Abstract
We developed a physics-informed neural network based on a mixture of Cartesian grid sampling and Latin hypercube sampling to solve forward and backward modified diffusion equations. We optimized the parameters in the neural networks and the mixed data sampling by considering the squeeze boundary condition and the mixture coefficient, respectively. Then, we used a given modified diffusion equation as an example to demonstrate the efficiency of the neural network solver for forward and backward problems. The neural network results were compared with the numerical solutions, and good agreement with high accuracy was observed. This neural network solver can be generalized to other partial differential equations.
Similar content being viewed by others
Introduction
Partial differential equations (PDEs), especially second-order PDEs, have been extensively used in physics, engineering, finance, and other fields. Some simple PDEs can be solved analytically, but most complex PDEs rely on numerical solutions, which are usually divided into forward and backward problems. Common numerical methods include finite difference1,2,3,4,5,6, finite element7,8,9,10,11,12,13, and Lagrange multiplier14,15,16,17,18. These conventional methods have been extensively applied to solve forward PDEs in various practical problems. However, the deep neural network (DNN) provides another solution for complex nonlinear PDEs without the domain discretization used in numerical methods and is thus suitable for forward and backward problems19,20,21,22,23.
DNN is exhibiting major advancement in solving PDEs and has attracted increasing attention in various research areas due to its universal approximations24,25,26,27. However, a large amount of labeled data is usually required for training DNN-based models to solve PDEs, and such data are often unavailable in many physical applications. To overcome this disadvantage, researchers have proposed a novel DNN-based neural network called the physics-informed neural network (PINN) help to reduce the needed training time in a physics-informed manner; in this network, physics-informed loss functions are constructed based on PDE residuals20,28,29,30,31. Generally, residual design plays an important role in PINN approximations, so residual DNN is a highly effective type of neural network32,33,34. PINN can encode any underlying physics law such that the differential operators in the governing PDEs can approximated by automatic differentiation35,36. With such advantages, PINNs have been applied extensively to solve complex PDEs in various application areas in recent years37,38,39,40,41,42,43,44,45,46,47. On the one hand, an increasing number of studies are being conducted to examine the approaches for building improved PINN models by incorporating such models into other methods. For example, Karniadakis et al. introduced a systematic PINN model for the first time and presented a series of PINN variants, including Bayesian PINN48, fractional PINN49, extended PINN50, parareal PINN51, non-Newtonian PINN52, hp-variational PINN53, and nonlocal PINN54. These extended PINN models were constructed to approximate various forward PDEs, either linear or nonlinear, in application areas ranging from engineering to finance. On the other hand, PINN models have also been extended to backward problems, such as advection-dispersion equations55, stochastic problems56, flow problems57, and conservation laws58. In these backward problems, the training data are inputted into DNNs to screen the unknown parameters in the PDEs by constructing PINN loss functions.
In DNN, data sampling is another important factor for solving second-order or higher-order PDEs21,59,60,61. Latin hypercube sampling (LHS) filters the variance associated with the additive components of a transformation, and it is a powerful sampling method for data analysis in nearly every field of computational science, engineering, and mathematics62,63,64,65. In LHS, because the sample space is divided into a series of subspaces that are randomly paired, the LHS algorithm iterates to determine optimal pairings on the basis of some specified criteria. To solve PDEs using neural networks, LHS28 and simple random sampling (RS)66,67 can be used to sample data sets. LHS and RS are usually employed to solve PDEs in regular domains. However, other sampling methods based on domain decomposition or irregular domains have also been developed to solve PDEs50,68,69.
In this study, we developed an improved LHS method, namely, GLHS, where LHS and Cartesian grid sampling (GS) are merged and optimized to deal with a data set under the periodic boundary condition that commonly appears in the theory and simulation of polymer chains under bulk conditions. We are aware that in polymer physics, especially in self-consistent field theory (SCFT), the modified diffusion equation (MDE), which is a parabolic-type second-order PDE, is a key equation in SCFT for Gaussian and wormlike chains70,71,72. Several classical numerical methods have been developed to solve MDEs in SCFT and have achieved great success in reproducing the microstructures and properties of self-assembled polymer chains73,74,75,76,77,78. Recently, Chen et al. trained the traditional DNN to solve the MDE in diblock copolymer systems and the static Schrodinger equation in quantum systems, and the efficiency of the solver was analyzed67,79. In this work, we developed a PINN with residual units, which combines with the GLHS, to solve the forward and backward MDEs used in polymer physics.
Several important issues are addressed in the current study. In the following section, we describe the PINN with residual units and mixed sampling method. Then, we solve the forward and backward problems in MDEs as examples to examine the PINN solver based on the mixed data sampling by optimizing the parameters in PINN and mixed data samplings. We also compare the PINN to the numerical results and the traditional NN to verify the accuracy and efficiency of the PINN. The research summary is presented in final section.
Neural network and data sampling
PINN with residual units
We describe PINN with residual units, as shown in Fig. 1. To solve the complex problems in network convergence caused by gradient disappearance or network degradation in the traditional neural network80,81, we apply the neural network based on residual units to solve second-order PDEs. We describe a residual unit in Fig. 1a, where the input layer (IL) is a neural network layer with weight, biases, and an activation function. For this network layer, the output of tensor \(\textbf{X}^{i-1}\) fed into the network is
where \(\textbf{W}^{i} \in \mathbb {R}^{\textbf{n}_{i-1} \times \textbf{n}_i}\) is the weight parameter in the network layer, \(\textbf{b}^{i} \in \mathbb {R}^{\textbf{n}_{i}}\) is the bias parameter in the network layer, and \(\textbf{n}^{i}\) represents the network width of the current layer, that is, the number of neurons. The activation function \(\sigma (\cdots )\), which is nonlinear, is the key factor in the universal approximation of the neural network. In general, the activation function selects nonlinear functions, such as sigmoid and tanh. Here, we choose tanh as the activation function22,66, i.e.,
The output layer (OL) is an ordinary network layer with weights and biases. For input tensor \(\textbf{X}^{i-1}\), its output can be expressed as
The residual unit is constructed by combining IL and OL, as illustrated in Fig. 1a. First, input tensor \(\textbf{X}^{i-1}\) in the residual unit goes through IL and OL to obtain
Second, input tensor \(\textbf{X}^{i-1}\) is connected by jump identity, and the output is obtained by the tanh activation function after accumulation with \(f(\textbf{X}^{i-1})\), namely,
This residual unit design enables us to transmit the input data directly from the lower network layer to the higher layer. This process differs from the common stacking of two neural network layers. Therefore, the residual unit can facilitate the optimization process of the neural network and solve the problem of network degradation to a certain extent.
Parameterized parabolic-type PDEs, such as MDE, can be expressed in a general form as follows:
where \(u(\textbf{r}, t)\) denotes the solution with respect to the time and space variables, \(N[u(\textbf{r}, t);\lambda ]\) represents the differential operator parameterized by \(\lambda \), \(\Omega \) is the spatial definition domain, and \(u_t\) denotes the partial derivative with respect to time t.
In accordance with the universal approximation of neural networks19,82, the solution \(u(\textbf{r}, t)\) in PDE can be equivalently expressed through DNN. In the current study, we use a PINN with residual units to solve PDE, as shown in Fig. 1b, instead of a common neural network. In such a PINN, the initial variables are inputted into the input layer and sent to several residual units (Fig. 1a), where the solutions are assessed by the physics-informed loss functions. Then, we obtain the solution \(u(\textbf{r}, t)\) in the output layer.
We illustrate the calculations for the partial derivatives with physical constraints in the solution process in Fig. 1c and d, and the chain derivative rule based on automatic differentiation is used35. In the PINNs, the forward and backward solutions in the PDEs are learned by optimizing the loss functions with physical information. The forward solution process for the PDE in the neural networks is shown in Fig. 1c. The physical constraints of the differential equation can be defined as
The representation network needs to learn that the judgment mode of the solution of the differential equation is \(\Gamma _e \rightarrow 0\), which simultaneously satisfies the physical constraints with the periodic boundary condition (PBC) \(\Gamma _p \rightarrow 0\), and the initial condition \(\Gamma _i \rightarrow 0\). Then, we can define the total loss function as
where \(\theta _0\) denotes an intermediate variable that includes the parameters appearing in \(\Gamma _e\), \(\Gamma _p\), and \(\Gamma _i\). The learning task stops in the forward process when \(J_0(\theta _0) \rightarrow 0\). For the reverse problem, as shown in Fig. 1d, the network needs to satisfy \(\Gamma _e \rightarrow 0\) and the constraints created by existing data in the network \(\Gamma _b \rightarrow 0\), which will be described in detail in solving MDE section. In the backward process, the gradient of a certain underlying output can be expanded as
Equation (9) shows that the gradient of the bottom output of the network can be decomposed into two terms. The first term indicates that the wrong signal can be directly transmitted to the bottom without any intermediate weight matrix transformation, thus alleviating the problem of gradient dispersion to a certain extent. The gradient will not disappear even if the weight of the intermediate layer matrix is small. The residual unit, which has been successfully applied to image recognition32,34, provides an efficient tool to solve the backward problems in PDEs in the current study.
Mixed GS and LHS method
The basic problem in solving PDEs by using a neural network is to produce results that satisfy the physical conditions in the differential equations, where the data points in the defined domain are fed to the neural network. Thus, selecting appropriate data points in the training process of DNNs is crucial. In the current work, we adopt a mixed sampling method (GLHS), i.e., mixture of Cartesian GS and LHS, in the PINN solver.
The data sampling distribution of GLHS. Take a total of 20 data, 2 dimensions, each dimension in the domain of [0,1] as an example. The blue dots represent the Cartesian grid data points and the crossed dots denote the data in LHS. (a) The data sampling on each dimension; (b) The overall data distribution after mixing.
Comparison of three types of sampling. The blue Y-axis on the left of the figure represents the number of data \(N_{1,ij}\) in 100 equal square data collections area of \(\Omega _1=[0,1;0,1]\), and the purple Y-axis on the right represents the number of data \(N_{2,ij}\) in 100 data collections area of \(\Omega _2=[0,0.1;0,0.1]\). The box represents the data distribution within the range of 25%-75% of \(N_{1,ij}\) and \(N_{2,ij}\). The solid line in the box is the median line of data distribution, the line cap represents the maximum and minimum value of data distribution. The red dotted line represents the ideal number of data distributions, 20,000 and 200 respectively.
For the GLHS method, we assume that the number of data points in Cartesian GS is \(\alpha {N}\), where N is the total number of data points and \(\alpha \) is a proportionality coefficient, i.e., \(\alpha \in [0,1]\). All the data are located in the domain \(\mathbb {R}^{\Omega }\). We allocate the \(\alpha {N}\) data points to the grid points where the n-dimensional Cartesian grids are equally divided into INT(\(\root n \of {\alpha {N}}\)) grids in one dimension. INT(\(\cdots \)) denotes the integer part of the number. This condition means that a total of \(N_G\) = [INT\((\root n \of {\alpha {N}})]^{n}\) data points are sent to the grid in an n-dimensional space, where one data point corresponds to one grid point. Then, the remaining data points, \(N_L\) are sampled by the LHS method, where \(N_L = N - [\)INT(\(\root n \of {\alpha {N}})]^{n}\). In LHS, the \(N_L\) data points are sent to \(N_L\) equal subdomains for RS, where each subdomain corresponds to one LHS data point62,64. To effectively describe the GLHS method, we show a simple example in Fig. 2, where \(\alpha =0.5\), \(N=20\), \(n=2\). For simplicity, we use the definition domain \(\Omega _1=[0, 1;0, 1]\) as an example (other examples are given in Fig. S1 of Supplementary information). The blue dots represent the Cartesian grid data points, and the crossed dots denote the data points in LHS. Clearly, 3 data points exist in the Cartesian grid sampling, and 11 data points are present in LHS, where the domain is divided into 11 subdomains in one dimension and each data point in a subdomain is randomly sampled, as shown in Fig. 2a. We expect that the extracted random data are evenly distributed in the definition domain, so a linear cumulative density function, CDF\((x)=\Omega ^{-1}{x}\), is used in each dimension for LHS. Then, the total data points in two dimensions are calculated by the Cartesian product, i.e., total of 9 data points in the Cartesian grids. Meanwhile, 11 data points are randomly paired into two dimensions via a cumulative linear density function, \(\Omega _1=[0, 1;0, 1]\), as shown in Fig. 2b. Finally, the Cartesian grid and LHS data are randomly mixed to obtain GLHS data as the final input data for PINN.
We present an example to illustrate the advantages of the GLHS method in Fig. 3, where three types of sampling, namely, RS, LHS, and GLHS, are compared. In each sampling type, two types of two-dimensional definition domains, i.e., \(\Omega _1=[0, 1;0, 1]\) and \(\Omega _2=[0, 0.1;0, 0.1]\), are used as examples. A total of 2,000,000 and 20,000 data points are randomly imported into the two definition domains. To examine the uniformity of the data distribution in \(\Omega _1\) and \(\Omega _2\), we divide the two definition domains into 100 equal square data collection areas and labeled the number of data points in each data collection area as \(N_{1,ij}\) and \(N_{2,ij}\) over all the regions in \(\Omega _1\) and \(\Omega _2\). The subscripts i and j denote the ij-th collection area in the two dimensions. We plot more detailed data distributions in the two-dimensional definition domains in Fig. S2 of Supplementary information. Thus, 100 data collection areas are distributed in each definition domain for RS, LHS, and GLHS. We sort \(N_{1,ij}\) and \(N_{2,ij}\) into a sequence on the basis of their values and label the \(N_{1,ij}\) and \(N_{2,ij}\) in the middle 25–75% of the sequence as a square box with dotted lines, as shown in Fig. 3a–c. Among the three sampling types, RS has the largest box and GLHS has the smallest one in the \(\Omega _1\) domains, indicating that GLHS possesses the most uniform data distributions in the definition domain.For the \(\Omega _2\) domains, GLHS also has the most uniform data distribution, but the data points at the middle of the sequence have values below 200, which is the average number of data points for the collection data in \(\Omega _2\) domains. This result may due to the reason that the amount of input data is not large enough in the \(\Omega _2\) domains. Recently, simple LHS with point transformation was used to increase the uniformity of data distribution83. In this study, we observe that GLHS has an obvious advantage over RS and LHS, although RS and LHS have similar sampling procedures in the uncertainty analysis62.
Solving for forward and backward MDE
In polymer physics, MDE is a key equation in self-consistent field theory and has been numerically solved by many methods70,71. We adopt MDE as an example to illustrate the use of PINN based on GLHS in solving forward and backward problems. First, we present the general form of forward and backward MDE. Second, we discuss the efficiency of GLHS and PBC loss functions in solving for MDE. Lastly, we discuss the forward and backward problems in MDE solutions by using PINN based on GLHS.
Problem setup
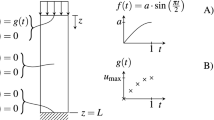
As an example, we take a simple form of forward and backward MDE with the initial conditions and PBCs, which can be expressed as
where the initial condition is \(u(x,0) = 1\) and the periodic boundary condition is \(u(0,t) = u(L,t)\). Here, only two dimensions are used; L is the period in the x dimension. And when \(\lambda \) is a given parameter, such as \(\lambda = 7\) used in this work, the problem becomes a forward problem. When \(\lambda \) is an unknown parameter, solving the equation becomes a backward problem. MDE is a linear, second-order, parabolic-type PDE in which the forward problem is addressed by DNN28,79. However, the backward problem in MDE still needs to be understood. Here, we use PINN based on GLHS to solve the forward and backward problems in MDE.
Optimization scheme for GLHS and PINN
The neural network and sampling method should be optimized when used to solve a special PDE. In this study, the core issue in GLHS is how to find the optimizing mixture coefficient \(\alpha \) as the special PDE; meanwhile, depth D and width W in the neural network are also important parameters in PINN and should be optimized. Given that the two types of parameters are independent, we adopt the independent variable method to optimize the GLHS and PINN parameters.
Comparison of training time (t) and standard error (\(\sigma \)) with two data sampling methods in the PINN with residual units. (a) The mixture of GS and RS at \(\alpha :(1-\alpha )\) ratio, where \(\alpha =0\) denotes simple random sampling; (b) The mixture of GS and LHS, i.e., GLHS, at \(\alpha :(1-\alpha )\) ratio, where \(\alpha =0\) denotes simple LHS sampling. The purple bars on the left represent the standard error of network training results, and the abscissa (0.001–0.5) is amplified by log\(_{10}\). The yellow bars on the right represent the time spent on network training.
The primary issue in GLHS, which is how to find optimizing mixture coefficient \(\alpha \) as the special PDE, is solved in PINN. We further describe the GLHS by comparing it with other mixture sampling methods in solving MDE using PINN, as shown in Fig. 4. For a comparison, we adopt two types of mixture samplings, namely, the mixture of random sampling and grid sampling, as shown in Fig. 4a, and GLHS in Fig. 4b, respectively. The data used are from solving for MDE in Eq. (10) with PBC of \(L=1.0\), where the number of residual units is 6 and the width of neural network layer is 20 in PINN. Then, we use a total of \(N=301 \times 301\) data points in the mixture of random and grid samplings as well as GLHS, where the number of data points in Cartesian GS is \(\alpha {N}\). To explain the efficiency of the sampling method, we employ sampling standard errors as follows:
where \(u_i\) is the PINN solving value and \(u_{0i}\) is the numerical value solved by the Crank-Nicholson method, which has been used for MDE in previous simulation calculations74,84,85,86. The sum takes over all the discrete data points in the definition domains used in the Crank-Nicholson method, and N is the total number of discrete data points.
In the mixture of random and grid sampling case, the results indicate that it is difficult to optimize \(\sigma \) and t as \(\alpha \) varies; t is the corresponding training time, as shown in Fig. 4a. In particular, when standard error \(\sigma \) has the minimum value, training time t is the longest. In the GLHS case, \(\sigma \) can reach its minimum value and the t minimum value simultaneously, as shown in Fig. 4b. We optimize GLHS with \(\alpha =0.5\) which indicates that the minimum error and training time can be achieved when the data numbers in GS are equal to those in LHS. Figure 4 also show the results of RS (\(\alpha =0.0\) in Fig. 4a), LHS (\(\alpha =0.0\) in Fig 4b) and GS (\(\alpha =1.0\) in Fig. 4a and b) by using PINN. We can find that in terms of t and \(\sigma \), these three sampling methods are not good choices for data samplings, comparing to the GLHS with \(\alpha =0.5\). These results agree with those data distributions described in Fig. 3. We note that the LHS has been used in PINN28. Here, we try to use the GLHS instead of LHS to enhance the efficiency for solving PDEs in PINN.
PBC is important in PDE, especially when handling a bulk polymer system. The boundary condition has been strengthened in previous studies when designing the structure of neural networks36,87,88. Thus, we consider PBC optimization when choosing the proper network parameters. As illustrated in Eq. (10), PBC in spatial period L can be regarded as \(u(0,t)=u(L,t)\). The PBC in numerical methods can be done by setting the calculation cell size to L, which satisfies \(u(0,t)=u(L,t)\)72,74. However, the PBC in neural networks has to satisfy the left condition \(u(x,t)=u(x-L,t)\), the right condition \(u(x,t)=u(x+L,t)\), or both, that is, the squeeze period condition where \(x \in [0,L]\). To identify the optimized PBC among the three PBCs, we construct the following types of loss functions.
where \(x_i \in [0,L]\) and \(t_i \in [0,T]\). Summation is performed over the two domains. Equation (12) is commonly used in numerical methods and listed here for comparison.
The training processes for the four PBC loss functions are shown in Fig. 5. The standard errors \((\sigma _p)\) from the results of PINN, i.e., u(x, t), and from the results of the Crank-Nicholson numerical method, i.e., \(u_0{(x,t)}\) are also listed on the right side. In Type 1, as shown in Fig. 5a, although loss function \(J_{p1}\) converges to a desired value, standard error \(\sigma _{p1}\) is too large to achieve correct solutions. For Types 2 and 3, as shown in Fig. 5b and c, loss functions \(J_{p2}\) and \(J_{p3}\) can converge to the desired values, and standard errors \(\sigma _{p2}\) and \(\sigma _{p3}\) are still able to converge to the desired values. However, when we use the squeeze period condition, loss function \(J_{p4}\) and standard error \(\sigma _{p4}\) can converge to the desired value in the training processes. Furthermore, we calculate the Pearson correlation coefficient, \(\rho _{X,Y}\), between standard error \(\sigma _p\) and loss function \(J_p\), which can be generally defined as
where X and Y denote standard error \(\sigma _p\) and loss function \(J_p\), respectively. \(\sigma _X\) denotes the standard error of X, and cov(X, Y) represents the covariance between X and Y. Then, we list the \(\rho _{X,Y}\) for the four types of PBC in Table 1. The data indicate that the first three types of PBC have a strongly correlation between \(\sigma _p\) and \(J_p\), but \(\sigma _p\) and \(J_p\) have almost no correlations in the squeeze boundary condition. This result indicates that the squeeze period method is feasible and can effectively improve the training accuracy of solving PDEs with PBC in PINN.
Then, we optimize the neural network parameters, namely, the depth and width in PINN(D and W). D is the number of residual units, and W is the number of neural units in the layer. D, W, and the amount of trained data(N) have a great influence on the output accuracy of the neural network. We train PINN in a multi-parameter space of D, W, and N, where \(D \in [3,4,5,6,7,8]\), \(W \in [10,15,20,25,30]\), and \(N \in [2000,4000,6000,8000,10000,20000]\), as indicated in Table 2, where only several typical combinations are shown. Indeed, \(D \times W \times N=180\) combinations exist in the full parameter space of [D, W, N]. Other detailed combinations can be found in Figs. S3 and S4 of Supplementary information.
Comparison of PINN results with numerical results. (a) The overall view in the two dimensional space of PINN results u(x, t); (b) Diagram of the difference between PINN results and numerical results; (c) The accuracy in one-dimensional space with given \(t={0.00,0.25,0.50,0.75,1.00}\); (d) The accuracy in one-dimensional space with given \(x={0.00,0.25,0.75}\). The blue dotted line represents the numerical results and the red line represents PINN results.
We use loss functions J and standard errors \(\sigma \) to screen the desired parameters. The optimal parameter corresponds to the minimum \(\sigma \), which is the combination with \(D=6\), \(W=20\), and \(N=20000\). Generally, an increase in N leads to a decrease in J and \(\sigma \), indicating a simple relationship. However, the optimal output relies on the complex combination of D and W. The data in Table 2 reveal that the best optimization combination is the parameter combination of \(D=6\), \(W=20\), and \(N=20000\). Generally, using a number of hidden layers during training results in a large precision loss, but the optimization combination here shows that this is not the case89.
Forward and backward solutions
First, we discuss the forward solutions for MDE by PINN. We use the optimal PINN parameters, \(D=6\), \(W=20\) and \(N=20000\), and the GLHS parameter, \(\alpha =0.5\), to solve the forward and backward problems in MDEs appearing in Eq. (10). We take the definition domains \(x \in [0,1]\) and \(t \in [0,1]\), and to evaluate the accuracy of the PINN solution u, we define the relative errors with respect to the numerical solutions \(u_0\) as follows:
where subscripts \(x_i\) and \(t_j\) correspond to definition domains x and t,respectively. We plot the neural network results and compare them with the numerical results in Fig. 6. We adopt an overall view in the two-dimensional space, as shown in Fig. 6a and b. The results indicate that the PINN results have high accuracy within \(10^{-3}\) distributed in the defintion domain. Then, we examine the accuracy in one-dimensional space with given x or t, as shown in Fig. 6c and d, respectively. The data confirm that our PINN results vary with the given x or t. Furthermore, we illustrate the data for the given x or t in Table 3, where the relative errors are also listed.
For the inverse problem in MDE, discovering the unknown parameters is difficult due to the complex physical constraints and gradient disappearance. Unlike in the case where a sparse regression method is employed to determine PDE by time series measurements in the spatial domain21, in this study, we design an interleaved training method with discontinuous double-loss functions \(\Gamma _e\) and \(\Gamma _b\). Loss function \(\Gamma _e\) is defined in Eq. 7, and \(\Gamma _b\) can be defined as
The sum over all the definition domains, \(u_0{(x_i,t_i)}\),is the numerical solution and taken as the standard value. In this method, we search for unknown parameter \(\lambda \) through loss function \(\Gamma _e\) and optimize the network solution through \(\Gamma _b\). That is, \(\Gamma _e\) is optimized to screen parameter \(\lambda \) at the first training stage. Then, we lock up the parameter \(\lambda \) to optimize the network solution \(\Gamma _b\) and obtain a high-accuracy network solution.
Specifically, we construct the PINN with four residual units, each of which has a full-connection layer width of 20 neural units, to solve the inverse problems. We optimize the network parameters and unknown parameters \(\theta _0{(w,b,\lambda )}\) via the loss function \(J_0{(\theta _0)}\) at the first stage. Then, we lock up parameter \(\lambda \) and optimize \(\theta _1{(w,b)}\) by using loss function \(J_1{(\theta _1)}\) at the second stage until the loss function reaches \(10^{-5}\), as shown in Fig. 7. We label the blue and brown dotted lines as \(\lambda =10\) and \(J_0{(\theta _0)}=0\), respectively. The results indicate that parameter \(\lambda =10.00111\) at \(t=108s\) when \(J_0{(\theta _0)}\le 10^{-5}\), as shown in Fig. 7a. At this point, we lock up parameter \(\lambda \), and loss function \(J_0{(\theta _0)}\) is automatically awitched to \(J_1{(\theta _1)}\). Then, network parameter \(\theta _1{(w,b)}\), which determines the network solution, is further optimized at the second stage. We observe that the loss function is discontinuous at this point, as shown in the inserted part in Fig. 7b. Generally, the loss function exhibits a sudden drop when the optimizer switches in the training process52. We also observe a discontinuous loss function in a small region of \(\Delta {J}=0.05\) when the loss function switches. In the current study, we obtain parameter \(\lambda \) with high accuracy. The absolute error of parameter \(\lambda \) is 0.0011, the relative error is \(\delta _\lambda =0.011\%\), and the standard error is \(\sigma _\lambda =0.0046\). In this work, we present a high-efficiency interleaved training method to search for the unknown parameters in the reverse problem in MDE, and it can be reasonably extended to other PDEs.
Comparison between PINN and NN
Comparison of training time (t) and standard error (\(\sigma \)) between PINN with residual units and traditional NN, Here, the width of neural network W is 20, data sampling number N is 20000, and the mixture coefficient of GLHS \(\alpha \) is 0.5 in both cases. The yellow and green bars denote the training time t in the PINN and traditional NN cases, respectively, while the blue dots in the bars represent the corresponding standard error \(\sigma \).
To illustrate the advantages of PINN with residual units, we compare the results from the PINN to the traditional NN by the standard error \(\sigma \) and training time t, as shown in Fig. 8. Here, the traditional NN is constructed without the residual units, which is similar to the previous works22,25. For convenient comparison, the other conditions are set to be the same as in both cases, including the data samplings (GLHS with \(\alpha \)=0.5), the input data number( \(N=20000\)), and the width of network (\(W=20\)). Since the residual unit consists of two neural network layers, we take two network layers in traditional NN as a layer for more convenient comparison, so that there are 2D network layers in the traditional NN. Then, we only change the depth of PINN D to output the training times and standard errors in both cases. The results indicate that with the increase of the number of neural network layers, the advantages of PINN with residual units become more and more obvious. In particular, when D = 6, the traditional neural network encounters a gradient explosion, leading to the meaningless output of loss and the other parameters, as shown in Fig. 8; when D = 7 and 8, the traditional NN can hardly be optimized bacause of the problem of gradient disapperance. This problem results in the extremely short training times t and large standard errors \(\sigma \). In addition, even in the shallow networks with small D, the training time of PINN with residual units is still superiority to those of traditional NN, as shown in Fig. 8. Furthermore, we have compared the efficiencies between the PINN and traditional NN with the other W, and the similar results were obtained, please see Tabel S1 in Supplementary information. We expect this PINN with residual untis based on the mixed data sampling to be applied to other works, especially to the three-dimensional PDEs, in the future.
Summary
We developed a PINN based on GLHS to solve the forward and backward MDEs by optimizing the corresponding parameters. This solver provides high efficiency and accuracy in solving forward and backward problems in one-dimensional MDEs, and it effectively avoids the problems of gradient disappearance and network degradation existing in traditional feedforward neural networks. For the neural network, we properly designed residual units for PINN to solve MDE, and squeeze PBC was considered. For data sampling, we considered the mixture of GS and LHS. We believe that this method can also used in other dimensional, and we will confirm the view in the future.
Then, we optimized the parameters used in PINN by considering the loss function of PBC. Specifically, the depth and width of the neural network, D and W, were optimized. The results indicated that the squeeze condition is suitable for MDE. We also optimized GLHS data sampling by adjusting the mixture coefficient \(\alpha \). The results revealed that the parameter combination \([D,W,N,\alpha ]\) should be optimized to [6, 20, 20000, 0.5] in the given MDE with high precision. We demonstrated how the hybrid solver deals with forward and backward problems in a special MDE. We compared the neural network solvers results to the numerical solutions and found good agreement. For the forward MDE, we obtained high-accuracy PINN solutions within \(10^{-3}\) by analyzing the errors between the PINN solutions and the numerical results. For the inverse problem in MDE, we designed the ITM method to screen the unknown parameters. Unknown parameter \(\lambda \) was locked up with relative error \(\delta _\lambda =0.011\%\) and standard error \(\sigma _\lambda =0.0046\). This PINN with residual units based on mixed data sampling GLHS can be generalized to other cases for other PDEs.
Data availability
The datasets used and/or analysed during the current study available from the corresponding author on reasonable request.
References
Meerschaert, M. M. & Tadjeran, C. Finite difference approximations for fractional advection-dispersion flow equations. J. Comput. Appl. Math. 172, 65–77 (2004).
Inan, B. & Bahadir, A. R. Numerical solution of the one-dimensional Burgers’ equation: Implicit and fully implicit exponential finite difference methods. Pramana 81, 547–556 (2013).
Alikhanov, A. A. A new difference scheme for the time fractional diffusion equation. J. Comput. Phys. 280, 424–438 (2015).
Gao, G., Sun, H. & Sun, Z. Stability and convergence of finite difference schemes for a class of time-fractional sub-diffusion equations based on certain superconvergence. J. Comput. Phys. 280, 510–528 (2015).
Moghaddam, B. P. & Machado, J. A stable three-level explicit spline finite difference scheme for a class of nonlinear time variable order fractional partial differential equations. Comput. Math. with Appl. 73, 1262–1269 (2017).
Elango, S. et al. Finite difference scheme for singularly perturbed reaction diffusion problem of partial delay differential equation with nonlocal boundary condition. Adv. Differ. Equ. 2021, 115 (2021).
Ying, L. Partial differential equations and the finite element method. Math. Comput. 76, 1693–1694 (2007).
Jiang, Y. & Ma, J. High-order finite element methods for time-fractional partial differential equations. J. Comput. Appl. Math. 235, 3285–3290 (2011).
Gunzburger, M. D., Webster, C. G. & Zhang, G. Stochastic finite element methods for partial differential equations with random input data. Acta Numer 23, 521–650 (2014).
Lehrenfeld, C., Olshanskii, M. A. & Xu, X. A stabilized trace finite element method for partial differential equations on evolving surfaces. SIAM J. Numer. Anal. 56, 1643–1672 (2018).
Li, C. & Wang, Z. The local discontinuous Galerkin finite element methods for Caputo-type partial differential equations: Mathematical analysis. Appl. Numer. Math. 150, 587–606 (2020).
Lai, J., Liu, F., Anh, V. V. & Liu, Q. A space-time finite element method for solving linear riesz space fractional partial differential equations. Numer Algorithms 88, 499–520 (2021).
Du, S. & Cai, Z. Adaptive finite element method for dirichlet boundary control of elliptic partial differential equations. J. Sci. Comput. 89, 36 (2021).
Xu, Y., Chen, Q. & Guo, Z. Optimization of heat exchanger networks based on Lagrange multiplier method with the entransy balance equation as constraint. Int. J. Heat Mass Transf. 95, 109–115 (2016).
Hamid, M., Usman, M., Zubair, T. & Mohyud-Din, S. T. Comparison of Lagrange multipliers for telegraph equations. Ain Shams Eng. J. 9, 2323–2328 (2017).
Antoine, X., Shen, J. & Tang, Q. Scalar Auxiliary Variable/Lagrange multiplier based pseudospectral schemes for the dynamics of nonlinear Schrödinger/Gross-Pitaevskii equations. J. Comput. Phys. 437, 110328 (2021).
Lee, H. G., Shin, J. & Lee, J.-Y. A high-order and unconditionally energy stable scheme for the conservative Allen-Cahn equation with a nonlocal Lagrange multiplier. J. Sci. Comput. 90, 51 (2022).
Yang, J. & Kim, J. Numerical simulation and analysis of the Swift-Hohenberg equation by the stabilized Lagrange multiplier approach. Comput. Appl. Math. 41, 20 (2022).
Lecun, Y., Bengio, Y. & Hinton, G. Deep learning. Nature 521, 436–444 (2015).
Raissi, M., Perdikaris, P. & Karniadakis, G. E. Machine learning of linear differential equations using Gaussian processes. J. Comput. Phys. 348, 683–693 (2017).
Rudy, S. H., Brunton, S. L., Proctor, J. L. & Kutz, J. N. Data-driven discovery of partial differential equations. Sci. Adv. 3, e1602614 (2017).
Brink, A. R., Najera-Flores, D. A. & Martinez, C. The neural network collocation method for solving partial differential equations. Neural Comput. Appl. 33, 5591–5608 (2021).
Chen, Z., Churchill, V., Wu, K. & Xiu, D. Deep neural network modeling of unknown partial differential equations in nodal space. J. Comput. Phys. 449, 110782 (2022).
Mistry, A., Franco, A. A., Cooper, S. J., Roberts, S. A. & Viswanathan, V. How machine learning will revolutionize electrochemical sciences. ACS Energy Lett. 6, 1422–1431 (2021).
Hauptmann, A. & Cox, B. Deep learning in photoacoustic tomography: current approaches and future directions. J. Biomed. Opt. 25, 112903 (2020).
Alipanahi, B., Delong, A., Weirauch, M. T. & Frey, B. J. Predicting the sequence specificities of DNA- and RNA-binding proteins by deep learning. Nat. Biotechnol. 33, 831–838 (2015).
Lake, B. M., Salakhutdinov, R. & Tenenbaum, J. B. Human-level concept learning through probabilistic program induction. Science 350, 1332–1338 (2015).
Raissi, M., Perdikaris, P. & Karniadakis, G. E. Physics-informed neural networks: A deep learning framework for solving forward and inverse problems involving nonlinear partial differential equations. J. Comput. Phys. 378, 686–707 (2019).
Owhadi, H. Bayesian numerical homogenization. Multiscale Model. Simul. 13, 812–828 (2015).
Raissi, M., Perdikaris, P. & Karniadakis, G. E. Inferring solutions of differential equations using noisy multi-fidelity data. J. Comput. Phys. 335, 736–746 (2017).
Karniadakis, G. E. et al. Physics-informed machine learning. Nat. Rev. Phys. 3, 422–440 (2021).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. IEEE Conf. Comput. Vis. Pattern Recognit Proc. 770–778 (2016).
Ruthotto, L. & Haber, E. Deep neural networks motivated by partial differential equations. J. Math. Imaging Vis. 62, 352–364 (2020).
Luo, Z., Sun, Z., Zhou, W., Wu, Z. & Kamata, S. I. Rethinking ResNets: Improved stacking strategies with high-order schemes for image classification. Complex. Intell. Syst. 8, 3395–3407 (2022).
Baydin, A. G., Pearlmutter, B. A., Radul, A. A. & Siskind, J. M. Automatic differentiation in machine learning: A survey. J. Mach. Learn. Res. 18, 5595–5637 (2018).
Lu, L., Meng, X., Mao, Z. & Karniadakis, G. E. Deepxde: A deep learning library for solving differential equations. SIAM Rev. 63, 208–228 (2021).
Cai, S., Mao, Z., Wang, Z., Yin, M. & Karniadakis, G. E. Physics-informed neural networks (PINNs) for fluid mechanics: A review. Acta. Mech. Sin. 37, 1727–1738 (2022).
Viana, F. A. & Subramaniyan, A. K. A survey of Bayesian calibration and physics-informed neural networks in scientific modeling. Arch. Comput. Methods Eng. 28, 3801–3830 (2021).
Mao, Z., Jagtap, A. D. & Karniadakis, G. E. Physics-informed neural networks for high-speed flows. Comput. Methods Appl. Mech. Eng. 360, 112789 (2020).
Zhang, E., Dao, M., Karniadakis, G. E. & Suresh, S. Analyses of internal structures and defects in materials using physics-informed neural networks. Sci. Adv. 8, eabk0644 (2022).
Cai, S., Wang, Z., Wang, S., Perdikaris, P. & Karniadakis, G. E. Physics-informed neural networks for heat transfer problems. J. Heat Transf. 143, 060801 (2021).
Chen, Z., Gao, J., Wang, W. & Yan, Z. Physics-informed generative neural network: An application to troposphere temperature prediction. Environ. Res. Lett. 16, 065003 (2021).
Bai, Y., Chaolu, T. & Bilige, S. The application of improved physics-informed neural network (IPINN) method in finance. Nonlinear Dyn. 107, 3655–3667 (2022).
Taghizadeh, E., Byrne, H. M. & Wood, B. D. Explicit physics-informed neural networks for nonlinear closure: The case of transport in tissues. J. Comput. Phys. 449, 110781 (2022).
Jiang, J. et al. Physics-informed deep neural network enabled discovery of size-dependent deformation mechanisms in nanostructures. Int. J. Solids Struct. 236–237, 111320 (2022).
Kissas, G. et al. Machine learning in cardiovascular flows modeling: Predicting arterial blood pressure from non-invasive 4D flow MRI data using physics-informed neural networks. Comput. Methods Appl. Mech. Eng. 358, 112623 (2020).
Riel, B., Minchew, B. & Bischoff, T. Data-driven inference of the mechanics of slip along glacier beds using physics-informed neural networks: Case study on rutford ice stream, Antarctica. J. Adv. Model. Earth Syst. 13, e2021MS002621 (2021).
Yang, L., Meng, X. & Karniadakis, G. E. B-pinns: Bayesian physics-informed neural networks for forward and inverse PDE problems with noisy data. J. Comput. Phys. 425, 109913 (2021).
Pang, G., Lu, L. & Karniadakis, G. E. Fpinns: Fractional physics-informed neural networks. SIAM J. Sci. Comput. 41, A2603–A2626 (2019).
Jagtap, A. D. & Karniadakis, G. E. Extended physics-informed neural networks (XPINNs): A generalized space-time domain decomposition based deep learning framework for nonlinear partial differential equations. Commun. Comput. Phys. 28, 2002–2041 (2020).
Meng, X., Li, Z., Zhang, D. & Karniadakis, G. E. PPINN: Parareal physics-informed neural network for time-dependent PDEs. Comput. Methods Appl. Mech. Eng. 370, 113250 (2020).
Mahmoudabadbozchelou, M., Karniadakis, G. E. & Jamali, S. nn-pinns: Non-Newtonian physics-informed neural networks for complex fluid modeling. Soft Matter. 18, 172–185 (2022).
Kharazmi, E., Zhang, Z. & Karniadakis, G. E. hp-VPINNs: Variational physics-informed neural networks with domain decomposition. Comput. Methods Appl. Mech. Engrg. 374, 113547 (2021).
Pang, G., D’Elia, M., Parks, M. & Karniadakis, G. E. nPINNs: Nonlocal physics-informed neural networks for a parametrized nonlocal universal laplacian operator. Algorithms Appl. J. Comput. Phys. 422, 109760 (2020).
He, Q. & Tartakovsky, A. M. Physics-informed neural network method for forward and backward advection-dispersion equations. Water Resour. Res. 57, e2020WR029479 (2021).
Zhang, D., Lu, L., Guo, L. & Karniadakis, G. E. Quantifying total uncertainty in physics-informed neural networks for solving forward and inverse stochastic problems. J. Comput. Phys. 397, 108850 (2019).
Lou, Q., Meng, X. & Karniadakis, G. E. Physics-informed neural networks for solving forward and inverse flow problems via the Boltzmann-BGK formulation. J. Comput. Phys. 447, 110676 (2021).
Jagtap, A. D., Kharazmi, E. & Karniadakis, G. E. Conservative physics-informed neural networks on discrete domains for conservation laws: Applications to forward and inverse problems. Comput. Methods Appl. Mech. Eng. 365, 113028 (2020).
Han, J., Jentzen, A. & Weinan, E. Solving high-dimensional partial differential equations using deep learning. Proc. Natl. Acad. Sci. USA 115, 8505–8510 (2018).
Ruthotto, L., Osher, S. J., Li, W., Nurbekyan, L. & Fung, S. W. A machine learning framework for solving high-dimensional mean field game and mean field control problems. Proc. Natl. Acad. Sci. USA 117, 9183–9193 (2020).
Bar-Sinai, Y., Hoyer, S., Hickey, J. & Brenner, M. P. Learning data-driven discretizations for partial differential equations. Proc. Natl. Acad. Sci. USA 116, 15344–15349 (2019).
Helton, J. C., Davis, F. J. & Johnson, J. D. A comparison of uncertainty and sensitivity analysis results obtained with random and Latin hypercube sampling. Reliab. Eng. Syst. Saf. 89, 305–330 (2005).
Navid, A., Khalilarya, S. & Abbasi, M. Diesel engine optimization with multi-objective performance characteristics by non-evolutionary Nelder-Mead algorithm: Sobol sequence and Latin hypercube sampling methods comparison in DoE process. Fuel 228, 349–367 (2018).
Shields, M. D. & Zhang, J. The generalization of Latin hypercube sampling. Reliab. Eng. Syst. Saf. 148, 96–108 (2016).
Chen, Y., Wen, J. & Cheng, S. Probabilistic load flow method based on nataf transformation and Latin hypercube sampling. IEEE Trans. Sustain. Energy 4, 294–301 (2013).
Sirignano, J. & Spiliopoulos, K. DGM: A deep learning algorithm for solving partial differential equations. J. Comput. Phys. 375, 1339–1364 (2018).
Li, H., Zhai, Q. & Chen, J. Z. Neural-network-based multistate solver for a static Schrödinger equation. Phys. Rev. A 103, 032405 (2021).
Gao, H., Sun, L. & Wang, J. X. PhyGeoNet: Physics-informed geometry-adaptive convolutional neural networks for solving parameterized steady-state PDEs on irregular domain. J. Comput. Phys. 428, 110079 (2021).
Dong, S. & Li, Z. Local extreme learning machines and domain decomposition for solving linear and nonlinear partial differential equations. Comput. Methods Appl. Mech. Eng. 387, 114129 (2021).
Matsen, M. W. The standard Gaussian model for block copolymer melts. J. Phys. Condens. Matter 14, R21–R47 (2002).
Fredrickson, G. H., Ganesan, V. & Drolet, F. Field-theoretic computer simulation methods for polymers and complex fluids. Macromolecules 35, 16–39 (2002).
Fredrickson, G. H. The Equilibrium Theory of Inhomogenous Polymers (Oxford University Press, Oxford, 2006).
Matsen, M. W. & Schick, M. Stable and unstable phases of a diblock copolymer melt. Phys. Rev. Lett. 72, 2660–2663 (1994).
Drolet, F. & Fredrickson, G. H. Combinatorial screening of complex block copolymer assembly with self-consistent field theory. Phys. Rev. Lett. 83, 4317–4320 (1999).
Guo, Z. et al. Discovering ordered phases of block copolymers: New results from a generic Fourier-space approach. Phys. Rev. Lett. 101, 028301 (2008).
Song, W., Tang, P., Qiu, F., Yang, Y. & Shi, A. C. Phase behavior of semiflexible-coil diblock copolymers: A hybrid numerical SCFT approach. Soft Matter 7, 929–938 (2011).
Jiang, Y. & Chen, J. Z. Self-consistent field theory and numerical scheme for calculating the phase diagram of wormlike diblock copolymers. Phys. Rev. E 88, 042603 (2013).
Jiang, Y. & Chen, J. Z. Influence of chain rigidity on the phase behavior of wormlike diblock copolymers. Phys. Rev. Lett. 110, 138305 (2013).
Wei, Q., Jiang, Y. & Chen, J. Z. Machine-learning solver for modified diffusion equations. Phys. Rev. E 98, 053304 (2018).
Glorot, X. & Bengio, Y. Understanding the difficulty of training deep feedforward neural networks. J. Mach. Learn. Res. 9, 249–256 (2010).
Balduzzi, D. et al. The shattered gradients problem if resnets are the answer, then what is the question?. ICLM Proc. 70, 342–350 (2017).
Li, X. Simultaneous approximations of multivariate functions and their by neural networks with one hidden layer. Neurocomputing 12, 327–343 (1996).
Bihlo, A. & Popovych, R. O. Physics-informed neural networks for the shallow-water equations on the sphere. J. Comput. Phys. 456, 111024 (2022).
Li, S., Chen, P., Wang, X., Zhang, L. & Liang, H. Surface-induced morphologies of lamella-forming diblock copolymers confined in nanorod arrays. J. Chem. Phys. 130, 014902 (2009).
Chen, P., Liang, H. & Shi, A. C. Origin of microstructures from confined asymmetric diblock copolymers. Macromolecules 40, 7329–7335 (2007).
Tang, P., Qiu, F., Zhang, H. & Yang, Y. Morphology and phase diagram of complex block copolymers: ABC linear triblock copolymers. Phys. Rev. E 69, 031803 (2004).
Lagaris, I. E., Likas, A. C. & Papageorgiou, D. G. Neural-network methods for boundary value problems with irregular boundaries. IEEE Trans. Neural Netw. 11, 1041–1049 (2000).
Sun, L., Gao, H., Pan, S. & Wang, J. X. Surrogate modeling for fluid flows based on physics-constrained deep learning without simulation data. Comput. Methods Appl. Mech. Eng. 361, 112732 (2020).
Berg, J. & Nyström, K. A unified deep artificial neural network approach to partial differential equations in complex geometries. Neurocomputing 317, 28–41 (2018).
Acknowledgements
We thank for the financial supports from the Program of National Natural Science Foundation of China (Grant No. 21973070).
Author information
Authors and Affiliations
Contributions
Methodology, M.X.; Supervision, L.S.; Validation, F.Q.; Writing-original draft, F.Q.; Writing-review and editing, L.S. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fang, Q., Mou, X. & Li, S. A physics-informed neural network based on mixed data sampling for solving modified diffusion equations. Sci Rep 13, 2491 (2023). https://doi.org/10.1038/s41598-023-29822-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-29822-3
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.