Abstract
Heart failure is the final destination of most cardiovascular diseases, and its complex molecular mechanisms remain largely uncertain. This study aimed to systematically investigate the underlying molecular mechanisms and diagnostic and therapeutic targets of heart failure using bioinformatics. We obtained 8 healthy samples and 8 heart failure samples from GSE8331 and GSE76701. After removing the batch effect, we performed a differential analysis on it and obtained 185 differentially expressed ID. The results of enrichment analysis showed that the molecular mechanisms of heart failure were mostly related to immune, inflammation, and metabolism-related pathways. Immune cell infiltration analysis showed that the degree of infiltration of Tgd cells and Neurons was significantly enriched in heart failure samples, whereas pDCs and NKTs were in healthy tissue samples. We obtained Hub genes including EGR1, EGR2, FOS and FOSB by PPI network analysis. We established a 4-gene diagnostic model with Hub gene, and validated it in GSE21610 and GSE57338, and evaluated the discriminative ability of Hub gene by ROC curve. The 4-gene diagnostic model has an AUC value of 0.775 in GSE21610 and 0.877 in GSE57338. In conclusion, we explored the underlying molecular mechanisms of heart failure and the immune cell infiltration environment of failing myocardium by performing bioinformatic analysis of the GEO dataset. In addition, we identified EGR1, EGR2, FOS and FOSB as potential diagnostic biomarkers and therapeutic targets for heart failure. More importantly, a diagnostic model of heart failure based on these 4 genes was developed, which leads to a new understanding of the pathogenesis of heart failure and may be an interesting target for future in-depth research.
Similar content being viewed by others
Introduction
Cardiovascular disease is one of the leading causes of death in humans and includes high blood pressure, arrhythmia, coronary heart disease, heart failure and other heart-related diseases. Heart failure is the ultimate outcome of most cardiovascular diseases, mostly due to structural changes in the heart or functional disorders that impair ventricular filling or ejection function1. An estimated 64.3 million people worldwide have heart failure2. According to the observation data from Europe, the one-year all-cause mortality rate of heart failure exceeds 20%, which brings a heavy economic and social burden to the family and society3. Even with guideline-guided drug therapy, the prognosis for heart failure remains poor. Advanced patients often require frequent hospitalizations and lifelong medical management4. Therefore, early diagnosis of heart failure is of great significance for prognosis.
The pathological hallmark of heart failure is ventricular remodeling. As far as the information that can be collected from peripheral blood is concerned, although it has the advantages of simplicity and convenience in the early diagnosis of heart failure, it is difficult to detect structural changes in the heart. Imaging tests often do not detect ventricular remodeling until after it has occurred.Therefore, exploring the changes of myocardial gene expression in patients with heart failure is helpful for the early diagnosis of ventricular remodeling in heart failure, and the underlying molecular mechanism behind it also has the value of intervening in the treatment of heart failure. In addition, immune response also plays an important role in the development of heart failure. Infiltrating immune cells release cytokines such as TGF-β1 and TNF-α, which promote heart remodeling5. Therefore, exploring immune responses in the progression of heart failure could also help develop new diagnostic and therapeutic approaches.
With the rapid development of high-throughput technologies and bioinformatics, an increasing number of biomarkers show potential in the diagnosis and prognosis of heart failure, such as middle regional preatrial natriuretic peptide (MR-proANP), middle regional adrenomedullin (MR-proADM), highly sensitive troponin, soluble ST2 (sST2), growth differentiation factor (GDF)-15, Galectin-3, copeptin, Cystatin C (Cys-C) and Sirtuin (SIRT), however, more clinical evidence is needed before these markers can be used in clinic6. In addition, even for the same disease, different data sets and different analysis methods may produce different analysis results. For example, the Vijayakrishna Kolur team concluded that HF is related to the adaptive immune system and neutrophil degranulation by analyzing GSE141910, and obtained 10 characteristic genes7. Yang et al.8 analyzed GSE21610 and believed that HF is related to G protein-coupled receptor binding, peroxisome and cAMP signaling pathways, and obtained 5 characteristic genes. Therefore, our work can complement theirs to gain a more comprehensive understanding of the molecular mechanisms of heart failure.
In this study, we analyzed 2 datasets from GEO databases. Through systematic bioinformatics analysis, we obtained differentially expressed genes (DEGs) between heart failure myocardial samples and healthy heart samples, and explored the underlying pathological mechanisms of heart failure through functional enrichment analysis and PPI network analysis. We analyzed immune infiltration in failing myocardium to help us understand the role of immunity in the development of heart failure. We also constructed a four-gene diagnostic model by logistic regression analysis and validated the diagnostic model's validity in two other datasets.This study explores the underlying pathogenesis of heart failure and hopes to provide potential targets for the diagnosis and treatment of heart failure in the future.
Materials and methods
GEO datasets
The expression dataset of cardiac RNAs was collected from the online GEO database (www.ncbi.nlm.nih.gov/geo/). The initial search used the keywords "Heart failure", "Homo sapiens", and "expression profiling by array". We choose GSE8331 and GSE76701 datasets of GPL570 platform as model sets. The validation set selects the same platform datasets, GSE21610 (from GPL570 platform), and other platforms datasets, GSE573389,10,11,12. GSE8331 contains 4 normal human heart samples and 4 failed heart samples with reduced ejection fraction. GSE76701 also contains 4 normal hearts and 4 failing hearts. GSE21610 contains 8 samples of normal hearts, 30 samples of failing hearts before ventricular assist devices (VAD) support and 30 samples of hearts after VAD support. We select 8 normal heart samples and 30 failing heart samples before VAD support as the validation set. GSE57338 is a dataset of 313 samples consisting of normal samples, ischemic left ventricle samples, and idiopathic dilated CMP left ventricle samples. We select 85 healthy samples and 72 failed heart samples among them as the validation set.
Data preprocessing and DEGs identification
We used R software (R version 4.2.0) for data preprocessing. Download GSE8331 and GSE76701 from the GEO database via the GEOquery package. Remove probes that correspond to multiple molecules for one probe. For multiple probes corresponding to the same molecule, only the probe with the largest signal value is retained. The filtered data will use the ComBat function of the sva package to remove batch differences. Check the standardization through the box plot, and check the clustering between the sample groups through the PCA plot and the UMAP plot. We used the limma package for differential analysis of heart failure samples and normal samples, and selected genes with p-value < 0.05 and |log2(FC)|≥ 1 as differential genes. Use the ggplot2 package and the pheatmap package to complete the drawing of the picture13,14,15,16.
Function and pathway enrichment analysis
To further analyze the function of myocardial expression profiles, we performed Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genes (KEGG) enrichment analysis of DEGs using the clusterProfiler package17,18,19,20. Use the GOplot package to combine the log2(FC) of the differential genes to calculate the Z-score value corresponding to each entry21. To avoid ignoring genes that are not significantly different but have important functions, we performed Gene Set Enrichment Analysis (GSEA) on all genes22.
Immune cell infiltration analysis
We used the xCell package (It was validated using extensive in-silico simulations and cytometry immunophenotyping.) to assess the content of immune and stromal cells in cardiac samples to delineate the cellular heterogeneity landscape of cardiac expression profiles23. Differences in cell distribution between the heart failure group and the normal group were compared by t-test, and the cut-off value was set at p < 0.1. In addition, we used Spearman's correlation coefficient to evaluate the strength of correlation between cells. The visualization part is done using the ggplot2 package.
Protein–protein interaction network and gene module identification
We uploaded the list of DEGs to the STRING database (string-db.org) to detect protein interactions with the following settings: Network type: full STRING network, meaning of network edges: evidence, minimum required interaction score: medium confidence (0.400)24. Subsequently, we imported the results into Cytoscape 3.6.1 for visualization and subsequent analysis25. In Cytoscape, we use the MCODE plugin to identify highly interconnected clusters with cutoff parameters set as follows: degree cutoff = 3, node score cutoff = 0. 2, k-Core = 2, Max. Depth = 100. The Hub gene was further screened with MCODE score > 4 as the standard26.
Hub genes verification and diagnosis model construction
We validated the Hub genes screened in the PPI network in GSE21610 (30 heart failure samples and 8 healthy samples) and GSE57338 (85 healthy samples and 72 heart failure samples). We used the pROC package to build diagnostic models of single genes and combined genes, and evaluated the discriminative ability of key genes and diagnostic models by ROC curves.
Results
Data preprocessing and DEGs identification
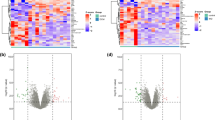
We combined the expression profiles of healthy heart samples and failing heart samples from GSE8331 and GSE76701 datasets of the GPL570 platform into a merged dataset containing 8 healthy samples and 8 disease samples. As shown in Fig. 1, we effectively remove batch effects between datasets by normalizing. After differential analysis, we obtained a total of 185 IDs with p-value < 0.05 and |log2(FC)|≥ 1, including 136 mRNAs, 46 LncRNAs and 3 unclassified genes. Figure 2 shows the PCA plot and UMAP plot of clustering among sample groups, as well as the volcano plot and heat map of differential IDs.
The PCA plot, UMAP plot, Volcano plot and Heatmap plot of gene expressions for 2 selected GEO Database with a screening criteria of |log2 FC|≥ 1 and adjust p value < 0.05. ((A) PCA plot; (B) UMAP plot; (C) Volcano plot; (D) Heatmap plot. PCA: Principal Component Analysis; UMAP: Uniform Manifold Approximation and Projection; Normal: Normal Group; HF: Heart Failure Group).
Functional enrichment analysis
GO and KEGG enrichment analysis
To evaluate the function of DEGs, we combined the log2(FC) of DEGs to perform GO and KEGG enrichment analysis, and the results are shown in Table 1. Categories of GO analysis include biological process (BP), cellular component (CC), and molecular function (MF). Figure 3 shows the enrichment results of GO and KEGG. The mainly enriched BP terms included response to molecule of bacterial origin, cell proliferation involved in metanephros development, response to lipopolysaccharide, and gliogenesis. The mainly enriched CC terms contained collagen-containing extracellular matrix, exocytic vesicle, haptoglobin-hemoglobin complex, and symmetric synapse. The mainly enriched MF terms were phosphatidylserine binding, neurexin family protein binding, protein tyrosine kinase activator activity, and modified amino acid binding. In KEGG pathway enrichment analysis, DEGs mainly enriched in Hematopoietic cell lineage, Glutathione metabolism, Drug metabolism—cytochrome P450, and Platinum drug resistance.
GSEA enrichment analysis
To assess the contribution of genes in the dataset in phenotypes, we performed GSEA enrichment analysis on them. The data set is C2.Cp, the random number seed is 2020, and the calculation is performed 1000 times. Each gene set contains between 10 and 500 genes. We visualized the top five datasets that met the criteria of FDR < 0.25 and p-value < 0.05, including Extracellular matrix organization, Genes encoding secreted soluble factors, Ensemble of genes encoding core extracellular matrix including ECM glycoproteins, collagens and proteoglycans, Cytokine-cytokine receptor interaction, and Diseases of metabolism. The visualization results are shown in Fig. 4, and the specific GSEA enrichment analysis information is shown in Table 2.
Immune cell infiltration analysis
xCell was used to assess cellular composition heterogeneity between heart failure samples and healthy samples. As shown in Fig. 5, 10 cell types were altered in heart failure heart tissue compared to healthy hearts, with 4 cell types significantly altered. Scores of cDC, HSC, Macrophages M2, Mast cells, Neurons, Erythrocytes, and Tgd cells were increased in heart failure heart tissue, whereas NKT, Myocytes, and pDC scores were decreased. In addition, we used Spearman's correlation coefficient to evaluate the strength of correlation between cells. As shown in Fig. 6, HSCs had the highest negative correlation with pDCs. Neurons had a high positive correlation with Macrophages M2, Mast cells, and HSC. Tgd cells had a high positive correlation with Mast cells.
PPI network construction and hub gene selection
We entered the DEGs into the STRING database for PPI network construction, and finally identified 64 nodes and 73 interactions. In addition, we identified a Hub gene module with 4 nodes and 6 edges by MCODE, and the results are shown in Fig. 7.
Hub genes verification and diagnosis model construction
We identified the composition of the Hub gene module as EGR1, EGR2, FOS and FOSB in the PPI network, all of which were up-regulated genes. We established ROC curve analysis to validate the diagnostic value of selected biomarkers. In the diagnostic model of a single gene, only EGR1 has high diagnostic value, and the AUC of other genes is not high. However, the combined genome has a high diagnostic value for heart failure. We constructed diagnostic models of Hub genes in different datasets by logistic regression and visualized them with ROC curves. The results are shown in Fig. 8.
Analysis of the disease predictive abilities of the four selected hub genes. (A) ROC curve analysis of diagnostic models for single and combined genes in merged dataset. (B) ROC curve analysis of diagnostic models for single and combined genes in GSE21610. (C) ROC curve analysis of diagnostic models for single and combined genes in GSE57338.
Discussion
HF is a major public health challenge. The use of drugs such as ARNI and dapagliflozin has also greatly improved the prognosis of heart failure. However, if the occurrence of heart failure can be detected earlier and the treatment can be intervened as soon as possible, the benefit of patients will be greater. In recent years, bioinformatics research has made great progress in the field of heart failure, and breakthroughs have been made in both the diagnosis of heart failure and the underlying molecular mechanisms. Integrating more biomarkers can comprehensively intervene in the diagnosis and treatment of heart failure. In addition, exploring related processes such as immune cells and pathways in the physiological and pathological processes of heart failure can also provide potential diagnostic and therapeutic targets to a certain extent.
In this study, we integrated 16 samples from 2 datasets and analyzed the data using bioinformatics.
In the GO enrichment analysis results, the main ones involved in heart failure were response to lipopolysaccharide, phosphatidylserine binding, and protein tyrosine kinase activator activity.
The cytokine hypothesis states that the progression of heart failure is attributable to sustained proinflammatory cytokine signaling. TNFα has been shown to induce cardiomyocyte dysfunction, cardiomyocyte fibrosis, and negative inotropic effects, and its elevation is associated with cardiac fibrosis, ventricular dilatation, and mortality27,28. Lipopolysaccharide is a cell wall component of Gram-negative bacteria and has a strong pro-inflammatory effect. Some researchers concluded that even a small concentration of lipopolysaccharide can effectively induce the release of TNFα in an in vitro whole blood model of heart failure patients29. Their study also demonstrated that response to lipopolysaccharide predicts survival in patients with chronic heart failure30. This suggests that maintaining the function of immune cells and the immune system is essential in the treatment of heart failure.
Phosphatidylserine is located on the cytoplasmic lobe of the plasma membrane bilayer. Energy loss during cellular injury or apoptosis results in disruption of plasma membrane asymmetry, resulting in surface exposure of phosphatidylserine. Subsequently, macrophages recognize damaged cells and complete the elimination of apoptotic cells. Modulation of membrane fluidity or phosphatidylserine binding can affect this process31. Cardiomyocyte apoptosis is a key regulator in the development of heart failure32. Therefore, phosphatidylserine can be employed as an early diagnostic indicator of heart failure. It may be also given potential therapeutic value in heart failure by modulating phosphatidylserine binding.
Tyrosine kinases are enzymes that catalyze the transfer of phosphate from ATP to tyrosine residues in polypeptides. The small molecule tyrosine kinase inhibitor imatinib is a targeted drug for the treatment of chronic myeloid leukemia, but its cardiotoxicity can cause heart failure. In cardiomyocytes, ABL functions to maintain endoplasmic reticulum homeostasis. Imatinib induces endoplasmic reticulum stress, activates the PERK signaling pathway, and produces mitochondrial dysfunction33. Therefore, maintaining the activity of protein tyrosine kinase activators may reduce the cardiotoxicity of antitumor drugs.
In the KEGG pathway enrichment analysis results, Glutathione metabolism and Drug metabolism—cytochrome P450 are mainly involved in heart failure. Abnormal intracellular glutathione metabolism is related to increased free radical production in chronic heart failure. Therefore, glutathione can be used as a marker for evaluating lipid peroxidation in heart failure34. Glutathione metabolism is also involved in endoplasmic reticulum stress and oxidative stress, and 5-oxoproline, the final product of Chac1 enzyme involved in metabolism, is also an important factor leading to heart failure35.In addition, in recent years, studies have found that ferroptosis is also associated with the occurrence of heart failure, which is related to the excessive production of ROS and the inhibition of glutathione synthesis by GPX436. The primary physiological importance of cytochrome P450 enzymes lies in their ability to metabolize arachidonic acid to epoxyeicosatrienoic acid and hydroxyeicosatetraenoic acid metabolites, which play a role in maintaining cardiac homeostasis37,38. In short, in the KEGG enrichment results, DEGs mainly interfere with heart failure by affecting metabolism-related pathways.
Among the GSEA enrichment results, extracellular matrix organization and Diseases of metabolism were mainly involved in heart failure. The cardiac extracellular matrix contains a dynamic molecular network that provides structural support to cardiac tissue. Its renewal is an important process of cardiac remodeling. Excessive activation of pro-inflammatory cytokines such as TNFα promotes this process39. Therefore, the extracellular matrix can not only be used as an indicator to evaluate cardiac homeostasis, but also can be used to design more effective anti-fibrotic cardiac regeneration strategies40. From the results of enrichment analysis, the molecular mechanism of heart failure is closely related to inflammation, metabolism, oxidative stress and immunity.
Immune cell infiltration analysis showed that the degree of infiltration of Tgd cells and Neurons was significantly enriched in heart failure samples, whereas pDCs and NKTs were in healthy tissue samples. Cytotoxicity is the main biological effect of Tgd cells. Studies have shown that IL-17A secreted by Tgd cells can induce cardiomyocyte apoptosis41. By regulating the IL-17A/Tgd cells axis, it can effectively regulate the level of inflammation and slow down the process of myocardial fibrosis42,43. The enrichment of Neurons in heart failure samples may be related to the increase of cardiac sympathetic nerve activity and the enhancement of N-type Ca2 + currents44. pDC and NKT are higher in healthy sample tissues, which is slightly different from previous studies45,46,47, and the specific mechanism behind it remains to be further studied.
According to PPI network analysis, we obtained Hub genes, including EGR1, EGR2, FOS and FOSB. EGR1 is widely present in human cells and has the ability to express rapidly, playing an important role in cell growth, differentiation, proliferation, and inflammatory responses48. Studies have shown that inhibiting the expression of EGR1 can slow myocardial fibrosis and thus treat heart failure49. EGR2, which belongs to the EGR family, has pro-inflammatory and pro-apoptotic effects. EGR2 can directly activate the expression of pro-apoptotic proteins such as BCL2 family and BINP3L, and can also upregulate inflammatory proteins such as IFNG and IL650. Studies have shown that inhibition of EGR2 expression via lncRNA and miRNA can alleviate cardiac apoptosis and protect the heart from ischemic injury51,52. FOS and FOSB belong to the FOS gene family and are regulators of cell proliferation, differentiation, and transformation. In the early stage of cardiac remodeling, c-fos and c-jun were up-regulated, and the two dimerized to form the transcription factor complex AP-1, followed by the up-regulation of inflammatory factors NF-kappa B and NFAT. In late stage, genes involved in cardiac hypertrophic response such as ANP and BNP are up-regulated53,54. Therefore, the FOS family can be used as an early diagnostic marker of cardiac remodeling and a target for early intervention in the treatment of heart failure.
The novelties of our study are as follows. First, we analyzed the molecular mechanisms of heart failure related to immunity, inflammation, and metabolism through bioinformatics. Second, we identified EGR1, EGR2, FOS and FOSB as potential diagnostic biomarkers and therapeutic targets for heart failure. The diagnostic model based on these 4 genes provides a new idea for our future research on the molecular mechanism of heart failure. Nevertheless, this study still has some shortcomings. First, we were unable to determine whether there was a causal relationship between differences in gene expression and the physiopathological mechanisms of heart failure. Secondly, myocardial tissue sampling is often ranked after peripheral blood examination and imaging examination in clinical practice, and the sampling cost is high and the clinical value is limited. Finally, the lack of medication and prognostic data in the GEO database limited further analysis of differential genes. Although our results were validated in two datasets, further clinical trials are needed to confirm our findings.
Conclusion
In summary, we explored the underlying molecular mechanisms of heart failure from the enrichment analysis and immune cell infiltration environment by bioinformatics analysis of the GEO dataset, which provided new ideas for the pathogenesis and treatment of heart failure. Through PPI network and logistic regression, we identified EGR1, EGR2, FOS and FOSB as potential diagnostic biomarkers and therapeutic targets for heart failure. More importantly, a diagnostic model of heart failure based on these 4 genes was developed, which leads to a new understanding of the pathogenesis of heart failure and may be an interesting target for future in-depth research.
Data availability
Publicly available datasets were analyzed in this study. This data can be found here: GSE8331; GSE76701; GSE21610; GSE57338.
References
Hunt, S. A. et al. ACC/AHA 2005 Guideline update for the diagnosis and management of chronic heart failure in the adult summary article. J. Am. Coll. Cardiol. 46(2005), 1116–1143. https://doi.org/10.1016/j.jacc2005.08023 (2005).
GBD Disease and Injury Incidence and Prevalence Collaborators. Global, regional, and national incidence, prevalence, and years lived with disability for 354 diseases and injuries for 195 countries and territories, 1990–2017: A systematic analysis for the global burden of disease study 2017. Lancet 392(10159), 1789–1858 (2017).
Maggioni, A. P. et al. EURObservational research programme: Regional differences and 1-year follow-up results of the heart failure pilot survey (ESC-HF Pilot). Eur. J. Heart Fail. 15, 808–817. https://doi.org/10.1093/eurjhf/hft050 (2013).
Yancy, C. W. et al. ACC/AHA/HFSA focused update of the 2013 ACCF/AHA guideline for the management of heart failure: A report of the American college of cardiology/American heart association task force on clinical practice guidelines and the heart failure society of America. Circulation 136(6), e137–e161. https://doi.org/10.1161/CIR.0000000000000509 (2017).
Schultheiss, H. P. et al. Dilated cardiomyopathy. Nat. Rev. Dis. Primers. 5(1), 32. https://doi.org/10.1038/s41572-019-0084-1 (2019).
Fan, S. & Hu, Y. 2022 Integrative analyses of biomarkers and pathways for heart failure. BMC Med. Genomics 15(1), 72. https://doi.org/10.1186/s12920-022-01221-z (2022).
Kolur, V., Vastrad, B., Vastrad, C., Kotturshetti, S. & Tengli, A. Identification of candidate biomarkers and therapeutic agents for heart failure by bioinformatics analysis. BMC Cardiovasc. Disord. 21(1), 329. https://doi.org/10.1186/s12872-021-02146-8 (2021).
Yang, Q., Bai, X., Li, X. & Hu, W. The identification of key genes and biological pathways in heart failure by integrated bioinformatics analysis. Comput. Math. Methods Med. 2021, 3859338. https://doi.org/10.1155/2021/3859338 (2021).
Kaneda, R. et al. Genome-wide histone methylation profile for heart failure. Genes Cells. 14(1), 69–77. https://doi.org/10.1111/j.1365-2443.2008.01252.x (2009).
Kim, E. H. et al. Differential protein expression and basal lamina remodeling in human heart failure. Proteomics Clin. Appl. 10(5), 585–596. https://doi.org/10.1002/prca.201500099 (2016).
Schwientek, P. et al. Global gene expression analysis in nonfailing and failing myocardium pre- and postpulsatile and nonpulsatile ventricular assist device support. Physiol Genomics. 42(3), 397–405. https://doi.org/10.1152/physiolgenomics.00030.2010 (2010).
Liu, Y. et al. RNA-Seq identifies novel myocardial gene expression signatures of heart failure. Genomics 105, 83–89. https://doi.org/10.1016/j.ygeno.2014.12.002 (2015).
Davis, S. & Meltzer, P. S. GEOquery: A bridge between the gene expression omnibus (GEO) and bioconductor. Bioinformatics 23(14), 1846–1847. https://doi.org/10.1093/bioinformatics/btm254 (2007).
Leek, J. T., Johnson, W. E., Parker, H. S., Jaffe, A. E. & Storey, J. D. The SVA package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics 28(6), 882–883. https://doi.org/10.1093/bioinformatics/bts034 (2012).
Gu, Z., Eils, R. & Schlesner, M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 32(18), 2847–2849. https://doi.org/10.1093/bioinformatics/btw313 (2016).
Smyth, G. K. Limma: Linear models for microarray data. In: Bioinformatics and computational biology solutions using R and Bioconductor. 2013.
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. Cluster profiler: An R package for comparing biological themes among gene clusters. OMICS 16(5), 284–287. https://doi.org/10.1089/omi.2011.0118 (2012).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28(1), 27–30. https://doi.org/10.1093/nar/28.1.27 (2000).
Kanehisa, M. Toward understanding the origin and evolution of cellular organisms. Protein Sci. 28(11), 1947–1951. https://doi.org/10.1002/pro.3715 (2019).
Kanehisa, M., Furumichi, M., Sato, Y., Kawashima, M. & Ishiguro-Watanabe, M. KEGG for taxonomy-based analysis of pathways and genomes. Nucleic Acids Res. 51(D1), D587–D592. https://doi.org/10.1093/nar/gkac963 (2023).
Walter, W., Sánchez-Cabo, F. & Ricote, M. GOplot: An R package for visually combining expression data with functional analysis. Bioinformatics 31(17), 2912–2914. https://doi.org/10.1093/bioinformatics/btv300 (2015).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. U. S. A. 102(43), 15545–15550. https://doi.org/10.1073/pnas.0506580102 (2005).
Aran, D., Hu, Z. & Butte, A. J. xCell: Digitally portraying the tissue cellular heterogeneity landscape. Genome Biol. 18(1), 220. https://doi.org/10.1186/s13059-017-1349-1 (2017).
Szklarczyk, D. et al. The STRING database in 2021: Customizable protein-protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Res. 49(D1), D605–D612. https://doi.org/10.1093/nar/gkaa1074 (2021).
Shannon, P. et al. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 13(11), 2498–2504. https://doi.org/10.1101/gr.1239303 (2003).
Bader, G. D. & Hogue, C. W. An automated method for finding molecular complexes in large protein interaction networks. BMC Bioinform. 13(4), 2. https://doi.org/10.1186/1471-2105-4-2 (2003).
Shirazi, L. F., Bissett, J., Romeo, F. & Mehta, J. L. Role of inflammation in heart failure. Curr. Atheroscler. Rep. 19(6), 27. https://doi.org/10.1007/s11883-017-0660-3 (2017).
Adamo, L., Rocha-Resende, C., Prabhu, S. D. & Mann, D. L. Reappraising the role of inflammation in heart failure. Nat. Rev. Cardiol. 17(5), 269–285. https://doi.org/10.1038/s41569-019-0315-x (2020).
von Haehling, S. et al. The relationship between age and production of tumour necrosis factor-alpha in healthy volunteers and patients with chronic heart failure. Int. J. Cardiol. 90(2–3), 197–204. https://doi.org/10.1016/s0167-5273(02)00566-1 (2003).
Ebner, N. et al. Lipopolysaccharide responsiveness is an independent predictor of death in patients with chronic heart failure. J. Mol. Cell Cardiol. 87, 48–53. https://doi.org/10.1016/j.yjmcc.2015.07.029 (2015).
Janko, C. et al. Cooperative binding of Annexin A5 to phosphatidylserine on apoptotic cell membranes. Phys. Biol. 10(6), 065006. https://doi.org/10.1088/1478-3975/10/6/065006 (2013).
van Empel, V. P. et al. Myocyte apoptosis in heart failure. Cardiovasc. Res. 67(1), 21–29. https://doi.org/10.1016/j.cardiores.2005.04.012 (2005).
Garcia-Alvarez, A., Garcia-Albeniz, X., Esteve, J., Rovira, M. & Bosch, X. Cardiotoxicity of tyrosine-kinase-targeting drugs. Cardiovasc. Hematol. Agents Med. Chem. 8(1), 11–21. https://doi.org/10.2174/187152510790796192 (2010).
Campolo, J. et al. Blood glutathione as independent marker of lipid peroxidation in heart failure. Int. J. Cardiol. 117(1), 45–50. https://doi.org/10.1016/j.ijcard.2006.04.065 (2007).
Bachhawat, A. K., Yadav, S., Jainarayanan, A. K. & Dubey, P. Heart failure and the glutathione cycle: An integrated view. Biochem. J. 477(17), 3123–3130. https://doi.org/10.1042/BCJ20200429 (2020).
Wu, X., Li, Y., Zhang, S. & Zhou, X. Ferroptosis as a novel therapeutic target for cardiovascular disease. Theranostics 11(7), 3052–3059. https://doi.org/10.7150/thno.54113 (2021).
Zordoky, B. N. & El-Kadi, A. O. Modulation of cardiac and hepatic cytochrome P450 enzymes during heart failure. Curr. Drug Metab. 9(2), 122–128. https://doi.org/10.2174/138920008783571792 (2008).
Aspromonte, N. et al. Modulation of cardiac cytochrome P450 in patients with heart failure. Expert Opin. Drug Metab. Toxicol. 10(3), 327–339. https://doi.org/10.1517/17425255.2014.872240 (2014).
Radauceanu, A. et al. Extracellular matrix turnover and inflammatory markers independently predict functional status and outcome in chronic heart failure. J. Card. Fail. 14(6), 467–474. https://doi.org/10.1016/j.cardfail.2008.02.014 (2008).
Perestrelo, A. R. et al. Multiscale analysis of extracellular matrix remodeling in the failing heart. Circ. Res. 128(1), 24–38. https://doi.org/10.1161/CIRCRESAHA.120.317685 (2021).
Liao, Y. H. et al. Interleukin-17A contributes to myocardial ischemia/reperfusion injury by regulating cardiomyocyte apoptosis and neutrophil infiltration. J. Am. Coll Cardiol. 59(4), 420–429. https://doi.org/10.1016/j.jacc.2011.10.863 (2012).
Blanco-Domínguez, R. et al. CD69 expression on regulatory T cells protects from immune damage after myocardial infarction. J. Clin. Invest. 132(21), e152418. https://doi.org/10.1172/JCI152418 (2022).
Yan, X. et al. Deleterious effect of the IL-23/IL-17A axis and γδT cells on left ventricular remodeling after myocardial infarction. J. Am. Heart Assoc. 1(5), e004408. https://doi.org/10.1161/JAHA.112.004408 (2012).
Tu, H. et al. Heart failure-induced changes of voltage-gated Ca2+ channels and cell excitability in rat cardiac postganglionic neurons. Am. J. Physiol. Cell Physiol. 306(2), C132–C142. https://doi.org/10.1152/ajpcell.00223.2013 (2014).
Kologrivova, I., Shtatolkina, M., Suslova, T. & Ryabov, V. Cells of the immune system in cardiac remodeling: Main players in resolution of inflammation and repair after myocardial infarction. Front. Immunol. 12, 664457. https://doi.org/10.3389/fimmu.2021.664457 (2021).
Vallejo, J., Cochain, C., Zernecke, A. & Ley, K. Heterogeneity of immune cells in human atherosclerosis revealed by scRNA-Seq. Cardiovasc. Res. 117(13), 2537–2543. https://doi.org/10.1093/cvr/cvab260 (2021).
Athanassopoulos, P. et al. Blood dendritic cell levels and phenotypic characteristics in relation to etiology of end-stage heart failure: Implications for dilated cardiomyopathy. Int. J. Cardiol. 131(2), 246–256. https://doi.org/10.1016/j.ijcard.2007.10.031 (2009).
Peng, J. & Xiang, Y. Value analysis of CD69 combined with EGR1 in the diagnosis of coronary heart disease. Exp. Ther. Med. 17(3), 2047–2052. https://doi.org/10.3892/etm.2019.7175 (2019).
Shen, J. et al. MiR-150–5p retards the progression of myocardial fibrosis by targeting EGR1. Cell Cycle 18(12), 1335–1348. https://doi.org/10.1080/15384101.2019.1617614 (2019).
Bo, Z. et al. EGR2 is a hub-gene in myocardial infarction and aggravates inflammation and apoptosis in hypoxia-induced cardiomyocytes. BMC Cardiovasc. Disord. 22(1), 373. https://doi.org/10.1186/s12872-022-02814-3 (2022).
Cao, X. et al. Silencing long non-coding RNA MIAT ameliorates myocardial dysfunction induced by myocardial infarction via MIAT/miR-10a-5p/EGR2 axis. Aging (Albany NY) 13(8), 11188–11206. https://doi.org/10.18632/aging.202785 (2021).
Tang, Y. et al. MicroRNA-150 protects the mouse heart from Ischaemic injury by regulating cell death. Cardiovasc. Res. 106(3), 387–397. https://doi.org/10.1093/cvr/cvv121 (2015).
Cheema, S. K., Tappia, P. S. & Dhalla, N. S. Modification of gene expression in rat cardiomyocytes by linoleic and docosahexaenoic acids 1. Can. J. Physiol. Pharmacol. 97(4), 320–327. https://doi.org/10.1139/cjpp-2018-0398 (2019).
Freire, G., Ocampo, C., Ilbawi, N., Griffin, A. J. & Gupta, M. Overt expression of AP-1 reduces alpha myosin heavy chain expression and contributes to heart failure from chronic volume overload. J. Mol. Cell Cardiol. 43(4), 465–478. https://doi.org/10.1016/j.yjmcc.2007.07.046 (2007).
Funding
Our work was supported by the National Natural Science Foundation of China [Grants nos. 81774247;81804045].
Author information
Authors and Affiliations
Contributions
Y.D.Y. conducted statistical analysis and drafted the article. Y.D.Y. and Y.L. were involved in the conception and design of the study. Y.D.Y. contributed to picture processing and article reviewing. Y.T.X. reviewed and proofread the article. Y.T.X. and Y.L. provided effective scientific suggestions and supervision and created the final revision of the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yu, Yd., Xue, Yt. & Li, Y. Identification and verification of feature biomarkers associated in heart failure by bioinformatics analysis. Sci Rep 13, 3488 (2023). https://doi.org/10.1038/s41598-023-30666-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-30666-0
This article is cited by
-
Exploration of potential shared gene signatures between periodontitis and multiple sclerosis
BMC Oral Health (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.