Abstract
Loss of seed shattering has been regarded as a key step during crop domestication. Mutagenesis contributes to the development of novel crop cultivars with a desired seed-shattering habit in a relatively short period of time, but also to uncovering the genetic architecture of seed shattering. ‘Minamiyutaka’, a non-shattering indica rice cultivar, was developed from the easy-shattering cultivar ‘Moretsu’ by mutation breeding via gamma-ray irradiation. In present study, we observed significant differences in shattering habit, breaking tensile strength, and abscission zone structure between ‘Moretsu’ and ‘Minamiyutaka’. Whole-genome mutation analysis of ‘Minamiyutaka’ newly identified a 13-bp deletion causing defective splicing in exon 3 of the OsSh1 gene which has previously been referred to as a candidate for controlling seed shattering. Using CRISPR/Cas9 gene editing, we demonstrated that loss-of-function mutation in OsSh1 causes non-shattering in rice. Furthermore, gene expression analysis suggests that OsSh1 may function downstream of qSH1, a known key gene involved in abscission zone differentiation. Nucleotide diversity analysis of OsSh1 in wild rice accessions and cultivars revealed that OsSh1 has been under strong selection during rice domestication, and a missense mutation might have contributed to the reduction of seed shattering from the wild progenitors to cultivated rice.
Similar content being viewed by others
Introduction
Loss or reduction of seed shattering is one of the key features in crop domestication, since it reduces yield losses from shattering and improves harvesting efficiency1,2. Asian cultivated rice (Oryza sativa L.), one of the most important cereal crops in the world, was domesticated from its Asian wild ancestor (O. rufipogon) more than ten thousand years ago3,4. In cultivated rice, the seed-shattering habit of wild rice was lost during domestication, and shattering degrees show a wide variation5. Generally, indica cultivars exhibit relatively easy shattering, whereas most japonica cultivars exhibit hard shattering5.
In rice, seeds shattering is implemented by an abscission zone in the junction of sterile lemma and pedicel6. The abscission zone, which is composed of one or two layers of small, isodiametrically shaped cells with thin cell walls, is formed at the young panicle development stage approximately 16–20 days before heading7, and gradually degrade after flowering8. The morphology and degradation behavior of the abscission zone differs in different rice varieties7,8,9. Seed shattering is controlled by a complex regulatory network6,9 and quantitative trait loci (QTLs) for seed shattering have been detected on almost all rice chromosomes10,11,12,13,14,15. Map-based cloning and genetic complementation experiments have revealed two domestication related mutations that facilitate the reduction of seed shattering. One mutation is a single amino acid substitution from Lysine residue to Asparagine at position 79 (K79N) in the gene SH4/SHA1, a transcription factor with an Myb3 DNA binding domain2,16. This mutation is responsible for the reduction of seed shattering from the wild progenitors to cultivated rice. Wild rice has a complete layer of abscission cells, while the SH4 domestication allele contributes to absent abscission cells near the vascular bundle, leading to incomplete development of the abscission zone in cultivated rice2. Another mutation is a single nucleotide polymorphism (SNP) in the 5′ regulatory region of the qSH1 gene, an ortholog of the Arabidopsis homeobox gene REPLUMLESS (RPL)5,17. This mutation results in the absence of abscission zone formation and thus loss of seed shattering in a subset of temperate japonica cultivar5,18. Except for SH4 and qSH1, many minor QTLs involved in seed shattering have not been cloned10,11,12,13,14,15.
Using artificial mutagenesis, some mutants with changed seed-shattering habit were obtained and the causal genes were identified or speculated19,20,21,22,23,24. An insertion of a > 4-kb fragment in YABBY-like gene OsSh1, an ortholog of Sh1 involved in seed shattering in sorghum, was thought to be the cause of the non-shattering phenotype in a rice mutant19,25. More recently, a genomic segment deletion containing ObSH3, an ortholog of Sh1, was revealed to cause the loss of seed dispersal in populations of African cultivated rice (O. glaberrima Steud.)26. Loss-of-function mutations in the APETALA2 (AP2) gene, SHAT122 or SHH1/SNB20, can inhibit the expression of qSH1, hence leading to loss of shattering. Furthermore, the natural variations in SHH1 were speculated to be associated with the domestication and improvement of seed shattering and yield-related traits in rice20. In wheat, domestication-related Q gene, involved in controlling seed shattering, is also an AP2-like gene27. These results suggest that the regulatory network controlling seed shattering is extensively conserved in grain crops, and that mutagenesis can facilitate the uncovering of the complex genetic architecture of seed shattering.
‘Minamiyutaka’ is a non-shattering indica rice cultivar for whole crop silage and is broadly cultivated in Japan. ‘Minamiyutaka’ was developed from a mutant obtained through gamma-ray irradiation of an easy-shattering cultivar ‘Moretsu’28. The heading and maturing time, and the morphological characteristics of both cultivars are almost the same28. In contrast, the reported mutants have pleiotropic phenotypes such as altered development of spikelet or inflorescence, besides the reduction of seed shattering. For example, shat1 shows spikelet and inflorescence developmental defects, shat2 shows smaller seeds22, and while ssh1 shows larger seeds20. We therefore hypothesized that a different causal gene was responsible for the non-shattering habit in ‘Minamiyutaka’. Identification of the causative mutation underlying the loss of shattering in ‘Minamiyutaka’ will be valuable for improving rice seed shattering without affecting other traits using molecular breeding strategies, and will also contribute to uncovering the complex regulatory pathways of seed shattering.
Recently, an approach combining bulked segregant analysis with whole-genome resequencing has dramatically accelerated the process of identifying candidate genes20,21,29. Alternatively, since gamma-ray-irradiation induces less than one hundred mutations in the whole genome of rice, and since, furthermore, most of them are in the intergenic and intronic regions, with only several mutations highly or moderately impacting the gene function30, we therefore supposed that the detection of the mutations in the whole genome and analysis of the effects of the mutations on gene function could directly identify the causal gene for an altered phenotype induced by gamma-ray-irradiation. In the present study, as expected, we successfully uncovered the putative causal gene for the loss of shattering in ‘Minamiyutaka’ with this approach and confirmed it by CRISPR (clustered regularly interspaced short palindromic repeats)/Cas9 gene editing.
Results
Seed-shattering characteristics in the parental cultivars
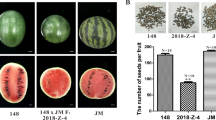
The cultivar ‘Moretsu’ has an easy-shattering habit, such that up to 40 of grains were shattered after grasping a panicle by hand at the maturity stage (Fig. 1a). In contrast, almost no grains were shattered in ‘Minamiyutaka’, confirming that it is of non-shattering. Subsequently, we evaluated the pulling strength (PS) and bending strength (BS), i.e., breaking tensile strength for detachment of a seed from the pedicel by pulling and bending, respectively. PS values of ‘Moretsu’ and ‘Minamiyutaka’ were 124.4 ± 29.6 and 211.2 ± 30.9 gf, respectively, showing a significant difference (P < 2.2 × 10–16) (Fig. 1b). BS values of ‘Minamiyutaka’ were 40.5 ± 18.9 gf, approximately three times that of ‘Moretsu’ (13.5 ± 6.2 gf) (Fig. 1c). These results revealed the strong resistance to shattering in ‘Minamiyutaka’. It should be noted that the PS and BS values may also be affected by the health status of a pedicel. If a pedicel was not well developed or was damaged, its PS or BS value would be small. While we endeavored to select spikelets that appeared healthy, standard deviation was still large for all evaluated spikelets of each cultivar.
Comparison of seed shattering characteristics in the rice cultivars ‘Moretsu’ and ‘Minamiyutaka’. (a) Seed shattering habits of rice panicles in ‘Moretsu’ (Left) and ‘Minamiyutaka’ (Right). Photos were taken after grasping rice panicles. Bars = 5 cm. (b) Comparison of pulling strength (PS). P values were determined by Student’s t test. In the violin plots, the violin shape indicates the kernel-density curve, the white node in the center indicates the average value, and the black box inside the violin indicates a box-and-whisker plot. Violin plots were created using R software62 and ggplot2 package63. (c) Comparison of bending strength (BS). P values were determined by Welch’s t test. (d) Longitudinal Sects. (2-μm) across the abscission zone of ‘Moretsu’ (Left) and ‘Minamiyutaka’ (Right). The sections were stained by toluidine blue. Bars = 100 μm. AZ abscission zone, P pedicel, SL sterile lemma, VB vascular bundle. (e) shows scanning electron microscopy photos of the fracture surface of the grain base of ‘Moretsu’ (Left) and ‘Minamiyutaka’ (Right) after detachment of grains. Bars = 100 μm. (f) shows close-up scanning electron microscopy photos corresponding to the yellow boxes in (e). Bars = 10 μm.
To distinguish precisely the differences in anatomical aspects of abscission zone between the two cultivars, longitudinal sections of spikelets at heading stage were observed. In ‘Moretsu’, isodiametrically shaped and thin-walled abscission zone cells were aligned completely and transversely in the basal area near sterile lemmas except in the region near the vascular bundle. In contrast, the alignment of abscission cells was discontinuous and incomplete in ‘Minamiyutaka’ (Fig. 1d).
The interface where a mature grain separates from the pedicel was observed using a scanning electron microscope (SEM). The microstructure of the surface of the grain base of ‘Moretsu’ was clearly different from that of ‘Minamiyutaka’ (Fig. 1e,f). ‘Moretsu’ was relatively smooth, but ‘Minamiyutaka’ was broken and rough. It is obvious that a stronger force was needed to remove grain from the pedicels in ‘Minamiyutaka’ than in ‘Moretsu’ and this led to a rougher surface in ‘Minamiyutaka’.
Segregation of non-shattering phenotype
The shattering degrees of F2 plants (n = 100) derived for the cross between ‘Moretsu’ and ‘Minamiyutaka’ were evaluated by grasping panicles by hand at the maturity stage. As a result, 27 and 73 plants showed a non-shattering and an easy-shattering phenotype, respectively (Fig. 2d). This segregation ratio fits a 1:3 ratio as determined by a chi-square test (χ2 = 0.64, P = 0.42), indicating that a single recessive locus was the cause of the non-shattering phenotype in ‘Minamiyutaka’.
Whole-genome resequencing identified a 13-bp deletion in OsSh1 as candidate causal mutation leading to loss of seed shattering in ‘Minamiyutaka’. (a) Integrative Genomics Viewer screenshot of the region containing the 13-bp deletion for ‘Moretsu’ (Upper Graph) and ‘Minamiyutaka’ (Lower Graph). Genome position and gene annotation are shown at the top and bottom, respectively. The 13-bp deletion causes defective splicing in exon 3. (b) Sanger sequencing chromatograms of the region containing the 13-bp deletion for ‘Moretsu’ (Upper Graph) and ‘Minamiyutaka’ (Lower Graph). (c) Sequence alignment of the deduced amino acid sequences of OsSh1 in ‘Moretsu’ and ‘Minamiyutaka’. The 13-bp deletion causes exon 3 skipping and a reading frameshift leading to a putative translational termination codon. Zinc finger domain and YABBY domain are underlined in red and green, respectively. (d) Seed shattering and genotype of the OsSh1 gene in an F2 population derived from the cross between ‘Moretsu’ and ‘Minamiyutaka’. W/W (n = 21) represents a homozygous allele for ‘Moretsu’, m/m (n = 27) represents a homozygous allele for ‘Minamiyutaka’, and W/m (n = 52) for a heterozygous allele.
Identification of a candidate causal mutation using whole-genome resequencing
Using next-generation sequencing, a total of 78.3, and 79.2 million sequence reads (each 150 bp) were obtained for ‘Moretsu’ and ‘Minamiyutaka’, respectively (Supplementary Table S1 online). After removing low-quality, unpaired and duplicated reads, about 84.0% of the clean reads were mapped to the ‘Nipponbare’ reference genome. The average coverages were 20.2 times.
Homozygous mutations including 62 SNPs, 7 deletions (< =3 bp), and 5 insertions (< =5 bp) unique to ‘Minamiyutaka’ were identified (Supplementary Table S2 online). No structural variation (SV) was detected. Mutation annotation analysis revealed only one mutation (Mutation ID: MN19, 13-bp deletion) with high impact on gene function (gene ID: Os03t0650000-01) (Fig. 2a). This mutation was validated by Sanger sequencing (Fig. 2b). In a previous study, this gene has been identified as an ortholog of Sh1 controlling seed shattering in sorghum, and has been named OsSh125. OsSh1, also called as OsYABBY231, belongs to the YABBY gene family composed of an N-terminal zinc finger domain and a C-terminal YABBY domain (helix-loop-helix motif)32. Analysis of the protein coding sequence (CDS) region revealed that the 13-bp deletion caused exon 3 (127 bp) skipping and a reading frameshift leading to a putative translational termination codon (PTC) which can result in a loss of YABBY domain in ‘Minamiyutaka’ (Fig. 2c, Supplementary Fig. S1 online). We therefore inferred the 13-bp deletion in OsSh1 to be the candidate causal mutation resulting in loss of seed shattering in ‘Minamiyutaka’.
A Cleaved Amplified Polymorphic Sequences (CAPS) marker designed based on the restriction enzyme site in the 13-bp deletion of OsSh1 was used to perform genotyping analysis of the F2 plants. As a result, 21, 52, and 27 plants were homozygous wild-type, heterozygous, and homozygous mutant-type, respectively (Fig. 2d). All the wild-type and heterozygous plants exhibited the easy-shattering phenotype, whereas all the mutant-type plants were of non-shattering phenotype (Fig. 2d). These results further suggested that OsSh1 is a strong candidate gene involved in seed shattering.
Genetic complementation using CRISPR/Cas9-based gene editing
‘Teqing’, a Chinese high-yielding indica rice cultivar with an easy-shattering phenotype, was used for OsSh1 gene editing. In T0 plants, 8 genotypes with 11 different mutant alleles and 10 genotypes with 11 different mutant alleles were detected in the target regions of the exon 1 and exon 2 of OsSh1, respectively (Supplementary Table S3 online). T0 transgenic plants were self-pollinated and only partial T1 lines were grown due to space limitation in the greenhouse. For seed-shattering evaluation, we selected three independent lines, each of which had more than four plants with homozygous Cas9-free mutation (Fig. 3a). All of them have one-base InDel (deletion/insertion) in exon1 or exon2, which may cause a reading frameshift and lead to loss of gene function. PS values of the three OsSh1-edited lines, i.e., T_Cas9-1, T_Cas9-2 and T_Cas9-3, were 180.0 ± 47.0, 195.9 ± 43.1, and 218.5 ± 46.5gf, respectively, significantly higher than that of ‘Teqing’, 110.1 ± 45.0 gf (P < 2.2 × 10–16) (Fig. 3b). The BS values of them were 26.6 ± 11.8, 38.4 ± 16.9, and 38.0 ± 15.4 gf, respectively, approximately 2 to 5 times higher than that of ‘Teqing’ (8.0 ± 5.9 gf) (P < 2.2 × 10–16) (Fig. 3c). The fracture surface of the grain base was investigated using an SEM. We found that ‘Teqing’ had a smooth fracture surface (Fig. 4a), whereas all the OsSh1-edited lines had a broken and rough fracture surface (Fig. 4b–d). These results indicated that OsSh1-edited lines had a non-shattering habit.
Comparison of shattering degrees among ‘Teqing’ (wild type) and OsSh1-edited lines in T1 generation at maturity stage. (a) Sequence alignments of the sgRNA target regions showing altered bases (in yellow) in different mutant lines. T_Cas9-1, T_Cas9-2, and T_Cas9-3 were of homozygous Cas9-free mutations derived from the T0 plants of 18–19-4b, 18–20-19a, and 18–26-14a (Supplementary Table S3 online), respectively. (b) Comparison of pulling strength (PS). P values were determined by pairwise comparisons using t tests with pooled SD and a Bonferroni correction. (c) Comparison of bending strength (BS). P values were determined by pairwise comparisons using Wilcoxon rank sum test and a Bonferroni correction. In the violin plots, the violin shape indicates the kernel-density curve, the white node in the center indicates the average value, and the black box inside the violin indicates a box-and-whisker plot. A total of 100 grains (n = 100) from four panicles were measured for each line. Violin plots were created using R software62 and ggplot2 package63.
OsSh1 expression profile
The expression profile of OsSh1 in various rice tissues deposited in the rice gene expression database RiceXPro33 showed that OsSh1 was strongly expressed in leaf blade, leaf sheath and stem rather than in root during vegetative stage (Supplementary Fig. S2 online). During reproductive stage, the OsSh1 was mainly expressed in pistil, lemma, palea, ovary, embryo and young endosperm rather than in anther and old endosperm.
OsSh1 functions downstream of qSH1
To examine the gene expression of OsSh1 in developing panicles of ‘Moretsu’ and ‘Minamiyutaka’, we performed reverse transcription quantitative PCR (RT-qPCR) analysis using three primer sets P1, P2, and P3 (Fig. 5a). All the results showed a significantly lower level of OsSh1 expression in ‘Minamiyutaka’ than in ‘Moretsu’ (Fig. 5b–d) (P < 0.01). Since qSH1 is the key gene involved in abscission layer formation5,22, its transcript level was analyzed. However, no significant differences were observed between these two cultivars (Fig. 5e). These results led us to suppose that OsSh1 might function downstream of qSH1, or alternatively, that OsSh1 and qSH1 were in different pathways, respectively. To address this, firstly, we developed a non-shattering chromosome segment substitution line (CSSL) ‘Takanari-qsh1’ which harbors a dysfunctional allele of qSH1 derived from a japonica rice cultivar ‘Koshihikari’ in the ‘Takanari’ genetic background34. In contrast, ‘Takanari’ is an easy-shattering indica rice cultivar with a functional allele at qSH121,34. PS and BS values of the ‘Takanari-qsh1’ were 188.5 ± 56.2 and 22.6 ± 13.0, respectively, significantly higher than those of ‘Takanari’, 119.1 ± 38.2 and 12.3 ± 10.5 (Supplementary Fig. S3 online), confirming the dysfunctional allele of qSH1 causing reduction of seed shattering. Hence, we investigated the gene expressions of OsSh1 and qSH1 in the young panicles of these two cultivars/lines. As a result, the transcript levels of qSH1 in ‘Takanari-qsh1’ were significantly lower than those in ‘Takanari’ (Fig. 6a), since an SNP in the 5′ regulatory region of the dysfunctional qSH1 can decrease its expression5. Interestingly, OsSh1 in ‘Takanari-qsh1’ also notably decreased as compared with ‘Takanari’ (Fig. 6b). These data suggest that OsSh1 functions downstream of qSH1.
RT-qPCR analysis of OsSh1 and qSH1 in young panicles of ‘Moretsu’ and ‘Minamiyutaka’. (a) The regions of OsSh1 were targeted using RT-qPCR with three primer sets, P1, P2, and P3, indicated by the black double-sided arrows. Note that the elements are not drawn to scale. Relative expression levels of OsSh1 and qSH1 are shown in (b)–(e). Data were normalized to ‘Moretsu’. Bars indicate mean values ± standard deviation (n = 4). Double asterisks denote a significant difference at P < 0.01 using Student's t test.
Nucleotide diversity in OsSh1
To investigate the nucleotide diversity of OsSh1 in wild and cultivated rice, we aligned the nucleotide sequences covering the entire OsSh1 gene (3,688 bp), a 1,653-bp 5′-flanking region, and a 1,354-bp 3′-flanking region, from 37 accessions of O. rufipogon, 84 indica varieties, 63 temperate japonica varieties, and 41 tropical japonica varieties (Supplementary Table S4 online). The nucleotide diversity (π) of OsSh1 was the highest in wild rice, moderate in indica rice, and lowest in japonica rice (Fig. 7a). The percentages of nucleotide diversity in indica/O. rufipogon and japonica/O. rufipogon were 23.2 and 1.5%, respectively, which are far below the percentages (53.3 and 20.0%, respectively) at the whole-genome level4,35. Furthermore, Tajima’s D based on the OsSh1 locus was negative and significantly (P < 0.01) different from neutral expectation in O. sativa (Supplementary Table S5 online). These results indicate that the OsSh1 was subjected to strong directional selection during rice domestication.
Nucleotide diversity analysis of OsSh1 in wild and cultivated rice. The consensus sequences from 40 accessions of O. rufipogon, 96 indica varieties, 146 temperate japonica varieties, and 46 tropical japonica varieties were downloaded from TASUKE+. (a) Sliding-window analysis of nucleotide polymorphism (π) of OsSh1. The values were calculated for each sliding window of 100 bp with an increment of 10 bp. (b) shows a missense variant (c to t) at position + 70 nearly fixed in rice varieties. Blue boxes represent exons, grey boxes represent UTRs, and thin black lines indicate flanking regions or introns. The transcription start site of exon 1 is taken as + 1 position. Note that the elements are not drawn to scale. Pie graphs show the allele frequencies of OsSh1-c70 and OsSh1-t70 in O. rufipogon, indica varieties, and japonica varieties.
Variation annotation analysis revealed that a single nucleotide variant, c to t, at position + 70 relative to the translation start site (c70t), caused a single amino acid substitution from a Leucine residue (L) to Phenylalanine (F) at position 24 (L24F) (Fig. 7b). We named these two alleles OsSh1-c70 and OsSh1-t70, respectively. No other variant causing amino acid change was identified. The OsSh1-c70 was the major allele in O. rufipogon (frequency = 80.8%), in contrast, OsSh1-t70 was the major allele in indica rice (frequency = 91.7%), and no OsSh1-c70 was found in japonica cultivars (Fig. 7b). To confirm this result, we checked the allele frequencies of OsSh1 in 436 accessions of O. rufipogon (Supplementary Table S6 online), and found the allele frequencies of OsSh1-c70 in Or-I, Or-II and Or-III to be 71.0 to 88.0%, or 81.7% overall. OsSh1-c70 was found to be distributed extensively in the original producing area, while OsSh1-t70 was not found in Western India, Western New Guinea, Papua New Guinea or Australia (Supplementary Fig. S4 online). Subsequently, we analyzed 1,774 indica cultivars and 844 japonica cultivars (Supplementary Table S7 online). The frequencies of OsSh1-t70 in indica and japonica cultivars was 99.1% and 97.8%, respectively. Furthermore, OsSh1-c70 was only found in the tropical subpopulation with a frequency at 5.0%, and was not found in subtropical and temperate subpopulations. The reason that no OsSh1-c70 was found in the japonica cultivars collected in TASUKE + may be due to the limited number of tropical japonica varieties.
Amino acid alignment of YABBY2 proteins in different monocot and dicot plants indicated that the Zinc finger domain and YABBY domain were highly conserved (Supplementary Fig. S5 online). Leucine residue at position 24 (L24) was almost completely conserved in YABBY2 proteins of all seed plants (Supplementary Fig. S5 online), and the other YABBY genes as well31. These results suggested that amino acid substitution L24F in OsSh1 protein might partially affect its function in controlling seed shattering.
Discussion
Non-shattering phenotype in ‘Minamiyutaka’ is defect in abscission zone formation
In the present study, we found remarkable improvement of resistance to shattering in ‘Minamiyutaka’ compared to its original cultivar ‘Moretsu’ (Fig. 1). This strong shattering resistance in ‘Minamiyutaka’ may be due to the defective abscission zone formation, as in the reported non-shattering mutants such as shat122, shat222, and ssh120. These results confirm that abscission zone formation is of key importance for controlling seed shattering.
Whole-genome resequencing may directly identify the causative mutation
Genetics analysis suggested that a single recessive locus was responsible for the non-shattering phenotype in ‘Minamiyutaka’ (Fig. 2), which may facilitate cloning of the candidate gene. Through whole-genome sequencing of ‘Moretsu’ and ‘Minamiyutaka’, we newly identified a 13-bp deletion in the OsSh1 gene of ‘Minamiyutaka’ (Fig. 2). The 13-bp deletion in OsSh1 caused exon 3 skipping and introduced a putative PTC in the coding regions of mRNA, which can lead to generation of a non-functional OsSh1 protein (Fig. 2). The PTC-containing mRNAs are known to be degraded via nonsense-mediated mRNA decay (NMD)36,37. We consistently observed a notable reduction in OsSh1 expression in ‘Minamiyutaka’ (Fig. 5), suggesting mRNAs with a PTC may be degraded. Furthermore, this 13-bp deletion was revealed to be completely associated with the non-shattering phenotype in an F2 population (Fig. 2). These results strongly indicate that OsSh1 was a candidate gene. These results also suggest that whole-genome sequencing of only the mutant generated by gamma-ray irradiation and its original wild type may directly identify the candidate causal gene. This strategy may be more cost-effective and time-saving than the bulked segregant analysis21.
Loss-of-function mutation in OsSh1 can result in non-shattering phenotype
The YABBY gene family, specific to seed plants, is expressed in the abaxial region of leaf primordia and floral organs, and promotes abaxial cell fate and lateral organ development in Arabidopsis38,39. YABBY genes in poaceae such as rice, maize and wheat are not expressed in a polar manner40,41,42, suggesting that the roles of YABBY genes have diversified during the evolution of plants. There are 8 YABBY genes in rice31, and they also show organ-specific expression patterns31,43. In the present study, OsSh1/OsYABBY2 was revealed to be expressed in all organs except for the roots and anthers, which is similar to a previous study31. In sorghum, Sh1, was identified as the gene controlling seed shattering using the mapping populations derived from the crosses between complete-shattering wild sorghum and non-shattering domesticated sorghum25. Interestingly, Lin et al.25 found that syntenic blocks containing Sh1 correspond to the seed-shattering related QTLs in rice44, maize45, and millet46, indicating that Sh1 genes for seed shattering were under parallel selection during rice, maize, and sorghum domestication. In the present study, we used the CRISPR-Cas9 genome editing system to induce loss-of-function mutations in OsSh1 in the easy-shattering indica rice cultivar ‘Teqing’. The results of breaking tensile strength and microstructure revealed a remarkable improvement in resistance to shattering in the OsSh1-edited lines compared to the wild type (Figs. 3, 4). We therefore provide direct experimental evidence for the first time that OsSh1 is involved in controlling seed shattering in Asian rice. Furthermore, the novel allele of OsSh1 in ‘Minamiyutaka’ may be applied in rice breeding program.
OsSh1 functions downstream of qSH1
Thus far, the pathway controlling the development of abscission zone located between the sterile lemma and the pedicel is still unclear. It has been revealed that qSH1 activity depends on SHAT1, SH4 and SHH1, indicating that qSH1 functions downstream of these genes20,22. In Arabidopsis, the YABBY genes such as FILAMENTOUS FLOWER (FIL) and YABBY3 (YAB3) play important regulatory roles in forming stripes of valve margin tissue that allow the fruit to shatter at maturity stage47. Furthermore, the expression pattern of FIL is regulated by RPL (the ortholog of qSH1 in rice)47. Our present study has revealed that the expression of qSH1 is not affected by loss-of-function of OsSh1 (Fig. 5), whereas the expression of OsSh1 decreases significantly when qSH1 is dysfunctional (Fig. 6). These results suggest that OsSh1 may function downstream of qSH1, similar to the pathway in Arabidopsis47 .
Artificial selection for OsSh1 in rice domestication
Resequencing 50 accessions of cultivated rice and wild rice (O. rufipogon and O. nivara) suggested that OsSh1 is one of the putative artificially selected genes48. We performed nucleotide diversity analysis and Tajima’s D test in many more wild rice accessions (O. rufipogon) and cultivars, and further confirmed very strong selection of OsSh1 during rice domestication (Fig. 7, Supplementary Table S5 online). Furthermore, only one missense variation c70t was identified in all accessions. Since the amino acid residue L24 harbored in the OsSh1-c70 allele was broadly conserved in seed plant species (Supplementary Fig. S5 online) as well as in the YABBY gene family31, OsSh1-c70 might be an ancestral allele. The OsSh1-c70 is the major allele in any ecotypes of O. rufipogon, and is randomly distributed geographically, whereas OsSh1-t70 is mainly distributed in Southern China, Southeast Asia and Eastern South Asia (Supplementary Table S6 online, Supplementary Fig. S4 online), indicating that the c70t mutation might have occurred spontaneously in one of these regions and then gradually spread out from there. Interestingly, OsSh1-c70 has become a rare allele in indica and tropical japonica cultivars which are cultivated in geographically similar regions where O. rufipogon is distributed, but has disappeared in subtropical and temperate japonica cultivars (Supplementary Table S7 online). Based on these results, we speculate possible evolutionary scenarios for OsSh1 whereby a common ancient rice with the OsSh1-t70 allele was first domesticated before the indica-japonica differentiation, while a few indica and tropical japonica rice accessions were crossed to local wild rice with the OsSh1-c70 allele after differentiation. To clarify this, it might be necessary to further investigate the detailed genomic variations and DNA polymorphisms among cultivated rice containing the OsSh1-c70 allele and wild rice.
Materials and methods
Materials and growth condition
‘Minamiyutaka’ and ‘Moretsu’ were crossed to generate an F2 population. A CSSL named ‘SL1303’, carrying a genomic region containing a dysfunctional allele of qSH1 and a functional allele of Semidwarf1 (SD1) from a japonica rice cultivar ‘Koshihikari’34 within the background of an indica rice cultivar ‘Takanari’, was kindly provided by Dr. Toshio Yamamoto (NARO)34. ‘Takanari’ is short-culm and easy-shattering, in contrast to which ‘Koshihikari’ and ‘SL1303’ show non-shattering and a long-culm phenotype. We backcrossed the ‘SL1303’ to ‘Takanari’ to develop a new non-shattering CSSL named ‘Takanari-qsh1’, whose morphological characteristics including the height of the culm is similar to that of ‘Takanari’.
Evaluation of shattering degree
To evaluate the seed shattering of ‘Moretsu’, ‘Minamiyutaka’, gene-edited plants (including wild types), ‘Takanari’, and ‘Takanari-qsh1’, panicles from the primary tillers were harvested at maturity stage and naturally air dried for more than two weeks in a room. PS and BS were measured according to Li et al.21 A total of 100 to 120 grains from four panicles were measured. The seed shattering of the F2 plants derived from the cross between ‘Moretsu’ and ‘Minamiyutaka’ were evaluated using a method based on grasping panicles by hand at the maturity stage. In brief, three panicles from a plant were grasped by hand at one time, then the number of shattered grains then being counted. If the number of shattered grains was less than or equal to 3, this plant was regarded as non-shattering. If the number of shattered grains was more than or equal to 20, this plant was regarded as easy-shattering.
Histological analysis
The pedicels were collected at the anthesis stage, followed by fixation in FAA (Formalin-Acetic-Alcohol) solution. After dehydration and embedding, the tissues were longitudinally sectioned into 2-μm-thick sections according to the methods described by Li et al.21. The sections were stained with 0.01% toluidine and were observed using an BX53 microscope (OLYMPUS, Tokyo, Japan).
Scanning electron microscopy
The bases of mature seeds were subjected to platinum sputter coating and observed using a SEM (JEOL JSM-5610 LV, Tokyo, Japan). High resolution images were obtained in high vacuum mode at 20 kV.
DNA extraction
The leaves were harvested from 30-day-old seedlings. For next generation sequencing, DNAs of ‘Moretsu’ and ‘Minamiyutaka’ were extracted using DNeasy Plant Maxi Kit (Qiagen Inc., Valencia, USA). For DNA marker analysis, DNAs of F2 plants and gene-edited plants were extracted using the simple DNA extraction method49.
Whole-genome resequencing
DNA samples of ‘Moretsu’ and ‘Minamiyutaka’ were subjected to paired-end (2 × 150 bp) sequencing using an Illumina HiSeq X Ten platform. The short reads were cleaned by removing low quality reads and unpaired reads using Trimmomatic (version 0.36)50 with the following parameters: LEADING:10, TRAILING:10, SLIDINGWINDOW:4:20, and MINLEN:36. Next, the clean reads were aligned to the Nipponbare reference sequence (IRGSP-1.0, http://rapdb.dna.affrc.go.jp)51 using the mapping tool Borrows Wheeler Aligner (version 0.7.17)52 and indexed as BAM files using SAMtools (version 1.3.1)53. Duplicate fragments were then marked and eliminated with MarkDuplicates tool in Picard-Tools (Version 2.7.1.0) (https://broadinstitute.github.io/picard/). SNP, insertion, and deletion calling was performed using the HaplotypeCaller tool in GATK (Version 3.7-0)54. Detection of SV was performed using Pindel55 and Manta56 with default parameters.
Homozygous variations between ‘Moretsu’ and ‘Minamiyutaka’ were called with settings described below: (1) the read depth of the variant site was more than five and less than 100, (2) allele frequencies of both cultivars at a site were higher than 0.8, (3) genotypes of the two cultivars were different. To ensure the accuracy of variation detection, the candidate variations were visually confirmed using the Integrative Genomics Viewer57.
Variation annotation analysis was conducted based on the gene annotation of the ‘Nipponbare’ using SnpEff v4.258.
Validation of the candidate causal mutation
The causal mutation inferred by variation annotation analysis was first verified by Sanger sequencing. The methods described below have been reproduced in part from Li et al.21. Briefly, primers (Supplementary Table S8 online) were designed using the Primer3 program (https://bioinfo.ut.ee/primer3-0.4.0/). Purified PCR product was used for the sequencing reaction using the BigDye Terminator V3.1 cycle sequencing kit (Thermo Fisher Scientific, MA, USA), followed by sequencing on a 3730xl Genetic Analyzer (Thermo Fisher Scientific).
For analysis of co-segregation of the causal mutation with non-shattering habit in the F2 population, a CAPS marker (Supplementary Table S8 online) was designed based on the restriction enzyme map analysis (https://www.restrictionmapper.org). PCR amplicons were digested by the restriction enzyme BspCNI (New England Biolabs, MA, USA) at 25 °C for 60 min, and then analyzed by electrophoresis on 2.0% PrimeGel Agarose PCR-Sieve HRS (Takara) gels for 40 min at 100 V.
Gene expression analyses
To determine the expression profile of OsSh1 in rice plants, we analyzed the microarray data from the rice gene expression database RiceXPro (https://ricexpro.dna.affrc.go.jp)33. RT-qPCR analysis was conducted to compare the gene expression levels between different cultivars or lines. Total RNA was extracted using the RNeasy Plant Mini Kit (Qiagen, Hilden, Germany), from the 70–90-mm long young panicles before heading, the stage when the abscission zone is developing59. Each cultivar had four replications. The first-strand cDNA was synthesized from 1.0 μg of total RNA using a SuperScript III First-Strand Synthesis SuperMix for qRT-PCR (Invitrogen by Life Technologies, Carlsbad, CA, USA). Ubiquitin gene was used as an endogenous control to normalize detected gene expression. RT-qPCR was performed using an SYBR Green Supermix Kit (Bio-Rad, Hercules, CA, USA) on a QuantStudio 1 System (Thermo Fisher Scientific). The PCR reaction mixture (15 μl) consisted of 0.2 μM forward and reverse primers, 1 × SYBR Advantage qPCR Premix and about 10 ng cDNA. The reactions were carried out using the following qPCR protocol: 2 min pre-incubation at 95 °C followed by 40 cycles of 95 °C for 15 s and 60 °C for 1 min, and finally a dissociation run from 60 to 95 °C. Primers used for RT-qPCR experiments are listed in Supplementary Table S8 online.
cDNA sequence analysis
A cDNA clone containing the full open reading frame (ORF) of OsSh1 was generated by PCR using the primers listed in Supplementary Table S8 online. The PCR products were cloned into pCR-BluntII-TOPO (Invitrogen) and subjected to Sanger sequence analysis as mentioned above. The mRNA sequences of the OsSh1 gene in ‘Moretsu’ and ‘Minamiyutaka’ have been deposited in the DNA Data Bank of Japan (DDBJ; https://www.ddbj.nig.ac.jp) with accession numbers LC522940 and LC522941, respectively. The deduced amino acid sequences were aligned using Clustal Omega with default parameters (https://www.ebi.ac.uk/Tools/msa/clustalo/). To align amino acid sequences of YABBY2 proteins in different plants, we downloaded sequences from the GenBank (https://www.ncbi.nlm.nih.gov) as follows: O. sativa L. ssp. japonica (AAX95527.1), O. barthii (AXM44150.1), O. brachyantha (XP_006650352.1), Aegilops tauschii ssp. tauschii (XP_020155668.1), Triticum aestivum (ABW80974.1), Brachypodium distachyon (XP_003561769.1), Arabidopsis thaliana (NP_001077490.1), Camelina sativa (XP_010458084.1), Gossypium hirsutum (XP_016683147.1), Brassica oleracea (XP_013638657.1). In addition, YABBY2 protein sequences in O. sativa L. ssp. indica (B8ANI4), O. rufipogon (A0A0E0NYX3), O. nivara (A0A0E0GR31), O. punctat (A0A0E0KGW2), O. glumipatula (A0A0D9ZB45) were from UniProt (https://www.uniprot.org).
Vector construction and transformation
The sgRNA-Cas9 plant expression vector pZH_OsCas9 and the guide RNA expression vector pZK_sgRNA were kindly provided by Dr. Masaki Endo and Dr. Seiichi Toki (NARO)60. The oligos used in constructing the sgRNA vectors for OsSh1 are shown in Supplementary Fig. S6a online. The easy-shattering indica rice cultivar ‘Teqing’ was subjected to Agrobacterium-mediated transformation using immature embryo-derived calli as described previously61. In brief, immature embryos of ‘Teqing’ from 10 to 14 days after flowering were infected by Agrobacterium carrying the pZH_OsU6gRNA_MMCas9 vector (Supplementary Fig. S6b online). After 5 days of co-cultivation, infected immature embryos were transferred to a fresh resting medium containing 400 mg/L carbenicillin disodium salt (Nakarai, Kyoto, Japan) to remove Agrobacterium. Following this, Hygromycin-resistant calli were selected over 4 weeks on a selection medium containing 400 mg/L carbenicillin disodium salt and hygromycin 30 mg/L (Wako Pure Chemicals, Osaka, Japan). Proliferating calli were then transferred to a fresh pre-regeneration medium containing 200 mg/L carbenicillin disodium salt and hygromycin 40 mg/L. After 8 days of culture, these calli were transferred to a fresh regeneration medium containing 30 mg/L hygromycin B and cultured for 2 weeks. The re-generated rice plants were grown in a closed greenhouse.
Analysis of CRISPR-induced mutations
To analyze the mutation induced in the regenerated plants, the first or second exon of OsSh1 were PCR amplified using the specified primers (Supplementary Table S8 online). PCR products were subjected to an Nsp I or MscI restriction enzyme reaction, followed by agarose gel electrophoresis. PCR products showing mutation by CAPS analysis were cloned into pCR-BluntII-TOPO (Invitrogen) and subjected to Sanger sequence analysis as mentioned above.
Statistical analysis
Statistical analyses were performed in R Software version 3.6.062. Violin plots were created in the R-package ggplot263. To determine the statistical significance, firstly, an F Test (two samples) or Bartlett’s test (> =3 samples) was used to test whether variances were equal for all samples. Then, if the variances were equal, Student’s t test (two samples) or pairwise comparisons using t tests with pooled SD (Standard Deviation) (> =3 samples) were performed, otherwise, Welch’s t test (two samples) or Wilcoxon rank sum test (> =3 samples) was performed. Finally, a Bonferroni correction was used to control for the family-wise type I error rate across the comparisons.
DNA polymorphism analysis of candidate gene
We exported the consensus sequences covering the coding region and the flanking regions of OsSh1 in wild and cultivated rice from TASUKE +64. The variant filter was set as follows: quality > =20 and depth > =4. The average number of reads that align the whole genome of an accession was set at more than eight. Nucleotide diversity analysis and test for neutral selection were performed using DnaSP (version 6.12.03)65. S, the number of polymorphic (segregating) sites; π, the average number of pairwise nucleotide differences per site66; θ, Watterson’s estimator of nucleotide polymorphism per site67, and Tajima’s D test68 were calculated. The alleles of OsSh1 were also checked in the accessions of O. rufipogon in the database OryzaGenome (https://viewer.shigen.info/oryzagenome2detail/about/about.xhtml), and O. sativa in Rice SNP-Seek Database (https://snp-seek.irri.org/index.zul).
Data availability
All NGS data files will be available in the DDBJ Sequenced Read Archive under the Accession Nos. DRA009647 and DRA009648, upon acceptance of this paper.
References
Di Vittori, V. et al. Convergent evolution of the seed shattering trait. Genes (Basel) https://doi.org/10.3390/genes10010068 (2019).
Li, C. B., Zhou, A. L. & Sang, T. Rice domestication by reducing shattering. Science 311, 1936–1939. https://doi.org/10.1126/science.1123604 (2006).
Kovach, M. J., Sweeney, M. T. & McCouch, S. R. New insights into the history of rice domestication. Trends Genet. 23, 578–587. https://doi.org/10.1016/j.tig.2007.08.012 (2007).
Huang, X. et al. A map of rice genome variation reveals the origin of cultivated rice. Nature 490, 497–501. https://doi.org/10.1038/nature11532 (2012).
Konishi, S. et al. An SNP caused loss of seed shattering during rice domestication. Science 312, 1392–1396. https://doi.org/10.1126/science.1126410 (2006).
Dong, Y. & Wang, Y. Z. Seed shattering: from models to crops. Front. Plant Sci. https://doi.org/10.3389/fpls.2015.00476 (2015).
Jin, I. D. On the formation and development of abscission layer in riceplants, Oryza sativa L. Jpn. J. Crop Sci. 55, 451–457 (1986).
Jin, I. D. & Inouye, J. Relation between grain shedding and pedicel morphology near the abscission layer of japonica-indica hybrid rices bred in Korea. Jpn. J. Crop Sci. 51, 271–275 (1982).
Thurber, C. S., Hepler, P. K. & Caicedo, A. L. Timing is everything: early degradation of abscission layer is associated with increased seed shattering in U.S. weedy rice. BMC Plant Biol. https://doi.org/10.1186/1471-2229-11-14 (2011).
Lee, G. H., Kang, I. K. & Kim, K. M. Mapping of novel QTL regulating grain shattering using doubled haploid population in rice (Oryza sativa L.). Int. J. Genom. https://doi.org/10.1155/2016/2128010 (2016).
Htun, T. M., Inoue, C., Chhourn, O., Ishii, T. & Ishikawa, R. Effect of quantitative trait loci for seed shattering on abscission layer formation in Asian wild rice Oryza rufipogon. Breed. Sci. 64, 199–205. https://doi.org/10.1270/jsbbs.64.199 (2014).
Qin, Y. et al. Identification for quantitative trait loci controlling grain shattering in rice. Genes Genom. 32, 173–180. https://doi.org/10.1007/s13258-009-0879-7 (2010).
Xiong, L. X., Liu, K. D., Dai, X. K., Xu, C. G. & Zhang, Q. F. Identification of genetic factors controlling domestication-related traits of rice using an F-2 population of a cross between Oryza sativa and O. rufipogon. Theor. Appl. Genet. 98, 243–251 (1999).
Tsujimura, Y. et al. Detection of a novel locus involved in non-seed-shattering behaviour of Japonica rice cultivar Oryza sativa “Nipponbare”. Theor. Appl. Genet. 132, 2615–2623. https://doi.org/10.1007/s00122-019-03376-3 (2019).
Subudhi, P. K. et al. Mapping of seed shattering loci provides insights into origin of weedy rice and rice domestication. J. Hered. 105, 276–287. https://doi.org/10.1093/jhered/est089 (2014).
Lin, Z. W. et al. Origin of seed shattering in rice (Oryza sativa L.). Planta 226, 11–20. https://doi.org/10.1007/s00425-006-0460-4 (2007).
Roeder, A. H., Ferrandiz, C. & Yanofsky, M. F. The role of the REPLUMLESS homeodomain protein in patterning the Arabidopsis fruit. Curr. Biol. 13, 1630–1635. https://doi.org/10.1016/j.cub.2003.08.027 (2003).
Konishi, S. et al. Molecular cloning of a major QTL, QSH-1, controlling seed shattering habit in rice. Plant Cell Physiol. 46, S198–S198 (2005).
Fukuta, Y. & Yagi, T. Mapping of a shattering resistance gene in a mutant line SR-5 induced from an indica rice variety, Nan-jing11. Breed. Sci 48, 345–348. https://doi.org/10.1270/jsbbs1951.48.345 (1998).
Jiang, L. et al. The APETALA2-Like transcription factor SUPERNUMERARY BRACT controls rice seed shattering and seed size. Plant Cell 31, 17–36. https://doi.org/10.1105/tpc.18.00304 (2019).
Li, F. et al. Identification of a locus for seed shattering in rice (Oryza sativa L.) by combining bulked segregant analysis with whole-genome sequencing. Mol. Breed. 39, 36. https://doi.org/10.1007/s11032-019-0941-3 (2019).
Zhou, Y. et al. Genetic control of seed shattering in rice by the APETALA2 transcription factor SHATTERING ABORTION1. Plant Cell 24, 1034–1048. https://doi.org/10.1105/tpc.111.094383 (2012).
Ji, H. et al. Inactivation of the CTD phosphatase-like gene OsCPL1 enhances the development of the abscission layer and seed shattering in rice. Plant J. 61, 96–106. https://doi.org/10.1111/j.1365-313X.2009.04039.x (2010).
Ji, H. S. et al. Characterization and mapping of a shattering mutant in rice that corresponds to a block of domestication genes. Genetics 173, 995–1005. https://doi.org/10.1534/genetics.105.054031 (2006).
Lin, Z. W. et al. Parallel domestication of the Shattering1 genes in cereals. Nat. Genet. 44, 720–724 (2012).
Lv, S. et al. Genetic control of seed shattering during African rice domestication. Nat. Plants 4, 331–337. https://doi.org/10.1038/s41477-018-0164-3 (2018).
Simons, K. J. et al. Molecular characterization of the major wheat domestication gene Q. Genetics 172, 547–555. https://doi.org/10.1534/genetics.105.044727 (2006).
Kato, H. et al. A newly bred rice variety “Minamiyutaka” for whole-crop silage. Bull. Miyazaki Agric. Exp. Stn. 41, 51–60 (2006).
Abe, A. et al. Genome sequencing reveals agronomically important loci in rice using MutMap. Nat. Biotechnol. 30, 174–178. https://doi.org/10.1038/nbt.2095 (2012).
Li, F., Shimizu, A., Nishio, T., Tsutsumi, N. & Kato, H. Comparison and characterization of mutations induced by gamma-ray and carbon-ion irradiation in rice (Oryza sativa L.) using whole-genome resequencing. G3 (Bethesda) 9, 3743–3751. https://doi.org/10.1534/g3.119.400555 (2019).
Toriba, T. et al. Molecular characterization the YABBY gene family in Oryza sativa and expression analysis of OsYABBY1. Mol. Genet. Genom. 277, 457–468. https://doi.org/10.1007/s00438-006-0202-0 (2007).
Bowman, J. L. & Smyth, D. R. CRABS CLAW, a gene that regulates carpel and nectary development in Arabidopsis, encodes a novel protein with zinc finger and helix-loop-helix domains. Development 126, 2387–2396 (1999).
Sato, Y. et al. RiceXPro Version 3.0: expanding the informatics resource for rice transcriptome. Nucleic Acids Res. 41, D1206–D1213. https://doi.org/10.1093/nar/gks1125 (2013).
Takai, T. et al. Genetic mechanisms underlying yield potential in the rice high-yielding cultivar Takanari, based on reciprocal chromosome segment substitution lines. BMC Plant Biol. 14, 295. https://doi.org/10.1186/s12870-014-0295-2 (2014).
Huang, X. et al. Genome-wide association studies of 14 agronomic traits in rice landraces. Nat. Genet. 42, 961–967. https://doi.org/10.1038/ng.695 (2010).
Frischmeyer, P. A. & Dietz, H. C. Nonsense-mediated mRNA decay in health and disease. Hum. Mol. Genet. 8, 1893–1900. https://doi.org/10.1093/hmg/8.10.1893 (1999).
Chang, Y. F., Imam, J. S. & Wilkinson, M. F. The nonsense-mediated decay RNA surveillance pathway. Annu. Rev. Biochem. 76, 51–74. https://doi.org/10.1146/annurev.biochem.76.050106.093909 (2007).
Bowman, J. L. The YABBY gene family and abaxial cell fate. Curr. Opin. Plant Biol. 3, 17–22. https://doi.org/10.1016/S1369-5266(99)00035-7 (2000).
Siegfried, K. R. et al. Members of the YABBY gene family specify abaxial cell fate in Arabidopsis. Development 126, 4117–4128 (1999).
Juarez, M. T., Twigg, R. W. & Timmermans, M. C. Specification of adaxial cell fate during maize leaf development. Development 131, 4533–4544. https://doi.org/10.1242/dev.01328 (2004).
Yamaguchi, T. et al. The YABBY gene DROOPING LEAF regulates carpel specification and midrib development in Oryza sativa. Plant Cell 16, 500–509. https://doi.org/10.1105/tpc.018044 (2004).
Zhao, W., Su, H. Y., Song, J., Zhao, X. Y. & Zhang, X. S. Ectopic expression of TaYAB1, a member of YABBY gene family in wheat, causes the partial abaxialization of the adaxial epidermises of leaves and arrests the development of shoot apical meristern in Arabidopsis. Plant Sci. 170, 364–371. https://doi.org/10.1016/j.plantsci.2005.09.008 (2006).
Liu, H. L., Xu, Y. Y., Xu, Z. H. & Chong, K. A rice YABBY gene, OsYABBY4, preferentially expresses in developing vascular tissue. Dev. Genes Evol. 217, 629–637. https://doi.org/10.1007/s00427-007-0173-0 (2007).
Onishi, K., Takagi, K., Kontani, M., Tanaka, T. & Sano, Y. Different patterns of genealogical relationships found in the two major QTLs causing reduction of seed shattering during rice domestication. Genome 50, 757–766. https://doi.org/10.1139/G07-051 (2007).
Doebley, J. & Stec, A. Genetic-analysis of the morphological differences between maize and teosinte. Genetics 129, 285–295 (1991).
Devos, K. M. & Gale, M. D. Genome relationships: the grass model in current research. Plant Cell 12, 637–646. https://doi.org/10.1105/tpc.12.5.637 (2000).
Dinneny, J. R., Weigel, D. & Yanofsky, M. F. A genetic framework for fruit patterning in Arabidopsis thaliana. Development 132, 4687–4696. https://doi.org/10.1242/dev.02062 (2005).
Xu, X. et al. Resequencing 50 accessions of cultivated and wild rice yields markers for identifying agronomically important genes. Nat. Biotechnol. 30, 105–111. https://doi.org/10.1038/nbt.2050 (2011).
Edwards, K., Johnstone, C. & Thompson, C. A simple and rapid method for the preparation of plant genomic DNA for PCR analysis. Nucleic Acids Res. 19, 1349–1349 (1991).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120. https://doi.org/10.1093/bioinformatics/btu170 (2014).
Kawahara, Y. et al. Improvement of the Oryza sativa Nipponbare reference genome using next generation sequence and optical map data. Rice 6, 4. https://doi.org/10.1186/1939-8433-6-4 (2013).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760. https://doi.org/10.1093/bioinformatics/btp324 (2009).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079. https://doi.org/10.1093/bioinformatics/btp352 (2009).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303. https://doi.org/10.1101/gr.107524.110 (2010).
Ye, K., Schulz, M. H., Long, Q., Apweiler, R. & Ning, Z. Pindel: a pattern growth approach to detect break points of large deletions and medium sized insertions from paired-end short reads. Bioinformatics 25, 2865–2871. https://doi.org/10.1093/bioinformatics/btp394 (2009).
Chen, X. et al. Manta: rapid detection of structural variants and indels for germline and cancer sequencing applications. Bioinformatics 32, 1220–1222. https://doi.org/10.1093/bioinformatics/btv710 (2016).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26. https://doi.org/10.1038/nbt.1754 (2011).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w(1118); iso-2; iso-3. Fly 6, 80–92. https://doi.org/10.4161/fly.19695 (2012).
Jin, I. D. On the formation and development of abscission layer in rice plants, Oryza sativa L. Jpn. J. Crop Sci. 55, 451–457 (1986).
Mikami, M., Toki, S. & Endo, M. Comparison of CRISPR/Cas9 expression constructs for efficient targeted mutagenesis in rice. Plant Mol. Biol. 88, 561–572. https://doi.org/10.1007/s11103-015-0342-x (2015).
Hiei, Y. & Komari, T. Improved protocols for transformation of indica rice mediated by Agrobacterium tumefaciens. Plant Cell Tissue Organ Cult. 85, 271–283 (2006).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, Vienna) https://www.R-project.org/ (2019).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer, New York) https://ggplot2.tidyverse.org (2016).
Kumagai, M. et al. TASUKE+: a web-based platform for exploring GWAS results and large-scale resequencing data. DNA Res. https://doi.org/10.1093/dnares/dsz022 (2019).
Rozas, J. et al. DnaSP 6: DNA sequence polymorphism analysis of large data sets. Mol. Biol. Evol. 34, 3299–3302. https://doi.org/10.1093/molbev/msx248 (2017).
Nei, M. Molecular Evolutionary Genetics (Columbia University Press, New York, 1987).
Watterson, G. A. On the number of segregating sites in genetical models without recombination. Theor. Popul. Biol. 7, 256–276 (1975).
Tajima, F. Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123, 585–595 (1989).
Acknowledgements
This work was supported by the Cabinet Office, Government of Japan, Cross-ministerial Strategic Innovation Promotion Program (SIP), “Technologies for creating next-generation agriculture, forestry and fisheries” (funding agency: Bio-oriented Technology Research Advancement Institution, NARO), and the Advanced Analysis Center Research Supporting Program of NARO. We gratefully acknowledge the Advanced Analysis Center of NARO for use of the high-speed processor system, Dr. Toshio Yamamoto for providing ‘SL1303’, Dr. Masaki Endo and Dr. Seiichi Toki for providing vectors, Dr. Yoshihiro Kawahara, Dr. Toshikazu Morishita, Dr. Hiroaki Yamanouchi and Dr. Nobuko Mase for suggestions for experiments, Dr. Jun-ichi Yonemaru for suggestions concerning manuscript revision. Ms. Izumi Odano for experiment support, and Mr. Masayoshi Tobita for support with planting materials.
Author information
Authors and Affiliations
Contributions
H.K., A.S., and F.L. conceived and coordinated the research plan. F.L. analyzed the next-generation sequencing data and conducted gene expression analysis. A.K. and M.O. conducted CRISPR/Cas9 gene editing. H.E. observed the fracture surface of the grain base using scanning electron microscopy. F.L. and A.K. wrote the manuscript. All authors reviewed and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, F., Komatsu, A., Ohtake, M. et al. Direct identification of a mutation in OsSh1 causing non-shattering in a rice (Oryza sativa L.) mutant cultivar using whole-genome resequencing. Sci Rep 10, 14936 (2020). https://doi.org/10.1038/s41598-020-71972-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-71972-1
This article is cited by
-
Genome sequencing-based coverage analyses facilitate high-resolution detection of deletions linked to phenotypes of gamma-irradiated wheat mutants
BMC Genomics (2022)
-
Meta-analysis of QTLs and candidate genes associated with seed germination in rice (Oryza sativa L.)
Physiology and Molecular Biology of Plants (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.