Abstract
Mutation breeding is a significant means of increasing breeding efficiency and accelerating breeding process. In present study, we explored a new method for mutations inducing in rice (Oryza sativa L.) by using direct current electrophoresis bath (DCEB). The results showed that 20 mM NaCl solution is the optimal buffer, and the mortality of rice seeds followed an upward trend with increasing voltage and processing time of DCEB. By exploring the mutagenic effects of γ-irradiation and DCEB on seed vigor and physiological damages, we found that the physiological damages induced by DCEB on seed vigor were significant compared with that by γ-irradiation. We screened two mutants with low filled grain percentage and one mutant with abnormal hull from the M2 generations. These three mutants were confirmed to be authentic mutants based on 48 SSR markers followed by the protocol NY/T 1433–2014. Whole-genome resequencing detected a total of 503 and 537 polymorphisms in the two mutants, respectively, and the DCEB mutagenesis induced mainly InDel variants, while the exon region of mutant genes occupied a large proportion, especially the SNP variants, which occupied about 20% of the mutation sites in the exon region.
Similar content being viewed by others
Introduction
With the spread of the 2019 coronavirus disease (COVID-19), in the latest edition of The State of Food Security and Nutrition in the World 2020, the World Food and Agriculture Organization (FAO) predicted that the world will not reach the goal of zero hunger by 20301, which means that the world will continue to face great challenges in food security. Rice (Oryza sativa L.) is the staple food for over half the world's population2. To ensure rice food security with changes in global climate and world population growth, there is an urgent need to develop high-yielding and high grain quality rice varieties at a greater pace3,4. Mutagenesis is a nearly 100-year-old technology, and mutation breeding currently plays a major role in the development of superior plant varieties worldwide5. However, spontaneous mutation rates for higher plants are low, ranging from 10–5 to 10–8, thus, mutagenesis is an important strategy to increase mutation frequencies6. The main mutagens are physical mutagens such as X-rays7,8,9, γ-irradiation10,11,12 and ion beams13,14,15,16, chemical mutagens such as ethyl methane sulfonate17 and sodium azide18,19. Scientists have developed rice varieties with various excellent traits, e.g. dwarfing, high yield and disease resistance20,21. Traditionally, mutations can be induced in a variety of ways, such as exposing plant propagules, including seeds, tissues and organs, to physical and chemical mutagens22. The most widely used method of mutation breeding, however, is the process of exposing seeds to chemical or physical mutagens to produce mutants that serve as new genetic resources with improved traits23. There are currently 3,365 registered mutant varieties in the International Atomic Energy Agency (FAO/IAEA) Mutant Variety Database, of which 2569 are seed propagated plants. Next-generation sequencing is increasingly being applied in single nucleotide polymorphism (SNP) detection and for assessing the effects of induced mutations24. Whole-genome resequencing (WGR) was employed to study the molecular characterization of mutations on a whole-genome level. Therefore, an increasing number of researchers have applied WGR technology to reveal mutagenic characteristics on the genome25.
Electrophoresis is a phenomenon in which charged molecules move in an electric field toward electrodes of opposite charge. The technique of using electrophoresis to separate substances is called electrophoretic technology, and an electrophoresis apparatus generally consists of a power supply, an electrophoresis bath, and a detection unit. In 1937, Tiselius, a Swedish scientist, designed the first electrophoresis apparatus, creating a new era of electrophoresis technology26. In recent years, electrophoresis has been widely used in various fields, such as biomedicine27,28, analytical chemistry29 and microbiology.
The present study provides a new method for mutations inducing in rice by using DC (direct current) electrophoresis bath (DCEB). Aimed to (1) explore the optimal treatment for DCEB mutagenesis; (2) compare the mutagenic effects induced by DCEB and by γ-irradiation, and (3) predict the mutation frequencies of DC electrophoretic bath mutagenesis by WGR. The present studies are presented as successful examples of this method which is simple, convenient, safe, flexible, and low costly in rice on mutation breeding.
Results
The optimal conditions for DC electrophoresis bath mutagenesis
Four buffer solutions, NaCl solution, NaOH solution, TAE buffer, and TBE buffer, were used to explore their effects on the germination of rice seeds, sterile water was used as the negative control. The results showed that the 20 mM NaCl solution was the most effective and accelerated the germination of rice seeds, followed by the NaOH solution, the germination rate (GR) showed a sharp downward trend with increasing concentrations of TAE and TBE buffers. It is worth noting that the effect of 20 mM NaCl solution on the germination rate of rice seeds was more pronounced than that of sterile water (Fig. 1). Therefore, 20 mM NaCl solution was selected as the buffer for DCEB.
Germination rate of rice seeds at different solution concentrations. (A) NaCl solution; (B) NaOH solution; (C) TAE buffer; and (D) TBE buffer. The error bars stand for standard deviations. Control: sterile water, different letters in each column indicate significant differences between different treatments (p < 0.05; Duncan’s test).
Seven voltages (20 V, 50 V, 80 V, 110 V, 140 V, 170 V and 200 V) and three treatment times (12 h, 24 h and 48 h) were used for a total of 21 treatment combinations, and 20 mM NaCl solution was used as the electrophoresis buffer. The results showed that the mortality of rice seeds followed an upward trend with increasing time and voltage, the increasing trend of mortality becomes flat as the voltage increases to 140 V, when the treatment time extend to 48 h, the mortality rate was all arrived 85% except 20 V, suggesting that the effect of voltage on mortality becomes insignificant when the time extend to 48 h. When treated under 110 v for 12 h, the mortality rate was 52.67%, when treated under 110 v for 50 V or 80 V, the mortality rates reached 52.5%, which were close to the LD50 (Fig. 2, Table 1).
Effect of γ-irradiation and DC electrophoresis bath mutagenesis on rice seed vigor
Finally, we selected six DCEB treatments with the three highest voltages (140 V, 170 V and 200 V), and six doses of γ-irradiation (50 Gy, 100 Gy, 150 Gy, 200 Gy, 250 Gy, and 300 Gy), as well as CK, consisting of thirteen treatments, were obtained to analyze seed vigor (Table 2). Compared with the control, γ-irradiation had a positive effect on rice seed vigor at 50 Gy and 100 Gy and a negative effect at other doses. The rice seed vigor index (VI) showed a trend of increasing and then decreasing with different doses of γ-irradiation (Fig. 3, Table 2). There were extremely significant differences between the positive and negative effects of the 50 Gy and 300 Gy doses, respectively (Table 3). This result indicates that when the dose of γ-irradiation reached 100 Gy, it had a positive effect on the seed vigor, and beyond this threshold, the seed vigor decreased. In contrast, DCEB mutagenesis had a negative effect on rice seed vigor under all six-level voltages treatments, the GR, GI and VI decreasing with increasing electrophoresis time and voltage (Fig. 2, Table 1). This result indicates that the both methods have effects on seed viability, and the DCEB mutagenesis has more negative significant effects on rice seed vigor than that γ-irradiation.
Effect of γ-irradiation and DCEB mutagenesis on agronomic traits
We investigated the agronomic traits of M1 generations induced by DCEBs and γ-irradiation, respective. In Table 4, the number of productive tillers of plants treated with γ-irradiation were significantly higher than that of control plants, while plant height and filled grain percentage were lower in treated plants than the control; no significant differences were found for other agronomic traits, such as thousand-grain weight and flag leaf length and width. In particular, the 250 Gy and 300 Gy doses produced more productive tillers and lower filled grain percentages than the control. There was almost no significant effect of DCEB treatment on the contemporary agronomic traits compared to the control (Table 4).
Physiological damage caused by γ-irradiation and DCEB mutagenesis
The combination of seed vigor performance and agronomic traits in the M1 generation suggested that γ-irradiation had a significant effect on the agronomic traits; while the DCEB had a significant effect on seed vigor index. When estimating the physiological damage caused by both mutagenesis methods, the physiological damages (germination rate, bud length, panicle weight and filled grain percentage) increased with increasing doses of γ-irradiation and showed a linear correlation. Moreover, the physiological damages caused by DCEB increased with higher voltage and a longer electrophoresis time.
Genetic background analysis of mutants
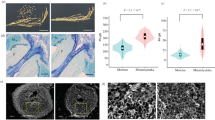
There were 116 seeds germinated with 102 healthy seedlings obtained from the DCEB treatment of 140 V, 48 h; 92 seeds germinated with 78 healthy seedlings and 72 seeds germinated with 60 healthy seedlings were obtained from the treatments of 170 V, 48 h and 200 V, 48 h, respectively. There was no apparent mutation from each M1 generations, however, from M2 generation (self-crossed from M1 generation)we screened two mutants (M1, from the treatment of 140 V, 48 h, and M2, from the treatment of 200 V, 48 h,) with low filled grain percentage and one mutant (M3, from the treatment of 170 V, 48 h) with abnormal hull. The fertile pollen percentage of M1, M2, M3 and WT was about 36.5%, 44.1%, 56.7% and 97.6%, respectively, and the filled grain percentage was 6.3%, 13.4%, 26.8% and 93.2%, respectively (Fig. 4). The genetic analysis of the mutants according to the 48 SSR markers showed that all three mutants shared the wild type bands (Fig. 5). These results clearly indicated that all three mutants are true mutants.
The Mutants of low filled grain percentage (M1 and M2) and abnormal hull (M3). (A) The plant type of M1 and M2 (right two) compared with WT (left); (B) The panicle shape of M1 and M2 (right two) compared with WT (left); (C) The plant type of M3 (right) compared with WT (left); (D) The panicle shape of M3 (right) compared with WT (left); and (E) The deformed glumes of M3. WT, wild type, M1, M2 and M3 were obtained from M2 generations which were self-crossed from M1 generations treated by DCEB of140 V, 48 h, 200 V, 48 h and 170 V, 48 h, respectively.
Parts of gel electropherograms profile of wild type and mutant lines. For each SSR maker, lanes from left to the right are: M, W, M1 and M2. M: DNA ladder. M: DNA ladder 1000; W: wild type; M1 and M2: low filled grain percentage mutants from 140 V, 48 h treatment and 2 from 140 V, 48 h treatment, respectively; M3: abnormal hull mutant from 110 V, 48 h treatment. The red boxes indicate the main amplified blots for each SSR, and the whole gel electropherograms profile and the original gels are presented in Supplementary Figure S2.
Evaluation of resequencing results
To further investigate the effects of DCEB mutagenesis at the genetic level, three experimental materials (WT, M1 and M2) were subjected to WGR in this study. Approximately 37.43 Gbp of clean data was produced, with a Q20 of 97.03%, Q30 of 92.25% and GC of 42.43% (averaged from WT, M1 and M2, respectively, Table 5). According to the results of WGR, the average sample-to-reference genome match was 98.70%, the average depth of coverage was 29X, and the genome coverage was 97.15%. The above results indicate that the sequencing data are reliable and can be used for SNP and InDel analysis.
Mutations that were identical to those of the wild type were filtered out. After filtering, the M1 mutant had a total of 101 SNPs and 402 InDels, and the M2 mutant had a total of 113 SNPs and 424 InDels. The mutation frequencies of M1 and M2 were calculated to be 2.39 × 10–6 and 2.56 × 10–6 (Calculated according to reference30), respectively. The chromosome with the highest number of mutations was Chr7, followed by Chr9 and Chr12, and the chromosomes with the lowest number of mutations were Chr8 and Chr6 (the distribution of the SNPs and InDels on each chromosome is shown in Supplementary Figure S1). SNP variants was present at the highest levels in downstream regions, followed by upstream regions, exon regions and intergenic regions. Meanwhile, a large proportion of the InDel variants were found in downstream and upstream regions and only about 7% was present in exon regions (Fig. 6A,B). SNP variants occupied 20% of the mutation sites in the exon region, which indicated that the effect of DCEB mutagenesis on the function of rice genes is relatively large.
SNP and InDel variants of the two mutants
In contrast to the wild type, among the two mutants M1 and M2, SNP mutations included 78 A to G transition (36.4%), 66 C to T transition (30.8%); and 23 A > C/T transversion (10.7%), 15 G > C/T transversion (7.0%), 13 C > A/G transversion (6.1%), and 19 T > A/G transversion (8.9%). In both mutants, A: G transitions were the most abundant type of mutations detected by DC bath, followed by the C:T transitions. The most frequent types of transversions in M1 mutants were A > C/T and T > A/G, and in M2 mutants, the most frequent type of transversions was A > C/T (Fig. 6C).
In both mutants, InDels mutations in the coding region were analyzed, and frame-shift were the most common type of mutation, accounting for approximately 37% of the entire coding region (Fig. 6D). The longest insertion mutation in the coding region was a frame-shift mutation at 97 bp, and the longest deletion mutation in the coding region was a stop-gained mutation at 99 bp. The most common mutations are 1–2 bp insertion and deletion mutations (Fig. 6E).
Discussion
Definition of the optimal median lethal dose for DCEB
According to the definition of median lethal dose (LD50), the result should be 50 V and 80 V for 24 h and 140 V for 12 h for DCEB, under which the seed germinations were close to 50%, however we didn’t screen the significant phenotypically mutants in the M2 generation of these treatments; while M1, M2, M3 and other types of suspected mutants appeared in 140 V, 200 V and 140 V for 48 h. The mutagenic effect on rice was further increased along with the increasing voltage and processing time in treatments above the LD50, so that we easily obtained significant phenotypically mutants. Therefore, LD50 is not necessarily suitable for DCEB mutagenesis, and a higher lethal dose (higher voltage or/and longer processing time) can be taken appropriately to be able to increase the probability of mutagenesis.
Effects of γ-irradiation and DCEB on physiological damage
This study investigated the effect of two methods (γ-irradiation and DCEB) on the physiological damages to rice seeds. The results showed that at low doses, γ-irradiation had a positive effect on the germination potential and germination index (GI), while high doses of γ-irradiation significantly reduced the shoot length and vigor index, which is consistent with the results of previous studies31,32,33. However, rice seeds treated with γ-irradiation in this study all had significantly higher panicle numbers and lower filled grain percentage than controls, especially after a dose of 250 Gy. These results may be due to the high doses of γ-irradiation causing pollen sterility34,35. In contrast, the rice seeds treated with the DCEB method all had extremely significantly lower values for rice seed viability, and the performance of the contemporary agronomic trait was essentially indistinguishable. The result suggests that γ-irradiation mainly affects contemporary agronomic traits, whereas the DCEB mainly affects rice seed vigor.
The effect of DCEB mutagenesis on gene function
In present study, SNPs were found to be the most frequent mutation type, with C: T > A: G being the most frequent type of transitions, which is consistent with the results of other common mutagens. Transitions occurred more frequently than transversions, with an observed transition/transversion ratio of 2.0. For InDels, 1–2 bps insertions or deletions were detected most frequently, and the longest insertion in the coding region was a 97 bps frameshift mutation, while the longest deletion was a 100 bps deletion with a stop codon gain mutation. The DCEB method produced a large proportion of mutations detected in the exon regions of genes, particularly for SNP variants, with 20 (19.8%) mutations on M1 and 23 (20.4%) on M2 were screened.
The advantages of DCEB
Compared with traditional irradiation mutagenesis methods36, DCEB only needs electrophoresis instrument to mutagenize, which has the advantages of low requirements for instruments and equipment, simple, convenient and flexible operation, safe experimental environment, low experimental cost, etc. According to the reported studies, the number of mutations in exon regions is less than 10% of the genome mutations caused by γ-radiation37,38, DCEB can cause about 20% exon mutation, this result suggests that the effect of DCEB mutagenesis on gene function in rice is relatively large, especially the mutagenic effect is apparent at a treatment time of 48 h. It provides a new method for future mutagenesis breeding in rice and even in other crops.
Conclusions
The present study provides a new mutagenesis method by using DCEB. We identified three mutants (M1, M2 and M3) from M2 generation and did the WGR for M1 and M2. M1 had a total of 101 SNPs and 402 InDels, and the M2 had a total of 113 SNPs and 424 InDels, indicating that insertional or deletion mutations were the main type of mutations as induced by DCEB. The mutation frequencies were calculated at 2.39 × 10–6 and 2.56 × 10–6 for M1 and M2, respectively. Meanwhile, the exon region of mutant genes occupied a large proportion, especially for SNP variants, which occupied about 20% of the mutation sites in the exon region. The DCEB may provide a simple, convenient, safe, flexible, and low-cost method for future mutagenesis breeding in rice and even in other crops.
Materials and methods
Experimental materials
In this study, XiangGeng 365, a cultivated variety rice (Oryza sativa L. ssp. japonica) was used as experimental material. We obtained the seeds from the National Engineering Research Center of Plant Space Breeding, South China Agricultural University (Guangzhou, China).
DCEB treatment of seeds
Plump seeds of XiangGeng 365 were soaked in water for 24 h and disinfected with 3–6% sodium hypochlorite for 30 min. For each DCEB treatment, 400 seeds were selected and placed in an electrophoresis tank lined with filter paper and buffered for various processing time, after which the buffer residue was removed by washing with water. Then, the seeds were placed on petri dishes lined with filter paper for germination, and physiological indexes such as the GR and GI were calculated. Plants, named as M1 generation that germinated normally were transplanted to the field (Fig. 7), harvested when mature and examined for agronomic traits including plant height, filled grain percentage and thousand grain weight; and around one thousand plants of each M2 generation (obtained from the self-crossed M1 plants) were transplanted in the field for screening of mutations.
Protocol for the DC electrophoresis bath. (A) Soak the rice seeds in clean water for 24 h and disinfected with 3–6% sodium hypochlorite for 30 min; (B) the soaked and disinfected seeds were placed in an electrophoresis tank lined with filter paper and buffer, and then start the DC electrophoresis for a certain period of time; (C) After electrophoresis, the buffer residue was removed by washing with water; (D) Seeds were placed on petri dishes lined with filter paper for germination, and (E) Seedlings from these germinated seeds were transplanted to the field.
γ-irradiation of seeds
In February 2021, rice seeds were irradiated in a γ-irradiator provided by Guangzhou Huada Biological Company. Each well of a 450-well plate filled with 1000 seeds were exposed to different doses of γ-irradiation ranging from 50 to 300 Gy with an interval of 50 Gy.
The mutagenic effects induced by γ-irradiation and DCEB on M1 generation
The VI including GR, GI, bud length, and agronomic traits of M1 generations including plant height, flag leaf length and width, panicle weight, panicle length, number of productive tillers, filled grain percentage and thousand-grain weight were compared between the two methods. The mutagenic effects of DCEB and γ-irradiation on the physiological damage to seeds were estimated by quantifying four parameters on M1 plants: GR, bud length, panicle weight and filled grain percentage. The formula for calculating each physiological index is as follows39:
Gt is the number of sprouts per day during the germination terminal period; Dt is the corresponding number of sprouting days; and S is the length of the seedling at germination time t.
Screening of mutants and analysis of its genetic background
In March 2021, rice seeds were treated by DCEB method with seven different voltages (20 V, 50 V, 80 V, 110 V, 140 V, 170 V and 200 V) for 12 h, 24 h and 48 h for each voltage, respectively. A total of 21 treatments, including a control (treating with sterile water and no voltage), were performed. The suspected mutants were obtained from the M2 generations, and were further analyzed for genetic background compared with its wild type (WT) to ascertain whether they are true mutations by using 48 SSR markers provided by the protocol for identification of rice varieties issued by China in 2014, NY/T 1433–201440.
Whole-genome resequencing of mutants
The constructed DNA libraries were sequenced by the Illumina sequence platform at Biomarker Technologies Co., Ltd. (Beijing, China).
Make quality evaluation and filtering for the raw reads (Paired ends) obtained from sequencing to get Clean Reads, which can be used for subsequent bioinformatics. Align Clean Reads with reference genome, perform variation detection and annotation for SNP, InDel of aligned result to perform DNA-level DEGs mining and DEGs function annotation.
Raw sequenced reads or raw reads were obtained from sequencing, which also have low quality reads with adaptors. To ensure bioinformatics quality, raw reads were filtered to get clean reads for subsequent bioinformatics. The raw reads (double-ended sequences) from the sequences were quality assessed and filtered to obtain clean reads, which were compared with the reference genome sequence (MSU 7.0) for subsequent bioinformatics analysis30. The main steps of data filtering are as follows: (1) remove the reads with adapter; (2) filter reads with N content over 10%; (3) remove reads whose base value (with quality value less than 10) is more than 50%.
Ethics approval
Experimental research and field studies on plants, including the collection of plant material, comply with relevant institutional, national, and international guidelines and legislation.
Data availability
All of the material is owned by the authors and/or no permissions are required. The data used to support the findings of this study are available from the corresponding author upon request.
References
Bongaarts, J. FAO, IFAD, UNICEF, WFP and WHO The state of food security and nutrition in the world 2020. Transforming food systems for affordable healthy diets FAO, 2020, 320 p. Popul. Dev. Rev. 47, 558. https://doi.org/10.1111/padr.12418 (2021)
Muthayya, S., Sugimoto, J. D., Montgomery, S. & Maberly, G. F. An overview of global rice production, supply, trade, and consumption. Ann. N. Y. Acad. Sci. 1324, 7–14. https://doi.org/10.1111/nyas.12540 (2014).
Javaria, T. et al. Applications and potential of genome-editing systems in rice improvement: Current and future perspectives. Agronomy 11, 1359. https://doi.org/10.3390/agronomy11071359 (2021).
Wang, W. et al. Genomic variation in 3,010 diverse accessions of Asian cultivated rice. Nature 557, 43–49. https://doi.org/10.1038/s41586-018-0063-9 (2018).
Raina, A. et al. Role of mutation breeding in crop improvement- past, present and future. Asian Res. J. Agricult. 2, 1–13. https://doi.org/10.9734/arja/2016/29334 (2016).
Viana, V. E., Pegoraro, C., Busanello, C. & de Oliveira, A. C. Mutagenesis in rice: The basis for breeding a new super plant. Front Plant Sci. 10, 1326. https://doi.org/10.3389/fpls.2019.01326 (2019).
Rahman, S. M., Takagi, Y., Kubota, K., Miyamoto, K. & Kawakita, T. High oleic acid mutant in soybean induced by X-ray irradiation. Taylor & Francis 58, 1070–1072. https://doi.org/10.1271/bbb.58.1070 (2014).
Stadler, L. J. Mutations in Barley induced by X-rays and radium. Science 68, 186–187. https://doi.org/10.1126/science.68.1756.186 (1928).
Stadler, L. J. Genetic effects of X-rays in maize. Proc. Natl. Acad. Sci. 14, 69–75. https://doi.org/10.1073/pnas.14.1.69 (1928).
Pan, X. et al. Chromatin states responsible for the regulation of differentially expressed genes under (60)Co~gamma ray radiation in rice. BMC Genom. 18, 778. https://doi.org/10.1186/s12864-017-4172-x (2017).
Peter, A., Ramchander, S., Kumar, K. K., Muthamilarasan, M. & Pillai, M. A. Assessment of efficacy of mutagenesis of gamma-irradiation in plant height and days to maturity through expression analysis in rice. PLoS ONE 16, e0245603. https://doi.org/10.1371/journal.pone.0245603 (2021).
Tan, C. et al. Characterization of genome-wide variations induced by gamma-ray radiation in barley using RNA-Seq. BMC Genom. 20, 783. https://doi.org/10.1186/s12864-019-6182-3 (2019).
Ichide, H., Morita, R., Shirakawa, Y., Hayashi, Y. & Abe, T. Targeted exome sequencing of unselected heavy-ion beam-irradiated populations reveals less-biased mutation characteristics in the rice genome. Plant J. 98, 301–314. https://doi.org/10.1111/tpj.14213 (2019).
Peng, X. et al. Characterization and fine mapping of a leaf wilt mutant, m3, induced by heavy ion irradiation of rice. Crop Sci. 59, 2679–2688. https://doi.org/10.2135/cropsci2019.03.0167 (2019).
Yamaguchi, H. Mutation breeding of ornamental plants using ion beams. Breed. Sci 68, 71–78. https://doi.org/10.1270/jsbbs.17086 (2018).
Zhang, J. et al. Time course analysis of genome-wide identification of mutations induced by and genes expressed in response to carbon ion beam irradiation in rice (Oryza sativa L.). Genes-Basel 1, 2. https://doi.org/10.3390/Genes12091391 (2021).
Henry, I. M. et al. Efficient genome-wide detection and cataloging of EMS-induced mutations using exome capture and next-generation sequencing. Plant Cell 26, 1382–1397. https://doi.org/10.1105/tpc.113.121590 (2014).
Sugihara, N., Higashigawa, T., Aramoto, D. & Kato, A. Haploid plants carrying a sodium azide-induced mutation (fdr1) produce fertile pollen grains due to first division restitution (FDR) in maize (Zea mays L.). Theor. Appl. Genet. 126, 2931–2941. https://doi.org/10.1007/s00122-013-2183-9 (2013).
Szurman-Zubrzycka, M. E. et al. HorTILLUS-A rich and renewable source of induced mutations for forward/reverse genetics and pre-breeding programs in Barley (Hordeum vulgare L.). Front. Plant Sci. 9, 216. https://doi.org/10.3389/fpls.2018.00216 (2018).
Futsuhara, Y., Toriyama, K. & Tsunoda, K. Breeding of a new rice variety “reimei” by gamma-ray irradiation. Jpn. Soc. Breed. 1, 7. https://doi.org/10.3389/fpls.2016.00262 (1967).
Ahloowalia, B. S., Maluszynski, M. & Nichterlein, K. Global impact of mutation-derived varieties. Euphytica 135, 187–204. https://doi.org/10.1023/B:EUPH.0000014914.85465.4f (2004).
Horn, L. N., Ghebrehiwot, H. M. & Shimelis, H. A. Selection of novel cowpea genotypes derived through gamma irradiation. Front Plant Sci. https://doi.org/10.3389/fpls.2016.00262 (2016).
Bai, S., Yu, H., Wang, B. & Li, J. Retrospective and perspective of rice breeding in China. J. Genet. Genom. 45, 603–612. https://doi.org/10.1016/j.jgg.2018.10.002 (2018).
Hardwick, S. A., Deveson, I. W. & Mercer, T. R. Reference standards for next-generation sequencing. Nat. Rev. Genet. 18, 473–484. https://doi.org/10.1038/nrg.2017.44 (2017).
Li, F., Shimizu, A., Nishio, T., Tsutsumi, N. & Kato, H. Comparison and characterization of mutations induced by gamma-ray and carbon-ion irradiation in rice (Oryza sativa L.) using whole-genome resequencing. G3: Genes Genomes Genet. 9, 3743–3751. https://doi.org/10.1534/g3.119.400555 (2019).
Arne, T. Electrophoresis of serum globulin: Electrophoretic analysis of normal and immune sera. Biochem. J. 31, 1464–1477. https://doi.org/10.1042/bj0311464 (1937).
Pereira, L., Hoffman, M. & Cremer, N. Electrophoretic analysis of polypeptides immune precipitated from cytomegalovirus-infected cell extracts by human sera. Infect. Immun. https://doi.org/10.1128/iai.36.3.933-942.1982 (1982).
Cunha, E. N., de Souza, M. F. B., Lanza, D. C. F. & Lima, J. A low-cost smart system for electrophoresis-based nucleic acids detection at the visible spectrum. PLoS ONE 15, e0240536. https://doi.org/10.1371/journal.pone.0240536 (2020).
Christoph, L., Jörg, B., Uta, W., Elisabeth, M. & Dominik, F. Clinical chemistry, vitamin, electrophoresis, and hematologic analytes of black-headed ibis (threskiornis melanocephalus). J. Zool. Wildl. Med. 51, 948–957. https://doi.org/10.1638/2020-0030 (2021).
Oono, Y. et al. Genome sequencing of ion-beam-induced mutants facilitates detection of candidate genes responsible for phenotypes of mutants in rice. Mutat. Res. 821, 111691. https://doi.org/10.1016/j.mrfmmm.2020.111691 (2020).
Jiya, M. et al. Effects of gamma irradiation on submergence tolerance of two selected varieties of lowland rice (Oryza sativa L.). GSC Biol. Pharm. Sci. 02, 031–037. https://doi.org/10.30574/gscbps.2018.2.3.0017 (2018).
Abdelnour-Esquivel, A., Perez, J., Rojas, M., Vargas, W. & Gatica-Arias, A. Use of gamma radiation to induce mutations in rice (Oryza sativa L.) and the selection of lines with tolerance to salinity and drought. In Vitro Cell. Dev. Biol. Plant 56, 88–97. https://doi.org/10.1007/s11627-019-10015-5 (2020).
Nurhidayah, S., Firmansyah, E. & Rahayu, S. in IOP Conference Series: Earth and Environmental Science Vol. 672 (2021).
Lalitha, R. et al. Radiation effect on germination and seedling traits in rice (Oryza sativa L.). Electron. J. Plant Breed. https://doi.org/10.5958/0975-928x.2019.00133.9 (2019).
Julia, T., Renuka, T., Nanita, H. & Jambhulkar, S. Mutagenic effectiveness and efficiency of Gamma rays in Indian mustard (Brassica juncea L. Czern and Coss). Int. J. Curr. Microbiol. Appl. Sci. 7, 3376–3386. https://doi.org/10.20546/ijcmas.2018.703.390 (2018).
Zhang, L. et al. Characterization of γ-radiation-induced DNA polymorphisms in the m1 population of the japonica rice variety gaogengnuo by whole-genome resequencing. Russ. J. Genet. 56, 693–705. https://doi.org/10.1134/S1022795420060149 (2020).
Li, S. et al. Frequency and type of inheritable mutations induced by gamma rays in rice as revealed by whole genome sequencing. J. Zhejiang Univ.-Sci. B 17, 905–915. https://doi.org/10.1631/jzus.B1600125 (2016).
Zhang, L. et al. Genome-wide analysis of radiation-induced mutations in rice mutant. Acta Laser Biol. Sin. 30, 59–66. https://doi.org/10.3969/j.issn.1007-7146.2021.01.008 (2021).
Wang, P., Li, D., Wang, L. J. & Adhikari, B. Effect of high temperature intermittent drying on rice seed viability and vigor. Int. J. Food Eng. https://doi.org/10.1515/ijfe-2016-0433 (2017).
Xu, Q. et al. Protocal for identification of rice varieties-SSR marker method. iN Edited by MOA China, NY/T 1433–2014 (2014).
Acknowledgements
This work was supported by the Key R&D Program of Guangzhou Science and Technology Project (Grant No. 202206010015). The authors would thank Biomarker Technologies (Guangzhou, China) for the resequencing service, and thank American Journal Experts for their valuable language service (verification code: CD97-A658-B3C1-2D62-355F).
Funding
This work was supported by Guangzhou Key Research and Development Project (Grant No. 202206010015).
Author information
Authors and Affiliations
Contributions
M.M.Z. and M.H. conceived and designed the experiments and wrote the paper. T.Z., X.Y.W., T.S. and L.X.W. conducted mutant screening, and characterization of mutants. J.F.W., WMX and M.H. conducted the experiments and performed data analysis. M.M.Z., M.H., T.G. and H.W. drafted proposals and corrected the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zou, M., Tong, S., Zou, T. et al. A new method for mutation inducing in rice by using DC electrophoresis bath and its mutagenic effects. Sci Rep 13, 6707 (2023). https://doi.org/10.1038/s41598-023-33742-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-33742-7
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.