoma; CESC, Cervical squamous cell carcinoma and endocervical adenocarcinoma; CHOL, Cholangiocarcinoma; COAD, Colon adenocarcinoma; COAD, Colon adenocarcinoma; READ, Rectum adenocarcinoma Esophageal carcinoma; DLBC, Lymphoid Neoplasm Diffuse Large B-cell Lymphoma; ESCA, Esophageal carcinoma; FPPP, FFPE Pilot Phase II; GBM, Glioblastoma multiforme; GBMLGG, Glioma; HNSC, Head and Neck squamous cell carcinoma; KICH, Kidney Chromophobe; KIRC, Kidney renal clear cell carcinoma; KIRP, Kidney renal papillary cell carcinoma; LAML, Acute Myeloid Leukemia; LGG, Brain Lower Grade Glioma; LIHC, Liver hepatocellular carcinoma; LUAD, Lung adenocarcinoma; LUSC, Lung squamous cell carcinoma; MESO, Mesothelioma; OV, Ovarian serous cystadenocarcinoma; PAAD, Pancreatic adenocarcinoma; PCPG, Pheochromocytoma and Paraganglioma; PRAD, Prostate adenocarcinoma; READ, Rectum adenocarcinoma; SARC, Sarcoma; SKCM, Skin Cutaneous Melanoma; STAD, Stomach adenocarcinoma; STES, Stomach and Esophageal carcinoma; TGCT, Testicular Germ Cell Tumors; THCA, Thyroid carcinoma; THYM, Thymoma; UCEC, Uterine Corpus Endometrial Carcinoma; UCS, Uterine Carcinosarcoma; UVM, Uveal Melanoma; AML, Acute Myeloid Leukemia; CCSK, Clear Cell Sarcoma of the Kidney; NBL, Neuroblastoma; OS, Osteosarcoma; RT, Rhabdoid Tumor; WT, High-Risk Wilms Tumor.
Immune cell infiltration correlation analysis
The correlation between key DEGs and immune infiltrate cells in total HNSC, HNSC-positive, and HNSC-negative samples was evaluated. As for total HNSC samples, we found that ACTN2, MYH1, MYH2, and MYH7 were negatively correlated with B cell, CD8+T cell, and positively correlated with CD4+T cell, macrophage, neutrophil, and dendritic cell (Fig. 7A–E). All five DEGs have a significant positive correlation with CD4+T cells and macrophages.
ACTN2 had positive correlation with CD4+T cell (partial correlation = 0.253; P = 2.03e−08), macrophage (partial correlation = 0.181; P = 6.03e−05), neutrophil (partial correlation = 0.157; P = 5.82e-04), and dendritic cell (partial correlation = 0.161; P = 4.02e−04) in HNSC samples. It showed an opposed correlation in HNSC-HPV positive and HNSC-HPV negative in immune cell infiltration of B cell and CD8+T cell, but not significant (Fig. 7A). Therefore, the HPV infection had no significant influence on immune cell infiltration of ACTN2.
MYH1 showed positive correlation with CD4+T cell (partial correlation = 0.201; P = 8.71e−06), macrophage (partial correlation = 0.146; P = 1.31e−03), neutrophil (partial correlation = 0.095; P = 3.76e−02), and dendritic cell (partial correlation = 0.104; P = 2.30e−02; Fig. 7B). The MYH1 showed a consistent correlation in the immune cells of HNSC-HPV positive and negative samples, except in neutrophils. Although MYH1 in HNSC-HPV positive sample is negatively correlated with neutrophils, but is not significant.
As for MYH2, MYH7, and NEB were significantly positively correlated with CD4+T cell (P < 0.001; Fig. 7C–E). They showed consistent correlation in the immune cells among the total HNSC, HNSC-HPV positive and negative samples; Some slight correlation opposition was observed in HNSC-HPV positive or negative with total HNSC, but not significant. Therefore, we concluded that no significant differences were identified in HNSC-HPV positive or negative with total HNSC in terms of immune cell infiltration of ACTN2, MYH1, MYH2, MYH7, and NEB.
Discussion
HNSC, a dreadful opponent, has become one of the world's most deadly diseases. Treatment for HNSC is costly, has a long cycle, and is not affordable. Therefore, the precise and early prediction of HNSC outcomes is critical for therapy options and prognosis improvement. Our present study focused on gene mutation and dysregulation expression in HNSC, expecting to identify essential biomarkers for HNSC.
In the present study, we explored the landscape of mutation and dysregulation in HNSC patients, showing that TP53, TTN, FAT1, CDKN2A, and MUC16 were the most predominant mutated genes. The mutation identification was consistent with Jiang et al.14. Among human genes, TP53 is a critical tumor suppressor gene, with low expression in normal cells and high expression in malignant tumors, regulating cell proliferation, apoptosis, angiogenesis, and DNA repair15. Mutated TP53 was found in various cancers and is correlated with reduced O.S. Furthermore; it showed that HNSC patients with TP53 mutations have a bleak prognosis than TP53-wildtype HNSC16. Our present study also found that mutated TP53 is observed in HNSC.
Then, the dysregulated genes were identified, with 187 DEGs having more mutation sites. We conducted the PPI analysis on these DEGs and identified two modules. We further analyzed these DEGs in these modules by GO-BP and KEGG analyses. The DEGs in module A are involved in the PI3K-Akt signaling pathway, a classic intracellular signaling pathway regulating cell apoptosis, proan essential pathway that regulates cell apoptosis, prognosis, and metastasis in HNSC17,18,19. The DEGs in module B participated in the cGMP-PKG signaling pathway. In addition, Tuttle et al.20 demonstrated that the cGMP-PKG signaling pathway is one of the therapeutic targets of HNSC.
The DEGs of ACTN2, MYH1, MYH2, MYH7, and NEB in module B were significantly associated with O.S. in HNSC. ACTN2 is a member of the spectrin gene superfamily that includes varying groups of cytoskeletal proteins. In breast cancer patients, mutated ACTN2 was related to invasive ductal carcinoma and suggested a worse O.S. than ductal carcinoma in situ21. Sun et al.22 demonstrated that ACTN2 is one of the hub genes selected by bioinformatics methods in PTEN mutation prostate cancer. Xu et al.23 revealed that negative ACTN2 expression contributed to a better O.S. in HNSC patients, consistent with our present study. Therefore, ACTN2 might be an essential biomarker for predicting O.S. of HNSC, which might be employed in clinical.
Through pan-cancer analysis, we found that MYH1, MYH2, and MYH7 are dysregulated in cancers, including HNSC. However, compared with HNSC, they were lower expressed in other cancers, revealing that these three genes can be considered single HNSC biomarkers. So far, different mutations in multiple members of the MYH gene family are associated with human hereditary diseases24. Among them, MYH2 mutations can cause a class of skeletal muscle diseases characterized by ophthalmoplegia. MYH7 mutations can cause skeletal muscle diseases, including myosin deposition myopathy, and Distal Laing myopathy is also closely related to hypertrophic cardiomyopathy. However, MYH1, MYH2, and MYH7 functions in HNSC are rarely mentioned. Further study into the interaction of the MYH gene family and HHSC appears to be an intriguing research topic.
As we all know, HPV infection is the most common causative factor for HNSC. Considering the significant influence of HPV on HNSC, some molecular research on HNSC have divided the samples into HNSC-HPV positive and negative groups. For example, Zhang et al.25 found that Dedicator of cytokinesis 8 (DOCK8) is identified as one prognosis biomarker associated with immune infiltration HPV-positive HNSC. Our present study identified five prognosis DEGs that are significantly related to the O.S. in HNSC. To determine whether the HPV infection had significant influences on the results. We further analyzed their expression in HPV-positive and -negative HNSCs, and no significant differences were observed. Moreover, we also tested the immune cell infiltration of DEGs in both HPV-positive and -negative HNSCs. Although studies have demonstrated that HPV influences the immune infiltration situation26, it is not observed in ACTN2, MYH1, MYH2, MYH7, and NEB.As genes highly expressed in the tumor cells are expected to affect tumor purity positively, the association between the expression of prognosis DEGs and six immune infiltrates was evaluated. We found that all five DEGs have a significant positive correlation with CD4+T cells and macrophages. Therefore, we speculate that genetic mutations and differential expression of ACTN2, MYH1, MYH2, MYH7, and NEB genes in HNSC cells may be important drivers of CD4+T cell and macrophage infiltration.
Conclusions
In our present study, we identified the biomarker of HNSC in terms of diagnosis and prognosis. TP53 was found as the most mutated gene in HNSC. ACTN2, MYH1, MYH2, MYH7, and NEB genes were significantly associated with poor prognosis. Moreover, MYH1, MYH2, and MYH7 were expected to be the specific diagnostic and prognostic biomarkers molecular biomarkers of HNSC.
Methods
Data collection
The clinical data and gene expression profiles of HNSC patients were downloaded from the UCSC Xena database (https://xenabrowser.net/datapages/). As calculated, a total of 506 tumors and 44 adjacent HNSC samples were included in our present study. The gene expression levels of examples were provided in Supplementary File 2. The somatic mutation data was from the TCGA database by Varscan27.
Screening of high-frequency mutation genes
The maftools28 was used to summarize the HNSC mutation information downloaded from the TCGA official website and screen the high-frequency mutation genes. The top 25 high-frequency genes with the mutation frequency were exhibited with a waterfall chart.
Differentially expressed gene (DEG) identification
The DEGs in tumor tissues were identified according to the FPKM value by limma package29 (Version 3.10.3), using P-value < 0.05 and log2 fold change (F.C.) > 2 as thresholds. The Pheatmap R package was employed for drawing the heatmap and volcano.
Then, the common DEGs with high-frequency mutation genes that exhibit a mutation frequency of more than 1% were retained for further study.
Protein–protein interaction (PPI) and module analyses
The PPI analysis was performed on all the common DEGs using STRING30 (Version: 10.0, http://www.string-db.org/) database, with a 0.4 (medium confidence) parameter. The network was constructed by Cytoscape (version: 3.2.0)31. The most significant clustered modules in the PPI network were analyzed using the Cytoscape plug-in MCODE32 (Version 1.4.2, http://apps.cytoscape.org/apps/MCODE) method, with a threshold of score ≥ 10.
Functional analysis
The biological process (B.P.) of Gene Ontology (G.O.) and the Kyoto Encyclopedia of Genes and Genomes (KEGG) analyses on DEGs were performed by clusterProfiler33, with adjusted P-value < 0.05 and gene number count ≥ 2. The top ten G.O. terms and KEGG pathways were visualized.
Key DEG expression validation and survival curve analysis
The key DEGs in the module were subjected to expression verification and survival analysis by GEPIA34. Survival analysis was grouped by median data and performed using OS survival information. The pan-cancer expression levels of key DEGs significantly associated with prognosis were assessed using UALCAN (http://ualcan.path.uab.edu/index.html). The expression levels of key DEGs in HNSC-positive and HNSC-negative samples were evaluated.
Immune cell infiltration analysis
As for the immune cell infiltration analysis, the HNSC samples were divided into HNSC-positive and HNSC-negative. Then, the infiltration of the key DEGs in six immune cells (B cells, CD4+ T cells, CD8+ T cells, neutrophils, macrophages, and dendritic cells) were identified through the TIMER database.
Ethics approval and consent to participate
UCSC Xena and TCGA belongs to public databases. The patients involved in the database have obtained ethical approval. All methods were carried out in accordance with relevant guidelines and regulations.
Data availability
The RNA-Seq and mutation data re-analyzed during the current study are available in the TCGA database with the links of https://xenabrowser.net/datapages/?dataset=TCGA-HNSC.htseq_fpkm.tsv&host=https%3A%2F%2Fgdc.xenahubs.net and https://xenabrowser.net/datapages/?dataset=TCGA-HNSC.varscan2_snv.tsv&host=https%3A%2F%2Fgdc.xenahubs.net.
References
Machiels, J.-P. et al. Advances in the management of squamous cell carcinoma of the head and neck. Fprime Rep. 6, 44 (2014).
Leemans, C. R., Braakhuis, B. J. & Brakenhoff, R. H. The molecular biology of head and neck cancer. Nat. Rev. Cancer 11, 9–22 (2011).
Vigneswaran, N. & Williams, M. D. Epidemiologic trends in head and neck cancer and aids in diagnosis. Oral Maxillofac. Surg. Clin. 26, 123–141 (2014).
Du, E. et al. Long-term survival in head and neck cancer: Impact of site, stage, smoking, and human papillomavirus status. Laryngoscope 129, 2506–2513 (2019).
Spence, T., Bruce, J., Yip, K. W. & Liu, F.-F. HPV associated head and neck cancer. Cancers 8, 75 (2016).
Leemans, C. R., Snijders, P. J. & Brakenhoff, R. H. The molecular landscape of head and neck cancer. Nat. Rev. Cancer 18, 269–282 (2018).
Fuller, C. D. et al. Conditional survival in head and neck squamous cell carcinoma: Results from the SEER dataset 1973–1998. Cancer Interdiscip. Int. J. Am. Cancer Soc. 109, 1331–1343 (2007).
Wood, D. E. et al. A machine learning approach for somatic mutation discovery. Sci. Transl. Med. 10, eaar7939 (2018).
Vougas, K. et al. Machine learning and data mining frameworks for predicting drug response in cancer: An overview and a novel in silico screening process based on association rule mining. Pharmacol. Therap. 203, 107395 (2019).
Ak, M. F. in Healthcare. 111 (MDPI).
Goyal, K., Sodhi, P., Aggarwal, P. & Kumar, M. in Proceedings of 2nd International Conference on Communication, Computing and Networking. 727–734 (Springer).
Leitheiser, M. et al. Machine learning models predict the primary sites of head and neck squamous cell carcinoma metastases based on DNA methylation. J. Pathol. 256, 378–387 (2022).
Rendleman, M. C. et al. Machine learning with the TCGA-HNSC dataset: Improving usability by addressing inconsistency, sparsity, and high-dimensionality. BMC Bioinform. 20, 1–9 (2019).
Jiang, A.-M. et al. Tumor mutation burden, immune cell infiltration, and construction of immune-related genes prognostic model in head and neck cancer. Int. J. Med. Sci. 18, 226 (2021).
Fischer, M., Grossmann, P., Padi, M. & DeCaprio, J. A. Integration of TP53, DREAM, MMB-FOXM1 and RB-E2F target gene analyses identifies cell cycle gene regulatory networks. Nucleic Acids Res. 44, 6070–6086 (2016).
Network, C. G. A. Comprehensive genomic characterization of head and neck squamous cell carcinomas. Nature 517, 576 (2015).
García-Carracedo, D. et al. Impact of PI3K/AKT/mTOR pathway activation on the prognosis of patients with head and neck squamous cell carcinomas. Oncotarget 7, 29780 (2016).
Yang, J. et al. Osthole induces cell cycle arrest and apoptosis in head and neck squamous cell carcinoma by suppressing the PI3K/AKT signaling pathway. Chem. Biol. Interact. 316, 108934 (2020).
Yang, S. et al. STC2 promotes head and neck squamous cell carcinoma metastasis through modulating the PI3K/AKT/Snail signaling. Oncotarget 8, 5976 (2017).
Tuttle, T. R., Mierzwa, M. L., Wells, S. I., Fox, S. R. & Ben-Jonathan, N. The cyclic GMP/protein kinase G pathway as a therapeutic target in head and neck squamous cell carcinoma. Cancer Lett. 370, 279–285 (2016).
Yang, M. et al. A breast one-patient panel of heterogeneous genomes reveals genetic alterations driving DCIS into invasive lesions. Future Oncol. 15, 1565–1576 (2019).
Sun, J., Li, S., Wang, F., Fan, C. & Wang, J. Identification of key pathways and genes in PTEN mutation prostate cancer by bioinformatics analysis. BMC Med. Genet. 20, 1–9 (2019).
Xu, Y. et al. A ceRNA-associated risk model predicts the poor prognosis for head and neck squamous cell carcinoma patients. Sci. Rep. 11, 1–18 (2021).
He, Y.-M. & Gu, M.-M. Research progress of myosin heavy chain genes in human genetic diseases. Yi chuan= Hereditas 39, 877–887 (2017).
Zhang, Z. et al. DOCK8 serves as a prognostic biomarker and is related to immune infiltration in patients with HPV positive head and neck squamous cell carcinoma. Cancer Control 28, 10732748211011952 (2021).
Qureshi, H. A. et al. Impact of HPV status on immune responses in head and neck squamous cell carcinoma. Oral Oncol. 127, 105774 (2022).
Koboldt, D. C. et al. VarScan 2: Somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res. 22, 568–576 (2012).
Mayakonda, A., Lin, D.-C., Assenov, Y., Plass, C. & Koeffler, H. P. Maftools: Efficient and comprehensive analysis of somatic variants in cancer. Genome Res. 28, 1747–1756. https://doi.org/10.1101/gr.239244.118 (2018).
Smyth, G. K. In Bioinformatics and Computational Biology Solutions Using R and Bioconductor (eds Gentleman, R. et al.) 397–420 (Springer, 2005).
Szklarczyk, D. et al. STRING v10: Protein–protein interaction networks, integrated over the tree of life. Nucleic Acids Res., gku1003 (2014).
Shannon, P. et al. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504. https://doi.org/10.1101/gr.1239303 (2003).
Bandettini, W. P. et al. MultiContrast delayed enhancement (MCODE) improves detection of subendocardial myocardial infarction by late gadolinium enhancement cardiovascular magnetic resonance: A clinical validation study. J. Cardiovasc. Magn. Reson. 14, 83. https://doi.org/10.1186/1532-429X-14-83 (2012).
Yumoto, R., Suzuka, S., Oda, K., Nagai, J. & Takano, M. Endocytic uptake of FITC-albumin by human alveolar epithelial cell line A549. Drug Metab. Pharmacokinet. 27, 336–343. https://doi.org/10.2133/dmpk.dmpk-11-rg-127 (2012).
Tang, Z. et al. GEPIA: A web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res. 45, W98–W102 (2017).
Funding
This work was supported by the General Program of National Natural Science Foundation of China (81972555 to Fang-Zhou Liu).
Author information
Authors and Affiliations
Contributions
G.J. all data analysis, and writing; Z.Y. data analysis, and writing; Y.Z. performed part of the analysis; X.Z. conceptualization, interpretation and discussion of results; F.L. conceptualization, advised all research, design of experiments, evaluation and discussion of results, revision of manuscript, and provided research funding. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ju, G., Yao, Z., Zhao, Y. et al. Data mining on identifying diagnosis and prognosis biomarkers in head and neck squamous carcinoma. Sci Rep 13, 10020 (2023). https://doi.org/10.1038/s41598-023-37216-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-37216-8
This article is cited by
-
Disulfidptosis features and prognosis in head and neck squamous cell carcinoma patients: unveiling and validating the prognostic signature across cohorts
Journal of Cancer Research and Clinical Oncology (2024)
-
Identification of Prognostic and Immune Characteristics of Two Lung Adenocarcinoma Subtypes Based on TRPV Channel Family Genes
The Journal of Membrane Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.